A Weakened Immune Response to Synthetic Exo-Peptides Predicts a Potential Biosecurity Risk in the Retrieval of Exo-Microorganisms
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Synthesis of Peptides
2.2. Assays and Reagents
2.3. Splenocytes Isolation and T Cell Response
2.4. Flow Cytometric Analyses
3. Results
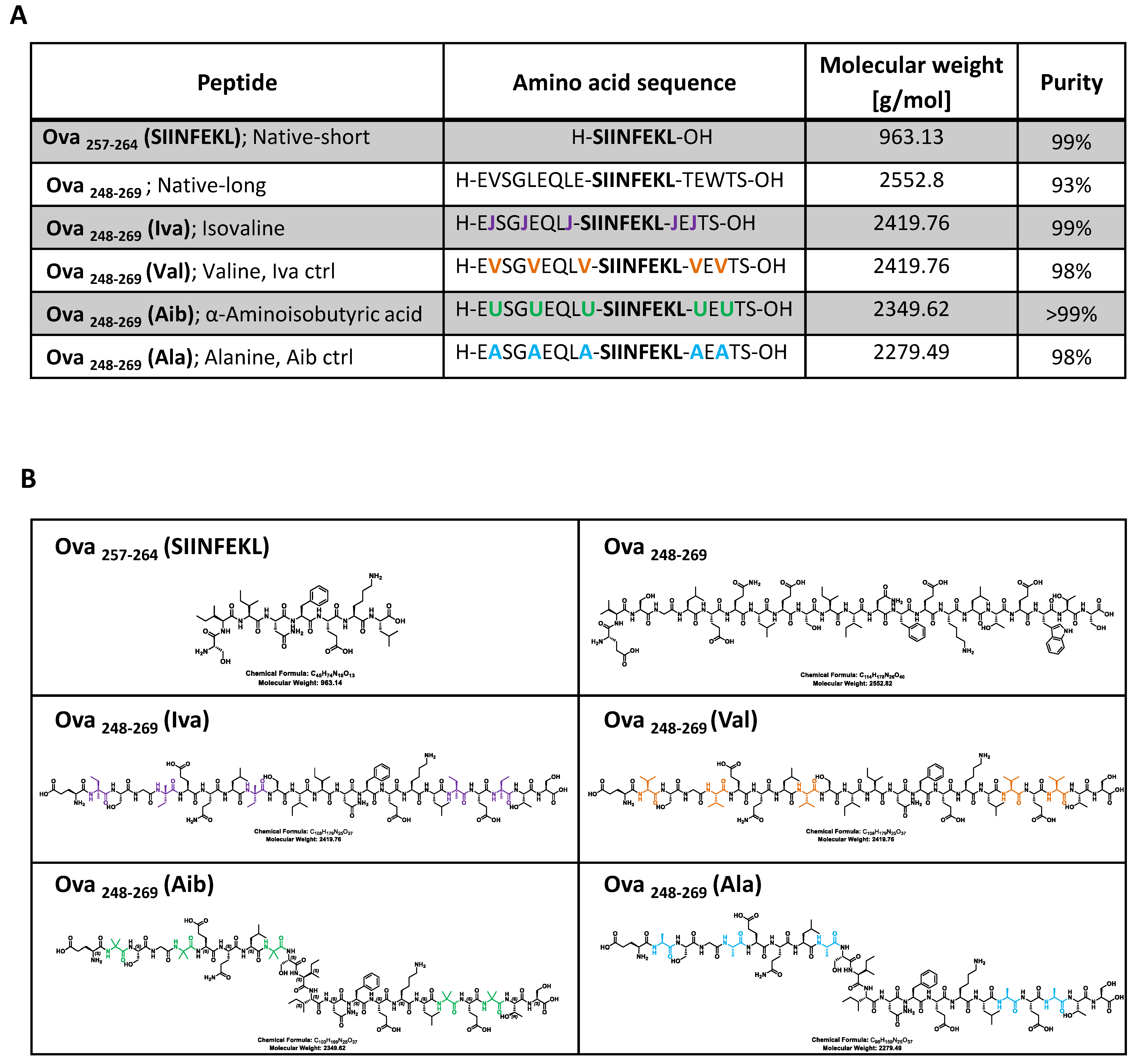
3.1. Peptide Synthesis to Generate High Purity Ova-Exo-Peptides
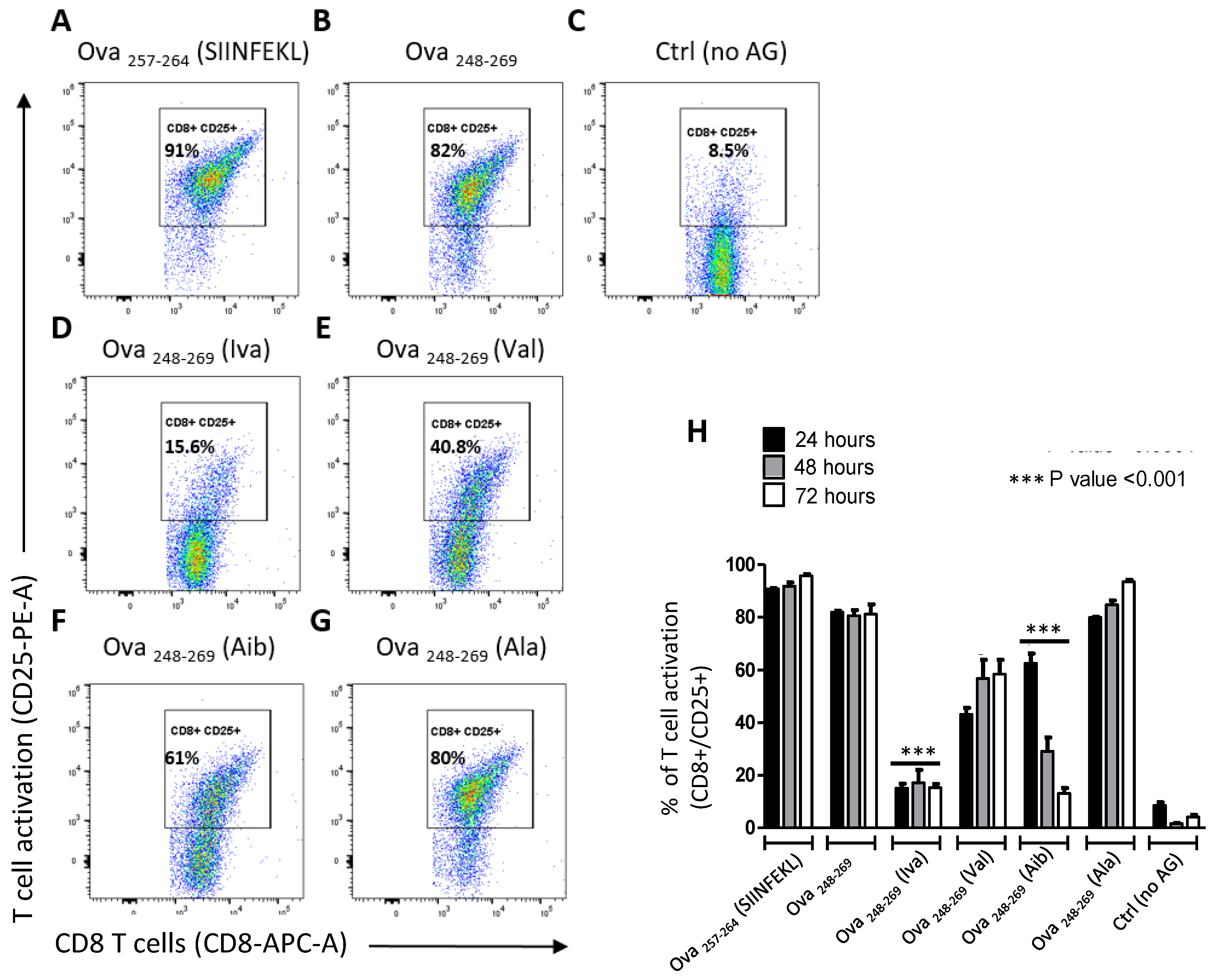
3.2. Iva- or Aib Containing Peptides Activate OT-I T Cells
3.3. Iva- or Aib Containing Peptides Induce OT-I Cell Proliferation
4. Discussion
5. Implications
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Des Marais, D.J.; Nuth, J.A., 3rd; Allamandola, L.J.; Boss, A.P.; Farmer, J.D.; Hoehler, T.M.; Jakosky, B.M.; Meadows, V.S.; Pohorille, A.; Runnegar, B.; et al. The NASA Astrobiology Roadmap. Astrobiology 2008, 8, 715–730. [Google Scholar] [CrossRef] [PubMed]
- Hays, L.E.; Graham, H.V.; Des Marais, D.J.; Hausrath, E.M.; Horgan, B.; McCollom, T.M.; Parenteau, M.N.; Potter-McIntyre, S.L.; Williams, A.J.; Lynch, K.L. Biosignature Preservation and Detection in Mars Analog Environments. Astrobiology 2017, 17, 363–400. [Google Scholar] [CrossRef] [PubMed]
- Baker, V.R.; Strom, R.G.; Gulick, V.C.; Kargel, J.S.; Komatsu, G.; Kale, V.S. Ancient oceans, ice sheets and the hydrological cycle on Mars. Nature 1991, 352, 589–594. [Google Scholar] [CrossRef]
- Orosei, R.; Lauro, S.E.; Pettinelli, E.; Cicchetti, A.; Coradini, M.; Cosciotti, B.; Di Paolo, F.; Flamini, E.; Mattei, E.; Pajola, M.; et al. Radar evidence of subglacial liquid water on Mars. Science 2018, 361, 490–493. [Google Scholar] [CrossRef]
- Carr, M.H.; Belton, M.J.; Chapman, C.R.; Davies, M.E.; Geissler, P.; Greenberg, R.; McEwen, A.S.; Tufts, B.R.; Greeley, R.; Sullivan, R.; et al. Evidence for a subsurface ocean on Europa. Nature 1998, 391, 363–365. [Google Scholar] [CrossRef] [PubMed]
- Parkinson, C.D.; Liang, M.C.; Yung, Y.L.; Kirschivnk, J.L. Habitability of enceladus: Planetary conditions for life. Orig. Life Evol. Biosph. 2008, 38, 355–369. [Google Scholar] [CrossRef]
- Petigura, E.A.; Howard, A.W.; Marcy, G.W. Prevalence of Earth-size planets orbiting Sun-like stars. Proc. Natl. Acad. Sci. USA 2013, 110, 19273–19278. [Google Scholar] [CrossRef]
- Fressin, F.; Torres, G.; Charbonneau, D.; Bryson, S.T.; Christiansen, J.; Dressing, C.D.; Jenkins, J.M.; Walkowicz, L.M.; Batalha, N.M. The False Positive Rate of Kepler and the Occurrence of Planets. Astrophys. J. 2013, 766, 81. [Google Scholar] [CrossRef]
- Dressing, C.D.; Charbonneau, D. The Occurrence of Potentially Habitable Planets Orbiting M Dwarfs Estimated from the Full Kepler Dataset and an Empirical Measurement of the Detection Sensitivity. Astrophys. J. 2015, 807, 45. [Google Scholar] [CrossRef]
- Tsiaras, A.; Waldmann, I.P.; Tinetti, G.; Tennyson, J.; Yurchenko, S.N. Water vapour in the atmosphere of the habitable-zone eight-Earth-mass planet K2-18 b. Nat. Astron. 2019. [Google Scholar] [CrossRef]
- Studier, M.H.; Hayatsu, R.; Anders, E. Organic Compounds in Carbonaceous Chondrites. Science 1965, 149, 1455–1459. [Google Scholar] [CrossRef] [PubMed]
- Pizzarello, S.; Shock, E. Carbonaceous Chondrite Meteorites: The Chronicle of a Potential Evolutionary Path between Stars and Life. Orig. Life Evol. Biosph. 2017, 47, 249–260. [Google Scholar] [CrossRef]
- Glavin, D.P.; Elsila, J.E.; Burton, A.S.; Callahan, M.P.; Dworkin, J.P.; Hilts, R.W.; Herd, C.D.K. Unusual nonterrestrial l-proteinogenic amino acid excesses in the Tagish Lake meteorite. Meteorit. Planet. Sci. 2012, 47, 1347–1364. [Google Scholar] [CrossRef]
- Burton, A.S.; Stern, J.C.; Elsila, J.E.; Glavin, D.P.; Dworkin, J.P. Understanding prebiotic chemistry through the analysis of extraterrestrial amino acids and nucleobases in meteorites. Chem. Soc. Rev. 2012, 41, 5459–5472. [Google Scholar] [CrossRef]
- Meierhenrich, U.J.; Munoz Caro, G.M.; Bredehoft, J.H.; Jessberger, E.K.; Thiemann, W.H. Identification of diamino acids in the Murchison meteorite. Proc. Natl. Acad. Sci. USA 2004, 101, 9182–9186. [Google Scholar] [CrossRef]
- Koga, T.; Naraoka, H. A new family of extraterrestrial amino acids in the Murchison meteorite. Sci. Rep. 2017, 7, 636. [Google Scholar] [CrossRef] [PubMed]
- Cronin, J.R.; Cooper, G.W.; Pizzarello, S. Characteristics and formation of amino acids and hydroxy acids of the Murchison meteorite. Adv. Space Res. 1995, 15, 91–97. [Google Scholar] [CrossRef]
- Martinez-Rodriguez, S.; Martinez-Gomez, A.I.; Rodriguez-Vico, F.; Clemente-Jimenez, J.M.; Las Heras-Vazquez, F.J. Natural occurrence and industrial applications of D-amino acids: An overview. Chem. Biodivers. 2010, 7, 1531–1548. [Google Scholar] [CrossRef] [PubMed]
- Glavin, D.P.; Callahan, M.; Dworkin, J.E.; Elsila, J. The Effects of Parent Body Processes on Amino Acids in Carbonaceous Chondrites. Meteorit. Planet. Sci. 2010, 45, 1948–1972. [Google Scholar] [CrossRef]
- Elsila, J.E.; Aponte, J.C.; Blackmond, D.G.; Burton, A.S.; Dworkin, J.P.; Glavin, D.P. Meteoritic Amino Acids: Diversity in Compositions Reflects Parent Body Histories. ACS Cent. Sci. 2016, 2, 370–379. [Google Scholar] [CrossRef]
- Cordova, A.; Engqvist, M.; Ibrahem, I.; Casas, J.; Sunden, H. Plausible origins of homochirality in the amino acid catalyzed neogenesis of carbohydrates. Chem. Commun. 2005, 36, 2047–2049. [Google Scholar] [CrossRef] [PubMed]
- Shimoyama, A.; Ogasawara, R. Dipeptides and diketopiperazines in the Yamato-791198 and Murchison carbonaceous chondrites. Orig. Life Evol. Biosph. 2002, 32, 165–179. [Google Scholar] [CrossRef]
- McGeoch, J.E.M.; McGeoch, M.W. Polymer amide in the Allende and Murchison meteorites. Meteorit. Planet. Sci. 2015, 50, 1971–1983. [Google Scholar] [CrossRef]
- McGeoch, J.E.M.; McGeoch, M.W. A 4641Da polymer of amino acids in Acfer 086 and Allende meteorites. arXiv 2017, arXiv:1707.09080 (accessed on 1 July 2017). [Google Scholar]
- McGeoch, M.W.; Dikler, S.; McGeoch, J.E.M. Hemolithin: A Meteoritic Protein containing Iron and Lithium. arXiv 2020, arXiv:2002.11688 (accessed on: 1 February 2020). [Google Scholar]
- Engel, M.H.; Macko, S.A.; Silfer, J.A. Carbon isotope composition of individual amino acids in the Murchison meteorite. Nature 1990, 348, 47–49. [Google Scholar] [CrossRef]
- Elsila, J.E.; Callahan, M.P.; Glavin, D.P.; Dworkin, J.P.; Bruckner, H. Distribution and stable isotopic composition of amino acids from fungal peptaibiotics: Assessing the potential for meteoritic contamination. Astrobiology 2011, 11, 123–133. [Google Scholar] [CrossRef]
- Netea, M.G.; Dominguez-Andres, J.; Eleveld, M.; Op den Camp, H.J.M.; van der Meer, J.W.M.; Gow, N.A.R.; de Jonge, M.I. Immune recognition of putative alien microbial structures: Host-pathogen interactions in the age of space travel. PLoS Pathog. 2020, 16, e1008153. [Google Scholar] [CrossRef]
- Townsend, A.; Bodmer, H. Antigen recognition by class I-restricted T lymphocytes. Annu. Rev. Immunol. 1989, 7, 601–624. [Google Scholar] [CrossRef]
- Townsend, A.R.; Gotch, F.M.; Davey, J. Cytotoxic T cells recognize fragments of the influenza nucleoprotein. Cell 1985, 42, 457–467. [Google Scholar] [CrossRef]
- Townsend, A.R.; Bastin, J.; Gould, K.; Brownlee, G.G. Cytotoxic T lymphocytes recognize influenza haemagglutinin that lacks a signal sequence. Nature 1986, 324, 575–577. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, S.; Kalia, V.; Haining, W.N.; Konieczny, B.T.; Subramaniam, S.; Ahmed, R. Functional and genomic profiling of effector CD8 T cell subsets with distinct memory fates. J. Exp. Med. 2008, 205, 625–640. [Google Scholar] [CrossRef]
- Hogquist, K.A.; Jameson, S.C.; Heath, W.R.; Howard, J.L.; Bevan, M.J.; Carbone, F.R. T cell receptor antagonist peptides induce positive selection. Cell 1994, 76, 17–27. [Google Scholar] [CrossRef]
- Goldberg, A.L.; Cascio, P.; Saric, T.; Rock, K.L. The importance of the proteasome and subsequent proteolytic steps in the generation of antigenic peptides. Mol. Immunol. 2002, 39, 147–164. [Google Scholar] [CrossRef]
- Craiu, A.; Akopian, T.; Goldberg, A.; Rock, K.L. Two distinct proteolytic processes in the generation of a major histocompatibility complex class I-presented peptide. Proc. Natl. Acad. Sci. USA 1997, 94, 10850–10855. [Google Scholar] [CrossRef] [PubMed]
- Waldmann, T.A. Disorders of the expression of the multichain IL-2 receptor in HTLV-I-associated adult T-cell leukemia. Haematol. Blood Transfus. 1989, 32, 293–298. [Google Scholar] [CrossRef]
- Yamagishi, A.; Kawaguchi, Y.; Hashimoto, H.; Yano, H.; Imai, E.; Kodaira, S.; Uchihori, Y.; Nakagawa, K. Environmental Data and Survival Data of Deinococcus aetherius from the Exposure Facility of the Japan Experimental Module of the International Space Station Obtained by the Tanpopo Mission. Astrobiology 2018, 18, 1369–1374. [Google Scholar] [CrossRef]
- Horneck, G.; Klaus, D.M.; Mancinelli, R.L. Space microbiology. Microbiol. Mol. Biol. Rev. 2010, 74, 121–156. [Google Scholar] [CrossRef]
- Levin, G.V.; Straat, P.A. The Case for Extant Life on Mars and Its Possible Detection by the Viking Labeled Release Experiment. Astrobiology 2016, 16, 798–810. [Google Scholar] [CrossRef]
- Merino, N.; Aronson, H.S.; Bojanova, D.P.; Feyhl-Buska, J.; Wong, M.L.; Zhang, S.; Giovannelli, D. Living at the Extremes: Extremophiles and the Limits of Life in a Planetary Context. Front. Microbiol. 2019, 10, 780. [Google Scholar] [CrossRef]
- Nicholson, W.L.; Krivushin, K.; Gilichinsky, D.; Schuerger, A.C. Growth of Carnobacterium spp. from permafrost under low pressure, temperature, and anoxic atmosphere has implications for Earth microbes on Mars. Proc. Natl. Acad. Sci. USA 2013, 110, 666–671. [Google Scholar] [CrossRef] [PubMed]
- Schuerger, A.C.; Nicholson, W.L. Twenty Species of Hypobarophilic Bacteria Recovered from Diverse Soils Exhibit Growth under Simulated Martian Conditions at 0.7 kPa. Astrobiology 2016, 16, 964–976. [Google Scholar] [CrossRef] [PubMed]
- Fajardo-Cavazos, P.; Morrison, M.D.; Miller, K.M.; Schuerger, A.C.; Nicholson, W.L. Transcriptomic responses of Serratia liquefaciens cells grown under simulated Martian conditions of low temperature, low pressure, and CO2-enriched anoxic atmosphere. Sci. Rep. 2018, 8, 14938. [Google Scholar] [CrossRef] [PubMed]
- Taubner, R.S.; Pappenreiter, P.; Zwicker, J.; Smrzka, D.; Pruckner, C.; Kolar, P.; Bernacchi, S.; Seifert, A.H.; Krajete, A.; Bach, W.; et al. Biological methane production under putative Enceladus-like conditions. Nat. Commun. 2018, 9, 748. [Google Scholar] [CrossRef]
- Akiyama, T.; Horie, K.; Hinoi, E.; Hiraiwa, M.; Kato, A.; Maekawa, Y.; Takahashi, A.; Furukawa, S. How does spaceflight affect the acquired immune system? NPJ Microgravity 2020, 6, 14. [Google Scholar] [CrossRef]
- Sonnenfeld, G.; Shearer, W.T. Immune function during space flight. Nutrition 2002, 18, 899–903. [Google Scholar] [CrossRef]
- Kaur, I.; Simons, E.R.; Castro, V.A.; Ott, C.M.; Pierson, D.L. Changes in monocyte functions of astronauts. Brain Behav. Immun. 2005, 19, 547–554. [Google Scholar] [CrossRef]
- Kaur, I.; Simons, E.R.; Castro, V.A.; Mark Ott, C.; Pierson, D.L. Changes in neutrophil functions in astronauts. Brain Behav. Immun. 2004, 18, 443–450. [Google Scholar] [CrossRef]
- Lavy, A.; Rusu, R.; Mates, A. Mycobacterium xenopi, a potential human pathogen. Isr J. Med. Sci. 1992, 28, 772–775. [Google Scholar]
- Singh, N.K.; Bezdan, D.; Checinska Sielaff, A.; Wheeler, K.; Mason, C.E.; Venkateswaran, K. Multi-drug resistant Enterobacter bugandensis species isolated from the International Space Station and comparative genomic analyses with human pathogenic strains. BMC Microbiol. 2018, 18, 175. [Google Scholar] [CrossRef]
- Crucian, B.; Stowe, R.P.; Mehta, S.; Quiriarte, H.; Pierson, D.; Sams, C. Alterations in adaptive immunity persist during long-duration spaceflight. NPJ Microgravity 2015, 1, 15013. [Google Scholar] [CrossRef] [PubMed]
- Gould, C.L.; Lyte, M.; Williams, J.; Mandel, A.D.; Sonnenfeld, G. Inhibited interferon-gamma but normal interleukin-3 production from rats flown on the space shuttle. Aviatspace Environ. Med. 1987, 58, 983–986. [Google Scholar]
- Sonnenfeld, G.; Mandel, A.D.; Konstantinova, I.V.; Taylor, G.R.; Berry, W.D.; Wellhausen, S.R.; Lesnyak, A.T.; Fuchs, B.B. Effects of spaceflight on levels and activity of immune cells. Aviatspace Environ. Med. 1990, 61, 648–653. [Google Scholar]
- Sonnenfeld, G.; Davis, S.; Taylor, G.R.; Mandel, A.D.; Konstantinova, I.V.; Lesnyak, A.; Fuchs, B.B.; Peres, C.; Tkackzuk, J.; Schmitt, D.A. Effect of space flight on cytokine production and other immunologic parameters of rhesus monkeys. J. Interferon Cytokine Res. 1996, 16, 409–415. [Google Scholar] [CrossRef] [PubMed]
- Sallusto, F.; Lenig, D.; Forster, R.; Lipp, M.; Lanzavecchia, A. Two subsets of memory T lymphocytes with distinct homing potentials and effector functions. Nature 1999, 401, 708–712. [Google Scholar] [CrossRef]
- Kaech, S.M.; Ahmed, R. Memory CD8+ T cell differentiation: Initial antigen encounter triggers a developmental program in naive cells. Nat. Immunol. 2001, 2, 415–422. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schaefer, K.; Dambuza, I.M.; Dall’Angelo, S.; Yuecel, R.; Jaspars, M.; Trembleau, L.; Zanda, M.; Brown, G.D.; Netea, M.G.; Gow, N.A.R. A Weakened Immune Response to Synthetic Exo-Peptides Predicts a Potential Biosecurity Risk in the Retrieval of Exo-Microorganisms. Microorganisms 2020, 8, 1066. https://doi.org/10.3390/microorganisms8071066
Schaefer K, Dambuza IM, Dall’Angelo S, Yuecel R, Jaspars M, Trembleau L, Zanda M, Brown GD, Netea MG, Gow NAR. A Weakened Immune Response to Synthetic Exo-Peptides Predicts a Potential Biosecurity Risk in the Retrieval of Exo-Microorganisms. Microorganisms. 2020; 8(7):1066. https://doi.org/10.3390/microorganisms8071066
Chicago/Turabian StyleSchaefer, Katja, Ivy M. Dambuza, Sergio Dall’Angelo, Raif Yuecel, Marcel Jaspars, Laurent Trembleau, Matteo Zanda, Gordon D. Brown, Mihai G. Netea, and Neil A. R. Gow. 2020. "A Weakened Immune Response to Synthetic Exo-Peptides Predicts a Potential Biosecurity Risk in the Retrieval of Exo-Microorganisms" Microorganisms 8, no. 7: 1066. https://doi.org/10.3390/microorganisms8071066
APA StyleSchaefer, K., Dambuza, I. M., Dall’Angelo, S., Yuecel, R., Jaspars, M., Trembleau, L., Zanda, M., Brown, G. D., Netea, M. G., & Gow, N. A. R. (2020). A Weakened Immune Response to Synthetic Exo-Peptides Predicts a Potential Biosecurity Risk in the Retrieval of Exo-Microorganisms. Microorganisms, 8(7), 1066. https://doi.org/10.3390/microorganisms8071066





