SARS-CoV-2 Inhibition by Sulfonated Compounds
Abstract
1. Introduction
2. Materials and Methods
2.1. Compounds and Synthesis of Materials
2.2. Cells and Virus
2.3. VSV-CoV-2 Production
2.4. VSV-CoV-2 Inhibition Assays
2.5. Plaque Assay on VERO-E6 Cells
2.6. Virucidal Assay
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pruijssers, A.J.; George, A.S.; Schafer, A.; Leist, S.R.; Gralinksi, L.E.; Dinnon, K.H., 3rd; Yount, B.L.; Agostini, M.L.; Stevens, L.J.; Chappell, J.D.; et al. Remdesivir Inhibits SARS-CoV-2 in Human Lung Cells and Chimeric SARS-CoV Expressing the SARS-CoV-2 RNA Polymerase in Mice. Cell Rep. 2020, 32, 107940. [Google Scholar] [CrossRef] [PubMed]
- Beigel, J.H.; Tomashek, K.M.; Dodd, L.E. Remdesivir for the Treatment of Covid-19-Preliminary Report. Reply. N. Engl. J. Med. 2020, 383. [Google Scholar] [CrossRef]
- Pan, H.; Peto, R.; Karim, Q.A.; Alejandria, M.; Henao-Restrepo, A.M.; García, C.H.; Kieny, M.-P.; Malekzadeh, R.; Murthy, S.; Preziosi, M.-P.; et al. Repurposed antiviral drugs for COVID-19 –interim WHO SOLIDARITY trial results. medRxiv 2020. [Google Scholar] [CrossRef]
- Hoffmann, M.; Mosbauer, K.; Hofmann-Winkler, H.; Kaul, A.; Kleine-Weber, H.; Kruger, N.; Gassen, N.C.; Muller, M.A.; Drosten, C.; Pohlmann, S. Chloroquine does not inhibit infection of human lung cells with SARS-CoV-2. Nature 2020. [Google Scholar] [CrossRef]
- Maisonnasse, P.; Guedj, J.; Contreras, V.; Behillil, S.; Solas, C.; Marlin, R.; Naninck, T.; Pizzorno, A.; Lemaitre, J.; Goncalves, A.; et al. Hydroxychloroquine use against SARS-CoV-2 infection in non-human primates. Nature 2020. [Google Scholar] [CrossRef] [PubMed]
- NIH. NIH Halts Clinical Trial of Hydroxychloroquine. Available online: https://www.nih.gov/news-events/news-releases/nih-halts-clinical-trial-hydroxychloroquine (accessed on 10 August 2020).
- WHO. “Solidarity” Clinical Trial for COVID-19 Treatments. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/global-research-on-novel-coronavirus-2019-ncov/solidarity-clinical-trial-for-covid-19-treatments (accessed on 20 October 2020).
- Group, R.C.; Horby, P.; Lim, W.S.; Emberson, J.R.; Mafham, M.; Bell, J.L.; Linsell, L.; Staplin, N.; Brightling, C.; Ustianowski, A.; et al. Dexamethasone in Hospitalized Patients with Covid-19-Preliminary Report. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- Xu, X.; Han, M.; Li, T.; Sun, W.; Wang, D.; Fu, B.; Zhou, Y.; Zheng, X.; Yang, Y.; Li, X.; et al. Effective treatment of severe COVID-19 patients with tocilizumab. Proc. Natl. Acad. Sci. USA 2020, 117, 10970–10975. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S.; Garg, I.; Bansal, A.; Kumar, B. COVID-19 infection and Thrombosis. Clin. Chim. Acta 2020. [Google Scholar] [CrossRef] [PubMed]
- Cagno, V.; Tseligka, E.D.; Jones, S.T.; Tapparel, C. Heparan Sulfate Proteoglycans and Viral Attachment: True Receptors or Adaptation Bias? Viruses 2019, 11, 596. [Google Scholar] [CrossRef]
- Stencel-Baerenwald, J.E.; Reiss, K.; Reiter, D.M.; Stehle, T.; Dermody, T.S. The sweet spot: Defining virus-sialic acid interactions. Nat. Rev. Microbiol. 2014, 12, 739–749. [Google Scholar] [CrossRef]
- Li, W.; Hulswit, R.J.G.; Widjaja, I.; Raj, V.S.; McBride, R.; Peng, W.; Widagdo, W.; Tortorici, M.A.; van Dieren, B.; Lang, Y.; et al. Identification of sialic acid-binding function for the Middle East respiratory syndrome coronavirus spike glycoprotein. Proc. Natl. Acad. Sci. USA 2017, 114, E8508–E8517. [Google Scholar] [CrossRef] [PubMed]
- Milewska, A.; Zarebski, M.; Nowak, P.; Stozek, K.; Potempa, J.; Pyrc, K. Human coronavirus NL63 utilizes heparan sulfate proteoglycans for attachment to target cells. J. Virol. 2014, 88, 13221–13230. [Google Scholar] [CrossRef]
- Lang, J.; Yang, N.; Deng, J.; Liu, K.; Yang, P.; Zhang, G.; Jiang, C. Inhibition of SARS pseudovirus cell entry by lactoferrin binding to heparan sulfate proteoglycans. PLoS ONE 2011, 6, e23710. [Google Scholar] [CrossRef]
- Mathewson, A.C.; Bishop, A.; Yao, Y.; Kemp, F.; Ren, J.; Chen, H.; Xu, X.; Berkhout, B.; van der Hoek, L.; Jones, I.M. Interaction of severe acute respiratory syndrome-coronavirus and NL63 coronavirus spike proteins with angiotensin converting enzyme-2. J. Gen. Virol. 2008, 89, 2741–2745. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Milanetti, E.; Miotto, M.; Rienzo, L.D.; Monti, M.; Gosti, G.; Ruocco, G. In-Silico evidence for two receptors based strategy of SARS-CoV-2. bioRxiv 2020. [Google Scholar] [CrossRef]
- Robson, B. Bioinformatics studies on a function of the SARS-CoV-2 spike glycoprotein as the binding of host sialic acid glycans. Comput. Biol. Med. 2020, 122, 103849. [Google Scholar] [CrossRef]
- Clausen, T.M.; Sandoval, D.R.; Spliid, C.B.; Pihl, J.; Perrett, H.R.; Painter, C.D.; Narayanan, A.; Majowicz, S.A.; Kwong, E.M.; McVicar, R.N.; et al. SARS-CoV-2 Infection Depends on Cellular Heparan Sulfate and ACE2. Cell 2020. [Google Scholar] [CrossRef]
- Kim, S.Y.; Jin, W.; Sood, A.; Montgomery, D.W.; Grant, O.C.; Fuster, M.M.; Fu, L.; Dordick, J.S.; Woods, R.J.; Zhang, F.; et al. Characterization of heparin and severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) spike glycoprotein binding interactions. Antivir. Res. 2020. [Google Scholar] [CrossRef]
- Mycroft-West, C.J.; Su, D.; Pagani, I.; Rudd, T.R.; Elli, S.; Guimond, S.E.; Miller, G.; Meneghetti, M.C.Z.; Nader, H.B.; Li, Y.; et al. Heparin inhibits cellular invasion by SARS-CoV-2: Structural dependence of the interaction of the surface protein (spike) S1 receptor binding domain with heparin. bioRxiv 2020. [Google Scholar] [CrossRef]
- Liu, L.; Chopra, P.; Li, X.; Wolfert, M.A.; Tompkins, S.M.; Boons, G.-J. SARS-CoV-2 spike protein binds heparan sulfate in a length- and sequence-dependent manner. bioRxiv 2020. [Google Scholar] [CrossRef]
- Tiwari, V.; Beer, J.C.; Sankaranarayanan, N.V.; Swanson-Mungerson, M.; Desai, U.R. Discovering small-molecule therapeutics against SARS-CoV-2. Drug. Discov. Today 2020, 25, 1535–1544. [Google Scholar] [CrossRef] [PubMed]
- Guimond, S.E.; Mycroft-West, C.J.; Gandhi, N.S.; Tree, J.A.; Buttigieg, K.R.; Coombes, N.; Nystrom, K.; Said, J.; Setoh, Y.X.; Amarilla, A.; et al. Pixatimod (PG545), a clinical-stage heparan sulfate mimetic, is a potent inhibitor of the SARS-CoV-2 virus. bioRxiv 2020. [Google Scholar] [CrossRef]
- Zhang, Q.; Chen, C.Z.; Swaroop, M.; Xu, M.; Wang, L.; Lee, J.; Wang, A.Q.; Pradhan, M.; Hagen, N.; Chen, L. Heparan sulfate assists SARS-CoV-2 in cell entry and can be targeted by approved drugs in vitro. Cell Discov. 2020, 6, 1–14. [Google Scholar] [CrossRef]
- Tree, J.A.; Turnbull, J.E.; Buttigieg, K.R.; Elmore, M.J.; Coombes, N.; Hogwood, J.; Mycroft-West, C.J.; Lima, M.A.; Skidmore, M.A.; Karlsson, R.; et al. Unfractionated heparin inhibits live wild-type SARS-CoV-2 cell infectivity at therapeutically relevant concentrations. Br. J. Pharm. 2020. [Google Scholar] [CrossRef]
- Cagno, V.; Andreozzi, P.; D’Alicarnasso, M.; Jacob Silva, P.; Mueller, M.; Galloux, M.; Le Goffic, R.; Jones, S.T.; Vallino, M.; Hodek, J.; et al. Broad-spectrum non-toxic antiviral nanoparticles with a virucidal inhibition mechanism. Nat. Mater. 2018, 17, 195–203. [Google Scholar] [CrossRef]
- Jones, S.T.; Cagno, V.; Janecek, M.; Ortiz, D.; Gasilova, N.; Piret, J.; Gasbarri, M.; Constant, D.A.; Han, Y.; Vukovic, L.; et al. Modified cyclodextrins as broad-spectrum antivirals. Sci. Adv. 2020, 6, eaax9318. [Google Scholar] [CrossRef]
- Berger Rentsch, M.; Zimmer, G. A vesicular stomatitis virus replicon-based bioassay for the rapid and sensitive determination of multi-species type I interferon. PLoS ONE 2011, 6, e25858. [Google Scholar] [CrossRef]
- Fukushi, S.; Mizutani, T.; Saijo, M.; Matsuyama, S.; Miyajima, N.; Taguchi, F.; Itamura, S.; Kurane, I.; Morikawa, S. Vesicular stomatitis virus pseudotyped with severe acute respiratory syndrome coronavirus spike protein. J. Gen. Virol. 2005, 86, 2269–2274. [Google Scholar] [CrossRef]
- Cagno, V.; Gasbarri, M.; Medaglia, C.; Gomes, D.; Clement, S.; Stellacci, F.; Tapparel, C. Sulfonated nanomaterials with broad-spectrum antiviral activity extending beyond heparan sulfate-dependent viruses. Antimicrob. Agents Chemother 2020. [Google Scholar] [CrossRef]
- Tandon, R.; Sharp, J.S.; Zhang, F.; Pomin, V.H.; Ashpole, N.M.; Mitra, D.; McCandless, M.G.; Jin, W.; Liu, H.; Sharma, P. Effective inhibition of SARS-CoV-2 entry by heparin and enoxaparin derivatives. J. Virol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Tseligka, E.D.; Sobo, K.; Stoppini, L.; Cagno, V.; Abdul, F.; Piuz, I.; Meylan, P.; Huang, S.; Constant, S.; Tapparel, C. A VP1 mutation acquired during an enterovirus 71 disseminated infection confers heparan sulfate binding ability and modulates ex vivo tropism. PLoS Pathog 2018, 14, e1007190. [Google Scholar] [CrossRef] [PubMed]
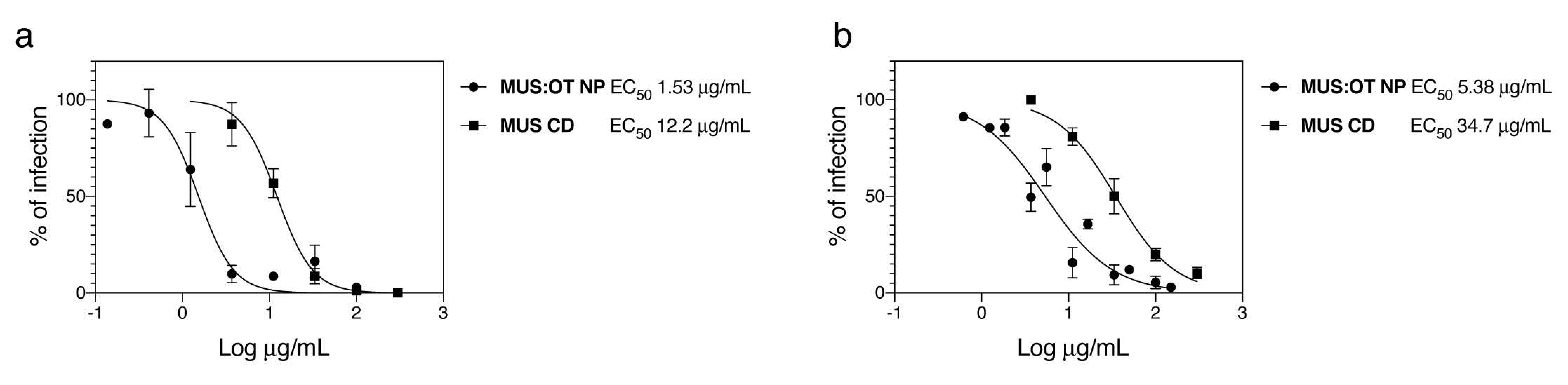
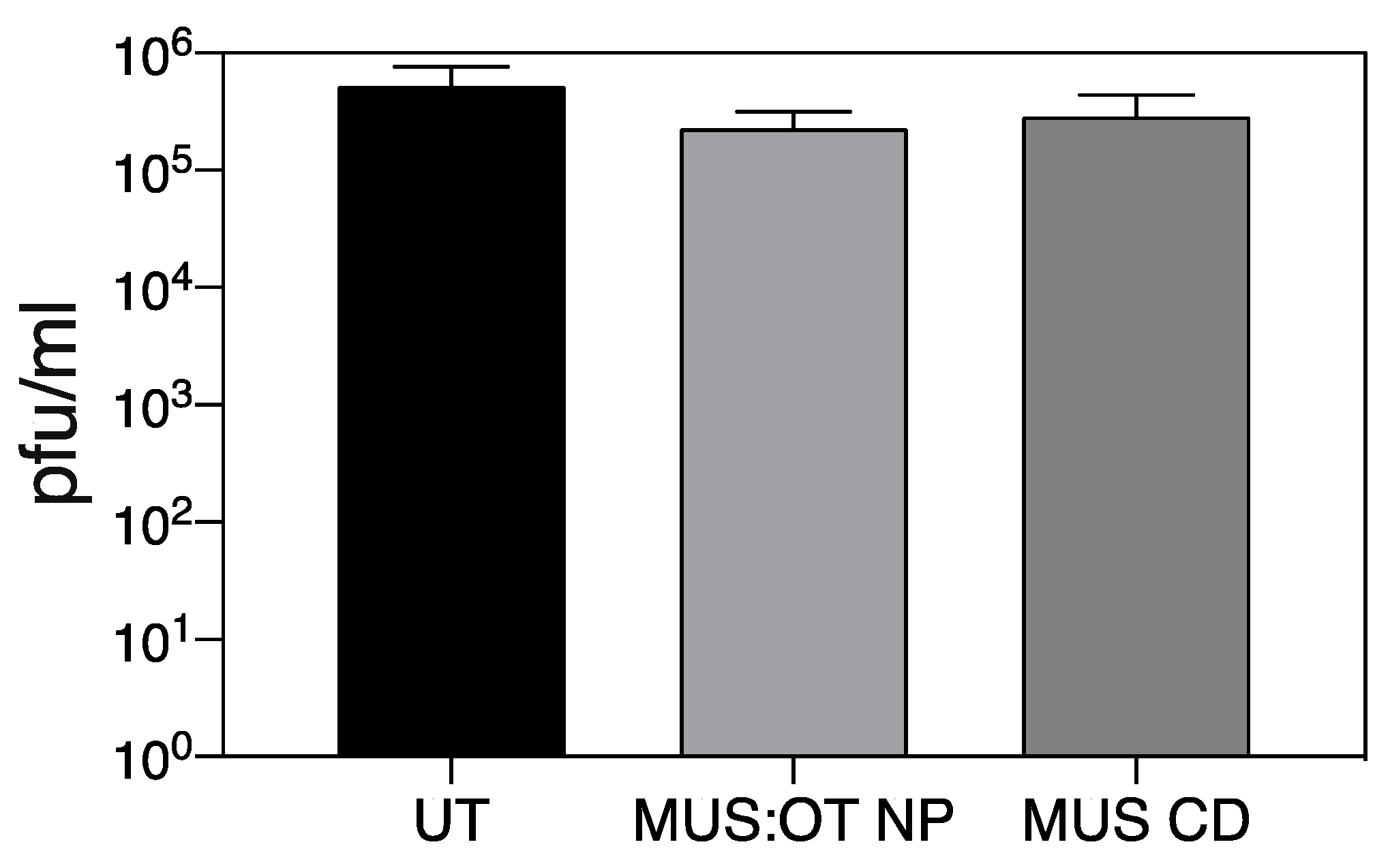
| Compound | EC50 (µg/mL) | EC50 µM | CC50 (µg/mL) | |
|---|---|---|---|---|
| VSV-CoV-2 | Carrageenan | >300 | >317 | >300 |
| K5N,OS(H) | >300 | >20 | >300 | |
| Sulfated ß-CD | >300 | >145 | >300 | |
| Resonium A | >300 | >300 | >300 | |
| MUS:OT NP | 1.5 | 0.005 | >300 | |
| MUS-CD | 12.2 | 4.0 | >300 | |
| Hydroxychloroquine | 3.9 | - | ||
| SARS-CoV-2 | Carrageenan | 267 | 282 | >300 |
| K5N,OS(H) | >300 | >20 | >300 | |
| Sulfated ß-CD | >1000 | >485 | >300 | |
| MUS:OT NP | 5.38 | 0.017 | >300 | |
| MUS-CD | 35.0 | 11.5 | >300 | |
| Heparin sodium salt (Tocris) | >1000 | >40 | ||
| Heparin Sodium Salt (Sigma) | >1000 | >55 | >300 | |
| Enoxaparin Sodium Salt | >1000 | >200 | ||
| Hydroxychloroquine | 2.92 | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gasbarri, M.; V’kovski, P.; Torriani, G.; Thiel, V.; Stellacci, F.; Tapparel, C.; Cagno, V. SARS-CoV-2 Inhibition by Sulfonated Compounds. Microorganisms 2020, 8, 1894. https://doi.org/10.3390/microorganisms8121894
Gasbarri M, V’kovski P, Torriani G, Thiel V, Stellacci F, Tapparel C, Cagno V. SARS-CoV-2 Inhibition by Sulfonated Compounds. Microorganisms. 2020; 8(12):1894. https://doi.org/10.3390/microorganisms8121894
Chicago/Turabian StyleGasbarri, Matteo, Philip V’kovski, Giulia Torriani, Volker Thiel, Francesco Stellacci, Caroline Tapparel, and Valeria Cagno. 2020. "SARS-CoV-2 Inhibition by Sulfonated Compounds" Microorganisms 8, no. 12: 1894. https://doi.org/10.3390/microorganisms8121894
APA StyleGasbarri, M., V’kovski, P., Torriani, G., Thiel, V., Stellacci, F., Tapparel, C., & Cagno, V. (2020). SARS-CoV-2 Inhibition by Sulfonated Compounds. Microorganisms, 8(12), 1894. https://doi.org/10.3390/microorganisms8121894






