Translation and Translational Control in Dinoflagellates
Abstract
1. Introduction
2. Post-Transcriptional Regulation of Gene Expression in Dinoflagellates
3. Translational Control of Gene Expression in Eukaryotes
3.1. Eukaryotic Translation
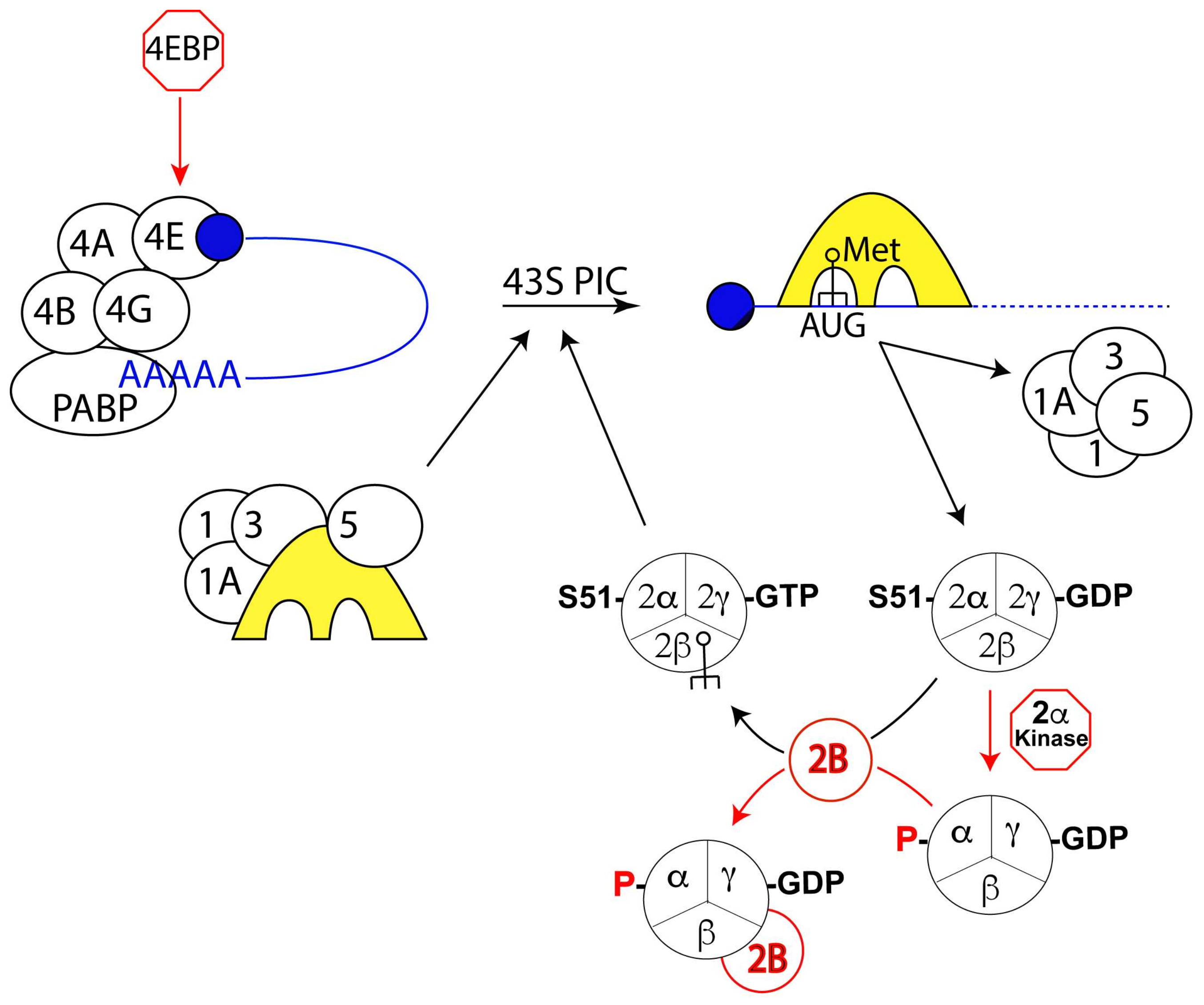
3.2. Initiation of Translation
4. The Translation Machinery in Dinoflagellates
4.1. Trans-Splicing of Mrna in Dinoflagellates
4.2. mRNA Codon Flanking Regions
4.3. Ribosomes in Dinoflagellates
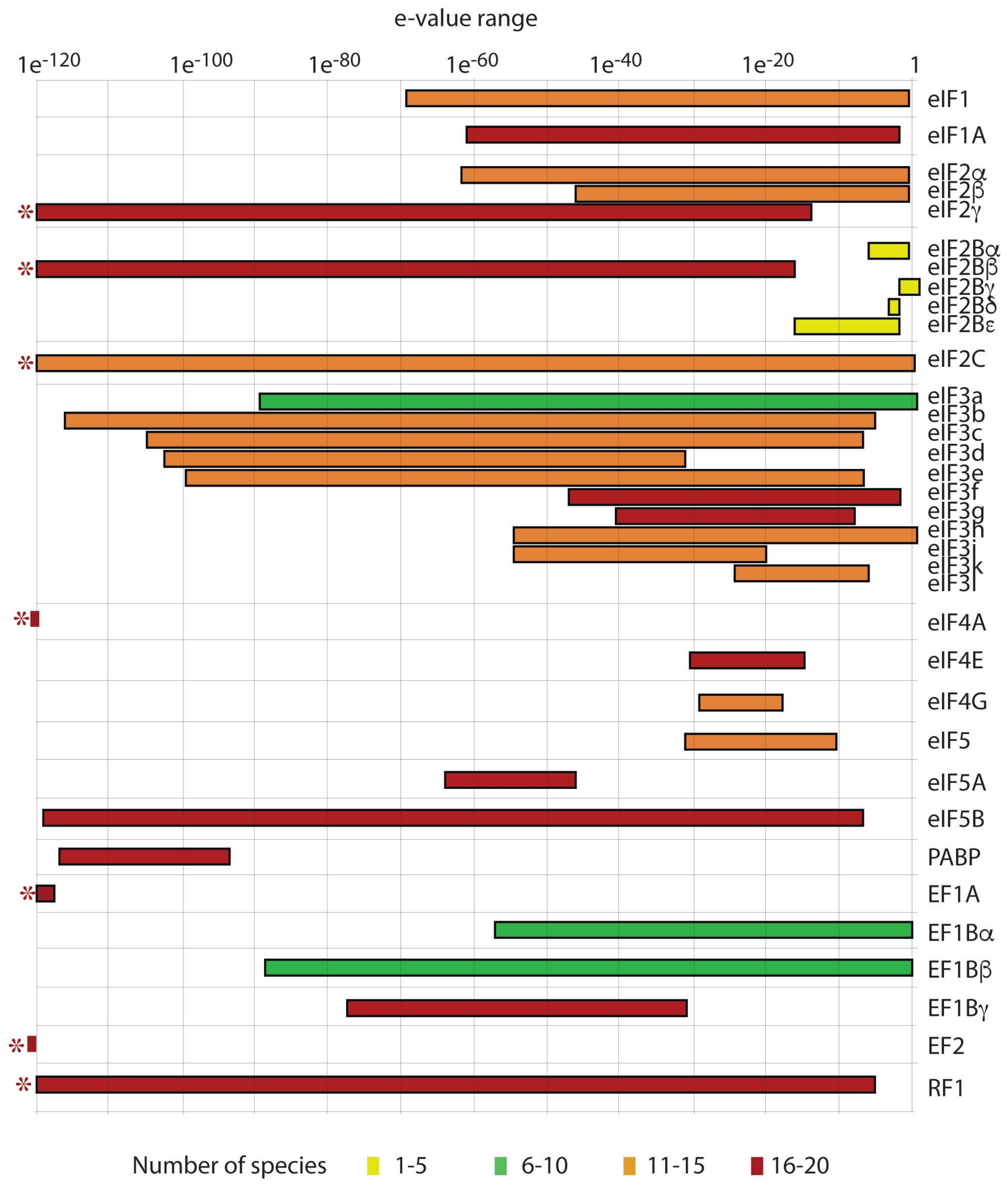
4.4. Translation Initiation Factors in Dinoflagellates
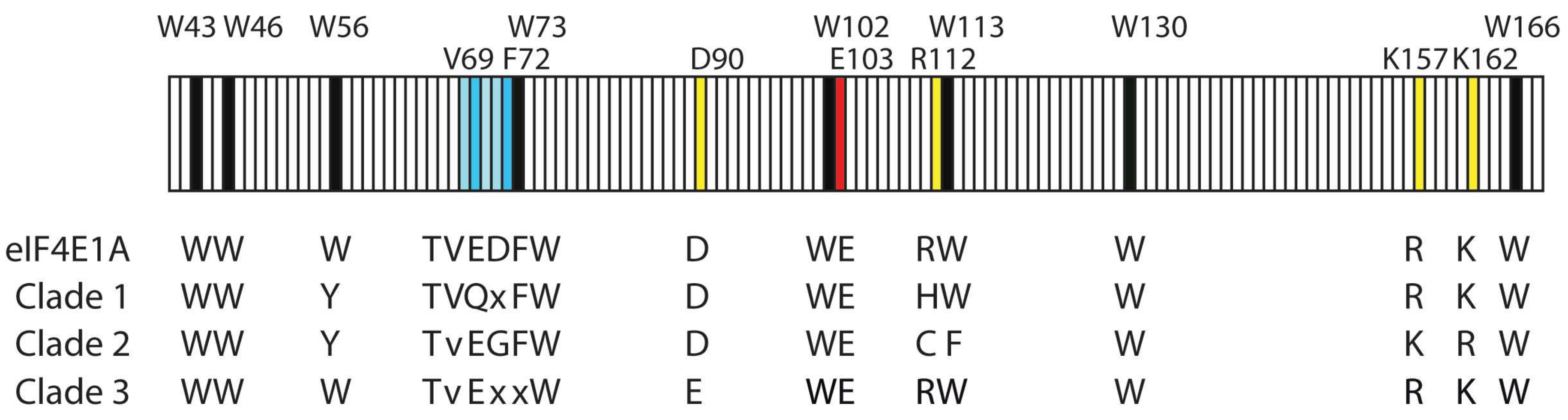
4.5. eIF4E
4.6. eIF4G
4.7. eIF2
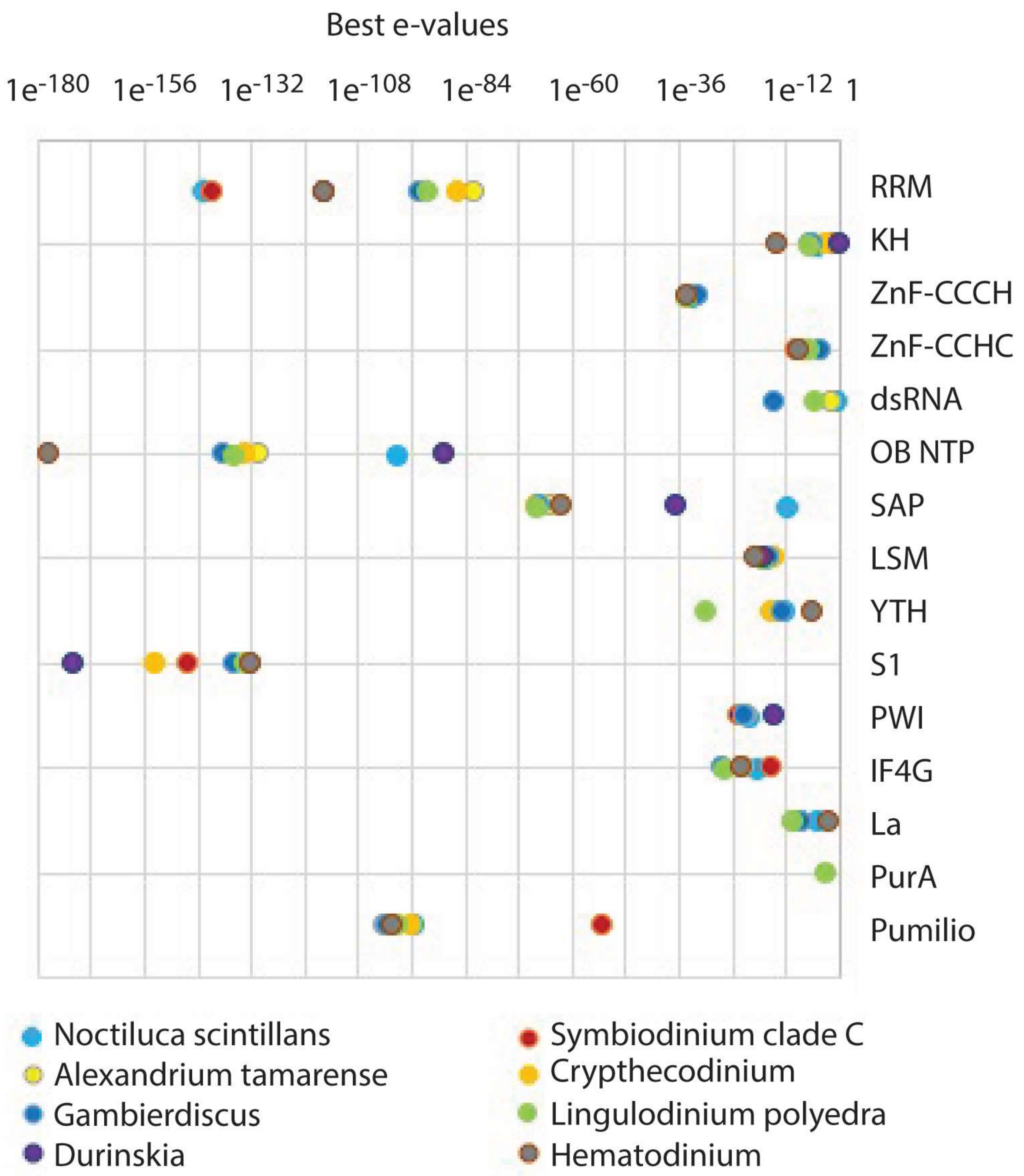
5. Translational Regulation by RNA-Binding Proteins
6. Translational Regulation by Small RNAs
7. Post-Translational Regulation
8. Conclusions
Acknowledgments
Conflicts of Interest
References
- Cavalier-Smith, T. The Biology of Free-Living Heterotrophic Flagellates; Oxford Universty Press: Oxford, UK, 1991. [Google Scholar]
- Taylor, F.J. Ultrastructure as a control for protistan molecular phylogeny. Am. Nat. 1999, 154, S125–S136. [Google Scholar] [CrossRef] [PubMed]
- Fast, N.M.; Xue, L.; Bingham, S.; Keeling, P.J. Re-examining alveolate evolution using multiple protein molecular phylogenies. J. Eukaryot. Microbiol. 2002, 49, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Saldarriaga, J.F.; McEwan, M.L.; Fast, N.M.; Taylor, F.J.; Keeling, P.J. Multiple protein phylogenies show that Oxyrrhis marina and Perkinsus marinus are early branches of the dinoflagellate lineage. Int. J. Syst. Evol. Microbiol. 2003, 53, 355–365. [Google Scholar] [CrossRef] [PubMed]
- Adl, S.M.; Simpson, A.G.; Farmer, M.A.; Andersen, R.A.; Anderson, O.R.; Barta, J.R.; Bowser, S.S.; Brugerolle, G.; Fensome, R.A.; Fredericq, S.; et al. The new higher level classification of eukaryotes with emphasis on the taxonomy of protists. J. Eukaryot. Microbiol. 2005, 52, 399–451. [Google Scholar] [CrossRef] [PubMed]
- Bachvaroff, T.R.; Gornik, S.G.; Concepcion, G.T.; Waller, R.F.; Mendez, G.S.; Lippmeier, J.C.; Delwiche, C.F. Dinoflagellate phylogeny revisited: Using ribosomal proteins to resolve deep branching dinoflagellate clades. Mol. Phylogenet. Evol. 2014, 70, 314–322. [Google Scholar] [CrossRef] [PubMed]
- Dagenais-Bellefeuille, S.; Morse, D. Putting the N in dinoflagellates. Front. Microbiol. 2013, 4, 369. [Google Scholar] [CrossRef] [PubMed]
- Taylor, F.J.R.; Hoppenrath, M.; Saldarriaga, J.F. Dinoflagellate diversity and distribution. Biodivers. Conserv. 2008, 17, 407–418. [Google Scholar] [CrossRef]
- Rae, P.M. Hydroxymethyluracil in eukaryote DNA: A natural feature of the pyrrophyta (dinoflagellates). Science 1976, 194, 1062–1064. [Google Scholar] [CrossRef] [PubMed]
- Steele, R.E.; Rae, P.M. Ordered distribution of modified bases in the DNA of a dinoflagellate. Nucleic Acids Res. 1980, 8, 4709–4725. [Google Scholar] [CrossRef] [PubMed]
- Moreno Diaz de la Espina, S.; Alverca, E.; Cuadrado, A.; Franca, S. Organization of the genome and gene expression in a nuclear environment lacking histones and nucleosomes: The amazing dinoflagellates. Eur. J. Cell Biol. 2005, 84, 137–149. [Google Scholar] [CrossRef] [PubMed]
- Spector, D.L. Dinoflagellate Nuclei. In Dinoflagellates; Spector, D.L., Ed.; Academic Press: New York, NY, USA, 1984; pp. 107–147. [Google Scholar]
- Bouligand, Y.; Norris, V. Chromosome separation and segregation in dinoflagellates and bacteria may depend on liquid crystalline states. Biochimie 2001, 83, 187–192. [Google Scholar] [CrossRef]
- Lin, S. Genomic understanding of dinoflagellates. Res. Microbiol. 2011, 162, 551–569. [Google Scholar] [CrossRef] [PubMed]
- Bachvaroff, T.R.; Place, A.R. From stop to start: Tandem gene arrangement, copy number and trans-splicing sites in the dinoflagellate Amphidinium carterae. PLoS ONE 2008, 3, e2929. [Google Scholar] [CrossRef] [PubMed]
- Bachvaroff, T.R.; Place, A.R.; Coats, D.W. Expressed sequence tags from Amoebophrya sp. infecting Karlodinium veneficum: Comparing host and parasite sequences. J. Eukaryot. Microbiol. 2009, 56, 531–541. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Cheng, S.; Song, B.; Zhong, X.; Lin, X.; Li, W.; Li, L.; Zhang, Y.; Zhang, H.; Ji, Z.; et al. The Symbiodinium kawagutii genome illuminates dinoflagellate gene expression and coral symbiosis. Science 2015, 350, 691–694. [Google Scholar] [CrossRef] [PubMed]
- Le, Q.H.; Markovic, P.; Jovine, R.; Hastings, J.; Morse, D. Structure and organization of the peridinin-chlorophyll a-protein gene in Gonyaulax polyedra. Mol. Gen. Genet. 1997, 255, 595–604. [Google Scholar] [PubMed]
- Lukes, J.; Leander, B.S.; Keeling, P.J. Cascades of convergent evolution: The corresponding evolutionary histories of euglenozoans and dinoflagellates. Proc. Natl. Acad. Sci. USA 2009, 106, 9963–9970. [Google Scholar] [CrossRef] [PubMed]
- Beauchemin, M.; Roy, S.; Daoust, P.; Dagenais-Bellefeuille, S.; Bertomeu, T.; Letourneau, L.; Lang, B.F.; Morse, D. Dinoflagellate tandem array gene transcripts are highly conserved and not polycistronic. Proc. Natl. Acad. Sci. USA 2012, 109, 15793–15798. [Google Scholar] [CrossRef] [PubMed]
- Guillebault, D.; Sasorith, S.; Derelle, E.; Wurtz, J.M.; Lozano, J.C.; Bingham, S.; Tora, L.; Moreau, H. A new class of transcription initiation factor, intermediate between TBPs and TLFs is present in the marine unicellular organism: The dinoflagellate Crypthecodinium cohnii. J. Biol. Chem. 2002, 277, 40881–40886. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, P. Biochemistry of the dinoflagellate nucleus. In The Biology of Dinoflagellates; Taylor, F.J.R., Ed.; Blackwell Scientific: Oxford, UK, 1987; pp. 143–173. [Google Scholar]
- Chan, Y.H.; Wong, J.T. Concentration-dependent organization of DNA by the dinoflagellate histone-like protein HCc3. Nucleic Acids Res. 2007, 35, 2573–2583. [Google Scholar] [CrossRef] [PubMed]
- Bayer, T.; Aranda, M.; Sunagawa, S.; Yum, L.K.; Desalvo, M.K.; Lindquist, E.; Coffroth, M.A.; Voolstra, C.R.; Medina, M. Symbiodinium transcriptomes: Genome insights into the dinoflagellate symbionts of reef-building corals. PLoS ONE 2012, 7, e35269. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Morse, D. A full suite of histone and histone modifying genes are transcribed in the dinoflagellate Lingulodinium. PLoS ONE 2012, 7, e34340. [Google Scholar] [CrossRef] [PubMed]
- Gornik, S.G.; Ford, K.L.; Mulhern, T.D.; Bacic, A.; McFadden, G.I.; Waller, R.F. Loss of nucleosomal DNA condensation coincides with appearance of a novel nuclear protein in dinoflagellates. Curr. Biol. 2012, 22, 2303–2312. [Google Scholar] [CrossRef] [PubMed]
- Janouskovec, J.; Gavelis, G.S.; Burki, F.; Dinh, D.; Bachvaroff, T.R.; Gornik, S.G.; Bright, K.J.; Imanian, B.; Strom, S.L.; Delwiche, C.F.; et al. Major transitions in dinoflagellate evolution unveiled by phylotranscriptomics. Proc. Natl. Acad. Sci. USA 2017, 114, E171–E180. [Google Scholar] [CrossRef] [PubMed]
- Cachon, J.; Cachon, M. Botanical Monographs. In The Biology of Dinoflagellates; Taylor, F.J.R., Ed.; Blackwell Scientific Publications: Hoboken, NJ, USA, 1987; Volume 21. [Google Scholar]
- Rizzo, P.J.; Jones, M.; Ray, S.M. Isolation and properties of isolated nuclei from the Florida red tide dinoflagellate Gymnodinium breve (Davis). J. Protozool. 1982, 29, 217–222. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.; Lin, S. Distinct gene number-genome size relationships for eukaryotes and non-eukaryotes: Gene content estimation for dinoflagellate genomes. PLoS ONE 2009, 4, e6978. [Google Scholar] [CrossRef] [PubMed]
- Shoguchi, E.; Shinzato, C.; Kawashima, T.; Gyoja, F.; Mungpakdee, S.; Koyanagi, R.; Takeuchi, T.; Hisata, K.; Tanaka, M.; Fujiwara, M.; et al. Draft assembly of the Symbiodinium minutum nuclear genome reveals dinoflagellate gene structure. Curr. Biol. 2013, 23, 1399–1408. [Google Scholar] [CrossRef] [PubMed]
- Wilson, T.; Hastings, J.W. Bioluminescence. Annu. Rev. Cell Dev. Biol. 1998, 14, 197–230. [Google Scholar] [CrossRef] [PubMed]
- Hastings, J.; Sweeney, B.M. The luminescent reaction in extracts of the marine dinoflagellate Gonyaulax polyedra. J. Cell. Physiol. 1957, 49, 209–218. [Google Scholar] [CrossRef]
- Hastings, J.W.; Astrachan, L.; Sweeney, B.M. A persistent daily rhythm in photosynthesis. J. Gen. Physiol. 1961, 45, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Sweeney, B.M.; Hastings, J.W. Effects of temperature upon diurnal rhythms. Cold Spring Harb. Symp. Quant. Biol. 1960, 25, 87–104. [Google Scholar] [CrossRef] [PubMed]
- Eppley, R.W.; Holm-Hansen, O.; Strickland, J.D.H. Some observations on the vertical migration of dinoflagellates. J. Phycol. 1968, 4, 333–340. [Google Scholar] [CrossRef] [PubMed]
- Morse, D.S.; Fritz, L.; Hastings, J.W. What is the clock? Translational regulation of circadian bioluminescence. Trends biochem. Sci. 1990, 15, 262–265. [Google Scholar] [CrossRef]
- Hastings, J.W. The Gonyaulax clock at 50: Translational control of circadian expression. Cold Spring Harb. Symp. Quant. Biol. 2007, 72, 141–144. [Google Scholar] [CrossRef] [PubMed]
- Sweeney, B.M.; Hastings, J.W. Characteristics of the diurnal rhythm of luminescence in Gonyaulax polyedra. J. Cell. Physiol. 1957, 49, 115–128. [Google Scholar] [CrossRef]
- Nakamura, H.; Kishi, Y.; Shimomura, O.; Morse, D.; Hastings, J.W. Structure of dinoflagellate luciferin and its enzymatic and non-enzymatic air oxidation products. J. Am. Chem. Soc. 1989, 111, 7607–7611. [Google Scholar] [CrossRef]
- Morse, D.; Pappenheimer, A.M., Jr.; Hastings, J.W. Role of a luciferin-binding protein in the circadian bioluminescent reaction of Gonyaulax polyedra. J. Biol. Chem. 1989, 264, 11822–11826. [Google Scholar] [PubMed]
- Morse, D.; Milos, P.M.; Roux, E.; Hastings, J.W. Circadian regulation of bioluminescence in Gonyaulax involves translational control. Proc. Natl. Acad. Sci. USA 1989, 86, 172–176. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.H.; Roeber, J.F.; Hastings, J.W. Circadian changes in enzyme concentration account for rhythm of enzyme activity in Gonyaulax. Science 1984, 223, 1428–1430. [Google Scholar] [CrossRef] [PubMed]
- Milos, P.; Morse, D.; Hastings, J.W. Circadian control over synthesis of many Gonyaulax proteins is at a translational level. Naturwiss 1990, 77, 87–89. [Google Scholar] [CrossRef] [PubMed]
- Markovic, P.; Roenneberg, T.; Morse, D. Phased protein synthesis at several circadian times does not change protein levels in Gonyaulax. J. Biol. Rhythm. 1996, 11, 57–67. [Google Scholar] [CrossRef] [PubMed]
- Schroder-Lorenz, A.; Rensing, L. Circadian changes in protein-synthesis rate and protein phosphorylation in cell-free extracts of Gonyaulax polyedra. Planta 1987, 170, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Akimoto, H.; Kinumi, T.; Ohmiya, Y. Circadian rhythm of a TCA cycle enzyme is apparently regulated at the translational level in the dinoflagellate Lingulodinium polyedrum. J. Biol. Rhythm. 2005, 20, 479–489. [Google Scholar] [CrossRef] [PubMed]
- Fagan, T.; Morse, D.; Hastings, J. Circadian synthesis of a nuclear encoded chloroplast Glyceraldehyde-3-phosphate dehydrogenase in the dinoflagellate Gonyaulax polyedra is translationally controlled. Biochemistry 1999, 38, 7689–7695. [Google Scholar] [CrossRef] [PubMed]
- Ten Lohuis, M.R.; Miller, D.J. Light-regulated transcription of genes encoding peridinin chlorophyll a proteins and the major intrinsic light-harvesting complex proteins in the dinoflagellate Amphidinium carterae Hulburt (Dinophycae). Changes In cytosine methylation accompany photoadaptation. Plant Physiol. 1998, 117, 189–196. [Google Scholar] [PubMed]
- Okamoto, O.K.; Robertson, D.L.; Fagan, T.F.; Hastings, J.W.; Colepicolo, P. Different regulatory mechanisms modulate the expression of a dinoflagellate iron-superoxide dismutase. J. Biol. Chem. 2001, 276, 19989–19993. [Google Scholar] [CrossRef] [PubMed]
- Black, A.R.; Azizkhan-Clifford, J. Regulation of E2F: A family of transcription factors involved in proliferation control. Gene 1999, 237, 281–302. [Google Scholar] [CrossRef]
- Lavia, P.; Jansen-Durr, P. E2F target genes and cell-cycle checkpoint control. Bioessays 1999, 21, 221–230. [Google Scholar] [CrossRef]
- Brunelle, S.A.; van Dolah, F.M. Post-transcriptional regulation of S-phase genes in the dinoflagellate, Karenia brevis. J. Eukaryot. Microbiol. 2011, 58, 373–382. [Google Scholar] [CrossRef] [PubMed]
- Morey, J.S.; Monroe, E.A.; Kinney, A.L.; Beal, M.; Johnson, J.G.; Hitchcock, G.L.; van Dolah, F.M. Transcriptomic response of the red tide dinoflagellate, Karenia brevis, to nitrogen and phosphorus depletion and addition. BMC Genom. 2011, 12, 346. [Google Scholar] [CrossRef] [PubMed]
- Van Dolah, F.M.; Lidie, K.B.; Morey, J.S.; Brunelle, S.A.; Ryan, J.C.; Monroe, E.A.; Haynes, B.L. Microarray analysis of diurnal and circadian regulated genes in the florida red-tide dinoflagellate Karenia brevis (Dinophyceae). J. Phycol. 2007, 43, 741–752. [Google Scholar] [CrossRef]
- Okamoto, O.K.; Hastings, J.W. Genome-wide analysis of redox-regulated genes in a dinoflagellate. Gene 2003, 321, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, O.K.; Hastings, J.W. Novel dinoflagellate circadian-clock genes identified through microarray analysis of a phase shifted clock. J. Phycol. 2003, 39, 1–9. [Google Scholar] [CrossRef]
- Yang, I.; Beszteri, S.; Tillmann, U.; John, U. Growth and nutrient-dependent gene expression in the toxigenic marine dinoflagellate Alexandrium minutum. Harmful Algae 2013, 12, 15–25. [Google Scholar] [CrossRef]
- Moustafa, A.; Evans, A.N.; Kulis, D.M.; Hackett, J.D.; Erdner, D.L.; Anderson, D.M.; Bhattacharya, D. Transcriptome profiling of a toxic dinoflagellate reveals a gene-rich protist and a potential impact on gene expression due to bacterial presence. PLoS ONE 2010, 5, e9688. [Google Scholar] [CrossRef] [PubMed]
- Erdner, D.L.; Anderson, D.M. Global transcriptional profiling of the toxic dinoflagellate Alexandrium fundyense using Massively Parallel Signature Sequencing. BMC Genom. 2006, 7, 88. [Google Scholar] [CrossRef] [PubMed]
- Lowe, C.D.; Mello, L.V.; Samatar, N.; Martin, L.E.; Montagnes, D.J.; Watts, P.C. The transcriptome of the novel dinoflagellate Oxyrrhis marina (Alveolata: Dinophyceae): Response to salinity examined by 454 sequencing. BMC Genom. 2011, 12, 519. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Beauchemin, M.; Dagenais-Bellefeuille, S.; Letourneau, L.; Cappadocia, M.; Morse, D. The Lingulodinium circadian system lacks rhythmic changes in transcript abundance. BMC Biol. 2014, 12, 107. [Google Scholar] [CrossRef] [PubMed]
- Tse, S.P.K.; Beauchemin, M.; Morse, D.; Lo, S.C.L. Refining transcriptome gene catalogs by MS-validation of expressed proteins. Proteomics 2018, 18. [Google Scholar] [CrossRef] [PubMed]
- Qin, X.; Ahn, S.; Speed, T.P.; Rubin, G.M. Global analyses of mRNA translational control during early Drosophila embryogenesis. Genome Biol. 2007, 8, R63. [Google Scholar] [CrossRef] [PubMed]
- Schwanhausser, B.; Busse, D.; Li, N.; Dittmar, G.; Schuchhardt, J.; Wolf, J.; Chen, W.; Selbach, M. Global quantification of mammalian gene expression control. Nature 2011, 473, 337–342. [Google Scholar] [CrossRef] [PubMed]
- Beilharz, T.H.; Preiss, T. Translational profiling: The genome-wide measure of the nascent proteome. Brief. Funct. Genom. Proteom. 2004, 3, 103–111. [Google Scholar] [CrossRef][Green Version]
- Pradet-Balade, B.; Boulme, F.; Beug, H.; Mullner, E.W.; Garcia-Sanz, J.A. Translation control: Bridging the gap between genomics and proteomics? Trends Biochem. Sci. 2001, 26, 225–229. [Google Scholar] [CrossRef]
- Maier, T.; Guell, M.; Serrano, L. Correlation of mRNA and protein in complex biological samples. FEBS Lett. 2009, 583, 3966–3973. [Google Scholar] [CrossRef] [PubMed]
- Thomas, A.; Lee, P.J.; Dalton, J.E.; Nomie, K.J.; Stoica, L.; Costa-Mattioli, M.; Chang, P.; Nuzhdin, S.; Arbeitman, M.N.; Dierick, H.A. A versatile method for cell-specific profiling of translated mRNAs in Drosophila. PLoS ONE 2012, 7, e40276. [Google Scholar] [CrossRef] [PubMed]
- Wisecaver, J.H.; Hackett, J.D. Dinoflagellate genome evolution. Annu. Rev. Microbiol. 2011, 65, 369–387. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Morse, D. Transcription and Maturation of mRNA in Dinoflagellates. Microorganisms 2013, 1, 71–99. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, J.E. Posttranscriptional control of gene expression in yeast. Microbiol. Mol. Biol. Rev. 1998, 62, 1492–1553. [Google Scholar] [PubMed]
- Jackson, R.J.; Hellen, C.U.; Pestova, T.V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell Biol. 2010, 11, 113–127. [Google Scholar] [CrossRef] [PubMed]
- Laursen, B.S.; Sorensen, H.P.; Mortensen, K.K.; Sperling-Petersen, H.U. Initiation of protein synthesis in bacteria. Microbiol. Mol. Biol. Rev. 2005, 69, 101–123. [Google Scholar] [CrossRef] [PubMed]
- Chang, B.; Halgamuge, S.; Tang, S.L. Analysis of SD sequences in completed microbial genomes: Non-SD-led genes are as common as SD-led genes. Gene 2006, 373, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Hering, O.; Brenneis, M.; Beer, J.; Suess, B.; Soppa, J. A novel mechanism for translation initiation operates in haloarchaea. Mol. Microbiol. 2009, 71, 1451–1463. [Google Scholar] [CrossRef] [PubMed]
- Tolstrup, N.; Sensen, C.W.; Garrett, R.A.; Clausen, I.G. Two different and highly organized mechanisms of translation initiation in the archaeon Sulfolobus solfataricus. Extremophiles 2000, 4, 175–179. [Google Scholar] [CrossRef] [PubMed]
- Slupska, M.M.; King, A.G.; Fitz-Gibbon, S.; Besemer, J.; Borodovsky, M.; Miller, J.H. Leaderless transcripts of the crenarchaeal hyperthermophile Pyrobaculum aerophilum. J. Mol. Biol. 2001, 309, 347–360. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Hu, G.Q.; She, Z.S.; Zhu, H. Leaderless genes in bacteria: Clue to the evolution of translation initiation mechanisms in prokaryotes. BMC Genom. 2011, 12, 361. [Google Scholar] [CrossRef] [PubMed]
- Benelli, D.; Londei, P. Begin at the beginning: Evolution of translational initiation. Res. Microbiol. 2009, 160, 493–501. [Google Scholar] [CrossRef] [PubMed]
- Benelli, D.; Londei, P. Translation initiation in Archaea: Conserved and domain-specific features. Biochem. Soc. Trans. 2011, 39, 89–93. [Google Scholar] [CrossRef] [PubMed]
- Londei, P. Evolution of translational initiation: New insights from the archaea. FEMS Microbiol. Rev. 2005, 29, 185–200. [Google Scholar] [CrossRef] [PubMed]
- Doi, N.; Zenno, S.; Ueda, R.; Ohki-Hamazaki, H.; Ui-Tei, K.; Saigo, K. Short-interfering-RNA-mediated gene silencing in mammalian cells requires Dicer and eIF2C translation initiation factors. Curr. Biol. 2003, 13, 41–46. [Google Scholar] [CrossRef]
- Hinnebusch, A.G. Molecular mechanism of scanning and start codon selection in eukaryotes. Microbiol. Mol. Biol. Rev. 2011, 75, 434–467. [Google Scholar] [CrossRef] [PubMed]
- Aitken, C.E.; Lorsch, J.R. A mechanistic overview of translation initiation in eukaryotes. Nat. Struct. Mol. Biol. 2012, 19, 568–576. [Google Scholar] [CrossRef] [PubMed]
- Hinnebusch, A.G.; Lorsch, J.R. The mechanism of eukaryotic translation initiation: New insights and challenges. Cold Spring Harb. Perspect. Biol. 2012, 4. [Google Scholar] [CrossRef] [PubMed]
- Valasek, L.S. ‘Ribozoomin’—Translation initiation from the perspective of the ribosome-bound eukaryotic initiation factors (eIFs). Curr. Protein Pept. Sci. 2012, 13, 305–330. [Google Scholar] [CrossRef] [PubMed]
- Yanagiya, A.; Svitkin, Y.V.; Shibata, S.; Mikami, S.; Imataka, H.; Sonenberg, N. Requirement of RNA binding of mammalian eukaryotic translation initiation factor 4GI (eIF4GI) for efficient interaction of eIF4E with the mRNA cap. Mol. Cell. Biol. 2009, 29, 1661–1669. [Google Scholar] [CrossRef] [PubMed]
- Kahvejian, A.; Svitkin, Y.V.; Sukarieh, R.; M’Boutchou, M.N.; Sonenberg, N. Mammalian poly(A)-binding protein is a eukaryotic translation initiation factor, which acts via multiple mechanisms. Genes Dev. 2005, 19, 104–113. [Google Scholar] [CrossRef] [PubMed]
- Lomakin, I.B.; Steitz, T.A. The initiation of mammalian protein synthesis and mRNA scanning mechanism. Nature 2013, 500, 307–311. [Google Scholar] [CrossRef] [PubMed]
- Lomakin, I.B.; Shirokikh, N.E.; Yusupov, M.M.; Hellen, C.U.; Pestova, T.V. The fidelity of translation initiation: Reciprocal activities of eIF1, IF3 and YciH. EMBO J. 2006, 25, 196–210. [Google Scholar] [CrossRef] [PubMed]
- Kolitz, S.E.; Takacs, J.E.; Lorsch, J.R. Kinetic and thermodynamic analysis of the role of start codon/anticodon base pairing during eukaryotic translation initiation. RNA 2009, 15, 138–152. [Google Scholar] [CrossRef] [PubMed]
- Pestova, T.V.; Lorsch, J.R.; Hellen, C.U.T. The mechanism of translation initiation in eukaryotes. In Translational Control in Biology and Medicine; Cold Spring Harbor Laboratory Press: Suffolk County, NY, USA, 2007; pp. 87–128. [Google Scholar]
- Ghosh, A.; Lima, C.D. Enzymology of RNA cap synthesis. Wiley Interdiscip. Rev. RNA 2010, 1, 152–172. [Google Scholar] [CrossRef] [PubMed]
- Lidie, K.B.; van Dolah, F.M. Spliced leader RNA-mediated trans-splicing in a dinoflagellate, Karenia brevis. J. Eukaryot. Microbiol. 2007, 54, 427–435. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Hou, Y.; Miranda, L.; Campbell, D.A.; Sturm, N.R.; Gaasterland, T.; Lin, S. Spliced leader RNA trans-splicing in dinoflagellates. Proc. Natl. Acad. Sci. USA 2007, 104, 4618–4623. [Google Scholar] [CrossRef] [PubMed]
- Hastings, K.E. SL trans-splicing: Easy come or easy go? Trends Genet. 2005, 21, 240–247. [Google Scholar] [CrossRef] [PubMed]
- Lasda, E.L.; Blumenthal, T. Trans-splicing. WIREs RNA 2011, 2, 417–434. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Campbell, D.A.; Sturm, N.R.; Dungan, C.F.; Lin, S. Spliced leader RNAs, mitochondrial gene frameshifts and multi-protein phylogeny expand support for the genus Perkinsus as a unique group of alveolates. PLoS ONE 2011, 6, e19933. [Google Scholar] [CrossRef] [PubMed]
- Hearne, J.L.; Pitula, J.S. Identification of two spliced leader RNA transcripts from Perkinsus marinus. J. Eukaryot. Microbiol. 2011, 58, 266–268. [Google Scholar] [CrossRef] [PubMed]
- Perry, K.L.; Watkins, K.P.; Agabian, N. Trypanosome mRNAs have unusual "cap 4" structures acquired by addition of a spliced leader. Proc. Natl. Acad. Sci. USA 1987, 84, 8190–8194. [Google Scholar] [CrossRef] [PubMed]
- Bangs, J.D.; Crain, P.F.; Hashizume, T.; McCloskey, J.A.; Boothroyd, J.C. Mass spectrometry of mRNA cap 4 from trypanosomatids reveals two novel nucleosides. J. Biol. Chem. 1992, 267, 9805–9815. [Google Scholar] [PubMed]
- Zamudio, J.R.; Mittra, B.; Campbell, D.A.; Sturm, N.R. Hypermethylated cap 4 maximizes Trypanosoma brucei translation. Mol. Microbiol. 2009, 72, 1100–1110. [Google Scholar] [CrossRef] [PubMed]
- Kozak, M. Regulation of translation via mRNA structure in prokaryotes and eukaryotes. Gene 2005, 361, 13–37. [Google Scholar] [CrossRef] [PubMed]
- Kozak, M. Mechanism of mRNA recognition by eukaryotic ribosomes during initiation of protein synthesis. Curr. Top. Microbiol. Immunol. 1981, 93, 81–123. [Google Scholar] [PubMed]
- Hamilton, R.; Watanabe, C.K.; de Boer, H.A. Compilation and comparison of the sequence context around the AUG startcodons in Saccharomyces cerevisiae mRNAs. Nucleic Acids Res. 1987, 15, 3581–3593. [Google Scholar] [CrossRef] [PubMed]
- Cigan, A.M.; Donahue, T.F. Sequence and structural features associated with translational initiator regions in yeast—A review. Gene 1987, 59, 1–18. [Google Scholar] [CrossRef]
- Joshi, C.P.; Zhou, H.; Huang, X.; Chiang, V.L. Context sequences of translation initiation codon in plants. Plant Mol. Biol. 1997, 35, 993–1001. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhuang, Y.; Gill, J.; Lin, S. Proof that dinoflagellate spliced leader (DinoSL) is a useful hook for fishing dinoflagellate transcripts from mixed microbial samples: Symbiodinium kawagutii as a case study. Protist 2013, 164, 510–527. [Google Scholar] [CrossRef] [PubMed]
- Bodyl, A.; Mackiewicz, P. Analysis of the targeting sequences of an iron-containing superoxide dismutase (SOD) of the dinoflagellate Lingulodinium polyedrum suggests function in multiple cellular compartments. Arch. Microbiol. 2007, 187, 281–296. [Google Scholar] [CrossRef] [PubMed]
- Bertucci, A.; Tambutte, E.; Tambutte, S.; Allemand, D.; Zoccola, D. Symbiosis-dependent gene expression in coral-dinoflagellate association: Cloning and characterization of a P-type H+-ATPase gene. Proc. Biol. Sci. 2010, 277, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Maroteaux, L.; Herzog, M.; Soyer-Gobillard, M.O. Molecular organization of dinoflagellate ribosomal DNA: evolutionary implications of the deduced 5.8 S rRNA secondary structure. Biosystems 1985, 18, 307–319. [Google Scholar] [CrossRef]
- Gressel, J.; Berman, T.; Cohen, N. Dinoflagellate ribosomal RNA; an evolutionary relic? J. Mol. Evol. 1975, 5, 307–313. [Google Scholar] [CrossRef] [PubMed]
- Jackson, K.E.; Habib, S.; Frugier, M.; Hoen, R.; Khan, S.; Pham, J.S.; Ribas de Pouplana, L.; Royo, M.; Santos, M.A.; Sharma, A.; et al. Protein translation in Plasmodium parasites. Trends Parasitol. 2011, 27, 467–476. [Google Scholar] [CrossRef] [PubMed]
- Zinoviev, A.; Shapira, M. Evolutionary conservation and diversification of the translation initiation apparatus in trypanosomatids. Comp. Funct. Genom. 2012, 2012, 813718. [Google Scholar] [CrossRef] [PubMed]
- De Melo Neto, O.P.; Reis, C.R.S.; Moura, D.M.N.; Freire, E.R.; Carrington, M. Unique and Conserved Features of the Protein Synthesis Apparatus in Parasitic Trypanosomatid (Trypanosoma and Leishmania) Species. In Evolution of the Protein Synthesis Machinery and Its Regulation, Hernández, G., Jagus, R., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 435–475. [Google Scholar]
- Eukaryotic Initiation Factors Gene Family. Available online: https://www.arabidopsis.org/browse/genefamily/eIF.jsp (accessed on 3 April 2018).
- Gingras, A.C.; Raught, B.; Sonenberg, N. eIF4 initiation factors: Effectors of mRNA recruitment to ribosomes and regulators of translation. Annu. Rev. Biochem. 1999, 68, 913–963. [Google Scholar] [CrossRef] [PubMed]
- Von der Haar, T.; Gross, J.D.; Wagner, G.; McCarthy, J.E. The mRNA cap-binding protein eIF4E in post-transcriptional gene expression. Nat. Struct. Mol. Biol. 2004, 11, 503–511. [Google Scholar] [CrossRef] [PubMed]
- Aravind, L.; Koonin, E.V. Eukaryote-specific domains in translation initiation factors: Implications for translation regulation and evolution of the translation system. Genome Res. 2000, 10, 1172–1184. [Google Scholar] [CrossRef] [PubMed]
- Marcotrigiano, J.; Gingras, A.C.; Sonenberg, N.; Burley, S.K. Cocrystal structure of the messenger RNA 5’ cap-binding protein (eIF4E) bound to 7-methyl-GDP. Cell 1997, 89, 951–961. [Google Scholar] [CrossRef]
- Matsuo, H.; Li, H.; McGuire, A.M.; Fletcher, C.M.; Gingras, A.C.; Sonenberg, N.; Wagner, G. Structure of translation factor eIF4E bound to m7GDP and interaction with 4E-binding protein. Nat. Struct. Biol. 1997, 4, 717–724. [Google Scholar] [CrossRef] [PubMed]
- Hernández, G.; Gillespie, K.M.; Bachvaroff, T.R.; Jagus, R.; Igreja, C.; Peter, D.; Bulfoni, M.; Cosson, B. Evolution of eIF4E-interacting proteins. In Evolution of the Protein Synthetic Machinery and Its Regulation; Hernandez, G., Jagus, R., Eds.; Spinger: Cham, Switzerland, 2016; pp. 165–186. [Google Scholar]
- Joshi, B.; Lee, K.; Maeder, D.L.; Jagus, R. Phylogenetic analysis of eIF4E-family members. BMC Evol. Biol. 2005, 5, 48. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, G.; Vazquez-Pianzola, P. Functional diversity of the eukaryotic translation initiation factors belonging to eIF4 families. Mech. Dev. 2005, 122, 865–876. [Google Scholar] [CrossRef] [PubMed]
- Jones, G.D.; Williams, E.P.; Place, A.R.; Jagus, R.; Bachvaroff, T.R. The alveolate translation initiation factor 4E family reveals a custom toolkit for translational control in core dinoflagellates. BMC Evol. Biol. 2015, 15, 14. [Google Scholar] [CrossRef] [PubMed]
- Jagus, R.; Bachvaroff, T.R.; Joshi, B.; Place, A.R. Diversity of Eukaryotic Translational Initiation Factor eIF4E in Protists. Comp. Funct. Genom. 2012, 2012, 134839. [Google Scholar] [CrossRef] [PubMed]
- Rhoads, R.E.; Dinkova, T.D.; Jagus, R. Approaches for analyzing the differential activities and functions of eIF4E family members. Methods Enzymol. 2007, 429, 261–297. [Google Scholar] [CrossRef] [PubMed]
- Rhoads, R.E. eIF4E: New family members, new binding partners, new roles. J. Biol. Chem. 2009, 284, 16711–16715. [Google Scholar] [CrossRef] [PubMed]
- Kamenska, A.; Simpson, C.; Standart, N. eIF4E-binding proteins: New factors, new locations, new roles. Biochem. Soc. Trans. 2014, 42, 1238–1245. [Google Scholar] [CrossRef] [PubMed]
- Lasko, P. Posttranscriptional regulation in Drosophila oocytes and early embryos. Wiley Interdiscip. Rev. RNA 2011, 2, 408–416. [Google Scholar] [CrossRef] [PubMed]
- Jung, M.Y.; Lorenz, L.; Richter, J.D. Translational control by neuroguidin, a eukaryotic initiation factor 4E and CPEB binding protein. Mol. Cell. Biol. 2006, 26, 4277–4287. [Google Scholar] [CrossRef] [PubMed]
- Wells, S.E.; Hillner, P.E.; Vale, R.D.; Sachs, A.B. Circularization of mRNA by eukaryotic translation initiation factors. Mol. Cell 1998, 2, 135–140. [Google Scholar] [CrossRef]
- Freire, E.R.; Sturm, N.R.; Campbell, D.A.; de Melo Neto, O.P. The Role of Cytoplasmic mRNA Cap-Binding Protein Complexes in Trypanosoma brucei and Other Trypanosomatids. Pathogens 2017, 6, 55. [Google Scholar] [CrossRef] [PubMed]
- Yoffe, Y.; Leger, M.; Zinoviev, A.; Zuberek, J.; Darzynkiewicz, E.; Wagner, G.; Shapira, M. Evolutionary changes in the Leishmania eIF4F complex involve variations in the eIF4E-eIF4G interactions. Nucleic Acids Res. 2009, 37, 3243–3253. [Google Scholar] [CrossRef] [PubMed]
- Park, E.H.; Walker, S.E.; Lee, J.M.; Rothenburg, S.; Lorsch, J.R.; Hinnebusch, A.G. Multiple elements in the eIF4G1 N-terminus promote assembly of eIF4G1*PABP mRNPs in vivo. EMBO J. 2011, 30, 302–316. [Google Scholar] [CrossRef] [PubMed]
- Jivotovskaya, A.V.; Valasek, L.; Hinnebusch, A.G.; Nielsen, K.H. Eukaryotic translation initiation factor 3 (eIF3) and eIF2 can promote mRNA binding to 40S subunits independently of eIF4G in yeast. Mol. Cell. Biol. 2006, 26, 1355–1372. [Google Scholar] [CrossRef] [PubMed]
- Wek, R.C.; Jiang, H.Y.; Anthony, T.G. Coping with stress: EIF2 kinases and translational control. Biochem. Soc. Trans. 2006, 34, 7–11. [Google Scholar] [CrossRef] [PubMed]
- Sonenberg, N.; Hinnebusch, A.G. Regulation of translation initiation in eukaryotes: Mechanisms and biological targets. Cell 2009, 136, 731–745. [Google Scholar] [CrossRef] [PubMed]
- Jennings, M.D.; Kershaw, C.J.; Adomavicius, T.; Pavitt, G.D. Fail-safe control of translation initiation by dissociation of eIF2 alpha phosphorylated ternary complexes. Elife 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Baird, T.D.; Wek, R.C. Eukaryotic initiation factor 2 phosphorylation and translational control in metabolism. Adv. Nutr. 2012, 3, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Donnelly, N.; Gorman, A.M.; Gupta, S.; Samali, A. The eIF2α kinases: Their structures and functions. Cell. Mol. Life Sci. 2013, 70, 3493–3511. [Google Scholar] [CrossRef] [PubMed]
- Mohrle, J.J.; Zhao, Y.; Wernli, B.; Franklin, R.M.; Kappes, B. Molecular cloning, characterization and localization of PfPK4, an eIF2 α kinase-related enzyme from the malarial parasite Plasmodium falciparum. Biochem. J. 1997, 328, 677–687. [Google Scholar] [CrossRef] [PubMed]
- Narasimhan, J.; Joyce, B.R.; Naguleswaran, A.; Smith, A.T.; Livingston, M.R.; Dixon, S.E.; Coppens, I.; Wek, R.C.; Sullivan, W.J., Jr. Translation regulation by eukaryotic initiation factor-2 kinases in the development of latent cysts in Toxoplasma gondii. J. Biol. Chem. 2008, 283, 16591–16601. [Google Scholar] [CrossRef] [PubMed]
- Vonlaufen, N.; Kanzok, S.M.; Wek, R.C.; Sullivan, W.J., Jr. Stress response pathways in protozoan parasites. Cell. Microbiol. 2008, 10, 2387–2399. [Google Scholar] [CrossRef] [PubMed]
- Joyce, B.R.; Konrad, C.; Wek, R.C.; Sullivan, W.J., Jr. Translation control is critical during acute and chronic stages of toxoplasmosis infection. Expert Rev. Anti-Infect. Ther. 2011, 9, 1–3. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, M.; Joyce, B.R.; Sullivan, W.J., Jr.; Nussenzweig, V. Translational control in Plasmodium and Toxoplasma parasites. Eukaryot. Cell 2013, 12, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Konrad, C.; Wek, R.C.; Sullivan, W.J., Jr. A GCN2-like eukaryotic initiation factor 2 kinase increases the viability of extracellular Toxoplasma gondii parasites. Eukaryot. Cell 2011, 10, 1403–1412. [Google Scholar] [CrossRef] [PubMed]
- Konrad, C.; Wek, R.C.; Sullivan, W.J., Jr. GCN2-like eIF2α kinase manages the amino acid starvation response in Toxoplasma gondii. Int. J. Parasitol. 2014, 44, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Dev, K.; Qiu, H.; Dong, J.; Zhang, F.; Barthlme, D.; Hinnebusch, A.G. The β/Gcd7 subunit of eukaryotic translation initiation factor 2B (eIF2B), a guanine nucleotide exchange factor, is crucial for binding eIF2 in vivo. Mol. Cell. Biol. 2010, 30, 5218–5233. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Shopan, J.; Mou, H.; Zhang, L.; Zhang, C.; Ma, W.; Walsh, J.A.; Hu, Z.; Yang, J.; Zhang, M. Eukaryotic translation initiation factor 2B-beta (eIF2B β), a new class of plant virus resistance gene. Plant J. Cell Mol. Biol. 2017, 90, 929–940. [Google Scholar] [CrossRef] [PubMed]
- Gomez, E.; Pavitt, G.D. Identification of domains and residues within the epsilon subunit of eukaryotic translation initiation factor 2B (eIF2B epsilon) required for guanine nucleotide exchange reveals a novel activation function promoted by eIF2B complex formation. Mol. Cell. Biol. 2000, 20, 3965–3976. [Google Scholar] [CrossRef] [PubMed]
- Glisovic, T.; Bachorik, J.L.; Yong, J.; Dreyfuss, G. RNA-binding proteins and post-transcriptional gene regulation. FEBS Lett. 2008, 582, 1977–1986. [Google Scholar] [CrossRef] [PubMed]
- Baltz, A.G.; Munschauer, M.; Schwanhausser, B.; Vasile, A.; Murakawa, Y.; Schueler, M.; Youngs, N.; Penfold-Brown, D.; Drew, K.; Milek, M.; et al. The mRNA-bound proteome and its global occupancy profile on protein-coding transcripts. Mol. Cell 2012, 46, 674–690. [Google Scholar] [CrossRef] [PubMed]
- Hentze, M.W.; Castello, A.; Schwarzl, T.; Preiss, T. A brave new world of RNA-binding proteins. Nat Rev Mol. Cell. Biol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Mittag, M.; Lee, D.-H.; Hastings, J.W. Circadian expression of the luciferin-binding protein correlates with the binding of a protein to the 3’ untranslated region of its mRNA. Proc. Natl. Acad. Sci. USA 1994, 91, 5257–5261. [Google Scholar] [CrossRef] [PubMed]
- Lapointe, M.; Morse, D. Reassessing the role of a 3’UTR-binding translational inhibitor in regulation of the circadian bioluminescence reaction of the dinoflagellate Gonyaulax. Biol. Chem. 2008, 389, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.C.; Ambros, V. An extensive class of small RNAs in Caenorhabditis elegans. Science 2001, 294, 862–864. [Google Scholar] [CrossRef] [PubMed]
- Reinhart, B.J.; Weinstein, E.G.; Rhoades, M.W.; Bartel, B.; Bartel, D.P. MicroRNAs in plants. Genes Dev. 2002, 16, 1616–1626. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Carmell, M.A.; Rivas, F.V.; Marsden, C.G.; Thomson, J.M.; Song, J.J.; Hammond, S.M.; Joshua-Tor, L.; Hannon, G.J. Argonaute2 is the catalytic engine of mammalian RNAi. Science 2004, 305, 1437–1441. [Google Scholar] [CrossRef] [PubMed]
- Vasudevan, S.; Tong, Y.; Steitz, J.A. Switching from repression to activation: MicroRNAs can up-regulate translation. Science 2007, 318, 1931–1934. [Google Scholar] [CrossRef] [PubMed]
- Sijen, T.; Fleenor, J.; Simmer, F.; Thijssen, K.L.; Parrish, S.; Timmons, L.; Plasterk, R.H.; Fire, A. On the role of RNA amplification in dsRNA-triggered gene silencing. Cell 2001, 107, 465–476. [Google Scholar] [CrossRef]
- Cerutti, H.; Casas-Mollano, J.A. On the origin and functions of RNA-mediated silencing: From protists to man. Curr. Genet. 2006, 50, 81–99. [Google Scholar] [CrossRef] [PubMed]
- Baum, J.; Papenfuss, A.T.; Mair, G.R.; Janse, C.J.; Vlachou, D.; Waters, A.P.; Cowman, A.F.; Crabb, B.S.; de Koning-Ward, T.F. Molecular genetics and comparative genomics reveal RNAi is not functional in malaria parasites. Nucleic Acids Res. 2009, 37, 3788–3798. [Google Scholar] [CrossRef] [PubMed]
- Gunasekera, A.M.; Patankar, S.; Schug, J.; Eisen, G.; Kissinger, J.; Roos, D.; Wirth, D.F. Widespread distribution of antisense transcripts in the Plasmodium falciparum genome. Mol. Biochem. Parasitol. 2004, 136, 35–42. [Google Scholar] [CrossRef] [PubMed]
- De Riso, V.; Raniello, R.; Maumus, F.; Rogato, A.; Bowler, C.; Falciatore, A. Gene silencing in the marine diatom Phaeodactylum tricornutum. Nucleic Acids Res. 2009, 37, e96. [Google Scholar] [CrossRef] [PubMed]
- Dagenais-Bellefeuille, S.; Beauchemin, M.; Morse, D. miRNAs Do Not Regulate Circadian Protein Synthesis in the Dinoflagellate Lingulodinium polyedrum. PLoS ONE 2017, 12, e0168817. [Google Scholar] [CrossRef] [PubMed]
- Hausser, J.; Landthaler, M.; Jaskiewicz, L.; Gaidatzis, D.; Zavolan, M. Relative contribution of sequence and structure features to the mRNA binding of Argonaute/EIF2C-miRNA complexes and the degradation of miRNA targets. Genome Res. 2009, 19, 2009–2020. [Google Scholar] [CrossRef] [PubMed]
- Gao, D.; Qiu, L.; Hou, Z.; Zhang, Q.; Wu, J.; Gao, Q.; Song, L. Computational Identification of MicroRNAs from the Expressed Sequence Tags of Toxic Dinoflagellate Alexandrium tamarense. Evol. Bioinform. Online 2013, 9, 479–485. [Google Scholar] [CrossRef] [PubMed]
- Baumgarten, S.; Bayer, T.; Aranda, M.; Liew, Y.J.; Carr, A.; Micklem, G.; Voolstra, C.R. Integrating microRNA and mRNA expression profiling in Symbiodinium microadriaticum, a dinoflagellate symbiont of reef-building corals. BMC Genom. 2013, 14, 704. [Google Scholar] [CrossRef] [PubMed]
- Walsh, C.T.; Garneau-Tsodikova, S.; Gatto, G.J., Jr. Protein posttranslational modifications: The chemistry of proteome diversifications. Angew. Chem. Int. Ed. Engl. 2005, 44, 7342–7372. [Google Scholar] [CrossRef] [PubMed]
- Karve, T.M.; Cheema, A.K. Small changes huge impact: The role of protein posttranslational modifications in cellular homeostasis and disease. J. Amino Acids 2011, 2011, 207691. [Google Scholar] [CrossRef] [PubMed]
- Khoury, G.A.; Baliban, R.C.; Floudas, C.A. Proteome-wide post-translational modification statistics: Frequency analysis and curation of the swiss-prot database. Sci Rep 2011, 1. [Google Scholar] [CrossRef] [PubMed]
- Cohen, P. The regulation of protein function by multisite phosphorylation—A 25 year update. Trends Biochem. Sci. 2000, 25, 596–601. [Google Scholar] [CrossRef]
- Holt, L.J.; Tuch, B.B.; Villen, J.; Johnson, A.D.; Gygi, S.P.; Morgan, D.O. Global analysis of Cdk1 substrate phosphorylation sites provides insights into evolution. Science 2009, 325, 1682–1686. [Google Scholar] [CrossRef] [PubMed]
- Kersten, B.; Agrawal, G.K.; Durek, P.; Neigenfind, J.; Schulze, W.; Walther, D.; Rakwal, R. Plant phosphoproteomics: An update. Proteomics 2009, 9, 964–988. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.Z.; Li, C.; Xie, Z.X.; Dong, H.P.; Lin, L.; Hong, H.S. Homology-Driven Proteomics of Dinoflagellates with Unsequenced Genomes Using MALDI-TOF/TOF and Automated De Novo Sequencing. Evid. Based Complement. Alternat. Med. 2011, 2011, 471020. [Google Scholar] [CrossRef] [PubMed]
- Vanselow, K.; Kramer, A. Role of phosphorylation in the mammalian circadian clock. Cold Spring Harb. Symp. Quant. Biol. 2007, 72, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Kusakina, J.; Dodd, A.N. Phosphorylation in the plant circadian system. Trends Plant Sci. 2012, 17, 575–583. [Google Scholar] [CrossRef] [PubMed]
- Schafmeier, T.; Kaldi, K.; Diernfellner, A.; Mohr, C.; Brunner, M. Phosphorylation-dependent maturation of Neurospora circadian clock protein from a nuclear repressor toward a cytoplasmic activator. Genes Dev. 2006, 20, 297–306. [Google Scholar] [CrossRef] [PubMed]
- Kurosawa, G.; Aihara, K.; Iwasa, Y. A model for the circadian rhythm of cyanobacteria that maintains oscillation without gene expression. Biophys. J. 2006, 91, 2015–2023. [Google Scholar] [CrossRef] [PubMed]
- Dodd, A.N.; Gardner, M.J.; Hotta, C.T.; Hubbard, K.E.; Dalchau, N.; Love, J.; Assie, J.M.; Robertson, F.C.; Jakobsen, M.K.; Goncalves, J.; et al. The Arabidopsis circadian clock incorporates a cADPR-based feedback loop. Science 2007, 318, 1789–1792. [Google Scholar] [CrossRef] [PubMed]
- Edwards, K.D.; Anderson, P.E.; Hall, A.; Salathia, N.S.; Locke, J.C.; Lynn, J.R.; Straume, M.; Smith, J.Q.; Millar, A.J. FLOWERING LOCUS C mediates natural variation in the high-temperature response of the Arabidopsis circadian clock. Plant Cell 2006, 18, 639–650. [Google Scholar] [CrossRef] [PubMed]
- Comolli, J.; Taylor, W.; Hastings, J.W. An inhibitor of protein phosphorylation stops the circadian oscillator and blocks light-induced phase shifting in Gonyaulax polyedra. J. Biol. Rhythm. 1994, 9, 13–26. [Google Scholar] [CrossRef] [PubMed]
- Comolli, J.C.; Hastings, J.W. Novel effects on the Gonyaulax circadian system produced by the protein kinase inhibitor staurosporine. J. Biol. Rhythm. 1999, 14, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Morse, D. The dinoflagellate Lingulodinium has predicted casein kinase 2 sites in many RNA binding proteins. Protist 2014, 165, 330–342. [Google Scholar] [CrossRef] [PubMed]
- Prosperi, E.; Scovassi, A.I.; Stivala, L.A.; Bianchi, L. Proliferating cell nuclear antigen bound to DNA synthesis sites: Phosphorylation and association with cyclin D1 and cyclin A. Exp. Cell Res. 1994, 215, 257–262. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Lo, S.C.; Matton, D.P.; Lang, B.F.; Morse, D. Daily changes in the phosphoproteome of the dinoflagellate Lingulodinium. Protist 2012, 163, 746–754. [Google Scholar] [CrossRef] [PubMed]
- Hochstrasser, M. Origin and function of ubiquitin-like proteins. Nature 2009, 458, 422–429. [Google Scholar] [CrossRef] [PubMed]
- Glickman, M.H.; Ciechanover, A. The ubiquitin-proteasome proteolytic pathway: Destruction for the sake of construction. Physiol. Rev. 2002, 82, 373–428. [Google Scholar] [CrossRef] [PubMed]
- Komander, D.; Rape, M. The ubiquitin code. Annu. Rev. Biochem. 2012, 81, 203–229. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Roy, S.; Jagus, R.; Morse, D. Translation and Translational Control in Dinoflagellates. Microorganisms 2018, 6, 30. https://doi.org/10.3390/microorganisms6020030
Roy S, Jagus R, Morse D. Translation and Translational Control in Dinoflagellates. Microorganisms. 2018; 6(2):30. https://doi.org/10.3390/microorganisms6020030
Chicago/Turabian StyleRoy, Sougata, Rosemary Jagus, and David Morse. 2018. "Translation and Translational Control in Dinoflagellates" Microorganisms 6, no. 2: 30. https://doi.org/10.3390/microorganisms6020030
APA StyleRoy, S., Jagus, R., & Morse, D. (2018). Translation and Translational Control in Dinoflagellates. Microorganisms, 6(2), 30. https://doi.org/10.3390/microorganisms6020030




