Fungal Biodiversity in the Alpine Tarfala Valley
Abstract
:1. Introduction
2. Experimental Section
2.1. Study Area
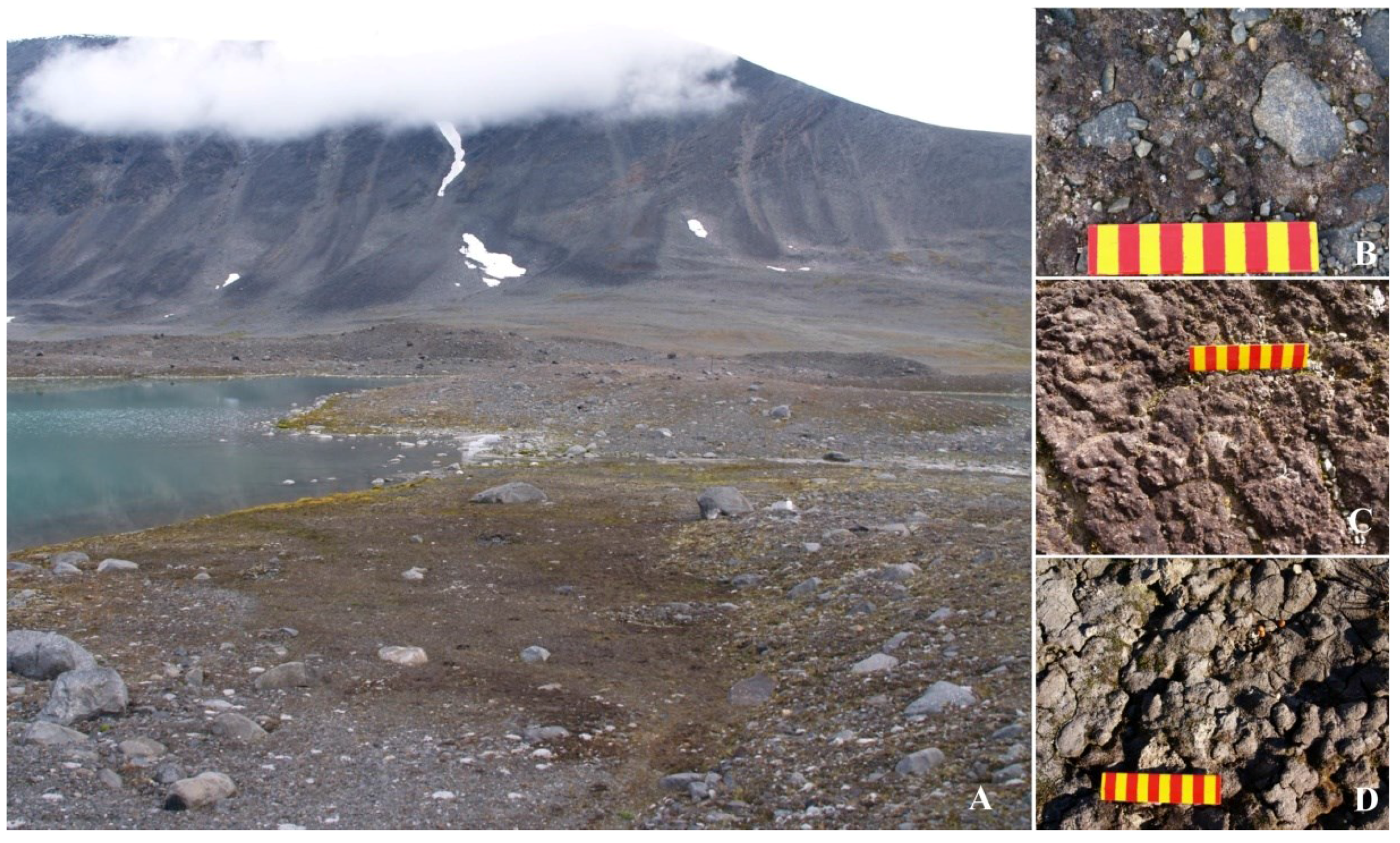

2.2. Sampling

| Moraine Locations | Sample Number | Position | Altitude | Date of Sampling |
|---|---|---|---|---|
| Isfallglaciären | C01 | 67°54′48.2″ N 18°35′19.8″ E | 1184 m | 25 August 2014 |
| Isfallglaciären | C02 | 67°54′50.8″ N 18°35′20.2″ E | 1178 m | 25 August 2014 |
| Isfallglaciären | C03 | 67°54′51.8″ N 18°35′23.4″ E | 1174 m | 25 August 2014 |
| Isfallglaciären | C04 | 67°54′52.1″ N 18°35′24.3″ E | 1175 m | 26 August 2014 |
| Isfallglaciären | C05 | 67°55′02.1″ N 18°35′22.3″ E | 1176 m | 26 August 2014 |
| Isfallglaciären | C06 | 67°55′03.3″ N 18°35′20.4″ E | 1179 m | 26 August 2014 |
| Isfallglaciären | C07 | 67°55′00.1″ N 18°35′19.5″ E | 1172 m | 26 August 2014 |
| Isfallglaciären | C08 | 67°55′08.2″ N 18°35′39.3″ E | 1164 m | 27 August 2014 |
| Isfallglaciären | C09 | 67°55′08.2″ N 18°35′39.3″ E | 1164 m | 27 August 2014 |
| Isfallglaciären | C10 | 67°55′02.1″ N 18°35′28.0″ E | 1175 m | 27 August 2014 |
| Isfallglaciären | C11 | 67°55′02.3″ N 18°35′27.9″ E | 1175 m | 28 August 2014 |
| Kaskasatjåkkaglaciären | C12 | 67°55′55.7″ N 18°36′06.4″ E | 1439 m | 30 August 2014 |
| Tarfalajaure Lake | C13 | 67°55′16.3″ N 18°36′28.2″ E | 1181 m | 30 August 2014 |
2.3. Soil Analyses
2.4. Isolation
2.5. DNA Extraction, Amplification and Sequencing
2.6. Denaturing Gradient Gel Electrophoresis Fingerprinting Statistics, Quantification and Analysis
3. Results and Discussion
| Samples | C (%) | Clay (%) | Silt (%) | Sand (%) | Skeleton (>2 mm) (%) | pH(H2O) 1:2.5 | T (°C) |
|---|---|---|---|---|---|---|---|
| C01 | 0.09 | 2 | 41 | 57 | 18 | 6.1 | 6.5 |
| C02 | 0.10 | 3 | 28 | 69 | 45 | 6.5 | 7.7 |
| C03 | 0.09 | 3 | 30 | 66 | 9 | 6.5 | 7.4 |
| C04 | 0.08 | 3 | 32 | 65 | 12 | 6.6 | 6.9 |
| C05 | 0.17 | 2 | 15 | 84 | 37 | 6.6 | 13.8 |
| C06 | 0.05 | 1 | 17 | 82 | 50 | 7.2 | 10.5 |
| C07 | 0.18 | 1 | 14 | 85 | 10 | 7.2 | 13.4 |
| C08 | 0.06 | 2 | 16 | 82 | 10 | 6.9 | 18.8 |
| C09 | 1.21 | 0 | 30 | 69 | 49 | 6.4 | 4.1 |
| C10 | 0.07 | 1 | 20 | 80 | 0 | 6.7 | 12.2 |
| C11 | 0.04 | 3 | 23 | 74 | 48 | 6.8 | 10.9 |
| C12 | 0.20 | 4 | 27 | 69 | 2 | 6.5 | 17.4 |
| C13 | 3.22 | 2 | 51 | 47 | 26 | 6.3 | 20.0 |
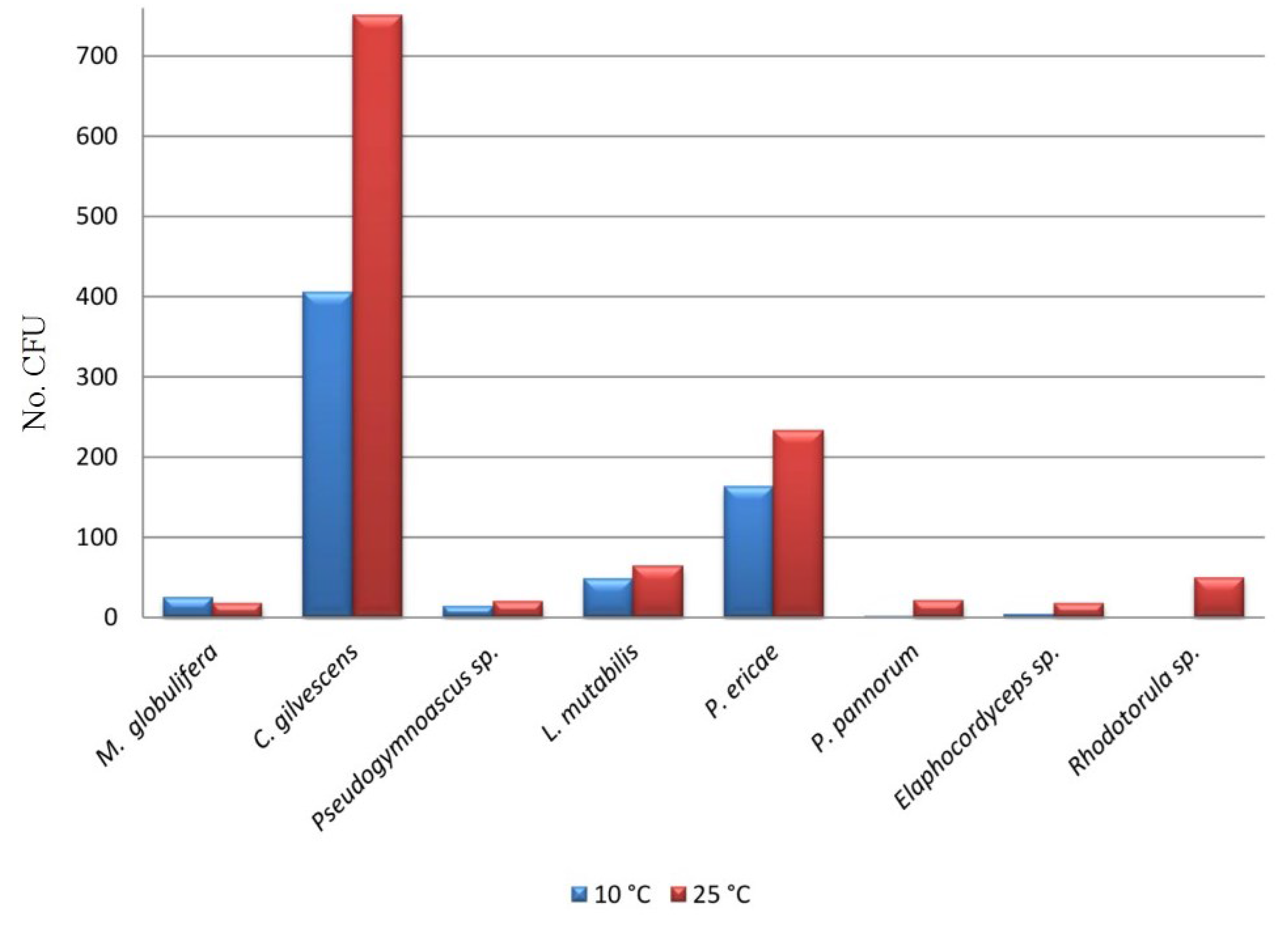
| Taxa | C01 | C02 | C03 | C04 | C05 | C06 | C07 | C08 | C09 | C10 | C11 | C12 | C13 | CFU (%) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Mortierella globulifera O. Rostr | 31 | - | - | - | - | - | - | - | - | 21 | - | - | - | 2.7 |
| Cryptococcus gilvescens Chernov & Babeva | 250 | 23 | 139 | 43 | 90 | - | 228 | 8 | 12 | - | 264 | 134 | - | 62.5 |
| Pseudogymnoascus sp. | - | 7 | 4 | - | - | - | - | - | 22 | - | - | - | - | 1.7 |
| Lecythophora mutabilis (J.F.H. Beyma) W. Gams & McGinnis | - | 131 | - | - | - | - | - | - | - | - | - | - | - | 6.9 |
| Pezoloma ericae (D.J. Read) Baral | 15 | - | 59 | 3 | 30 | 25 | - | 11 | - | 140 | 42 | 10 | 63 | 20.9 |
| Pseudogymnoascus pannorum (Link) Minnis & D.L. Lindner | - | - | - | - | 1 | 21 | - | - | - | - | - | - | - | 1.2 |
| Elaphocordyceps sp. | - | - | - | - | - | - | - | - | 12 | 17 | - | - | - | 1.5 |
| Rhodotorula sp. | 33 | - | - | - | - | - | 13 | - | 3 | - | - | - | - | 2.6 |
| CFU Total | 329 | 161 | 202 | 46 | 121 | 46 | 241 | 19 | 49 | 178 | 306 | 144 | 63 | 1905 |
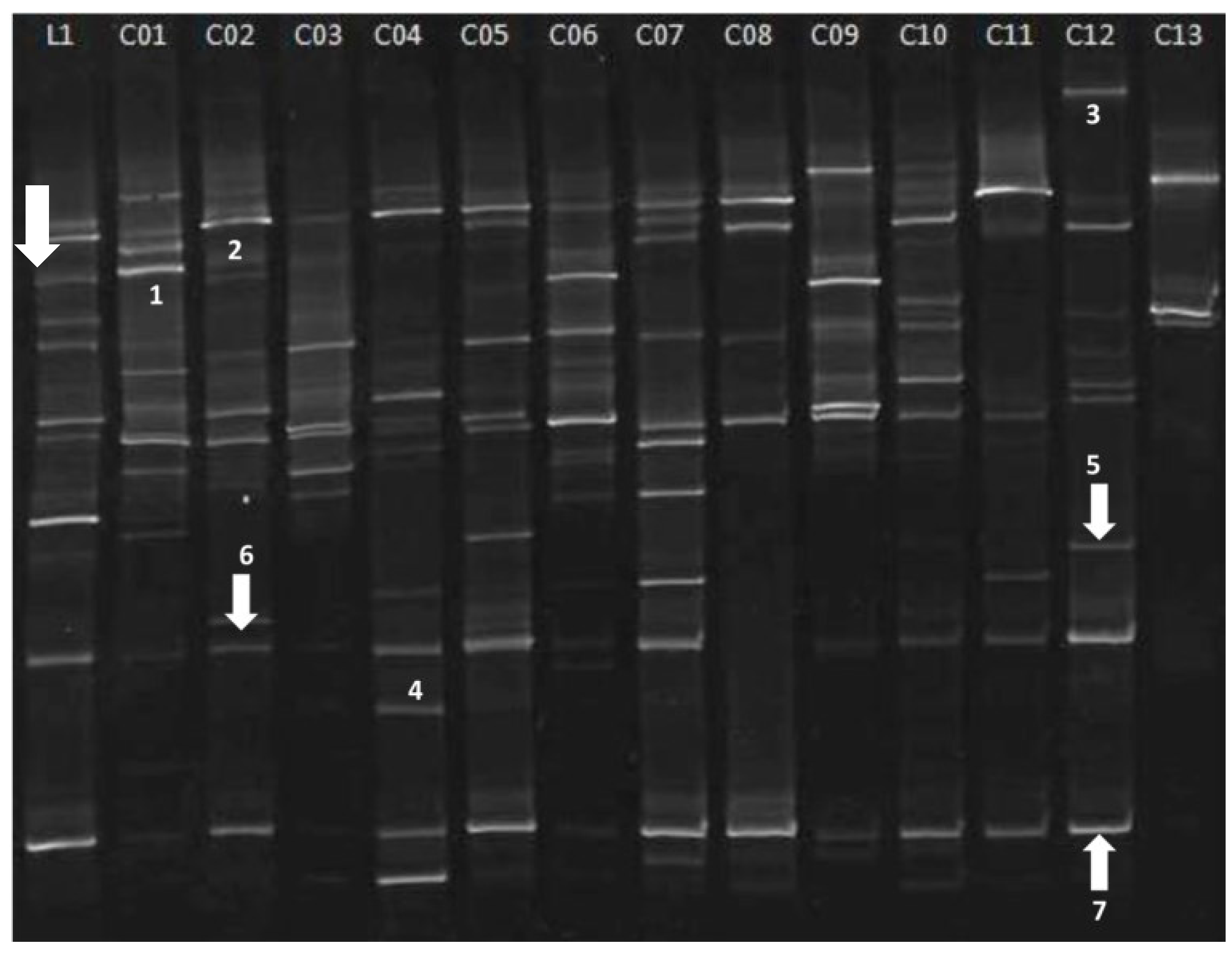
| Band a | Highest match (NCBI Accession No.) | Identity (%) b |
|---|---|---|
| 5 | Rhodotorula sp. (JF805370.1) | 100% |
| 6 | Cryptococcus gilvescens (AB032678.1) | 99% |
| 7 | Pezoloma ericae (AM887700.1) | 99% |
| Sample | C01 | C02 | C03 | C04 | C05 | C06 | C07 | C08 | C09 | C10 | C11 | C12 | C13 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Richness (S) | 12 | 8 | 9 | 11 | 12 | 11 | 11 | 6 | 9 | 14 | 6 | 10 | 4 |
| Shannon (Hʹ) | 1.06 | 0.9 | 0.55 | 1.01 | 1.06 | 1.02 | 1.01 | 0.75 | 0.9 | 1.0 | 0.7 | 0.87 | 0.57 |
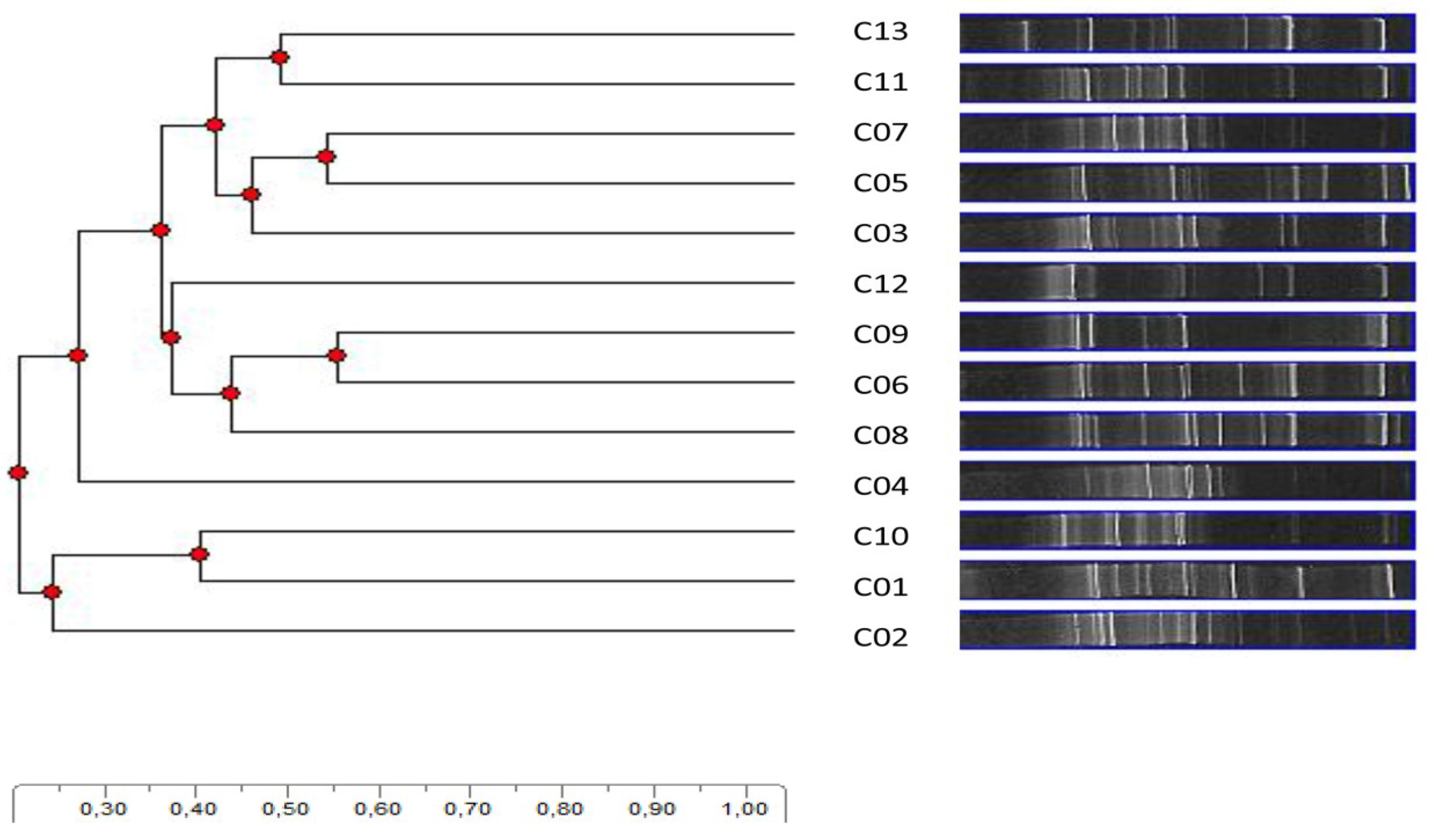
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Belnap, J.; Lange, O.L. Biological Soil Crusts: Structure, Function and Management; Springer-Verlag: Berlin, Germany, 2001; p. 503. [Google Scholar]
- Bates, S.T.; Garcia-Pichel, F.; Nash, T.H. Fungal components of biological soil crusts: Insights from culture-dependent and culture-independent studies. In Biology of Lichens—Symbiosis, Ecology, Environmental Monitoring, Systematics, Cyber Applications; Nash, T., III, Geiser, L., McCune, B., Triebel, D., Tornescu, A.M., Sanders, W., Eds.; Bibliotheca Lichenologica: Stuttgard, Genermy, 2010; Volume 105, pp. 197–210. [Google Scholar]
- Soule, T.; Anderson, I.J.; Johnson, S.L.; Bates, S.T.; Garcia-Pichel, F. Archaeal populations in biological soil crusts from arid lands in North America. Soil Biol. Biochem. 2009, 41, 2069–2074. [Google Scholar] [CrossRef]
- Cameron, R.E.; Blank, G.B. Desert Algae: Soil Crusts and Diaphanous Substrata as Algal Habitats; Jet Propulsion Laboratory, California Institute of Technology: Pasadena, CA, USA, 1966; pp. 32–971. [Google Scholar]
- Friedmann, E.I.; Galun, M. Desert algae, lichens and fungi. In Desert Biology; Brown, G.W., Ed.; Academic Press: New York, NY, USA, 1974; pp. 165–212. [Google Scholar]
- Belnap, J.; Gardner, J.S. Soil microstructure of the Colorado Plateau: The role of the cyanobacterium Microcoleus vaginatus. Great Basin Nat. 1993, 53, 40–47. [Google Scholar]
- Borin, S.; Ventura, S.; Tambone, F.; Mapelli, F.; Schubotz, F.; Brusetti, L.; Scaglia, B.; Acqui, L.P.D.; Solheim, B.; Turicchia, S.; et al. Rock weathering creates oases of life in a high Arctic desert. Environ. Microbiol. 2010, 12, 293–303. [Google Scholar] [CrossRef] [PubMed]
- Eldridge, D.; Tozer, M.E. A practical guide to soil lichens and bryophytes of Australia’s dry country. Lichenologist 1998, 30, 303–304. [Google Scholar]
- Evans, R.D.; Johansen, J.R. Microbiotic crusts and ecosystem processes. Crit. Rev. Plant Sci. 1999, 18, 183–225. [Google Scholar] [CrossRef]
- Harper, K.T.; Marble, J.R. A role for nonvascular plants in management of arid and semiarid rangeland. In Vegetation Science Applications for Rangeland Analysis and Management; Tueller, P.T., Ed.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 1988; pp. 135–169. [Google Scholar]
- States, J.S.; Christensen, M. Fungi associated with biological soil crusts in desert grasslands of Utah and Wyoming. Mycologia 2001, 93, 432–439. [Google Scholar] [CrossRef]
- Grishkan, I.; Zaady, E.; Nevo, E. Soil crust microfungi along a southward rainfall gradient in desert ecosystems. Eur. J. Soil. Biol. 2006, 42, 33–42. [Google Scholar] [CrossRef]
- Bates, S.T.; Garcia-Pichel, F. A culture-independent study of free-living fungi in biological soil crusts of the Colorado Plateau: Their diversity and relative contribution to microbial biomass. Environ. Microbiol. 2009, 11, 56–67. [Google Scholar] [CrossRef] [PubMed]
- Porras-Alfaro, A.; Herrera, J.; Natvig, D.O.; Lipinski, K.; Sinsabaugh, R.L. Diversity and distribution of soil fungal communities in a semiarid grassland. Mycologia 2011, 103, 10–21. [Google Scholar] [CrossRef] [PubMed]
- Holmlund, P.; Jansson, P. Glaciological Research at Tarfala Research Station; Stockholm University: Stockholm, Sweden, 2002; p. 48. [Google Scholar]
- Società Italiana della Scienza del Suolo (SISS). Metodi Normalizzati di Analisi del Suolo; Edagricole: Bologna, Italy, 1985; p. 100. (In Italian) [Google Scholar]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols, a Guide to Methods and Applications; Innis, M.A., Gelfe, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Larena, I.; Salazar, O.; Gonzales, V.; Julian, M.C.; Rubio, V. Design of a primer for ribosomal DNA internal transcribed spacer with enhanced specifity for Ascomycetes. J. Biotechnol. 1999, 75, 187–194. [Google Scholar] [CrossRef]
- Sanger, F.; Nicklen, S.; Coulson, A.R. DNA sequencing with chain-terminating inhibitors. Proc. Nat. Acad. Sci. 1977, 74, 5463–5467. [Google Scholar] [CrossRef] [PubMed]
- Muyzer, G.; de Waal, E.C.; Uitterlinden, A.G. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl. Environ. Microbiol. 1993, 59, 695–700. [Google Scholar] [PubMed]
- Tao, G.; Liu, Z.Y.; Hyde, K.D.; Lui, X.Z.; Yu, Z.N. Whole rDNA analysis reveals novel and endophytic fungi in Bletilla ochracea (Orchidaceae). Fungal Divers 2008, 33, 101–122. [Google Scholar]
- Nakatsubo, T.; Bekk, Y.S.; Uchida, M.; Muraoka, H.; Kume, A.; Ohtsuka, T.; Masuzawa, T.; Kanda, H.; Koizumi, H. Ecosystem development and carbon cycle on a glacier foreland in the high Arctic, Ny-Ǻlesund, Svalbard. J. Plant Res. 2005, 118, 173–179. [Google Scholar] [CrossRef] [PubMed]
- Yoshitake, S.; Uchida, M.; Koizumi, H.; Nakatsubo, T. Carbon and nitrogen limitation of soil microbial respiration in a High Arctic successional glacier foreland near Ny-Ǻlesund, Svalbard. Polar Res. 2007, 26, 22–30. [Google Scholar] [CrossRef]
- Zucconi, L.; Pagano, S.; Fenice, M.; Selbmann, L.; Tosi, S.; Onofri, S. Growth temperature preferences of fungal strains from Victoria Land, Antarctica. Polar Biol. 1996, 16, 53–61. [Google Scholar] [CrossRef]
- Zucconi, L.; Selbmann, L.; Buzzini, P.; Turchetti, B.; Guglielmin, M.; Frisvad, J.C.; Onofri, S. Searching for eukaryotic life preserved in Antarctic permafrost. Polar Biol. 2012, 35, 749–757. [Google Scholar] [CrossRef]
- Buzzini, P.; Branda, E.; Goretti, M.; Turchetti, B. Psychrophilic yeasts from worldwide glacial habitats: diversity, adaptation strategies and biotechnological potential. FEMS Microbiol. Ecol. 2012, 82, 217–241. [Google Scholar] [CrossRef] [PubMed]
- Turchetti, B.; Buzzini, P.; Goretti, M.; Branda, E.; Diolaiuti, G.; D’Agata, C.; Smiraglia, C.; Vaughan-Martini, A. Psychrophilic yeasts in glacial environments of Alpine glaciers. FEMS Microbiol. Ecol. 2008, 63, 73–83. [Google Scholar] [CrossRef] [PubMed]
- Carrasco, M.; Rozas, J.M.; Barahona, S.; Alcaíno, J.; Cifuentes, V.; Baeza, M. Diversity and extracellular enzymatic activities of yeasts isolated from King George Island, the sub-Antarctic region. BMC Microbiol. 2012, 12, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Zalar, P.; Gunde-Cimerman, N. Cold adapted yeasts in Arctic habitats. In Cold-Adapted Yeasts. Biodiversity, Adaptation Strategies and Biotechnological Significance; Buzzini, P., Margesin, R., Eds.; Springer-Verlag: Berlin, Germany, 2014; pp. 49–74. [Google Scholar]
- Singh, P.; Singh, S.M. Characterization of yeast and filamentous fungi isolated from cryoconite holes of Svalbard, Arctic. Polar Biol. 2011, 35, 575–583. [Google Scholar] [CrossRef]
- Sieber, T.N.; Grünig, C.R. Fungal root endophytes. In Plant Roots: The Hidden Half, 4th ed.; Eshel, A., Beeckman, T., Eds.; CRC Press: Boca Raton, FL, USA, 2013; pp. 38–49. [Google Scholar]
- Bent, E.; Kiekel, P.; Brenton, R.; Taylor, D.L. Root-associated ectomycorrhizal fungi shared by various boreal forest seedlings naturally regenerating after a fire in interior Alaska and correlation of different fungi with host growth responses. Appl. Environ. Microbiol. 2011, 77, 3351–3359. [Google Scholar] [CrossRef] [PubMed]
- Panikov, N.S.; Sizova, M.V. Growth kinetics of microorganisms isolated from Alaskan soil and permafrost in solid media frozen down to −35 °C. FEMS Microbiol. Ecol. 2007, 59, 500–512. [Google Scholar] [CrossRef] [PubMed]
- Hayes, M.A. The Geomyces Fungi: Ecology and Distribution. BioScience 2012, 62, 819–823. [Google Scholar]
- Timling, I.; Taylor, D.L. Peeking through a frosty window: Molecular insights into the ecology of Arctic soil fungi. Fungal Ecol. 2012, 30, 1–11. [Google Scholar] [CrossRef]
- Bates, S.T.; Nash, T.N.; Garcia-Pichel, F. Patterns of diversity for fungal assemblages of biological soil crusts from the southwestern United States. Mycologia 2012, 104, 353–361. [Google Scholar] [CrossRef] [PubMed]
- Gundlapally, S.; Garcia-Pichel, F. The community and phylogenetic diversity of biological soil crusts in the Colorado Plateau studied by molecular fingerprinting and intensive cultivation. Microb. Ecol. 2006, 52, 345–357. [Google Scholar] [CrossRef] [PubMed]
- Collins, S.L.; Sinsabaugh, R.L.; Crenshaw, C.; Green, L.; Porras-Alfaro, A.; Stursova, M.; Zeglin, L.H. Pulse dynamics and microbial processes in aridland ecosystems. J. Ecol. 2008, 96, 413–420. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Coleine, C.; Selbmann, L.; Ventura, S.; D’Acqui, L.P.; Onofri, S.; Zucconi, L. Fungal Biodiversity in the Alpine Tarfala Valley. Microorganisms 2015, 3, 612-624. https://doi.org/10.3390/microorganisms3040612
Coleine C, Selbmann L, Ventura S, D’Acqui LP, Onofri S, Zucconi L. Fungal Biodiversity in the Alpine Tarfala Valley. Microorganisms. 2015; 3(4):612-624. https://doi.org/10.3390/microorganisms3040612
Chicago/Turabian StyleColeine, Claudia, Laura Selbmann, Stefano Ventura, Luigi Paolo D’Acqui, Silvano Onofri, and Laura Zucconi. 2015. "Fungal Biodiversity in the Alpine Tarfala Valley" Microorganisms 3, no. 4: 612-624. https://doi.org/10.3390/microorganisms3040612
APA StyleColeine, C., Selbmann, L., Ventura, S., D’Acqui, L. P., Onofri, S., & Zucconi, L. (2015). Fungal Biodiversity in the Alpine Tarfala Valley. Microorganisms, 3(4), 612-624. https://doi.org/10.3390/microorganisms3040612










