Proteogenomic Characterization of Pseudomonas veronii SM-20 Growing on Phenanthrene as Only Carbon and Energy Source
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals
2.2. Microorganisms
2.3. Genome Sequencing, Assembly, and Annotation
2.4. Phenotype Microarray Assay
2.5. Phenanthrene Biodegradation Assays
2.6. Comparative Proteomics Analysis of PHE-Exposed SM-20 Cells
3. Results and Discussion
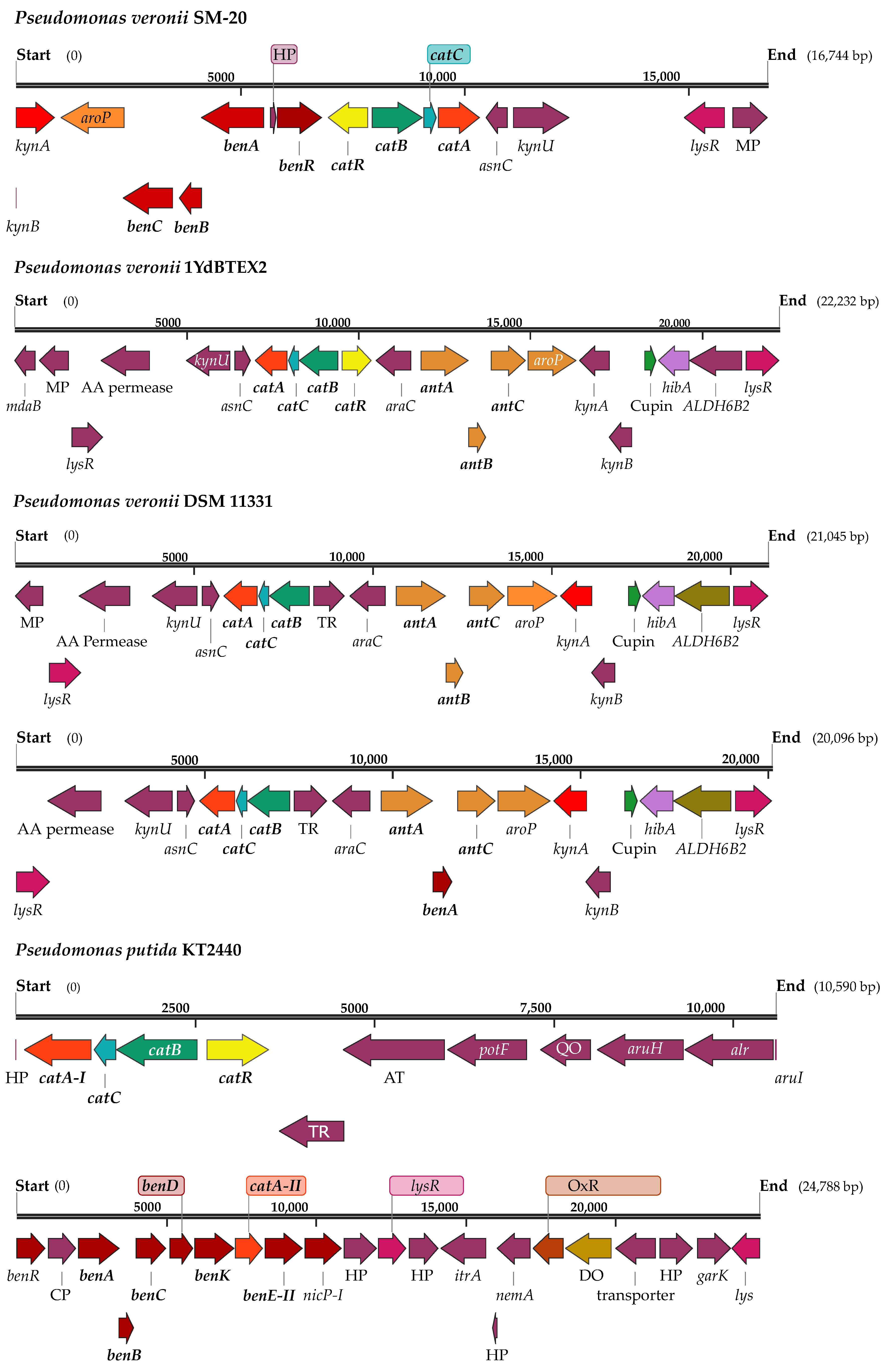
3.1. Phylogenomic and Metabolic Potential of the Newly Isolated Strain SM-20 as Compared with Other Pseudomonas Strains
3.2. Phenotypic Analysis of P. veronii Strain SM-20
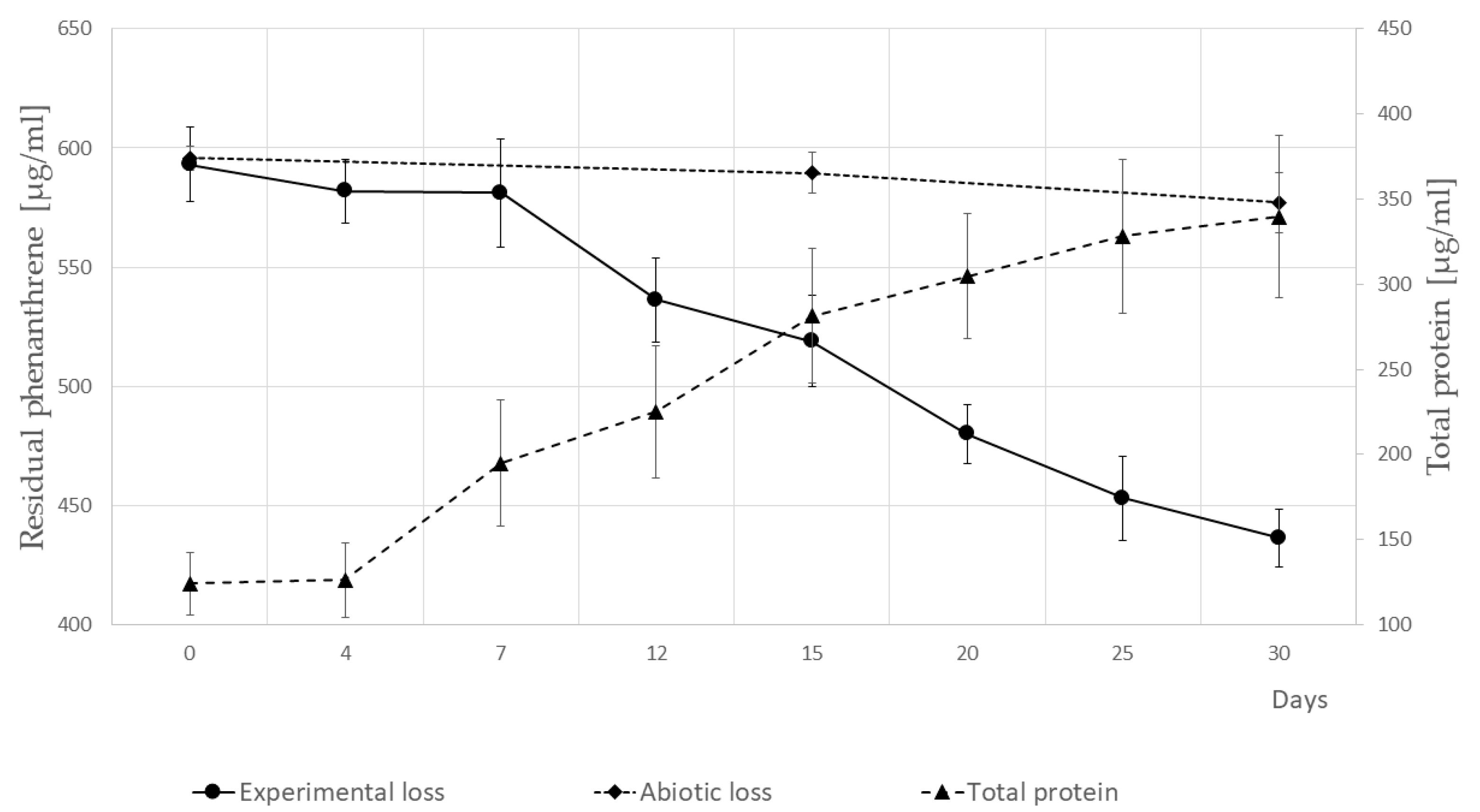
3.3. Growth of P. veronii Isolate SM-20 in the Presence of PHE as the Only Carbon and Energy Source and Identification of PHE Catabolic Intermediates
3.4. Proteomic Profiling of P. veronii SM-20 in Response to PHE Exposure
3.5. Enzymes Involved in Aromatic Hydrocarbon Metabolism and Transportation
3.6. Enzymatic Pathway for PHE Catabolism
3.7. Proteins Involved in Bacterial Surface Remodeling and Stress Response
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Guarino, C.; Zuzolo, D.; Marziano, M.; Conte, B.; Baiamonte, G.; Morra, L.; Benotti, D.; Gresia, D.; Stacul, E.R.; Cicchella, D.; et al. Investigation and assessment for an effective approach to the reclamation of polycyclic aromatic hydrocarbon (PAHs) contaminated site: SIN Bagnoli, Italy. Sci. Rep. 2019, 9, 11522. [Google Scholar] [CrossRef]
- Lu, C.; Hong, Y.; Liu, J.; Gao, Y.; Ma, Z.; Yang, B.; Ling, W.; Waigi, M.G. A PAH-degrading bacterial community enriched with contaminated agricultural soil and its utility for microbial bioremediation. Environ. Pollut. 2019, 251, 773–782. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S.; Kumar, M. Biodegradation of polycyclic aromatic hydrocarbons (PAHs): A sustainable approach. In Sustainable Green Technologies for Environmental Management; Shah, S., Venkatramanan, V.P.R., Eds.; Springer: Singapore, 2019; pp. 111–139. [Google Scholar]
- Ghosal, D.; Ghosh, S.; Dutta, T.K.; Ahn, Y. Current state of knowledge in microbial degradation of polycyclic aromatic hydrocarbons (PAHs): A review. Front. Microbiol. 2016, 7, 208111. [Google Scholar] [CrossRef] [PubMed]
- Michalska, J.; Piński, A.; Zur, J.; Mrozik, A. Analysis of the bioaugmentation potential of Pseudomonas putida OR45a and Pseudomonas putida KB3 in the sequencing batch reactors fed with the phenolic landfill leachate. Water 2020, 12, 906. [Google Scholar] [CrossRef]
- Holmes, D.E.; Dang, Y.; Smith, J.A. Nitrogen cycling during wastewater treatment. Adv. Appl. Microbiol. 2019, 106, 113–192. [Google Scholar] [CrossRef]
- Jun, S.R.; Wassenaar, T.M.; Nookaew, I.; Hauser, L.; Wanchai, V.; Land, M.; Timm, C.M.; Lu, T.Y.S.; Schadt, C.W.; Doktycz, M.J.; et al. Diversity of Pseudomonas genomes, including populus-associated isolates, as revealed by comparative genome analysis. Appl. Environ. Microbiol. 2015, 82, 375–383. [Google Scholar] [CrossRef]
- Zhou, Z.; Liu, Y.; Zanaroli, G.; Wang, Z.; Xu, P.; Tang, H. Enhancing bioremediation potential of Pseudomonas putida by developing its acid stress tolerance with glutamate decarboxylase dependent system and global regulator of extreme radiation resistance. Front. Microbiol. 2019, 10, 2033. [Google Scholar] [CrossRef] [PubMed]
- Anayo, O.F.; Scholastica, E.C.; Peter, O.C.; Nneji, U.G.; Obinna, A.; Mistura, L.O. The beneficial roles of Pseudomonas in medicine, industries, and environment: A review. In Pseudomonas aeruginosa—An Armory Within; Sriramulu, D., Ed.; IntechOpen: London, UK, 2019; pp. 1–13. [Google Scholar] [CrossRef]
- Muriel-Millán, L.F.; Rodríguez-Mejía, J.L.; Godoy-Lozano, E.E.; Rivera-Gómez, N.; Gutierrez-Rios, R.-M.; Morales-Guzmán, D.; Trejo-Hernández, M.R.; Estradas-Romero, A.; Pardo-López, L. Functional and genomic characterization of a Pseudomonas aeruginosa strain isolated from the southwestern gulf of mexico reveals an enhanced adaptation for long-chain alkane degradation. Front. Mar. Sci. 2019, 6, 572. [Google Scholar] [CrossRef]
- Morales, M.; Sentchilo, V.; Bertelli, C.; Komljenovic, A.; Kryuchkova-Mostacci, N.; Bourdilloud, A.; Linke, B.; Goesmann, A.; Harshman, K.; Segers, F.; et al. The genome of the toluene-degrading Pseudomonas veronii strain 1YdBTEX2 and its differential gene expression in contaminated sand. PLoS ONE 2016, 11, e0165850. [Google Scholar] [CrossRef] [PubMed]
- Mullaeva, S.A.; Delegan, Y.A.; Streletskii, R.A.; Sazonova, O.I.; Petrikov, K.V.; Ivanova, A.A.; Dyatlov, I.A.; Shemyakin, I.G.; Bogun, A.G.; Vetrova, A.A. Pseudomonas veronii strain 7-41 degrading medium-chain n-alkanes and polycyclic aromatic hydrocarbons. Sci. Rep. 2022, 12, 20527. [Google Scholar] [CrossRef] [PubMed]
- Havryliuk, O.; Hovorukha, V.; Patrauchan, M.; Youssef, N.H.; Tashyrev, O. Draft whole genome sequence for four highly copper resistant soil isolates Pseudomonas lactis strain UKR1, Pseudomonas panacis strain UKR2, and Pseudomonas veronii strains UKR3 and UKR4. Curr. Res. Microb. Sci. 2020, 1, 44–52. [Google Scholar] [CrossRef] [PubMed]
- Brennerova, M.V.; Zavala-Meneses, S.G.; Josefiova, J.; Branny, P.; Buriankova, K.; Vetrovsky, T.; Junca, H. A global survey reveals a divergent extradiol dioxygenase clade as a widespread complementary contributor to the biodegradation of mono- and polycyclic aromatic hydrocarbons. Environ. Res. 2022, 204, 111954. [Google Scholar] [CrossRef] [PubMed]
- Lee, P.Y.; Costumbrado, J.; Hsu, C.Y.; Kim, Y.H. Agarose gel electrophoresis for the separation of DNA fragments. JoVE 2012, 62, 3923. [Google Scholar] [CrossRef]
- Wick, R.R.; Judd, L.M.; Gorrie, C.L.; Holt, K.E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 2017, 13, e1005595. [Google Scholar] [CrossRef] [PubMed]
- Aziz, R.K.; Bartels, D.; Best, A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef] [PubMed]
- Lobb, B.; Jean-Marie Tremblay, B.; Moreno-Hagelsieb, G.; Doxey, A.C. An assessment of genome annotation coverage across the bacterial tree of life. Microb. Genom. 2020, 6, e000341. [Google Scholar] [CrossRef] [PubMed]
- Meier-Kolthoff, J.P.; Göker, M. TYGS Is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat. Commun. 2019, 10, 2182. [Google Scholar] [CrossRef] [PubMed]
- Checcucci, A.; Borruso, L.; Petrocchi, D.; Perito, B. Diversity and metabolic profile of the microbial communities inhabiting the darkened white marble of Florence cathedral. Int. Biodeterior. Biodegrad. 2022, 171, 105420. [Google Scholar] [CrossRef]
- Galardini, M.; Mengoni, A.; Biondi, E.G.; Semeraro, R.; Florio, A.; Bazzicalupo, M.; Benedetti, A.; Mocali, S. DuctApe: A suite for the analysis and correlation of genomic and OMNILOGTM phenotype microarray data. Genomics 2014, 103, 1–10. [Google Scholar] [CrossRef]
- Masák, J.; Čejková, A.; Jirků, V. Isolation of acetone/ethylene glycol utilizing and biofilm forming strains of bacteria. J. Microbiol. Methods 1997, 30, 133–139. [Google Scholar] [CrossRef]
- Yin, C.; Xiong, W.; Qiu, H.; Peng, W.; Deng, Z.; Lin, S.; Liang, R. Characterization of the phenanthrene-degrading Sphingobium yanoikuyae SJTF8 in heavy metal co-existing liquid medium and analysis of its metabolic pathway. Microorganisms 2020, 8, 946. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Liu, S.; Sun, K.; Sheng, Y.; Gu, Y.; Gao, Y. Colonization on root surface by a phenanthrene-degrading endophytic bacterium and its application for reducing plant phenanthrene contamination. PLoS ONE 2014, 9, e108249. [Google Scholar] [CrossRef] [PubMed]
- Cajthaml, T.; Erbanová, P.; Sasek, V.; Moeder, M. Breakdown products on metabolic pathway of degradation of benz[a]anthracene by a ligninolytic fungus. Chemosphere 2006, 64, 560–564. [Google Scholar] [CrossRef] [PubMed]
- Nováková, S.; Šubr, Z.; Kováč, A.; Fialová, I.; Beke, G.; Danchenko, M. Cucumber mosaic virus resistance: Comparative proteomics of contrasting Cucumis sativus cultivars after long-term infection. J. Proteom. 2020, 214, 103626. [Google Scholar] [CrossRef] [PubMed]
- Goedhart, J.; Luijsterburg, M.S. VolcaNoseR is a web app for creating, exploring, labeling and sharing volcano plots. Sci. Rep. 2020, 10, 20560. [Google Scholar] [CrossRef] [PubMed]
- Yu, N.Y.; Wagner, J.R.; Laird, M.R.; Melli, G.; Rey, S.; Lo, R.; Dao, P.; Cenk Sahinalp, S.; Ester, M.; Foster, L.J.; et al. PSORTb 3.0: Improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 2010, 26, 1608. [Google Scholar] [CrossRef] [PubMed]
- Imai, K.; Asakawa, N.; Tsuji, T.; Akazawa, F.; Ino, A.; Sonoyama, M.; Mitaku, S. SOSUI-GramN: High performance prediction for sub-cellular localization of proteins in gram-negative bacteria. Bioinformation 2008, 2, 417. [Google Scholar] [CrossRef]
- Cantalapiedra, C.P.; Hernández-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. EggNOG-Mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef] [PubMed]
- Imperato, V.; Portillo-Estrada, M.; McAmmond, B.M.; Douwen, Y.; Van Hamme, J.D.; Gawronski, S.W.; Vangronsveld, J.; Thijs, S. Genomic diversity of two hydrocarbon-degrading and plant growth-promoting Pseudomonas species isolated from the oil field of Bóbrka (Poland). Genes 2019, 10, 443. [Google Scholar] [CrossRef]
- Montes, C.; Altimira, F.; Canchignia, H.; Castro, Á.; Sánchez, E.; Miccono, M.; Tapia, E.; Sequeida, Á.; Valdés, J.; Tapia, P.; et al. A draft genome sequence of Pseudomonas veronii R4: A grapevine (Vitis vinifera L.) root-associated strain with high biocontrol potential. Stand. Genom. Sci. 2016, 11, 1–10. [Google Scholar] [CrossRef]
- Valderrama, J.A.; Durante-Rodríguez, G.; Blázquez, B.; García, J.L.; Carmona, M.; Díaz, E. Bacterial degradation of benzoate: Cross-regulation between aerobic and anaerobic pathways. J. Biol. Chem. 2012, 287, 10494–10508. [Google Scholar] [CrossRef] [PubMed]
- Phale, P.S.; Malhotra, H.; Shah, B.A. Degradation strategies and associated regulatory mechanisms/features for aromatic compound metabolism in bacteria. Adv. Appl. Microbiol. 2020, 112, 1–65. [Google Scholar] [CrossRef] [PubMed]
- Belda, E.; van Heck, R.G.A.; José Lopez-Sanchez, M.; Cruveiller, S.; Barbe, V.; Fraser, C.; Klenk, H.P.; Petersen, J.; Morgat, A.; Nikel, P.I.; et al. The revisited genome of Pseudomonas putida KT2440 enlightens its value as a robust metabolic chassis. Environ. Microbiol. 2016, 18, 3403–3424. [Google Scholar] [CrossRef] [PubMed]
- Elomari, M.; Coroler, L.; Hoste, B.; Gillis, M.; Izard, D.; Leclerc, H. DNA relatedness among Pseudomonas strains isolated from natural mineral waters and proposal of Pseudomonas veronii sp. nov. Int. J. Syst. Bacteriol. 1996, 46, 1138–1144. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Lu, C.D. The cryptic dsda gene encodes a functional D-serine dehydratase in Pseudomonas aeruginosa PAO1. Curr. Microbiol. 2016, 72, 788–794. [Google Scholar] [CrossRef] [PubMed]
- Remus-Emsermann, M.N.P.; Schmid, M.; Gekenidis, M.T.; Pelludat, C.; Frey, J.E.; Ahrens, C.H.; Drissner, D. Complete genome sequence of Pseudomonas citronellolis P3B5, a candidate for microbial phyllo-remediation of hydrocarbon-contaminated sites. Stand. Genom. Sci. 2016, 11, 75. [Google Scholar] [CrossRef] [PubMed]
- Saati-Santamaría, Z.; Baroncelli, R.; Rivas, R.; García-Fraile, P. Comparative genomics of the genus Pseudomonas reveals host- and environment-specific evolution. Microbiol. Spectr. 2022, 10, e0237022. [Google Scholar] [CrossRef]
- Ramsey, C.; MacGowan, A.P. A review of the pharmacokinetics and pharmacodynamics of aztreonam. J. Antimicrob. Chemother. 2016, 71, 2704–2712. [Google Scholar] [CrossRef]
- Alav, I.; Kobylka, J.; Kuth, M.S.; Pos, K.M.; Picard, M.; Blair, J.M.A.; Bavro, V.N. Structure, assembly, and function of tripartite efflux and type 1 secretion systems in gram-negative bacteria. Chem. Rev. 2021, 121, 5479–5596. [Google Scholar] [CrossRef]
- Nag, A.; Mehra, S. A major facilitator superfamily (MFS) efflux pump, SCO4121, from Streptomyces coelicolor with roles in multidrug resistance and oxidative stress tolerance and its regulation by a MARR regulator. Appl. Environ. Microbiol. 2021, 87, e02238-20. [Google Scholar] [CrossRef]
- Pasqua, M.; Grossi, M.; Scinicariello, S.; Aussel, L.; Barras, F.; Colonna, B.; Prosseda, G. The MFS efflux pump EmrKY contributes to the survival of Shigella within macrophages. Sci. Rep. 2019, 9, 2906. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Huang, H.; Wu, M.; Yu, X.; Chen, Y.; Liu, P.; Li, X. Pseudomonas sp. LZ-Q continuously degrades phenanthrene under hypersaline and hyperalkaline condition in a membrane bioreactor system. Biophys. Rep. 2015, 1, 156–167. [Google Scholar] [CrossRef] [PubMed]
- Bisht, S.; Pandey, P.; Bhargava, B.; Sharma, S.; Kumar, V.; Sharma, K.D. Bioremediation of polyaromatic hydrocarbons (PAHs) using rhizosphere technology. Braz. J. Microbiol. 2015, 46, 7. [Google Scholar] [CrossRef] [PubMed]
- Puntus, I.F.; Filonov, A.E.; Akhmetov, L.I.; Karpov, A.V.; Boronin, A.M. Phenanthrene degradation by bacteria of the genera Pseudomonas and Burkholderia in model soil systems. Mikrobiologiia 2008, 77, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Isaac, P.; Martínez, F.L.; Bourguignon, N.; Sánchez, L.A.; Ferrero, M.A. Improved PAHs removal performance by a defined bacterial consortium of indigenous Pseudomonas and Actinobacteria from Patagonia, Argentina. Int. Biodeterior. Biodegrad. 2015, 101, 23–31. [Google Scholar] [CrossRef]
- Zhao, H.P.; Wu, Q.S.; Wang, L.; Zhao, X.T.; Gao, H.W. Degradation of phenanthrene by bacterial strain isolated from soil in oil refinery fields in Shanghai China. J. Hazard. Mater. 2009, 164, 863–869. [Google Scholar] [CrossRef] [PubMed]
- Mawad, A.; Albasri, H.; Shalkami, A.G.; Alamri, S.; Hashem, M. Synergistic degradation of phenanthrene by constructed Pseudomonas spp. consortium compared with pure strains. Environ. Technol. Innov. 2021, 24, 101942. [Google Scholar] [CrossRef]
- Nwinyi, O.C.; Ajayi, O.O.; Amund, O.O. Degradation of polynuclear aromatic hydrocarbons by two strains of Pseudomonas. Braz. J. Microbiol. 2016, 47, 551. [Google Scholar] [CrossRef] [PubMed]
- Volkering, F.; Breure, A.M.; Sterkenburg, A.; van Andel, J.G. Microbial degradation of polycyclic aromatic hydrocarbons: Effect of substrate availability on bacterial growth kinetics. Appl. Microbiol. Biotechnol. 1992, 36, 548–552. [Google Scholar] [CrossRef]
- Hua, F.; Wang, H.Q. Uptake and trans-membrane transport of petroleum hydrocarbons by microorganisms. Biotechnol. Biotechnol. Equip. 2014, 28, 165–175. [Google Scholar] [CrossRef]
- Pozdnyakova, N.; Muratova, A.; Turkovskaya, O. Degradation of polycyclic aromatic hydrocarbons by co-culture of Pleurotus ostreatus Florida and Azospirillum brasilense. Appl. Microbiol. 2022, 2, 735–748. [Google Scholar] [CrossRef]
- Hadibarata, T.; Tachibana, S.; Askari, M. Identification of metabolites from phenanthrene oxidation by phenoloxidases and dioxygenases of Polyporus sp. S133. J. Microbiol. Biotechnol. 2011, 21, 299–304. [Google Scholar] [CrossRef] [PubMed]
- Hidayat, A.; Yanto, D.H.Y. Biodegradation and metabolic pathway of phenanthrene by a new tropical fungus, Trametes hirsuta D7. J. Environ. Chem. Eng. 2018, 6, 2454–2460. [Google Scholar] [CrossRef]
- Hammel, K.; Gai, W.; Green, B.; Moen, M. Oxidative degradation of phenanthrene by the ligninolytic fungus Phanerochaete chrysosporium. Appl. Environ. Microbiol. 1992, 58, 1832–1838. [Google Scholar] [CrossRef] [PubMed]
- Teng, C.; Wu, S.; Gong, G. Bio-removal of phenanthrene, 9-fluorenone and anthracene-9,10-dione by laccase from Aspergillus niger in waste cooking oils. Food Control 2019, 105, 219–225. [Google Scholar] [CrossRef]
- Muratova, A.; Pozdnyakova, N.; Makarov, O.; Baboshin, M.; Baskunov, B.; Myasoedova, N.; Golovleva, L.; Turkovskaya, O. Degradation of phenanthrene by the rhizobacterium Ensifer meliloti. Biodegradation 2014, 25, 787–795. [Google Scholar] [CrossRef]
- Kim, Y.H.; Freeman, J.P.; Moody, J.D.; Engesser, K.H.; Cerniglia, C.E. Effects of pH on the degradation of phenanthrene and pyrene by Mycobacterium vanbaalenii PYR-1. Appl. Microbiol. Biotechnol. 2005, 67, 275–285. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.-S.; Keum, Y.-S.; Li, Q.X. Mycobacterium aromativorans JS19b1T degrades phenanthrene through C-1,2, C-3,4 and C-9,10 dioxygenation pathways. Int. Biodeterior. Biodegrad. 2012, 70, 96. [Google Scholar] [CrossRef] [PubMed]
- Barone, R.; Nastro, R.A.; Gambino, E.; Toscanesi, M.; Picciall, G.; Napoli, L.D.; Trifuoggi, M.; Piccialli, V.; Guida, M. Pseudomonas anguilliseptica strain-A1 degradation of polycyclic aromatic hydrocarbons in soil microcosms: Focus on detoxification activity and free water-soluble protein extracts kinetics and efficiency. J. Bioremediat. Biodegrad. 2017, 8, 1–7. [Google Scholar] [CrossRef]
- Guzik, U.; Greń, I.; Hupert-Kocurek, K.; Wojcieszyńska, D. Catechol 1,2-dioxygenase from the new aromatic compounds–degrading Pseudomonas putida strain n6. Int. Biodeterior. Biodegrad. 2011, 65, 504–512. [Google Scholar] [CrossRef]
- Borowski, T.; Georgiev, V.; Siegbahn, P.E.M. Catalytic reaction mechanism of homogentisate dioxygenase: A hybrid DFT study. J. Am. Chem. Soc. 2005, 127, 17303–17314. [Google Scholar] [CrossRef] [PubMed]
- Setlhare, B.; Kumar, A.; Mokoena, M.P.; Olaniran, A.O. Catechol 1,2-dioxygenase is an analogue of homogentisate 1,2-dioxygenase in Pseudomonas chlororaphis Strain UFB2. Int. J. Mol. Sci. 2019, 20, 61. [Google Scholar] [CrossRef] [PubMed]
- Kamimura, N.; Masai, E. The protocatechuate 4,5-cleavage pathway: Overview and new findings. In Biodegradative Bacteria; Nojiri, H., Tsuda, M., Fukuda, M., Kamagata, Y., Eds.; Springer: Tokyo, Japan, 2013; pp. 207–226. [Google Scholar] [CrossRef]
- Guzik, U.; Hupert-Kocurek, K.; Wojcieszysk, D. Intradiol dioxygenases—The key enzymes in xenobiotics degradation. In Biodegradation of Hazardous and Special Products; Chamy, R., Rosenkranz, F., Eds.; IntechOpen: London, UK, 2013. [Google Scholar] [CrossRef]
- Ladino-Orjuela, G.; Gomes, E.; da Silva, R.; Salt, C.; Parsons, J.R. Metabolic pathways for degradation of aromatic hydrocarbons by bacteria. Rev. Environ. Contam. Toxicol. 2016, 237, 105–121. [Google Scholar] [CrossRef] [PubMed]
- Cafaro, V.; Izzo, V.; Scognamiglio, R.; Notomista, E.; Capasso, P.; Casbarra, A.; Pucci, P.; Di Donato, A. Phenol hydroxylase and toluene/o-xylene monooxygenase from Pseudomonas stutzeri OX1: Interplay between two enzymes. Appl. Environ. Microbiol. 2004, 70, 2211–2219. [Google Scholar] [CrossRef] [PubMed]
- Robrock, K.R.; Mohn, W.W.; Eltis, L.D.; Alvarez-Cohen, L. Biphenyl and ethylbenzene dioxygenases of Rhodococcus jostii RHA1 transform PBDEs. Biotechnol. Bioeng. 2011, 108, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Xie, F.; Zhang, G.; Shi, Y.; Qian, S. Purification, characterization, and substrate specificity of two 2,3-dihydroxybiphenyl 1,2-dioxygenase from Rhodococcus sp. R04, showing their distinct stability at various temperature. Biochimie 2008, 90, 1530–1538. [Google Scholar] [CrossRef] [PubMed]
- Perez-Dominguez, F.; Carrillo-Beltrán, D.; Blanco, R.; Muñoz, J.P.; León-Cruz, G.; Corvalan, A.H.; Urzúa, U.; Calaf, G.M.; Aguayo, F. Role of pirin, an oxidative stress sensor protein, in epithelial carcinogenesis. Biology 2021, 10, 116. [Google Scholar] [CrossRef] [PubMed]
- Widiatningrum, T.; Maeda, S.; Kataoka, K.; Sakurai, T. A pirin-like protein from Pseudomonas stutzeri and its quercetinase activity. Biochem. Biophys. Rep. 2015, 3, 144. [Google Scholar] [CrossRef] [PubMed]
- Soo, P.C.; Horng, Y.T.; Lai, M.J.; Wei, J.R.; Hsieh, S.C.; Chang, Y.L.; Tsai, Y.H.; Lai, H.C. Pirin regulates pyruvate catabolism by interacting with the pyruvate dehydrogenase E1 subunit and modulating pyruvate dehydrogenase activity. J. Bacteriol. 2007, 189, 109–118. [Google Scholar] [CrossRef]
- Hickman, S.J.; Cooper, R.E.M.; Bellucci, L.; Paci, E.; Brockwell, D.J. Gating of TonB-dependent transporters by substrate-specific forced remodelling. Nat. Commun. 2017, 8, 14804. [Google Scholar] [CrossRef]
- Akhtar, A.A.; Turner, D.P.J. The role of bacterial ATP-binding cassette (ABC) transporters in pathogenesis and virulence: Therapeutic and vaccine potential. Microb. Pathog. 2022, 171, 105734. [Google Scholar] [CrossRef] [PubMed]
- Fujita, M.; Mori, K.; Hara, H.; Hishiyama, S.; Kamimura, N.; Masai, E. A TonB-dependent receptor constitutes the outer membrane transport system for a lignin-derived aromatic compound. Commun. Biol. 2019, 2, 432. [Google Scholar] [CrossRef]
- Samantarrai, D.; Sagar, A.L.; Gudla, R.; Siddavattam, D. TonB-dependent transporters in Sphingomonads: Unraveling their distribution and function in environmental adaptation. Microorganisms 2020, 8, 359. [Google Scholar] [CrossRef] [PubMed]
- Tang, K.; Jiao, N.; Liu, K.; Zhang, Y.; Li, S. Distribution and functions of TonB-dependent transporters in marine bacteria and environments: Implications for dissolved organic matter utilization. PLoS ONE 2012, 7, e41204. [Google Scholar] [CrossRef] [PubMed]
- Michalska, K.; Chang, C.; Mack, J.C.; Zerbs, S.; Joachimiak, A.; Collart, F.R. characterization of transport proteins for aromatic compounds derived from lignin: Benzoate derivative binding proteins. J. Mol. Biol. 2012, 423, 555–575. [Google Scholar] [CrossRef] [PubMed]
- Jerina, D.M. The 1982 Bernard B. Brodie award lecture. Metabolism of aromatic hydrocarbons by the cytochrome P-450 system and epoxide hydrolase. Drug Metab. Dispos. 1983, 11, 1–4. [Google Scholar] [PubMed]
- Zeinali, M.; Vossoughi, M.; Ardestani, S.K. Degradation of phenanthrene and anthracene by Nocardia otitidiscaviarum strain TSH1, a moderately thermophilic bacterium. J. Appl. Microbiol. 2008, 105, 398–406. [Google Scholar] [CrossRef] [PubMed]
- Haroune, N.; Combourieu, B.; Besse, P.; Sancelme, M.; Reemtsma, T.; Kloepfer, A.; Diab, A.; Knapp, J.S.; Baumberg, S.; Delort, A.M. Benzothiazole degradation by Rhodococcus pyridinovorans strain PA: Evidence of a catechol 1,2-dioxygenase activity. Appl. Environ. Microbiol. 2002, 68, 6114–6120. [Google Scholar] [CrossRef] [PubMed]
- Meulenberg, R.; Rijnaarts, H.H.M.; Doddema, H.J.; Field, J.A. Partially oxidized polycyclic aromatic hydrocarbons show an increased bioavailability and biodegradability. FEMS Microbiol. Lett. 1997, 152, 45–49. [Google Scholar] [CrossRef]
- Suman, J.; Sredlova, K.; Fraraccio, S.; Jerabkova, M.; Strejcek, M.; Kabickova, H.; Cajthaml, T.; Uhlik, O. Transformation of hydroxylated polychlorinated biphenyls by bacterial 2-hydroxybiphenyl 3-monooxygenase. Chemosphere 2024, 349, 140909. [Google Scholar] [CrossRef]
- Setlhare, B.; Kumar, A.; Mokoena, M.P.; Pillay, B.; Olaniran, A.O. Phenol hydroxylase from Pseudomonas sp. KZNSA: Purification, characterization and prediction of three-dimensional structure. Int. J. Biol. Macromol. 2020, 146, 1000–1008. [Google Scholar] [CrossRef] [PubMed]
- Marx, D.C.; Plummer, A.M.; Faustino, A.M.; Devlin, T.; Roskopf, M.A.; Leblanc, M.J.; Lessen, H.J.; Amann, B.T.; Fleming, P.J.; Krueger, S.; et al. SurA is a cryptically grooved chaperone that expands unfolded outer membrane proteins. Proc. Natl. Acad. Sci. USA 2020, 117, 28026–28035. [Google Scholar] [CrossRef] [PubMed]
- Bugg, T.; Foght, J.M.; Pickard, M.A.; Gray, M.R. Uptake and active efflux of polycyclic aromatic hydrocarbons by Pseudomonas fluorescens LP6a. Appl. Environ. Microbiol. 2000, 66, 5387. [Google Scholar] [CrossRef] [PubMed]
- Broniatowski, M.; Binczycka, M.; Wójcik, A.; Flasiński, M.; Wydro, P. Polycyclic aromatic hydrocarbons in model bacterial membranes–Langmuir monolayer studies. Biochim. Biophys. Acta-Biomembr. 2017, 1859, 2402–2412. [Google Scholar] [CrossRef] [PubMed]
- Scott, C.C.; Finnerty, W.R. Characterization of intracytoplasmic hydrocarbon inclusions from the hydrocarbon-oxidizing Acinetobacter species HO1-N. J. Bacteriol. 1976, 127, 481. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Jiang, R.; Kong, D.; Liu, Z.; Wu, X.; Xu, J.; Li, Y. Transmembrane transport of polycyclic aromatic hydrocarbons by bacteria and functional regulation of membrane proteins. Front. Environ. Sci. Eng. 2020, 14, 9. [Google Scholar] [CrossRef]
- Misra, H.S.; Rajpurohit, Y.S.; Khairnar, N.P. Pyrroloquinoline-quinone and its versatile roles in biological processes. J. Biosci. 2012, 37, 313–325. [Google Scholar] [CrossRef] [PubMed]
- Poole, K.; Neshat, S.; Krebes, K.; Heinrichs, D.E. Cloning and nucleotide sequence analysis of the ferripyoverdine receptor gene FpvA of Pseudomonas aeruginosa. J. Bacteriol. 1993, 175, 4597. [Google Scholar] [CrossRef] [PubMed]
- Ghysels, B.; Dieu, B.T.M.; Beatson, S.A.; Pirnay, J.P.; Ochsner, U.A.; Vasil, M.L.; Cornelis, P. FpvB, an alternative type i ferripyoverdine receptor of Pseudomonas aeruginosa. Microbiology 2004, 150, 1671–1680. [Google Scholar] [CrossRef]
- Holden, V.I.; Breen, P.; Houle, S.; Dozois, C.M.; Bachman, M.A. Klebsiella pneumoniae siderophores induce inflammation, bacterial dissemination, and HIF-1ɑ stabilization during pneumonia. mBio 2016, 7, e01397-16. [Google Scholar] [CrossRef]

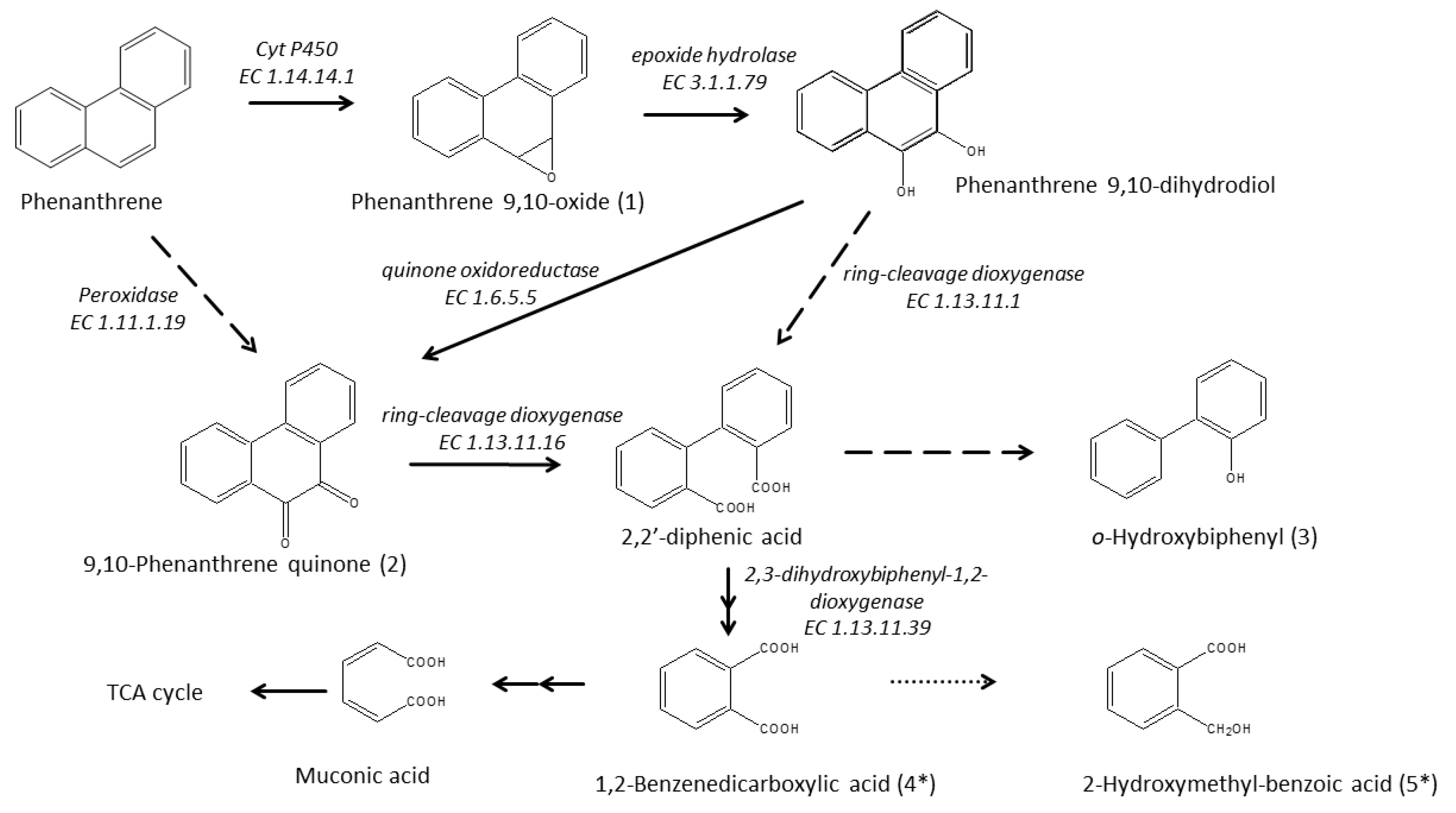
| Product No. | tR [min] | MW | m/z of Fragment Ions (Relative Intensity in %) | Structural Suggestion |
|---|---|---|---|---|
| 1 | 13.1 | 194 | 194 (22.6), 178 (7.5), 165 (99.9), 139 (4.1), 115 (2.2) | Phenanthrene 9,10-oxide |
| 2 | 12.49 | 208 | 208 (99.9), 180 (82.3), 152 (59.2), 126 (9.8), 76 (19.4) | 9,10-Phenanthrene quinone |
| 3 | 7.06 | 170 | 170 (99.9), 141 (31.7), 115 (22.4), 89 (3.8), 63 (4.1) | o-Hydroxybiphenyl |
| 4 | 6.19 | 166 | 148 (26.2), 104 (99.9), 76 (63.9), 50 (38.0) | Phthalic anhydride |
| 5 | 5.44 | 152 | 134 (99.9), 106 (44.4), 78 (90.1), 51 (14.6) | 2-Coumaranone |
| Accession ID | Gene ID | EC Number | Name |
|---|---|---|---|
| Oxidoreductases | |||
| A0A1D3JSA5 | peg.2743 | 1.13.11.5 | Homogentisate 1,2-dioxygenase |
| A0A1D3JXN6 | peg.644 | 1.13.11.27 | 4-hydroxyphenylpyruvate dioxygenase |
| A0A1D3K3P0 | peg.4964 | 1.14.12.1 | Anthranilate 1,2-dioxygenase small subunit |
| A0A1D3K3R6 | peg.4971 | 1.13.11.1 | Catechol 1,2-dioxygenase |
| A0A1D3JYJ0 | peg.977 | 1.2.1.85 | 2-hydroxymuconic semialdehyde dehydrogenase |
| A0A1D3K906 | peg.984,3084,3088 | 1.13.11.2 | CatO2ase |
| A0A1D3K930 | peg.4843 | 1.14.12.18 | Isopropylbenzene dioxygenase iron-sulfur protein small subunit |
| A0A1D3JYR3 | peg.1053 | 1.14.12.19 | 3-phenylpropionate dioxygenase |
| A0A1D3K1V5 | peg.2936 | 1.13.11.39 | Extradiol dioxygenase |
| A0A1D3JYZ4 | peg.979 | 1.14.13.7 | Phenol hydroxylase |
| A0A1D3JYN7 | peg.982 | 1.14.13.7 | Phenol hydroxylase P1 protein |
| A0A1D3JWX5 | peg.342 | 1.3.5.1 | Acetoin:2,6-dichlorophenolindophenol oxidoreductase subunit alpha |
| A0A1D3JX27 | peg.343 | 1.3.5.1 | Acetoin:2,6-dichlorophenolindophenol oxidoreductase subunit beta |
| A0A1D3JRD8 | peg.4571 | 1.14.14.1 | Monooxygenase |
| A0A1D3JTA7 | 1.13.11.3 | Protocatechuate 3, 4-dioxygenase subunit alpha | |
| A0A1D3JSV4 | peg.2577 | 1.18.1.2 | Ferredoxin--NADP reductase |
| A0A1D3JVD3 | peg.6254,4787 | 1.2.1.3 | Aldehyde dehydrogenase |
| A0A1D3K5B8 | peg.3524 | 1.2.1.3 | Aldehyde dehydrogenase |
| A0A1D3JYP8 | peg.1022, 3705 | 1.2.1.3 | Aldehyde dehydrogenase PuuC |
| A0A1D3K973 | peg.1047,4821 | 1.2.1.10 | Acetaldehyde dehydrogenase |
| A0A1D3K0Y9 | peg.1742 | 1.6.5.5 | Quinone oxidoreductase |
| A0A1D3JVV0 | peg.6390 | 7.1.1.- | NADH-quinone oxidoreductase |
| A0A1D3JVY4 | peg. 6394 | 7.1.1.- | NADH-quinone oxidoreductase subunit B |
| A0A1D3JVV9 | peg.6393 | 7.1.1.- | NADH-quinone oxidoreductase subunit C/D |
| A0A1D3JVX6 | peg.6391 | 7.1.1.2 | NADH-quinone oxidoreductase subunit F |
| A0A1D3JW33 | peg.6376 | 1.11.1.19 | DyP type peroxidase |
| A0A1D3JPQ1 | peg.3915 | 1.14.14.1 | Cytochrome P450 |
| A0A1D3JYG5 | peg.937 | 1.13.11.24 | Quercetin 2,3-dioxygenase |
| A0A1D3K115 | peg.1907 | 1.13.11.24 | Pirin |
| A0A1D3JZS8 | peg.1440 | 1.13.11.16 | 2,3-dihydroxyphenyl propionate/2,3-dihydroxicinnamic acid 1,2-dioxygenase |
| A0A1D3K298 | peg.5553 | 1.1.1.100 | 3-oxoacyl-[acyl-c arrier-protein] reductase |
| A0A1D3JWU0 | peg.304 | Oxidoreductase | |
| A0A1D3K7I1 | peg.3568 | 1.14.11.- | Fe2OG dioxygenase domain-containing protein |
| Lyases | |||
| A0A1D3K2B2 | peg.5665 | 4.1.3.17 | 4-hydroxy-4-methyl-2-oxoglutarate aldolase |
| A0A1D3K4Q4 | peg.3296 | 4.1.3.27 | Anthranilate synthase component 1 |
| A0A1D3K4Q5 | peg.3295 | 4.1.3.27 | Anthranilate synthase component 2 |
| A0A1D3K908 | peg.1046,1445,4823 | 4.1.3.39 | 4-hydroxy-2-oxovalerate aldolase |
| A0A1D3JYJ6 | peg.976,3081 | 4.2.1.132 | 2-hydroxyhexa-2,4-dienoate hydratase |
| A0A1D3K0Q9 | peg.1455 | 4.1.2.61 | Hydroxycinnamoyl-CoA hydratase-lyase |
| Isomerases | |||
| A0A1D3JYR4 | peg.970 | 5.3.2.6 | 2-hydroxymuconate tautomerase |
| A0A1D3K3L8 | peg.4970 | 5.3.3.4 | Muconolactone Delta-isomerase |
| A0A1D3JX36 | peg.428 | 5.5.1.5 | 3-carboxymuconate cyclase |
| Transferases | |||
| A0A1D3K077 | peg.1572 | 2.8.3.6 | 3-oxoadipate CoA-transferase |
| A0A1D3K0A2 | peg.1573 | 2.8.3.6 | 3-oxoadipate CoA-transferase |
| Hydrolases | |||
| A0A1D3JTI3 | peg.2407 | 3.1.1.24 | 3-oxoadipate enol-lactonase |
| A0A1D3JXH6 | peg.538 | 3.1.1.79 | Alpha/beta hydrolase |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zavala-Meneses, S.G.; Firrincieli, A.; Chalova, P.; Pajer, P.; Checcucci, A.; Skultety, L.; Cappelletti, M. Proteogenomic Characterization of Pseudomonas veronii SM-20 Growing on Phenanthrene as Only Carbon and Energy Source. Microorganisms 2024, 12, 753. https://doi.org/10.3390/microorganisms12040753
Zavala-Meneses SG, Firrincieli A, Chalova P, Pajer P, Checcucci A, Skultety L, Cappelletti M. Proteogenomic Characterization of Pseudomonas veronii SM-20 Growing on Phenanthrene as Only Carbon and Energy Source. Microorganisms. 2024; 12(4):753. https://doi.org/10.3390/microorganisms12040753
Chicago/Turabian StyleZavala-Meneses, Sofía G., Andrea Firrincieli, Petra Chalova, Petr Pajer, Alice Checcucci, Ludovit Skultety, and Martina Cappelletti. 2024. "Proteogenomic Characterization of Pseudomonas veronii SM-20 Growing on Phenanthrene as Only Carbon and Energy Source" Microorganisms 12, no. 4: 753. https://doi.org/10.3390/microorganisms12040753
APA StyleZavala-Meneses, S. G., Firrincieli, A., Chalova, P., Pajer, P., Checcucci, A., Skultety, L., & Cappelletti, M. (2024). Proteogenomic Characterization of Pseudomonas veronii SM-20 Growing on Phenanthrene as Only Carbon and Energy Source. Microorganisms, 12(4), 753. https://doi.org/10.3390/microorganisms12040753









