Examination on the Occurrence of Coinfections in Diagnostic Transmittals in Cases of Stillbirth, Mummification, Embryonic Death, and Infertility (SMEDI) Syndrome in Germany
Abstract
1. Introduction
2. Material and Methods
2.1. Sample Material
2.2. Sampling
2.3. Sample Collection
2.4. Molecular Biological Examinations
2.4.1. Viral Pathogens
2.4.2. Leptospira spp.
2.5. Statistical Analysis
3. Results
3.1. Study Population
3.2. Molecular Biological Examinations
3.2.1. Detection of Single Pathogens
3.2.2. Detection of (Co)infections in Terms of SMEDI
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Sanchez, R.E., Jr.; Nauwynck, H.J.; McNeilly, F.; Allan, G.M.; Pensaert, M.B. Porcine circovirus 2 infection in swine foetuses inoculated at different stages of gestation. Vet. Microbiol. 2001, 83, 169–176. [Google Scholar] [CrossRef] [PubMed]
- Streck, A.F.; Truyen, U. Porcine Parvovirus. Curr. Issues Mol. Biol. 2020, 37, 33–46. [Google Scholar] [CrossRef] [PubMed]
- Vereecke, N.; Kvisgaard, L.K.; Baele, G.; Boone, C.; Kunze, M.; Larsen, L.E.; Theuns, S.; Nauwynck, H. Molecular epidemiology of Porcine Parvovirus Type 1 (PPV1) and the reactivity of vaccine-induced antisera against historical and current PPV1 strains. Virus Evol. 2022, 8, veac053. [Google Scholar] [CrossRef] [PubMed]
- Reif, J.; Renzhammer, R.; Brunthaler, R.; Weissenbacher-Lang, C.; Auer, A.; Kreutzmann, H.; Fux, R.; Ladinig, A.; Unterweger, C. Reproductive failure in an Austrian piglet-producing farm due to porcine circovirus genotype 2d. Acta Vet. Hung. 2022, 70, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Sydler, T.; Brugnera, E.; Weilenmann, R.; Zimmermann, D.; Engels, M.; Sidler, X. First description of PCV-2-induced SMEDI-syndrome in Switzerland. Tierarztl. Prax. Ausg. G Grosstiere Nutztiere 2011, 39, 231–236. [Google Scholar] [CrossRef][Green Version]
- Rose, N.; Blanchard, P.; Cariolet, R.; Grasland, B.; Amenna, N.; Oger, A.; Durand, B.; Balasch, M.; Jestin, A.; Madec, F. Vaccination of porcine circovirus type 2 (PCV2)-infected sows against porcine Parvovirus (PPV) and Erysipelas: Effect on post-weaning multisystemic wasting syndrome (PMWS) and on PCV2 genome load in the offspring. J. Comp. Pathol. 2007, 136, 133–144. [Google Scholar] [CrossRef]
- Madson, D.M.; Patterson, A.R.; Ramamoorthy, S.; Pal, N.; Meng, X.J.; Opriessnig, T. Reproductive failure experimentally induced in sows via artificial insemination with semen spiked with porcine circovirus type 2. Vet. Pathol. 2009, 46, 707–716. [Google Scholar] [CrossRef]
- Truyen, U.; Streck, A.F. Parvoviruses. In Diseases of Swine; John Wiley & Sons: Hoboken, NJ, USA, 2019; pp. 611–621. [Google Scholar]
- Pozzi, P.; Alborali, G.L.; Etinger, M.; Hadani, Y. Epidemiological Investigation of the Prevalence of Leptospira spp. in Pigs in Israel. Isr. J. Vet. Med. 2020, 75, 72–79. [Google Scholar]
- Handke, M.; Engels, M.; Prohaska, S.; Keller, C.; Brugnera, E.; Sydler, T.; Sidler, X. Infection related fertility disorders in Swiss pig breeding farms at the end of the postweaning multisystemic wasting syndrome (PMWS) epizooty. Schweiz. Arch. Tierheilkde 2012, 154, 437–444. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Oropeza-Moe, M.; Oropeza Delgado, A.J.; Framstad, T. Porcine circovirus type 2 associated reproductive failure in a specific pathogen free (SPF) piglet producing herd in Norway: A case report. Porcine Health Manag. 2017, 3, 25. [Google Scholar] [CrossRef]
- Pittman, J.S. Reproductive failure associated with porcine circovirus type 2 in gilts. J Swine Health Prod 2008, 16, 144–148. [Google Scholar]
- O’Connor, B.; Gauvreau, H.; West, K.; Bogdan, J.; Ayroud, M.; Clark, E.G.; Konoby, C.; Allan, G.; Ellis, J.A. Multiple porcine circovirus 2-associated abortions and reproductive failure in a multisite swine production unit. Can. Vet. J. 2001, 42, 551–553. [Google Scholar]
- Segales, J. Porcine circovirus type 2 (PCV2) infections: Clinical signs, pathology and laboratory diagnosis. Virus Res. 2012, 164, 10–19. [Google Scholar] [CrossRef]
- Strutzberg-Minder, K.; Tschentscher, A.; Beyerbach, M.; Homuth, M.; Kreienbrock, L. Passive surveillance of Leptospira infection in swine in Germany. Porcine Health Manag. 2018, 4, 10. [Google Scholar] [CrossRef] [PubMed]
- Tochetto, C.; Lima, D.A.; Varela, A.P.M.; Loiko, M.R.; Paim, W.P.; Scheffer, C.M.; Herpich, J.I.; Cerva, C.; Schmitd, C.; Cibulski, S.P.; et al. Full-Genome Sequence of Porcine Circovirus type 3 recovered from serum of sows with stillbirths in Brazil. Transbound. Emerg. Dis. 2018, 65, 5–9. [Google Scholar] [CrossRef] [PubMed]
- Serena, M.S.; Cappuccio, J.A.; Barrales, H.; Metz, G.E.; Aspitia, C.G.; Lozada, I.; Perfumo, C.J.; Quiroga, M.A.; Pineyro, P.; Echeverria, M.G. First detection and genetic characterization of porcine circovirus type 3 (PCV3) in Argentina and its association with reproductive failure. Transbound. Emerg. Dis. 2020, 68, 1761–1766. [Google Scholar] [CrossRef]
- Molossi, F.A.; de Cecco, B.S.; de Almeida, B.A.; Henker, L.C.; da Silva, M.S.; Mosena, A.C.S.; Canal, C.W.; Brandalise, L.; Simao, G.M.R.; Vanucci, F.; et al. PCV3-associated reproductive failure in pig herds in Brazil. Trop. Anim. Health Prod. 2022, 54, 293. [Google Scholar] [CrossRef]
- Saporiti, V.; Franzo, G.; Sibila, M.; Segales, J. Porcine circovirus 3 (PCV-3) as a causal agent of disease in swine and a proposal of PCV-3 associated disease case definition. Transbound. Emerg. Dis. 2021, 68, 2936–2948. [Google Scholar] [CrossRef]
- Krakowka, S.; Ellis, J.A.; Meehan, B.; Kennedy, S.; McNeilly, F.; Allan, G. Viral wasting syndrome of swine: Experimental reproduction of postweaning multisystemic wasting syndrome in gnotobiotic swine by coinfection with porcine circovirus 2 and porcine parvovirus. Vet. Pathol. 2000, 37, 254–263. [Google Scholar] [CrossRef] [PubMed]
- Allan, G.M.; Kennedy, S.; McNeilly, F.; Foster, J.C.; Ellis, J.A.; Krakowka, S.J.; Meehan, B.M.; Adair, B.M. Experimental reproduction of severe wasting disease by co-infection of pigs with porcine circovirus and porcine parvovirus. J. Comp. Pathol. 1999, 121, 1–11. [Google Scholar] [CrossRef]
- Ellis, J.; Krakowka, S.; Lairmore, M.; Haines, D.; Bratanich, A.; Clark, E.; Allan, G.; Konoby, C.; Hassard, L.; Meehan, B.; et al. Reproduction of lesions of postweaning multisystemic wasting syndrome in gnotobiotic piglets. J. Vet. Diagn. Investig. 1999, 11, 3–14. [Google Scholar] [CrossRef]
- Althouse, G.C.; Kauffold, J.; Rossow, S. Diseases of the Reproductive System. In Diseases of Swine; John Wiley & Sons: Hoboken, NJ, USA, 2019; pp. 373–392. [Google Scholar]
- Chen, H.Y.; Li, X.K.; Cui, B.A.; Wei, Z.Y.; Li, X.S.; Wang, Y.B.; Zhao, L.; Wang, Z.Y. A TaqMan-based real-time polymerase chain reaction for the detection of porcine parvovirus. J. Virol. Methods 2009, 156, 84–88. [Google Scholar] [CrossRef] [PubMed]
- Eddicks, M.; Koeppen, M.; Willi, S.; Fux, R.; Reese, S.; Sutter, G.; Stadler, J.; Ritzmann, M. Low prevalence of porcine circovirus type 2 infections in farrowing sows and corresponding pre-suckling piglets in southern German pig farms. Vet. Microbiol. 2016, 187, 70–74. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, A.S.; Costa, P.; Rocha, T.; Amaro, A.; Vieira, M.L.; Ahmed, A.; Thompson, G.; Hartskeerl, R.A.; Inacio, J. Direct detection and differentiation of pathogenic Leptospira species using a multi-gene targeted real time PCR approach. PLoS ONE 2014, 9, e112312. [Google Scholar] [CrossRef] [PubMed]
- Mora-Diaz, J.; Pineyro, P.; Shen, H.; Schwartz, K.; Vannucci, F.; Li, G.; Arruda, B.; Gimenez-Lirola, L. Isolation of PCV3 from Perinatal and Reproductive Cases of PCV3-Associated Disease and In Vivo Characterization of PCV3 Replication in CD/CD Growing Pigs. Viruses 2020, 12, 219. [Google Scholar] [CrossRef]
- Pensaert, M.B.; Sanchez, R.E., Jr.; Ladekjaer-Mikkelsen, A.S.; Allan, G.M.; Nauwynck, H.J. Viremia and effect of fetal infection with porcine viruses with special reference to porcine circovirus 2 infection. Vet. Microbiol. 2004, 98, 175–183. [Google Scholar] [CrossRef]
- Eddicks, M.; Beuter, B.; Stuhldreier, R.; Nolte, T.; Reese, S.; Sutter, G.; Ritzmann, M.; Fux, R. Cross-sectional study on viraemia and shedding of porcine circovirus type 2 in a subclinically infected multiplier sow herd. Vet. Rec. 2019, 184, 189. [Google Scholar] [CrossRef]
- Segales, J.; Sibila, M. Revisiting Porcine Circovirus Disease Diagnostic Criteria in the Current Porcine Circovirus 2 Epidemiological Context. Vet. Sci. 2022, 9, 110. [Google Scholar] [CrossRef]
- Eddicks, M.; Maurer, R.; Deffner, P.; Eddicks, L.; Sipos, W.; Reese, S.; Cvjetkovic, V.; Krejci, R.; Opriessnig, T.; Ritzmann, M.; et al. Cross-Sectional Study on the Prevalence of PCV Types 2 and 3 DNA in Suckling Piglets Compared to Grow-Finish Pigs in Downstream Production. Pathogens 2022, 11, 671. [Google Scholar] [CrossRef]
- Brunborg, I.M.; Moldal, T.; Jonassen, C.M. Quantitation of porcine circovirus type 2 isolated from serum/plasma and tissue samples of healthy pigs and pigs with postweaning multisystemic wasting syndrome using a TaqMan-based real-time PCR. J. Virol. Methods 2004, 122, 171–178. [Google Scholar] [CrossRef]
- Olvera, A.; Sibila, M.; Calsamiglia, M.; Segales, J.; Domingo, M. Comparison of porcine circovirus type 2 load in serum quantified by a real time PCR in postweaning multisystemic wasting syndrome and porcine dermatitis and nephropathy syndrome naturally affected pigs. J. Virol. Methods 2004, 117, 75–80. [Google Scholar] [CrossRef] [PubMed]
- Unterweger, C.; Brunthaler, R.; Auer, A.; Fux, R.; Weissenbacher-Lang, C.; Ladinig, A. Reconsideration of the diagnostic criteria required for PCV2 reproductive disease. Vet. J. 2021, 272, 105660. [Google Scholar] [CrossRef]
- Opriessnig, T.; Langohr, I. Current state of knowledge on porcine circovirus type 2-associated lesions. Vet. Pathol. 2013, 50, 23–38. [Google Scholar] [CrossRef]
- Nickoll, I.; Striegl, J.; Fux, R.; Sutter, G.; Ritzmann, M.; Eddicks, M. Porzines circovirus typ 2 (PCV2)-infektionen in oberbayerischen schweinemastbeständen mit routinemäßiger impfung gegen PCV2 -bedeutung für das auftreten von lungenveränderungen bei schlachttierkörpern. Tieraerztl. Umsch. 2016, 71, 360–370. [Google Scholar]
- Harms, P.A.; Halbur, P.G.; Sorden, S.D. Three cases of porcine respiratory disease complex associated with porcine circovirus type 2 infection. J. Swine Health Prod. 2002, 10, 27–30. [Google Scholar]
- Hansen, M.S.; Pors, S.E.; Jensen, H.E.; Bille-Hansen, V.; Bisgaard, M.; Flachs, E.M.; Nielsen, O.L. An investigation of the pathology and pathogens associated with porcine respiratory disease complex in Denmark. J. Comp. Pathol. 2010, 143, 120–131. [Google Scholar] [CrossRef] [PubMed]
- Serena, M.S.; Dibárbora, M.; Olivera, V.; Metz, G.E.; Aspitia, C.G.; Pereda, A.; Echeverría, M.G.; Cappuccio, J. Evidence of porcine circovirus type 2 and co-infection with ungulate protoparvovirus 1 (porcine parvovirus) in mummies and stillborn piglets in subclinically infected farm. Infect. Genet. Evol. 2021, 89, 104735. [Google Scholar] [CrossRef]
- Pescador, C.A.; Bandarra, P.M.; Castro, L.A.; Antoniassi, N.A.; Ravazzolo, A.P.; Sonne, L.; Cruz, C.E.; Driemeier, D. Co-infection by porcine circovirus type 2 and porcine parvovirus in aborted fetuses and stillborn piglets in southern Brazil. Pesqui. Veterinária Bras. 2007, 27, 425–429. [Google Scholar] [CrossRef]
- de Castro, A.M.M.G.; Cruz, T.F.; Salgado, V.R.; Kanashiro, T.M.; Ferrari, K.L.; Araujo, J.P.; Brandão, P.E.; Richtzenhain, L.J. Detection of porcine circovirus genotypes 2a and 2b in aborted foetuses from infected swine herds in the State of São Paulo, Brazil. Acta Vet. Scand. 2012, 54, 29. [Google Scholar] [CrossRef]
- Sanchez, R.E., Jr.; Meerts, P.; Nauwynck, H.J.; Pensaert, M.B. Change of porcine circovirus 2 target cells in pigs during development from fetal to early postnatal life. Vet. Microbiol. 2003, 95, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Zeeuw, E.; Leinecker, N.; Herwig, V.; Selbitz, H.-J.; Truyen, U. Study of the virulence and cross-neutralization capability of recent porcine parvovirus field isolates and vaccine viruses in experimentally infected pregnant gilts. J. Gen. Virol. 2007, 88, 420–427. [Google Scholar] [CrossRef]
- Jozwik, A.; Manteufel, J.; Selbitz, H.J.; Truyen, U. Vaccination against porcine parvovirus protects against disease, but does not prevent infection and virus shedding after challenge infection with a heterologous virus strain. J. Gen. Virol. 2009, 90 Pt 10, 2437–2441. [Google Scholar] [CrossRef]
- Sharma, R.; Saikumar, G. Porcine parvovirus-and porcine circovirus 2-associated reproductive failure and neonatal mortality in crossbred Indian pigs. Trop. Anim. Health Prod. 2010, 42, 515–522. [Google Scholar] [CrossRef] [PubMed]
- Dunne, H.; Gobble, J.; Hokanson, J.; Kradel, D.; Bubash, G. Porcine reproductive failure associated with a newly identified “SMEDI” group of picorna viruses. Am. J. Vet. Res. 1965, 26, 1284–1297. [Google Scholar]
- Ellis, J.; Bratanich, A.; Clark, E.; Allan, G.; Meehan, B.; Haines, D.; Harding, J.; West, K.; Krakowka, S.; Konoby, C. Coinfection by porcine circoviruses and porcine parvovirus in pigs with naturally acquired postweaning multisystemic wasting syndrome. J. Vet. Diagn. Investig. 2000, 12, 21–27. [Google Scholar] [CrossRef]
- Opriessnig, T.; Fenaux, M.; Yu, S.; Evans, R.; Cavanaugh, D.; Gallup, J.; Pallares, F.; Thacker, E.; Lager, K.; Meng, X. Effect of porcine parvovirus vaccination on the development of PMWS in segregated early weaned pigs coinfected with type 2 porcine circovirus and porcine parvovirus. Vet. Microbiol. 2004, 98, 209–220. [Google Scholar] [CrossRef]
- Baxter, E.; Jarvis, S.; D’eath, R.; Ross, D.; Robson, S.; Farish, M.; Nevison, I.; Lawrence, A.; Edwards, S. Investigating the behavioural and physiological indicators of neonatal survival in pigs. Theriogenology 2008, 69, 773–783. [Google Scholar] [CrossRef]
- Evans, H.E.; Sack, W.O. Prenatal development of domestic and laboratory mammals: Growth curves, external features and selected references. Zent. Vet. C 1973, 2, 11–45. [Google Scholar] [CrossRef] [PubMed]
- Palinski, R.; Pineyro, P.; Shang, P.; Yuan, F.; Guo, R.; Fang, Y.; Byers, E.; Hause, B.M. A Novel Porcine Circovirus Distantly Related to Known Circoviruses Is Associated with Porcine Dermatitis and Nephropathy Syndrome and Reproductive Failure. J. Virol. 2017, 91, e01879-16. [Google Scholar] [CrossRef] [PubMed]
- Faccini, S.; Barbieri, I.; Gilioli, A.; Sala, G.; Gibelli, L.R.; Moreno, A.; Sacchi, C.; Rosignoli, C.; Franzini, G.; Nigrelli, A. Detection and genetic characterization of Porcine circovirus type 3 in Italy. Transbound. Emerg. Dis. 2017, 64, 1661–1664. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, R.C. Fetal mummification in the major domestic species: Current perspectives on causes and management. Vet. Med. 2015, 6, 233–244. [Google Scholar] [CrossRef] [PubMed]
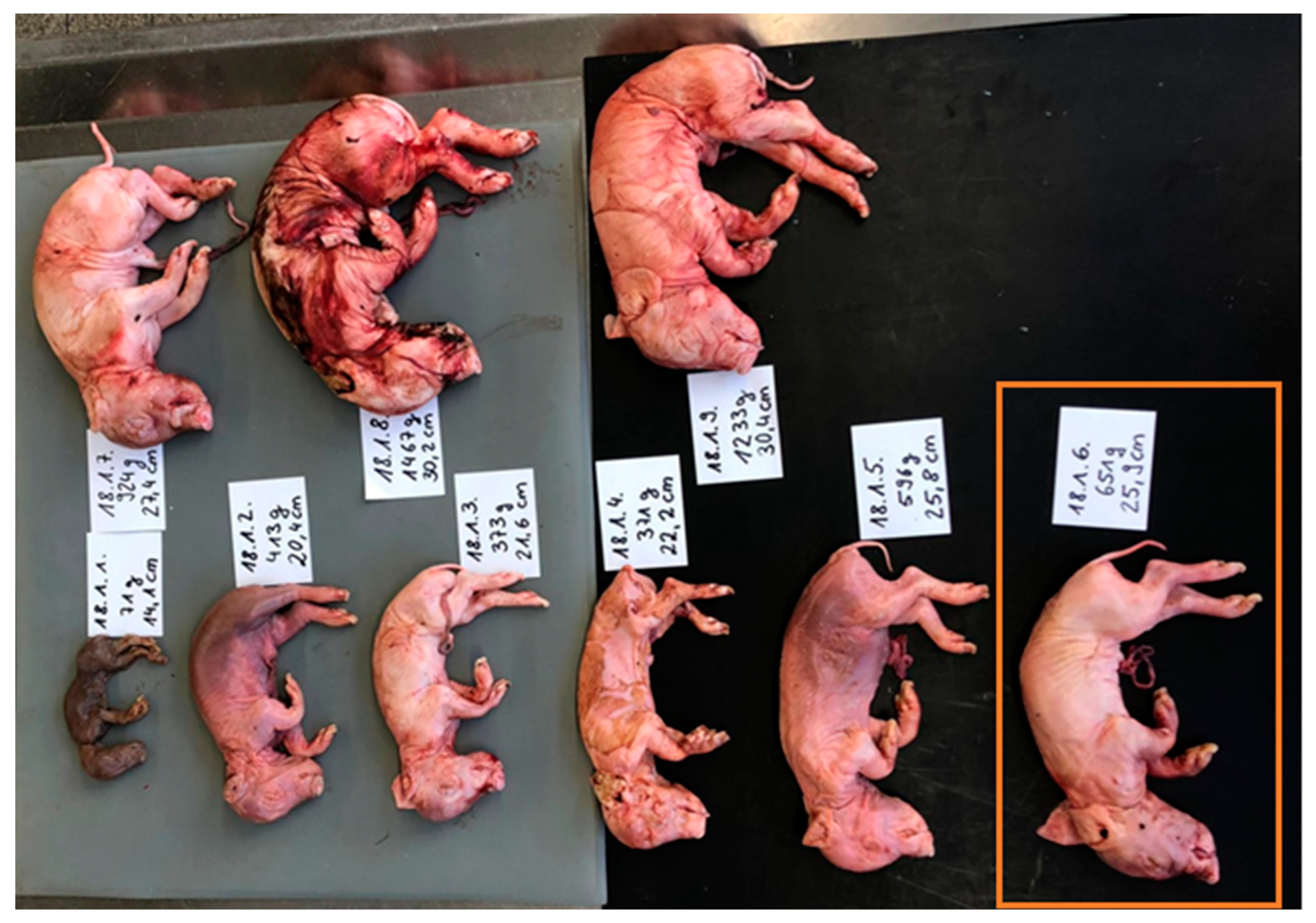


| Appearance of Fetuses | Total Population | Randomly Selected Study Population | |
|---|---|---|---|
| mummified | 30.4% (109/358) | 30.4% (48/158) | |
| stillborn | macerated | 21.5% (77/358) | 20.9% (33/158) |
| fresh | 44.7% (160/358) | 41.8% (66/158) | |
| total | 66.2% (237/358) | 62.6% (99/158) | |
| weak-born * | 3.4% (12/358) | 7% (11/158) |
| Total Detected | PPV1 | PCV2 | PCV3 | Leptospira spp. |
|---|---|---|---|---|
| Farms (n = 18) | 50% (27.8–72.2%) | 61.1% (38.9–83.3%) | 27.8% (11.1–50.0%) | 11.1% (0.0–27.8%) |
| Litters (n = 40) | 27.5% (15.0–40.0%) | 50.0% (35.0–67.5%) | 17.5% (7.5–30.0%) | 17.5% (7.5–30.0%) |
| Fetuses (n = 158) | 14.6% (9.5–20.9%) | 28.5% (21.5–36.1%) | 11.4 (6.3–17.1%) | 12.7% (7.6–17.7%) |
| Independent Variable | Dependent Variable | p-Value Chi2 Test | p-Value Binary Logistic Regression | OR | Lower CI | Upper CI |
|---|---|---|---|---|---|---|
| Fetus PCV2 DNA-positive | Leptospira spp. DNA-positive | <0.001 | <0.001 | 26.301 | 4.911 | 140.870 |
| Fetus stillborn-macerated | 0.004 | 0.003 | 8.673 | 2.119 | 35.488 | |
| Gilt | 0.023 | <0.001 | 0.006 | 0.001 | 0.060 | |
| Fetus PPV1 DNA-positive | 0.047 | <0.001 | 0.114 | 0.014 | 0.490 | |
| Sow Lepto.-vacc. | 0.045 | Not included in the multivariable analysis due to redundant data as no Leptospira spp.-vaccinated herds had positive fetuses. | ||||
| Fetus Leptospira spp. DNA-positive | PCV2 DNA-positive | <0.001 | <0.001 | 9.058 | 2.772 | 29.595 |
| PCV3 DNA-positive | 0.025 | 0.008 | 0.078 | 0.008 | 0.742 | |
| Sow PCV2-vaccinated | 0.001 | 0.001 | 0.051 | 0.007 | 0.351 | |
| Gilt | PPV1 DNA-positive | 0.002 | - | - | - | - |
| PCV2 Sow vaccination | 0.047 | - | - | - | - | |
| Lepto Sow vaccination | 0.010 | - | - | - | - | |
| Fetus Leptospira spp. DNA-positive | 0.047 | - | - | - | - | |
| PCV2 DNA-positive | PCV3 DNA-positive | - | - | - | - | - |
| Gilt | - | - | - | - | - | |
| (Co) Infections | PPV1 | PCV2 | PCV3 | Lepto. 1 | PCV2 + PPV1 | PCV2 + Lepto. 1 | PCV3 + Lepto. 1 | PCV2 + PCV3 + Lepto. 1 | No Pathogen |
|---|---|---|---|---|---|---|---|---|---|
| Farms (n = 18) | 16.7% (0.0– 33.3%) | 16.7% (0.0– 33.3%) | 16.7% (0.0– 33.3%) | - | 33.3% (11.1– 55.6%) | - | - | 11.1% (0.0– 27.8%) | 5.6% (0.0– 16.7%) |
| Litters (n = 40) | 12.5% (2.5– 22.5%) | 22.5% (10.0– 35.0%) | 12.5% (2.5– 24.9%) | - | 15.0% (5.0– 27.5%) | 10.0% (2.5– 20.0%) | 2.5% (0.0– 7.5%) | 2.5% (0.0– 7.5%) | 22.5% (10.0– 37.5%) |
| Fetuses (n = 158) | 8.9% (4.4– 13.3%) | 14.6% (9.5– 20.3%) | 9.5% (5.1– 14.6%) | 2.5% (0.6– 5.1%) | 5.7% (2.5– 9.5%) | 8.2% (4.4– 12.7%) | 1.3% (0.0– 3.2%) | 0.6% (0.0– 2.5%) | 48.7% (40.5– 55.7%) |
| Mean Cq-Value of Single Pathogens in the Tissue Pools | ||||
|---|---|---|---|---|
| (Co)infections | PPV1 | PCV2 | PCV3 | Leptospira spp. |
| PPV1 (n = 14) | 27.21 (±1.04) | - | - | - |
| PCV2 (n = 23) | - | 27.41 (±8.28) | - | - |
| PCV3 (n = 15) | - | - | 26.07 (±7.57) | - |
| Leptospira spp. (n = 4) | - | - | - | 36.75 (±2.07) |
| PPV1 + PCV2 (n = 9) | 21.96 (±11.23) | 32.77 (±1.77) | - | - |
| PCV2 + Leptospira spp. (n = 13) | - | 31.21 (±1.98) | - | 36.38 (±1.44) |
| PCV3 + Leptospira spp. (n = 2) | - | - | 33.69 (±3.49) | 33.50 (±0.70) |
| PCV2 + PCV3 + Leptospira spp. (n = 1) | - | 30.09 (±0) | 32.43 (±0) | 37.00 (±0) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Eddicks, M.; Gründl, J.; Seifert, A.; Eddicks, L.; Reese, S.; Tabeling, R.; Swam, H.; Strutzberg-Minder, K.; Ritzmann, M.; Fux, R. Examination on the Occurrence of Coinfections in Diagnostic Transmittals in Cases of Stillbirth, Mummification, Embryonic Death, and Infertility (SMEDI) Syndrome in Germany. Microorganisms 2023, 11, 1675. https://doi.org/10.3390/microorganisms11071675
Eddicks M, Gründl J, Seifert A, Eddicks L, Reese S, Tabeling R, Swam H, Strutzberg-Minder K, Ritzmann M, Fux R. Examination on the Occurrence of Coinfections in Diagnostic Transmittals in Cases of Stillbirth, Mummification, Embryonic Death, and Infertility (SMEDI) Syndrome in Germany. Microorganisms. 2023; 11(7):1675. https://doi.org/10.3390/microorganisms11071675
Chicago/Turabian StyleEddicks, Matthias, Julia Gründl, Annika Seifert, Lina Eddicks, Sven Reese, Robert Tabeling, Hanny Swam, Katrin Strutzberg-Minder, Mathias Ritzmann, and Robert Fux. 2023. "Examination on the Occurrence of Coinfections in Diagnostic Transmittals in Cases of Stillbirth, Mummification, Embryonic Death, and Infertility (SMEDI) Syndrome in Germany" Microorganisms 11, no. 7: 1675. https://doi.org/10.3390/microorganisms11071675
APA StyleEddicks, M., Gründl, J., Seifert, A., Eddicks, L., Reese, S., Tabeling, R., Swam, H., Strutzberg-Minder, K., Ritzmann, M., & Fux, R. (2023). Examination on the Occurrence of Coinfections in Diagnostic Transmittals in Cases of Stillbirth, Mummification, Embryonic Death, and Infertility (SMEDI) Syndrome in Germany. Microorganisms, 11(7), 1675. https://doi.org/10.3390/microorganisms11071675






