Transcriptome and Secretome Analyses of Endophyte Methylobacterium mesophilicum and Pathogen Xylella fastidiosa Interacting Show Nutrient Competition
Abstract
:1. Introduction
2. Materials and Methods
2.1. Xf and Mm Cultivation Conditions
2.2. Bacterial Growth Evaluation
2.3. Transcriptomic Analyses
2.3.1. Experimental Design
2.3.2. qPCR Quantification of Xf and Mm
2.3.3. Total RNA Isolation, cDNA Library Preparation, and Sequencing
2.3.4. Bioinformatic Analyses and Statistics
2.4. Secretomic Analyses
2.4.1. Protein Extraction from Xf, Mm, and Co-Culture Supernatants
2.4.2. Shotgun Proteomics
2.5. Integration of Transcriptomics and Proteomics Analyses
3. Results
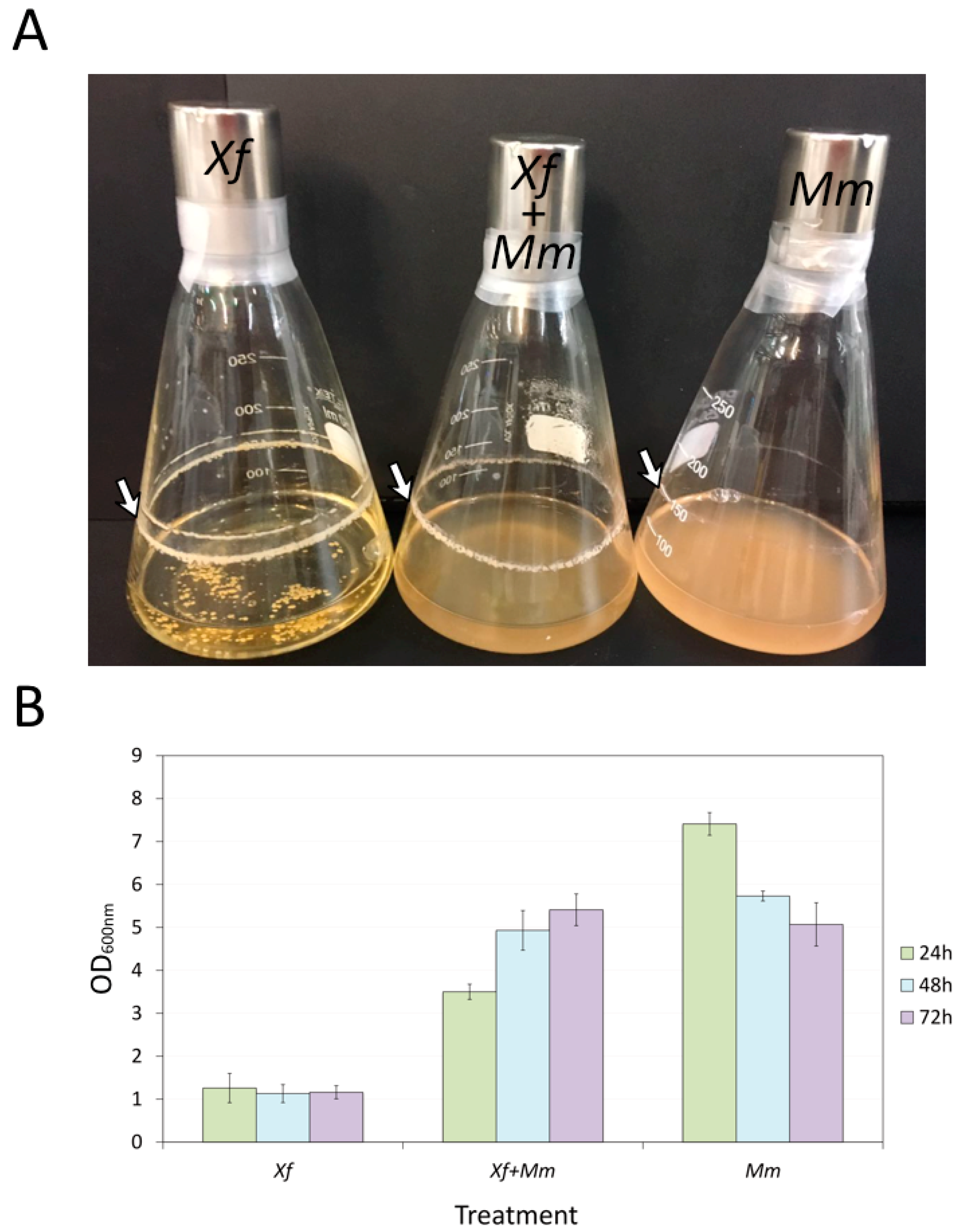
3.1. Growth of Mm and Xf in Co-Culture and Xf Biofilm Formation
3.2. Bacteria Quantification by qPCR
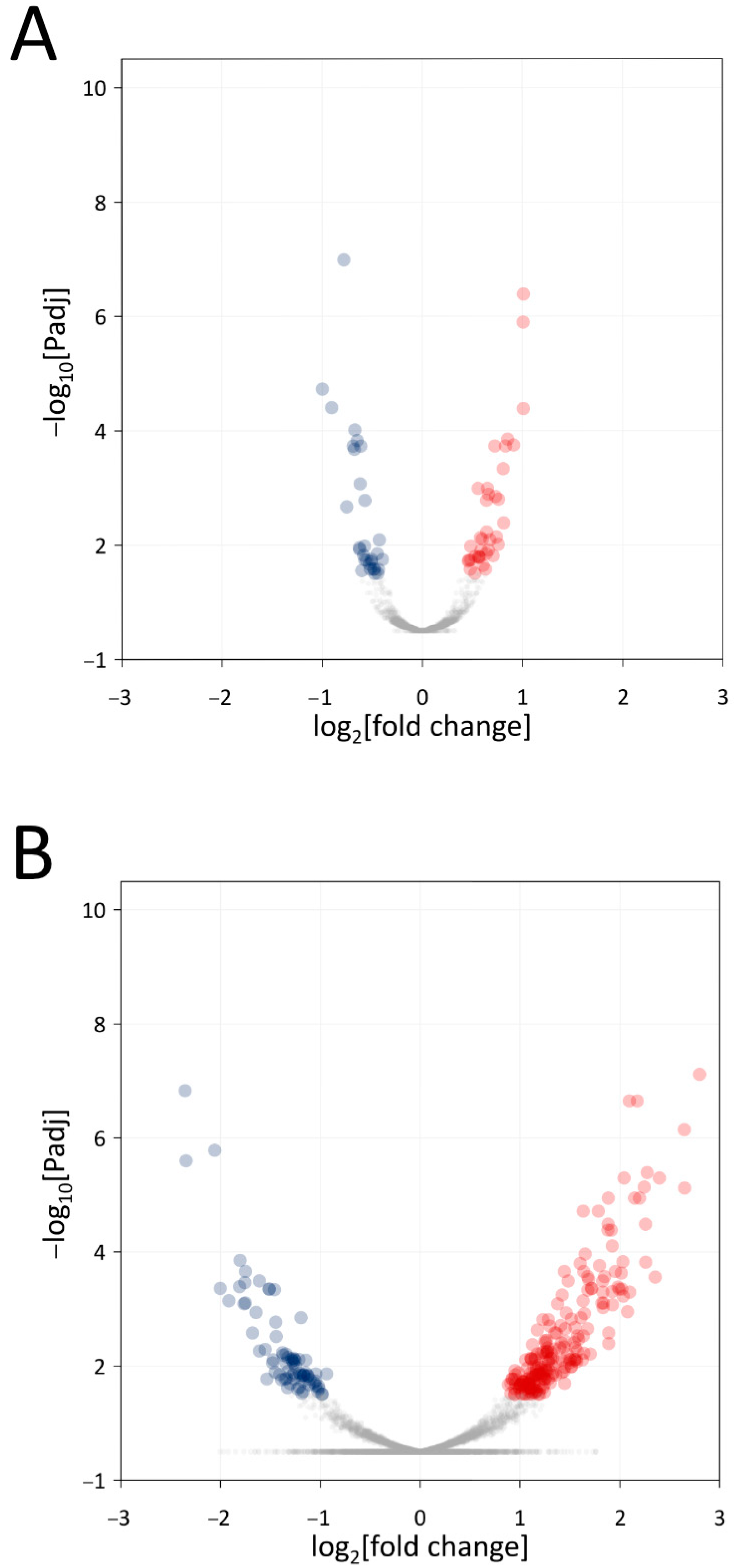
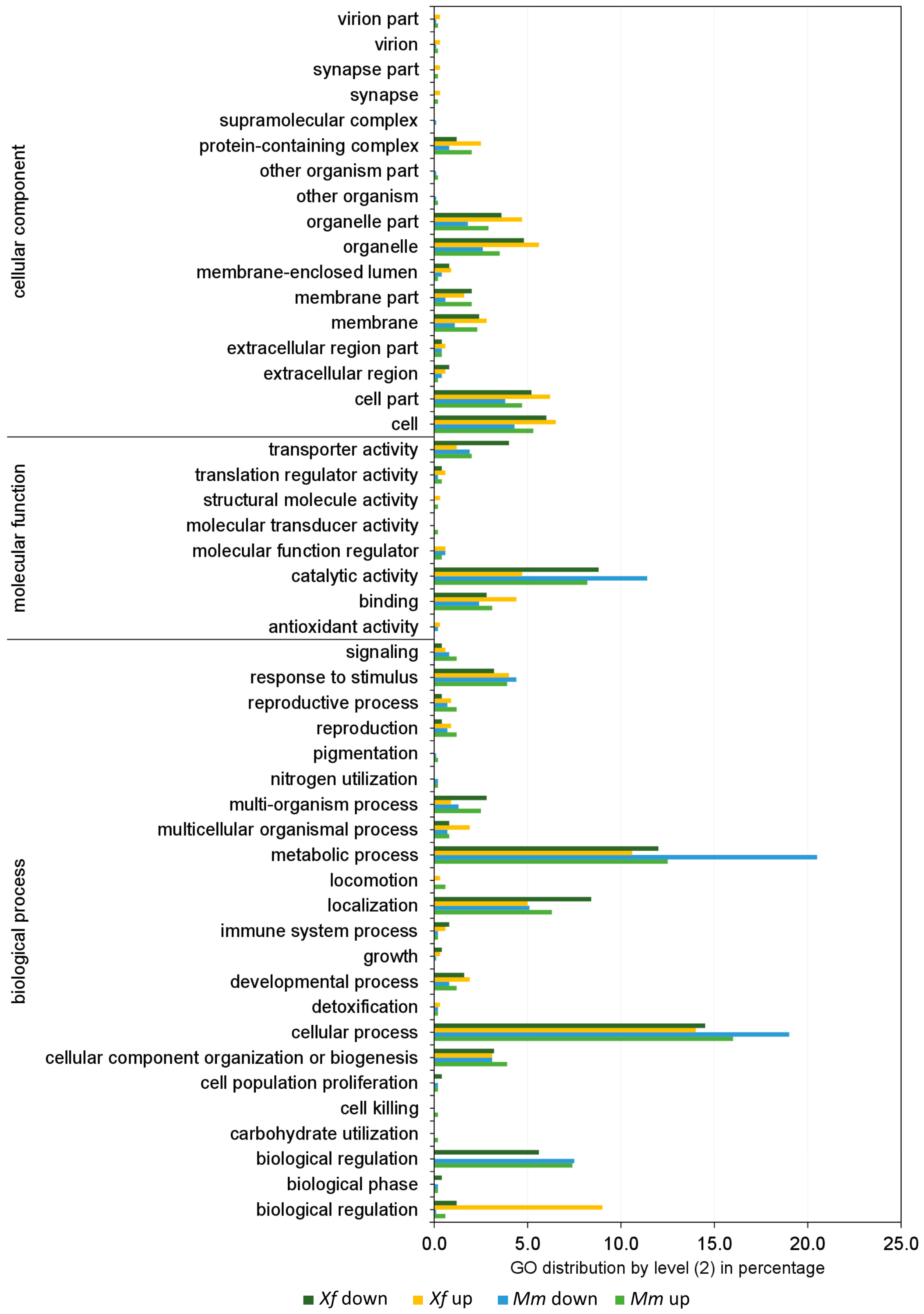
3.3. Transcriptomic Analyses
3.4. Transcriptome Analysis of Xf+Mm Co-Culture Compared to Xf Monoculture
3.4.1. Macromolecule Metabolism
3.4.2. Transport
3.4.3. Stress-Related Genes
3.4.4. Pathogenicity
3.5. Transcriptome Analysis of Xf+Mm Co-Culture Compared to Mm Monoculture
3.5.1. DNA, RNA, and Protein Metabolism
3.5.2. Lipid, Sugar, Amino Acid and Nucleotide Metabolism
3.5.3. Transport
3.5.4. Stress-Related Genes
3.5.5. Defense Mechanism-Related Genes
3.6. Secretome Analyses
3.6.1. Total Secretome of Xf and Mm
3.6.2. Xf Secretome
3.6.3. Mm Secretome
3.6.4. Secretome Predictions and Functional Analyses
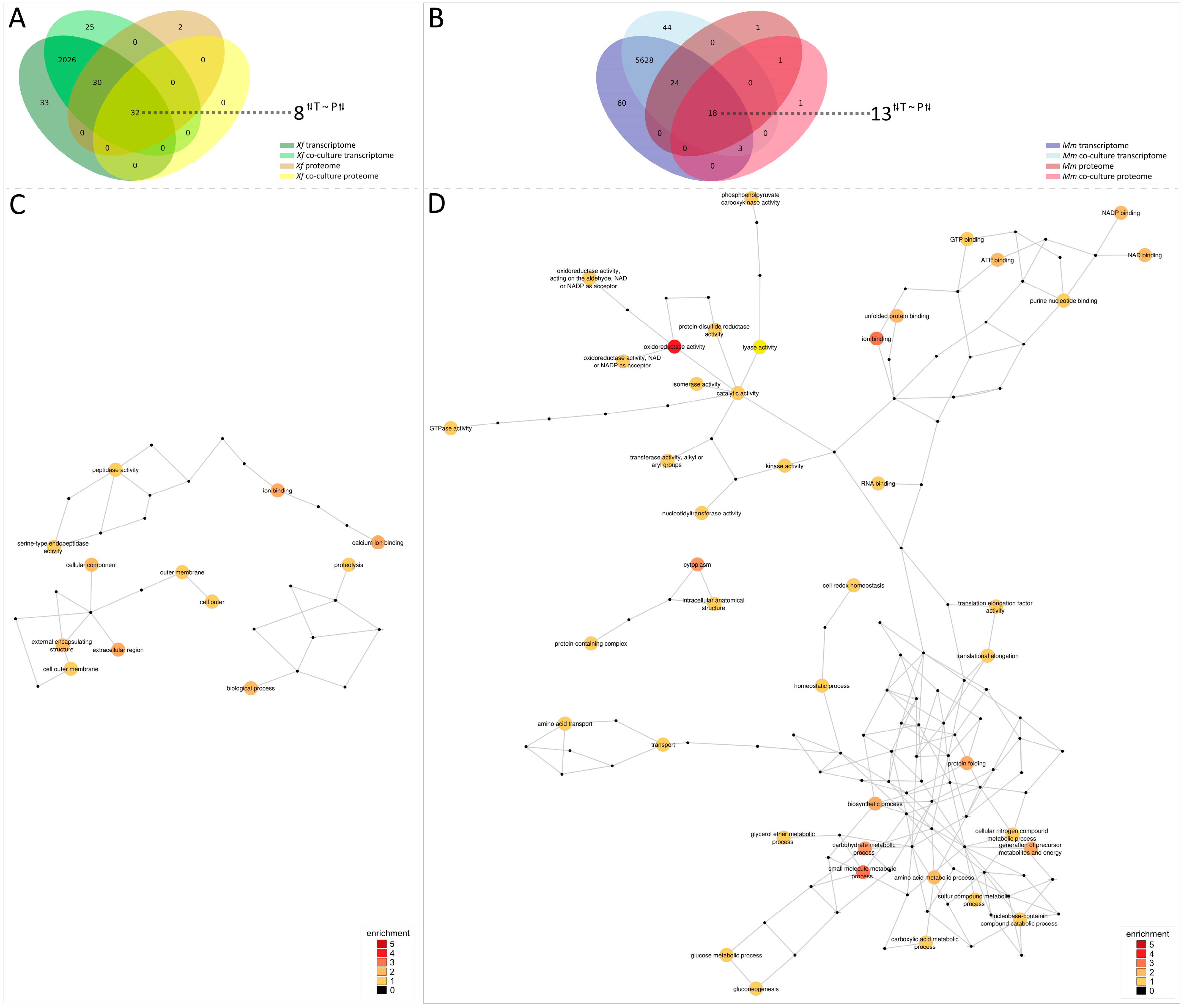
3.7. Integrating Transcriptome and Secretome Data
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wells, J.M.; Raju, B.C.; Hung, H.-Y.; Weisburg, W.G.; Mandelco-Paul, L.; Brenner, D.J. Xylella fastidiosa gen. nov., sp. nov: Gram-negative, xylem-limited, fastidious plant bacteria related to Xanthomonas spp. Int. J. Syst. Bacteriol. 1987, 37, 136–143. [Google Scholar] [CrossRef]
- Hopkins, D.L. Xylella fastidiosa: Xylem-limited bacterial pathogen of plants. Ann. Rev. Phytopathol. 1989, 27, 271–290. [Google Scholar] [CrossRef]
- Chang, C.J.; Garnier, M.; Zreik, L.; Rossetti, V.; Bove, J.M. Culture and serological detection of the xylem-limited bacterium causing citrus variegated chlorosis and its identification as a strain ofXylella fastidiosa. Curr. Microbiol. 1993, 27, 137–142. [Google Scholar] [CrossRef] [PubMed]
- Hopkins, D.L.; Purcell, A.H. Xylella fastidiosa: Cause of Pierce’s disease of grapevine and other emergent diseases. Plant Dis. 2002, 86, 1056–1066. [Google Scholar] [CrossRef]
- Cariddi, C.; Saponari, M.; Boscia, D.; De Stradis, A.; Loconsole, G.; Nigro, F.; Porcelli, F.; Potere, O.; Martelli, G.P. Isolation of a Xylella fastidiosa strain infecting olive and oleander in Apulia, Italy. J. Plant Pathol. 2014, 96, 425–429. [Google Scholar]
- Sicard, A.; Zeilinger, A.R.; Vanhove, M.; Schartel, T.E.; Beal, D.J.; Daugherty, M.P.; Almeida, R.P.P. Xylella fastidiosa: Insights into an Emerging Plant Pathogen. Annu. Rev. Phytopathol. 2018, 56, 181–202. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, S.; Almeida, R.P.; Lindow, S. Living in two worlds: The plant and insect lifestyles of Xylella fastidiosa. Annu. Rev. Phytopathol. 2008, 46, 243–271. [Google Scholar] [CrossRef] [PubMed]
- Janse, J.D.; Obradovic, A. Xylella fastidiosa: Its biology, diagnosis, control and risks. J. Plant Pathol. 2010, 92, S1.35–S31.48. [Google Scholar]
- Backus, E.A.; Morgan, D.J. Spatiotemporal colonization of Xylella fastidiosa in its vector supports the role of egestion in the inoculation mechanism of foregut-borne plant pathogens. Phytopathology 2011, 101, 912–922. [Google Scholar] [CrossRef]
- de Souza, J.B.; Almeida-Souza, H.O.; Zaini, P.A.; Alves, M.N.; de Souza, A.G.; Pierry, P.M.; da Silva, A.M.; Goulart, L.R.; Dandekar, A.M.; Nascimento, R. Xylella fastidiosa subsp. pauca Strains Fb7 and 9a5c from Citrus Display Differential Behavior, Secretome, and Plant Virulence. Int. J. Mol. Sci. 2020, 21, 6769. [Google Scholar] [CrossRef] [PubMed]
- Zaini, P.A.; Fogaca, A.C.; Lupo, F.G.; Nakaya, H.I.; Vencio, R.Z.; da Silva, A.M. The iron stimulon of Xylella fastidiosa includes genes for type IV pilus and colicin V-like bacteriocins. J. Bacteriol. 2008, 190, 2368–2378. [Google Scholar] [CrossRef]
- Ciraulo, M.B.; Santos, D.S.; Rodrigues, A.C.; de Oliveira, M.V.; Rodrigues, T.; de Oliveira, R.C.; Nunes, L.R. Transcriptome analysis of the phytobacterium Xylella fastidiosa growing under xylem-based chemical conditions. J. Biomed. Biotechnol. 2010, 2010, 781365. [Google Scholar] [CrossRef] [PubMed]
- Giampetruzzi, A.; Morelli, M.; Saponari, M.; Loconsole, G.; Chiumenti, M.; Boscia, D.; Savino, V.N.; Martelli, G.P.; Saldarelli, P. Transcriptome profiling of two olive cultivars in response to infection by the CoDiRO strain of Xylella fastidiosa subsp. pauca. BMC Genom. 2016, 17, 475. [Google Scholar] [CrossRef]
- Gouran, H.; Gillespie, H.; Nascimento, R.; Chakraborty, S.; Zaini, P.A.; Jacobson, A.; Phinney, B.S.; Dolan, D.; Durbin-Johnson, B.P.; Antonova, E.S.; et al. The Secreted Protease PrtA Controls Cell Growth, Biofilm Formation and Pathogenicity in Xylella fastidiosa. Sci. Rep. 2016, 6, 31098. [Google Scholar] [CrossRef]
- Parker, J.K.; Chen, H.; McCarty, S.E.; Liu, L.Y.; De La Fuente, L. Calcium transcriptionally regulates the biofilm machinery of Xylella fastidiosa to promote continued biofilm development in batch cultures. Env. Microbiol. 2016, 18, 1620–1634. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; De La Fuente, L. Calcium transcriptionally regulates movement, recombination and other functions of Xylella fastidiosa under constant flow inside microfluidic chambers. Microb. Biotechnol. 2020, 13, 548–561. [Google Scholar] [CrossRef]
- Smolka, M.B.; Martins-de-Souza, D.; Winck, F.V.; Santoro, C.E.; Castellari, R.R.; Ferrari, F.; Brum, I.J.; Galembeck, E.; Della Coletta Filho, H.; Machado, M.A.; et al. Proteome analysis of the plant pathogen Xylella fastidiosa reveals major cellular and extracellular proteins and a peculiar codon bias distribution. Proteomics 2003, 3, 224–237. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.S.; De Souza, A.A.; Takita, M.A.; Labate, C.A.; Machado, M.A. Analysis of the biofilm proteome of Xylella fastidiosa. Proteome Sci. 2011, 9, 58. [Google Scholar] [CrossRef] [PubMed]
- Mendes, J.S.; Santiago, A.S.; Toledo, M.A.; Horta, M.A.; de Souza, A.A.; Tasic, L.; de Souza, A.P. In vitro Determination of Extracellular Proteins from Xylella fastidiosa. Front. Microbiol. 2016, 7, 2090. [Google Scholar] [CrossRef] [PubMed]
- Nascimento, R.; Gouran, H.; Chakraborty, S.; Gillespie, H.W.; Almeida-Souza, H.O.; Tu, A.; Rao, B.J.; Feldstein, P.A.; Bruening, G.; Goulart, L.R.; et al. The Type II Secreted Lipase/Esterase LesA is a Key Virulence Factor Required for Xylella fastidiosa Pathogenesis in Grapevines. Sci. Rep. 2016, 6, 18598. [Google Scholar] [CrossRef]
- Ryan, R.P.; Dow, J.M. Communication with a growing family: Diffusible signal factor (DSF) signaling in bacteria. Trends Microbiol. 2011, 19, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Voegel, T.M.; Warren, J.G.; Matsumoto, A.; Igo, M.M.; Kirkpatrick, B.C. Localization and characterization of Xylella fastidiosa haemagglutinin adhesins. Microbiology 2010, 156, 2172–2179. [Google Scholar] [CrossRef] [PubMed]
- Ionescu, M.; Zaini, P.A.; Baccari, C.; Tran, S.; da Silva, A.M.; Lindow, S.E. Xylella fastidiosa outer membrane vesicles modulate plant colonization by blocking attachment to surfaces. Proc. Natl. Acad. Sci. USA 2014, 111, E3910–E3918. [Google Scholar] [CrossRef] [PubMed]
- Santiago, A.D.; Mendes, J.S.; Dos Santos, C.A.; de Toledo, M.A.; Beloti, L.L.; Crucello, A.; Horta, M.A.; Favaro, M.T.; Munar, D.M.; de Souza, A.A.; et al. The Antitoxin Protein of a Toxin-Antitoxin System from Xylella fastidiosa Is Secreted via Outer Membrane Vesicles. Front. Microbiol. 2016, 7, 2030. [Google Scholar] [CrossRef]
- Feitosa-Junior, O.R.; Stefanello, E.; Zaini, P.; Nascimento, R.; Pierry, P.M.; Dandekar, A.; Lindow, S.; da Silva, A.M. Proteomic and metabolomic analyses of Xylella fastidiosa OMV-enriched fractions reveal association with virulence factors and signaling molecules of the DSF family. Phytopathology 2019, 109, 1344–1353. [Google Scholar] [CrossRef]
- Merfa, M.V.; Niza, B.; Takita, M.A.; De Souza, A.A. The MqsRA Toxin-Antitoxin System from Xylella fastidiosa Plays a Key Role in Bacterial Fitness, Pathogenicity, and Persister Cell Formation. Front. Microbiol. 2016, 7, 904. [Google Scholar] [CrossRef]
- Anguita-Maeso, M.; Ares-Yebra, A.; Haro, C.; Roman-Ecija, M.; Olivares-Garcia, C.; Costa, J.; Marco-Noales, E.; Ferrer, A.; Navas-Cortes, J.A.; Landa, B.B. Xylella fastidiosa Infection Reshapes Microbial Composition and Network Associations in the Xylem of Almond Trees. Front. Microbiol. 2022, 13, 866085. [Google Scholar] [CrossRef]
- Araujo, W.L.; Marcon, J.; Maccheroni, W., Jr.; Van Elsas, J.D.; Van Vuurde, J.W.; Azevedo, J.L. Diversity of endophytic bacterial populations and their interaction with Xylella fastidiosa in citrus plants. Appl. Env. Microbiol. 2002, 68, 4906–4914. [Google Scholar] [CrossRef]
- Lacava, P.T.; Araujo, W.L.; Marcon, J.; Maccheroni, W., Jr.; Azevedo, J.L. Interaction between endophytic bacteria from citrus plants and the phytopathogenic bacteria Xylella fastidiosa, causal agent of citrus-variegated chlorosis. Lett. Appl. Microbiol. 2004, 39, 55–59. [Google Scholar] [CrossRef]
- Dourado, M.N.; Santos, D.S.; Nunes, L.R.; Costa de Oliveira, R.L.; de Oliveira, M.V.; Araujo, W.L. Differential gene expression in Xylella fastidiosa 9a5c during co-cultivation with the endophytic bacterium Methylobacterium mesophilicum SR1.6/6. J. Basic. Microbiol. 2015, 55, 1357–1366. [Google Scholar] [CrossRef]
- Lacava, P.T.; Li, W.B.; Araujo, W.L.; Azevedo, J.L.; Hartung, J.S. Rapid, specific and quantitative assays for the detection of the endophytic bacterium Methylobacterium mesophilicum in plants. J. Microbiol. Methods 2006, 65, 535–541. [Google Scholar] [CrossRef]
- Andreote, F.D.; Lacava, P.T.; Gai, C.S.; Araujo, W.L.; Maccheroni, W., Jr.; van Overbeek, L.S.; van Elsas, J.D.; Azevedo, J.L. Model plants for studying the interaction between Methylobacterium mesophilicum and Xylella fastidiosa. Can. J. Microbiol. 2006, 52, 419–426. [Google Scholar] [CrossRef] [PubMed]
- Gai, C.S.; Lacava, P.T.; Quecine, M.C.; Auriac, M.C.; Lopes, J.R.; Araujo, W.L.; Miller, T.A.; Azevedo, J.L. Transmission of Methylobacterium mesophilicum by Bucephalogonia xanthophis for paratransgenic control strategy of citrus variegated chlorosis. J. Microbiol. 2009, 47, 448–454. [Google Scholar] [CrossRef] [PubMed]
- Rogers, E.E.; Backus, E.A. Anterior foregut microbiota of the glassy-winged sharpshooter explored using deep 16S rRNA gene sequencing from individual insects. PLoS ONE 2014, 9, e106215. [Google Scholar] [CrossRef] [PubMed]
- Dourado, M.N.; Camargo Neves, A.A.; Santos, D.S.; Araujo, W.L. Biotechnological and agronomic potential of endophytic pink-pigmented methylotrophic Methylobacterium spp. Biomed. Res. Int. 2015, 2015, 909016. [Google Scholar] [CrossRef] [PubMed]
- Bogas, A.C.; Aguilar-Vildoso, C.I.; Camargo Neves, A.A.; Araujo, W.L. Effects of growth-promoting endophytic Methylobacterium on development of Citrus rootstocks. Afr. J. Microbiol. Res. 2016, 10, 646–653. [Google Scholar]
- Madhaiyan, M.; Suresh Reddy, B.V.; Anandham, R.; Senthilkumar, M.; Poonguzhali, S.; Sundaram, S.P.; Sa, T. Plant growth-promoting Methylobacterium induces defense responses in groundnut (Arachis hypogaea L.) compared with rot pathogens. Curr. Microbiol. 2006, 53, 270–276. [Google Scholar] [CrossRef]
- Tani, A.; Sahin, N.; Kimbara, K. Methylobacterium oxalidis sp. nov., isolated from leaves of Oxalis corniculata. Int. J. Syst. Evol. Microbiol. 2012, 62, 1647–1652. [Google Scholar] [CrossRef] [PubMed]
- Jirakkakul, J.; Khoiri, A.N.; Duangfoo, T.; Dulsawat, S.; Sutheeworapong, S.; Petsong, K.; Wattanachaisaereekul, S.; Paenkaew, P.; Tachaleat, A.; Cheevadhanarak, S.; et al. Insights into the genome of sp. NMS14P, a novel bacterium for growth promotion of maize, chili, and sugarcane. PloS ONE 2023, 18, e0281505. [Google Scholar] [CrossRef]
- Vera, R.T.; García, A.J.B.; Alvarez, F.J.C.; Ruiz, J.M.; Martín, F.F. Application and effectiveness of as a biological inoculant in maize and strawberry crops. Folia Microbiol. 2023. [Google Scholar] [CrossRef]
- Trotsenko, Y.A.; Ianova, E.G.; Doronina, N.V. Aerobic methylotrophic bacteria as phytosymbionts. Microbiology 2001, 70, 623–632. [Google Scholar] [CrossRef]
- Meena, K.K.; Kumar, M.; Kalyuzhnaya, M.G.; Yandigeri, M.S.; Singh, D.P.; Saxena, A.K.; Arora, D.K. Epiphytic pink-pigmented methylotrophic bacteria enhance germination and seedling growth of wheat (Triticum aestivum) by producing phytohormone. Antonie Van. Leeuwenhoek 2012, 101, 777–786. [Google Scholar] [CrossRef] [PubMed]
- Cervantes-Martínez, J.; Lopez-Díaz, S.; Rodriguez-Garay, B. Detection of the effects of Methylobacterium in Agave tequilana Weber var. azul by laser-induced fluorescence. Plant Sci. 2004, 166, 889–892. [Google Scholar] [CrossRef]
- Lee, H.S.; Madhaiyan, C.W.K.; Choi, S.J.; Chung, K.Y.; Sa, T.M. Physiological enhancement of early growth of rice seedlings (Oryza sativa L.) by production of phytohormone of N2-fixing methylotrophic isolates. Biol. Fertil. Soils 2006, 42, 402–408. [Google Scholar] [CrossRef]
- Van Dien, S.J.; Marx, C.J.; O’brien, B.N.; Listrom, M. Genetic characterization of the carotenoid biosynthetic pathway in Methylobacterium extorquens AM1 and isolation of a colorless mutant. Appl. Environ. Microbiol. 2003, 69, 7563–7566. [Google Scholar] [CrossRef]
- Toyama, H.; Anthony, C.; Lidstrom, M.E. Construction of insertion and deletion mxa mutants of Methylobacterium extorquens AM1 by electroporation. FEMS Microbiol. Lett. 1998, 166, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Dourado, M.N.; Bogas, A.C.; Pomini, A.M.; Andreote, F.D.; Quecine, M.C.; Marsaioli, A.J.; Araujo, W.L. Methylobacterium-plant interaction genes regulated by plant exudate and quorum sensing molecules. Braz. J. Microbiol. 2013, 44, 1331–1339. [Google Scholar] [CrossRef]
- Araujo, W.L.; Santos, D.S.; Dini-Andreote, F.; Salgueiro-Londono, J.K.; Camargo-Neves, A.A.; Andreote, F.D.; Dourado, M.N. Genes related to antioxidant metabolism are involved in Methylobacterium mesophilicum-soybean interaction. Antonie Van. Leeuwenhoek 2015, 108, 951–963. [Google Scholar] [CrossRef] [PubMed]
- Li, W.B.; Zreik, L.; Fernandes, N.G.; Miranda, V.S.; Teixeira, D.C.; Ayres, A.J.; Garnier, M.; Bov, J.M. A triply cloned strain of xylella fastidiosa multiplies and induces symptoms of citrus variegated chlorosis in sweet orange. Curr. Microbiol. 1999, 39, 106–108. [Google Scholar] [CrossRef]
- Davis, M.J.; French, W.J.; Schaad, N.W. Axenic culture of the bacteria associated with phony disease of peach and plum leaf scald. Curr. Microbiol. 1981, 6, 309–314. [Google Scholar] [CrossRef]
- Pooler, M.R.; Hartung, J.S. Specific PCR detection and identification of Xylella fastidiosa strains causing citrus variegated chlorosis. Curr. Microbiol. 1995, 31, 377–381. [Google Scholar] [CrossRef] [PubMed]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glockner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Andrews, S.; Lindenbaum, P.; Howard, B.; Ewels, P. FastQC High Throughput Sequence QC Report; 0.11.2; Babraham Bioinformatics: Cambridge, UK, 2014. [Google Scholar]
- Mortazavi, A.; Williams, B.A.; McCue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Conesa, A.; Gotz, S.; Garcia-Gomez, J.M.; Terol, J.; Talon, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef]
- Craig, R.; Beavis, R.C. TANDEM: Matching proteins with tandem mass spectra. Bioinformatics 2004, 20, 1466–1467. [Google Scholar] [CrossRef] [PubMed]
- Petersen, T.N.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef] [PubMed]
- Bendtsen, J.D.; Kiemer, L.; Fausboll, A.; Brunak, S. Non-classical protein secretion in bacteria. BMC Microbiol. 2005, 5, 58. [Google Scholar] [CrossRef]
- Zhao, S.; Ye, Z.; Stanton, R. Misuse of RPKM or TPM normalization when comparing across samples and sequencing protocols. RNA 2020, 26, 903–909. [Google Scholar] [CrossRef] [PubMed]
- Python-Core-Team. Python: A Dynamic, Open Source Programming Language; Python-Core-Team: Beaverton, OR, USA, 2015. [Google Scholar]
- Mantel, N. The detection of disease clustering and a generalized regression approach. Cancer Res. 1967, 27, 209–220. [Google Scholar]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Ludvigsen, M.; Honoré, B. Transcriptomics and Proteomics: Integration? In Encyclopedia of Life Sciences; John Wiley: Hoboken, NJ, USA, 2018; pp. 1–7. [Google Scholar]
- Rinschen, M.M.; Gödel, M.; Grahammer, F.; Zschiedrich, S.; Helmstädter, M.; Kretz, O.; Zarei, M.; Braun, D.A.; Dittrich, S.; Pahmeyer, C.; et al. A Multi-layered Quantitative Expression Atlas of the Podocyte Unravels Kidney Disease Candidate Genes. Cell Rep. 2018, 23, 2495–2508. [Google Scholar] [CrossRef]
- Stubbendieck, R.M.; Straight, P.D. Multifaceted Interfaces of Bacterial Competition. J. Bacteriol. 2016, 198, 2145–2155. [Google Scholar] [CrossRef]
- Alteri, C.J.; Mobley, H.L.T. The Versatile Type VI Secretion System. Microbiol. Spectr. 2016, 4, 337–356. [Google Scholar] [CrossRef] [PubMed]
- Kazmierczak, B.I. The Enemy of my Enemy: Bacterial Competition in the Cystic Fibrosis Lung. Cell Host Microbe 2020, 28, 502–504. [Google Scholar] [CrossRef] [PubMed]
- Lefevre, P.; Braibant, M.; de Wit, L.; Kalai, M.; Roeper, D.; Grotzinger, J.; Delville, J.P.; Peirs, P.; Ooms, J.; Huygen, K.; et al. Three different putative phosphate transport receptors are encoded by the Mycobacterium tuberculosis genome and are present at the surface of Mycobacterium bovis BCG. J. Bacteriol. 1997, 179, 2900–2906. [Google Scholar] [CrossRef]
- Roberts, I.S. The biochemistry and genetics of capsular polysaccharide production in bacteria. Annu. Rev. Microbiol. 1996, 50, 285–315. [Google Scholar] [CrossRef]
- Wang, N.; Li, J.L.; Lindow, S.E. RpfF-dependent regulon of Xylella fastidiosa. Phytopathology 2012, 102, 1045–1053. [Google Scholar] [CrossRef] [PubMed]
- Gambetta, G.A.; Matthews, M.A.; Syvanen, M. The Xylella fastidosa RTX operons: Evidence for the evolution of protein mosaics through novel genetic exchanges. BMC Genom. 2018, 19, 329. [Google Scholar] [CrossRef]
- Linhartova, I.; Bumba, L.; Masin, J.; Basler, M.; Osicka, R.; Kamanova, J.; Prochazkova, K.; Adkins, I.; Hejnova-Holubova, J.; Sadilkova, L.; et al. RTX proteins: A highly diverse family secreted by a common mechanism. FEMS Microbiol. Rev. 2010, 34, 1076–1112. [Google Scholar] [CrossRef]
- Kato, S.; Kowashi, Y.; Demuth, D.R. Outer membrane-like vesicles secreted by Actinobacillus actinomycetemcomitans are enriched in leukotoxin. Microb. Pathog. 2002, 32, 1–13. [Google Scholar] [CrossRef]
- Kulp, A.; Kuehn, M.J. Biological functions and biogenesis of secreted bacterial outer membrane vesicles. Annu. Rev. Microbiol. 2010, 64, 163–184. [Google Scholar] [CrossRef]
- Siljamaki, P.; Varmanen, P.; Kankainen, M.; Sukura, A.; Savijoki, K.; Nyman, T.A. Comparative exoprotein profiling of different Staphylococcus epidermidis strains reveals potential link between nonclassical protein export and virulence. J. Proteome Res. 2014, 13, 3249–3261. [Google Scholar] [CrossRef]
- Perez-Cruz, C.; Delgado, L.; Lopez-Iglesias, C.; Mercade, E. Outer-inner membrane vesicles naturally secreted by gram-negative pathogenic bacteria. PLoS ONE 2015, 10, e0116896. [Google Scholar] [CrossRef]
- Schwechheimer, C.; Kuehn, M.J. Outer-membrane vesicles from Gram-negative bacteria: Biogenesis and functions. Nat. Rev. Microbiol. 2015, 13, 605–619. [Google Scholar] [CrossRef] [PubMed]
- Huberts, D.H.; van der Klei, I.J. Moonlighting proteins: An intriguing mode of multitasking. Biochim. Biophys. Acta 2010, 1803, 520–525. [Google Scholar] [CrossRef] [PubMed]
- Jeffery, C.J. Why study moonlighting proteins? Front. Genet. 2015, 6, 211. [Google Scholar] [CrossRef]
- Wang, W.; Jeffery, C.J. An analysis of surface proteomics results reveals novel candidates for intracellular/surface moonlighting proteins in bacteria. Mol. Biosyst. 2016, 12, 1420–1431. [Google Scholar] [CrossRef]
- Caldas, T.D.; El Yaagoubi, A.; Richarme, G. Chaperone properties of bacterial elongation factor EF-Tu. J. Biol. Chem. 1998, 273, 11478–11482. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Wardhan, V.; Kumar, A.; Rathi, D.; Pandey, A.; Chakraborty, S.; Chakraborty, N. Secretome analysis of chickpea reveals dynamic extracellular remodeling and identifies a Bet v1-like protein, CaRRP1 that participates in stress response. Sci. Rep. 2015, 5, 18427. [Google Scholar] [CrossRef]
- Neckers, L. Molecular Chaperones in Pathogen Virulence: Emerging New Targets for Therapy. Cell Host Microbe 2009, 4, 519–527. [Google Scholar] [CrossRef] [PubMed]
- Ghazaei, C. Molecular chaperones impacts in growth, metabolism, pathogenicity and production of virulence factors in bacterial pathogen. Rev. Med. Microbiol. 2018, 29, 24–29. [Google Scholar] [CrossRef]
- Evans, A.G.L.; Davey, H.M.; Cookson, A.; Currinn, H.; Cooke-Fox, G.; Stanczyk, P.J.; Whitworth, D.E. Predatory activity of Myxococcus xanthus outer-membrane vesicles and properties of their hydrolase cargo. Microbiology 2012, 158, 2742–2752. [Google Scholar] [CrossRef] [PubMed]
- Ingel, B.; Castro, C.; Burbank, L.; Her, N.; De Anda, N.I.; Way, H.; Wang, P.; Roper, C. Xylella fastidiosa requires the Type II secretion system for pathogenicity and survival in grapevine. Mol. Plant Microbe Interact. 2023, 36. [Google Scholar] [CrossRef]



| Treatment | Number of Reads | Mapped Reads in Pairs | % Mapped Reads | Unique Fragments |
|---|---|---|---|---|
| Xf monoculture | 4,400,994 | 3,523,819 | 80.16 | 1,759,574 |
| Co-culture mapped in Xf genome | 4,410,865 | 2,284,762 | 51.05 | 1,140,964 |
| Co-culture mapped in Mm genome | 4,410,865 | 1,158,502 | 26.41 | 578,282 |
| Mm monoculture | 3,981,063 | 2,892,542 | 71.45 | 1,444,234 |
| MC | CC | CDS Number IMG Xf * | CDS Number NCBI Xf * | Description and Symbol ** | Predicted Molecular Mass (kDa) | Sequence Coverage (%) | SignalP | SecP Score |
|---|---|---|---|---|---|---|---|---|
| 1 | 3 | XF9a_01687 | XF_1803 | Omp1X | 21 | 26 | 20|21 | |
| 2 | 1 | XF9a_01416 | XF_1547 | Membrane lipoprotein Lpp | 16 | 73 | 17|18 | |
| 3 | 2 | XF9a_00315 | XF_0343 | Outer membrane protein mopB | 42 | 28 | 0.94 | |
| 4 | 11 | XF9a_00326 | XF_0363 | Outer membrane protein OmpA | 26 | 33 | 27|28 | |
| 5 | 4 | XF9a_00948 | XF_1026 | Serine protease PspB | 95 | 18 | 0.95 | |
| 6 | 7 | XF9a_01869 | XF_1981 | Adhesin XadA3 | 118 | 23 | 0.95 | |
| 7 | 12 | XF9a_00900 | XF_0975 | PorinO OprP | 44 | 35 | 21|22 | |
| 8 | 6 | XF9a_00602 | XF_0668 | Hemolysin toxin protein RTX | 128 | 22 | 0.74 | |
| 9 | 13 | XF9a_00831 | XF_0898 | Membrane lipoprotein | 13 | 53 | ||
| 10 | 9 | XF9a_02226 | XF_2349 | Autotransporter beta-domain | 81 | 26 | 0.85 | |
| 11 | 5 | XF9a_01123 | XF_1219 | Protein of Unknown Function | 9 | 23 | ||
| 12 | 10 | XF9a_02129 | XF_2237 | TonB-dependent receptor | 103 | 21 | 31|32 | |
| 13 | - | XF9a_01121 | XF_1217 | Protein of Unknown Function | 9 | 17 | ||
| 14 | 8 | XF9a_00936 | XF_1011 | Hemolysin toxin protein RTX | 173 | 5 | 0.57 | |
| 15 | 14 | XF9a_00892 | XF_0964 | Membrane lipoprotein | 19 | 24 | 0.56 | |
| 16 | 15 | XF9a_01736 | XF_1851 | Serine protease | 105 | 10 | 0.95 | |
| 17 | - | XF9a_01786 | XF_1896 | Outer membrane protein OmpA | 20 | 10 | 27|28 | |
| 18 | 21 | XF9a_00582 | XF_0644 | Peptidylprolyl isomerase Fkbp | 25 | 18 | 19|20 | |
| 19 | 19 | XF9a_00514 | XF_0565 | Lipid-binding SYLF | 32 | 13 | 22|23 | |
| 20 | 16 | XF9a_02272 | XF_2407 | Hemolysin toxin protein RTX | 219 | 5 | 0.69 | |
| 21 | 18 | XF9a_02555 | XF_2713 | Porin CirA | 96 | 15 | 24|25 | |
| 22 | 29 | XF9a_02447 | XF_2586 | Outer membrane export factor TolC | 49 | 23 | 21|22 | |
| 23 | 17 | XF9a_00557 | XF_0615 | Chaperonin GroEL | 58 | 16 | ||
| 24 | 20 | XF9a_01776 | XF_1887 | Cysteine/serine peptidase PS-46 | 79 | 16 | 22|23 | |
| 25 | 26 | XF9a_01787 | XF_1897 | protein TolB | 48 | 17 | 0.78 | |
| 26 | - | XF9a_02412 | XF_2548 | Succinyl-CoA ligase SucD | 30 | 21 | 0.64 | |
| 27 | 30 | XF9a_00323 | XF_0357 | Lipase/Esterase LesA | 42 | 15 | 0.90 | |
| 28 | - | XF9a_01186 | XF_1297 | Gluconolactonase | 37 | 5 | 25|26 | |
| 29 | 23 | XF9a_01725 | XF_1840 | Protein of Unknown Function | 25 | 7 | 0.90 | |
| 30 | - | XF9a_01712 | XF_1827 | Organic hydroperoxide reductase OsmC/OhrA | 15 | 20 | ||
| 31 | - | XF9a_00558 | XF_0616 | Chaperonin GroES | 10 | 39 | ||
| 32 | - | XF9a_01475 | XF_1604 | Glutathione peroxidase | 21 | 16 | 27|28 | |
| 33 | - | XF9a_00607 | XF_0672 | Acyl carrier protein AcpP | 9 | 24 | ||
| 34 | 22 | XF9a_00073 | XF_0082 | Fimbrial chaperone protein PapD | 29 | 15 | 30|31 | |
| 35 | - | XF9a_00777 | XF_0855 | Lipoprotein NlpD | 26 | 8 | 26|27 | |
| 36 | - | XF9a_00319 | XF_0353 | Translation initiation inhibitor | 14 | 18 | ||
| 37 | 25 | XF9a_02217 | XF_2340 | Chaperone protein DnaK | 68 | 15 | ||
| 38 | - | XF9a_01697 | XF_1811 | Outer membrane protein Slp | 18 | 17 | 0.91 | |
| 39 | - | XF9a_01043 | XF_1133 | Tryptophan repressor binding protein WrbA | 20 | 10 | 0.92 | |
| 40 | 24 | XF9a_01514 | XF_1649 | Protein of Unknown Function | 30 | 11 | ||
| 41 | - | XF9a_00742 | XF_0816 | Zn-dependent peptidase | 108 | 4 | ||
| 42 | - | XF9a_00954 | XF_1036 | Porin | 111 | 9 | 33|34 | |
| 43 | 27 | XF9a_00221 | XF_0239 | Polyribonucleotide nucleotidyltransferase Pnp | 76 | 7 | ||
| 44 | 28 | XF9a_00502 | XF_0550 | TonB-dependent receptor | 114 | 4 | 34|35 | |
| 45 | - | XF9a_02478 | XF_2622 | Cold shock protein, CspA | 9 | 21 | 0.89 | |
| 46 | - | XF9a_02496 | XF_2640 | Elongation factor Tu tufA | 43 | 16 | ||
| 47 | - | XF9a_01153 | XF_1253 | Acetyl esterase/lipase | 35 | 6 | 0.59 | |
| 48 | - | XF9a_00069 | XF_0078 | Fimbrial protein MrkD | 37 | 6 | 37|38 | |
| 49 | - | XF9a_02168 | XF_2283 | Beta-lactamase-like | 34 | 8 | 23|24 | |
| 50 | - | XF9a_01476 | XF_1605 | Fkbp-type peptidyl-prolyl cis-trans isomerase FkpA | 32 | 7 | 0.74 | |
| 51 | - | XF9a_00181 | XF_0196 | DUF2059 | 20 | 20 | 26|27 | |
| 52 | - | XF9a_00120 | XF_0138 | Aminopeptidase PepA | 52 | 12 | ||
| 53 | - | XF9a_00760 | XF_0838 | Chaperone SurA | 51 | 5 | 25|26 | |
| 54 | 32 | XF9a_00744 | XF_0820 | Zn-dependent amino- or carboxypeptidase | 58 | 9 | 0.52 | |
| 55 | - | XF9a_01399 | XF_1527 | Type II protein GspD/PulD | 81 | 3 | 0.95 | |
| 56 | - | XF9a_02411 | XF_2547 | Succinate--CoA ligase subunit beta SucC | 41 | 7 | ||
| 57 | - | XF9a_01116 | XF_1211 | Malate dehydrogenase Mdh | 35 | 8 | ||
| 58 | - | XF9a_01785 | XF_1895 | Tol-pal System protein YbgF | 30 | 8 | 23|24 | |
| 59 | - | XF9a_00964 | XF_1046 | Outer membrane protein BamA | 88 | 6 | 26|27 | |
| 60 | - | XF9a_01449 | XF_1577 | DUF2184 | 37 | 6 | 0.53 | |
| 61 | 31 | XF9a_02619 | XF_2773 | Hemagglutinin HxfA | 361 | 1 | 0.96 | |
| 62 | - | XF9a_00008 | XF_0007 | Protein of Unknown Function | 44 | 6 | 0.75 | |
| 63 | - | XF9a_00264 | XF_0290 | Aconitase | 98 | 3 | 0.78 | |
| 64 | - | XF9a_00072 | XF_0081 | Outer membrane FimD | 98 | 2 | 0.91 |
| MC | CC | CDS Number Mb * | CDS Number “Old NCBI Locustag” Mb * | Description and Symbol ** | Predicted Molecular Mass kDa | Sequence Coverage (%) | SignalP | SecP Score |
|---|---|---|---|---|---|---|---|---|
| 1 | - | MMSR116_RS18845 | MMSR116_19085 | Flagellin | 41 | 16 | 0.97 | |
| 2 | 2 | MMSR116_RS23965 | MMSR116_24295 | Cytochrome c class I | 13 | 15 | 22|23 | |
| 3 | - | MMSR116_RS28515 | MMSR116_28855 | Uncharacterized protein | 12 | 10 | 24|25 | |
| 4 | 3 | MMSR116_RS16505 | MMSR116_16705 | Formaldehyde-activating enzyme | 18 | 8 | 0.82 | |
| 5 | 10 | MMSR116_RS16515 | MMSR116_16715 | Chaperonin GroES | 11 | 21 | ||
| 6 | - | MMSR116_RS22410 | MMSR116_22715 | Uncharacterized protein | 17 | 19 | 25|26 | |
| 7 | - | MMSR116_RS23130 | MMSR116_23445 | Peptidyl-prolyl cis-trans isomerase | 19 | 19 | 22|23 | |
| 8 | 9 | MMSR116_RS12040 | MMSR116_12205 | Thioredoxin Trx | 12 | 20 | 0.73 | |
| 9 | 5 | MMSR116_RS00465 | MMSR116_00470 | Porin | 29 | 19 | 20|21 | |
| 10 | - | MMSR116_RS24205 | MMSR116_24535 | Uncharacterized protein | 15 | 14 | 23|24 | |
| 11 | 8 | MMSR116_RS12140 | MMSR116_12305 | Uncharacterized protein | 10 | 12 | 23|24 | |
| 12 | 22 | MMSR116_RS25265 | MMSR116_25605 | Elongation factor Tuf1 | 43 | 6 | ||
| 13 | - | MMSR116_RS13860 | MMSR116_14035 | Signal peptide protein | 19 | 20 | 21|22 | |
| 14 | 6 | MMSR116_RS20185 | MMSR116_20450 | Extracellular solute-binding protein | 33 | 23 | 27|28 | |
| 15 | 12 | MMSR116_RS27490 | MMSR116_27830 | Multiple sugar-binding periplasmic receptor ChvE | 38 | 15 | 27|28 | |
| 16 | 7 | MMSR116_RS07030 | MMSR116_07145 | Glyceraldehyde-3-phosphate dehydrogenase | 36 | 13 | ||
| 17 | 4 | MMSR116_RS11970 | MMSR116_12135 | Cysteine synthase A | 34 | 14 | ||
| 18 | - | MMSR116_RS17175 | MMSR116_17365 | Peptidase PepSY | 20 | 21 | 21|22 | |
| 19 | 13 | MMSR116_RS09940 | MMSR116_10105 | Chaperone protein DnaK | 69 | 10 | ||
| 20 | - | MMSR116_RS16510 | MMSR116_16710 | Chaperone protein GroEL | 58 | 20 | ||
| 21 | 1 | MMSR116_RS07550 | MMSR116_07665 | Extracellular ligand-binding receptor | 39 | 26 | 20|21 | |
| 22 | 14 | MMSR116_RS09540 | MMSR116_09680 | NAD-binding 6-phosphogluconate dehydrogenase | 30 | 14 | ||
| 23 | - | MMSR116_RS26210 | MMSR116_26565 | Catalase-related peroxidase | 35 | 8 | 19|20 | |
| 24 | - | MMSR116_RS09370 | MMSR116_09510 | Superoxide dismutase | 23 | 9 | 0.51 | |
| 25 | - | MMSR116_RS25335 | MMSR116_25675 | Transcription elongation factor GreA | 16 | 13 | ||
| 26 | - | MMSR116_RS18790 | MMSR116_19035 | Uncharacterized protein | 8 | 28 | 23|24 | |
| 27 | - | MMSR116_RS03745 | MMSR116_03785 | Uncharacterized protein | 23 | 7 | 23|24 | |
| 28 | - | MMSR116_RS24405 | MMSR116_24735 | Transcriptional regulator, MucR family | 18 | 10 | 0.96 | |
| 29 | - | MMSR116_RS12930 | MMSR116_13100 | Uncharacterized protein | 22 | 9 | 27|28 | |
| 30 | - | MMSR116_RS09955 | MMSR116_10120 | Redoxin | 17 | 11 | ||
| 31 | - | MMSR116_RS23810 | MMSR116_24135 | Citrate synthase | 48 | 3 | ||
| 32 | - | MMSR116_RS14870 | MMSR116_15050 | Uncharacterized protein | 17 | 16 | 36|37 | |
| 33 | - | MMSR116_RS01755 | MMSR116_01770 | Alanine racemase domain-containing protein | 40 | 12 | ||
| 34 | 15 | MMSR116_RS14980 | MMSR116_15160 | Phosphoenolpyruvate carboxykinase PckA | 59 | 6 | ||
| 35 | 18 | MMSR116_RS20370 | MMSR116_20635 | Malate dehydrogenase Mdh | 34 | 7 | ||
| 36 | - | MMSR116_RS20190 | MMSR116_20455 | branched-chain amino acid ABC transporter substrate-binding protein | 43 | 9 | 25|26 | |
| 37 | 20 | MMSR116_RS00865 | MMSR116_00880 | Methanol/ethanol family PQQ-dependent dehydrogenase | 69 | 3 | 24|25 | |
| 38 | 19 | MMSR116_RS20480 | MMSR116_20745 | Polyribonucleotide nucleotidyltransferase Pnp | 80 | 3 | ||
| 39 | 23 | MMSR116_RS29630 | MMSR116_29970 | Ketol-acid reductoisomerase IlvC | 37 | 5 | ||
| 40 | - | MMSR116_RS11180 | MMSR116_11330 | Adenylosuccinate synthetase PurA | 48 | 6 | ||
| 41 | - | MMSR116_RS21010 | MMSR116_21290 | Methylenetetrahydrofolate dehydrogenase | 29 | 8 | 0.81 | |
| 42 | - | MMSR116_RS05635 | MMSR116_05745 | 30S ribosomal protein S1 RpsA | 63 | 3 | ||
| 43 | - | MMSR116_RS03750 | MMSR116_03790 | Aconitate hydratase | 97 | 3 | 0.60 | |
| - | 11 | MMSR116_RS21075 *** | MMSR116_21360 | Inorganic pyrophosphatase Ppa | 20 | 8 | ||
| 44 | - | MMSR116_RS13495 | MMSR116_13665 | Pyruvate, phosphate dikinase | 97 | 2 | ||
| - | 16 | MMSR116_RS19390 *** | MMSR116_19640 | Adenosylhomocysteinase AhcY | 51 | 5 | ||
| - | 17 | MMSR116_RS19735 *** | MMSR116_19990 | Phosphate-binding protein PstS | 37 | 6 | 23|24 | |
| - | 22 | **** | MMSR116_18145 *** | Porin | 59 | 3 | 0.89 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dourado, M.N.; Pierry, P.M.; Feitosa-Junior, O.R.; Uceda-Campos, G.; Barbosa, D.; Zaini, P.A.; Dandekar, A.M.; da Silva, A.M.; Araújo, W.L. Transcriptome and Secretome Analyses of Endophyte Methylobacterium mesophilicum and Pathogen Xylella fastidiosa Interacting Show Nutrient Competition. Microorganisms 2023, 11, 2755. https://doi.org/10.3390/microorganisms11112755
Dourado MN, Pierry PM, Feitosa-Junior OR, Uceda-Campos G, Barbosa D, Zaini PA, Dandekar AM, da Silva AM, Araújo WL. Transcriptome and Secretome Analyses of Endophyte Methylobacterium mesophilicum and Pathogen Xylella fastidiosa Interacting Show Nutrient Competition. Microorganisms. 2023; 11(11):2755. https://doi.org/10.3390/microorganisms11112755
Chicago/Turabian StyleDourado, Manuella Nobrega, Paulo Marques Pierry, Oseias Rodrigues Feitosa-Junior, Guillermo Uceda-Campos, Deibs Barbosa, Paulo A. Zaini, Abhaya M. Dandekar, Aline Maria da Silva, and Welington Luiz Araújo. 2023. "Transcriptome and Secretome Analyses of Endophyte Methylobacterium mesophilicum and Pathogen Xylella fastidiosa Interacting Show Nutrient Competition" Microorganisms 11, no. 11: 2755. https://doi.org/10.3390/microorganisms11112755
APA StyleDourado, M. N., Pierry, P. M., Feitosa-Junior, O. R., Uceda-Campos, G., Barbosa, D., Zaini, P. A., Dandekar, A. M., da Silva, A. M., & Araújo, W. L. (2023). Transcriptome and Secretome Analyses of Endophyte Methylobacterium mesophilicum and Pathogen Xylella fastidiosa Interacting Show Nutrient Competition. Microorganisms, 11(11), 2755. https://doi.org/10.3390/microorganisms11112755








