Molecular Identification and Pathogenicity of Diaporthe eres and D. hongkongensis (Diaporthales, Ascomycota) Associated with Cherry Trunk Diseases in China
Abstract
:1. Introduction
2. Materials and Methods
2.1. Field Surveys and Isolation of Fungi
2.2. Molecular Identification of Fungi
2.2.1. Fungal DNA Sequencing
2.2.2. Phylogenetic Analysis
2.3. Morphological Characterization
2.4. Temperature Growth Studies
2.5. Pathogenicity Study
3. Results
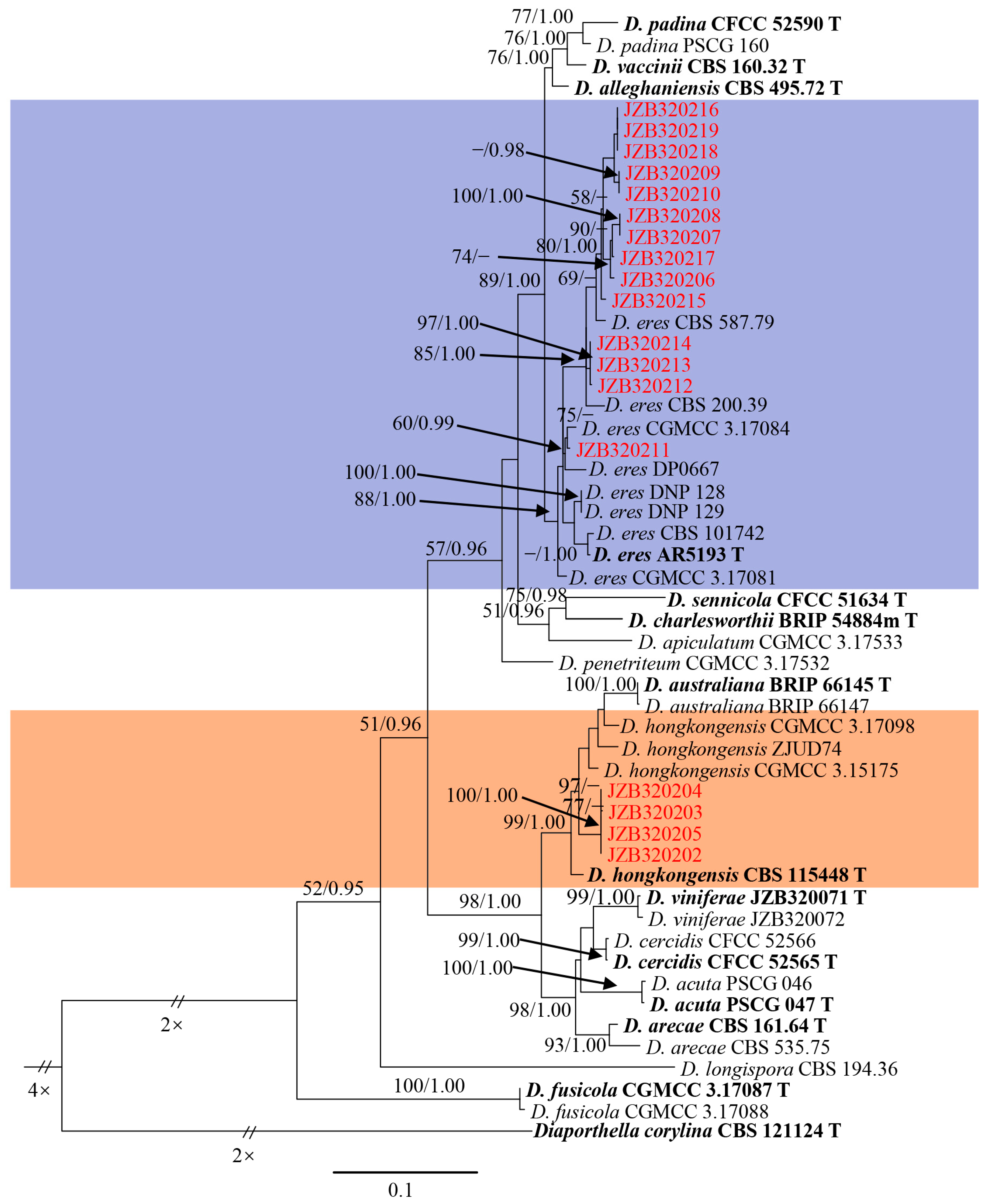
3.1. Molecular Identification and Phylogenetic Analysis
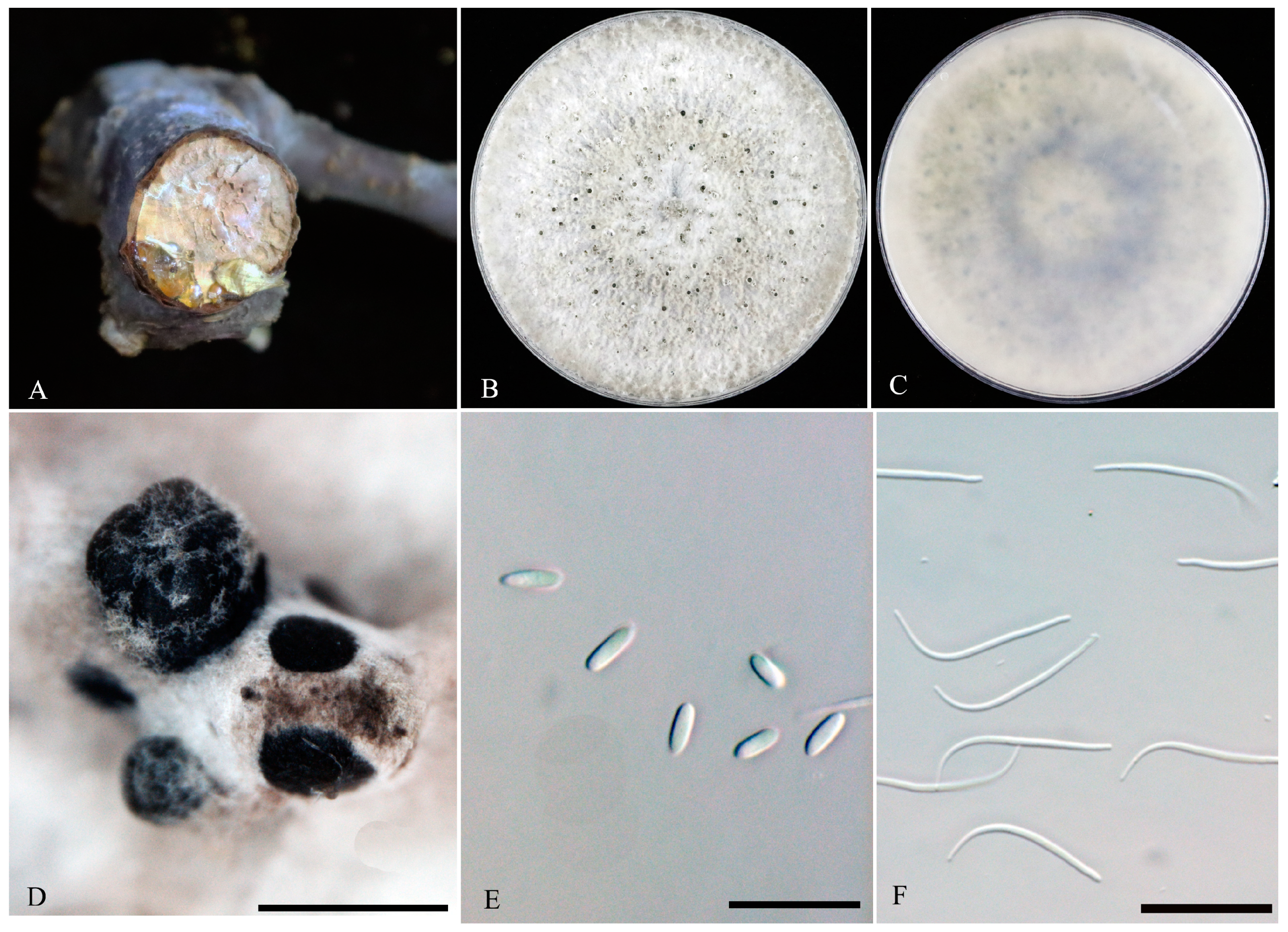
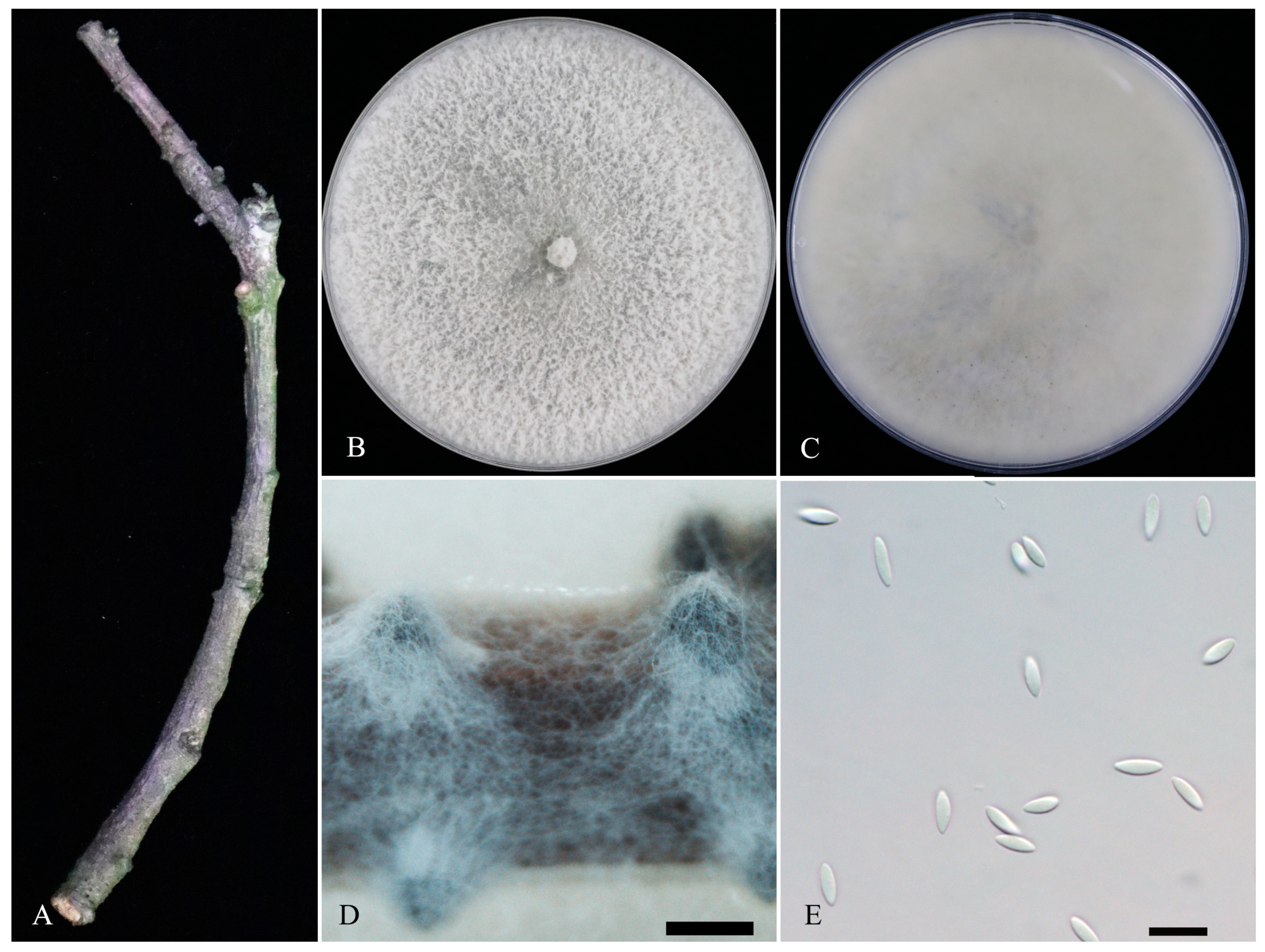
3.2. Morphological Characterization
3.3. Taxonomy
3.4. Pathogenicity Study
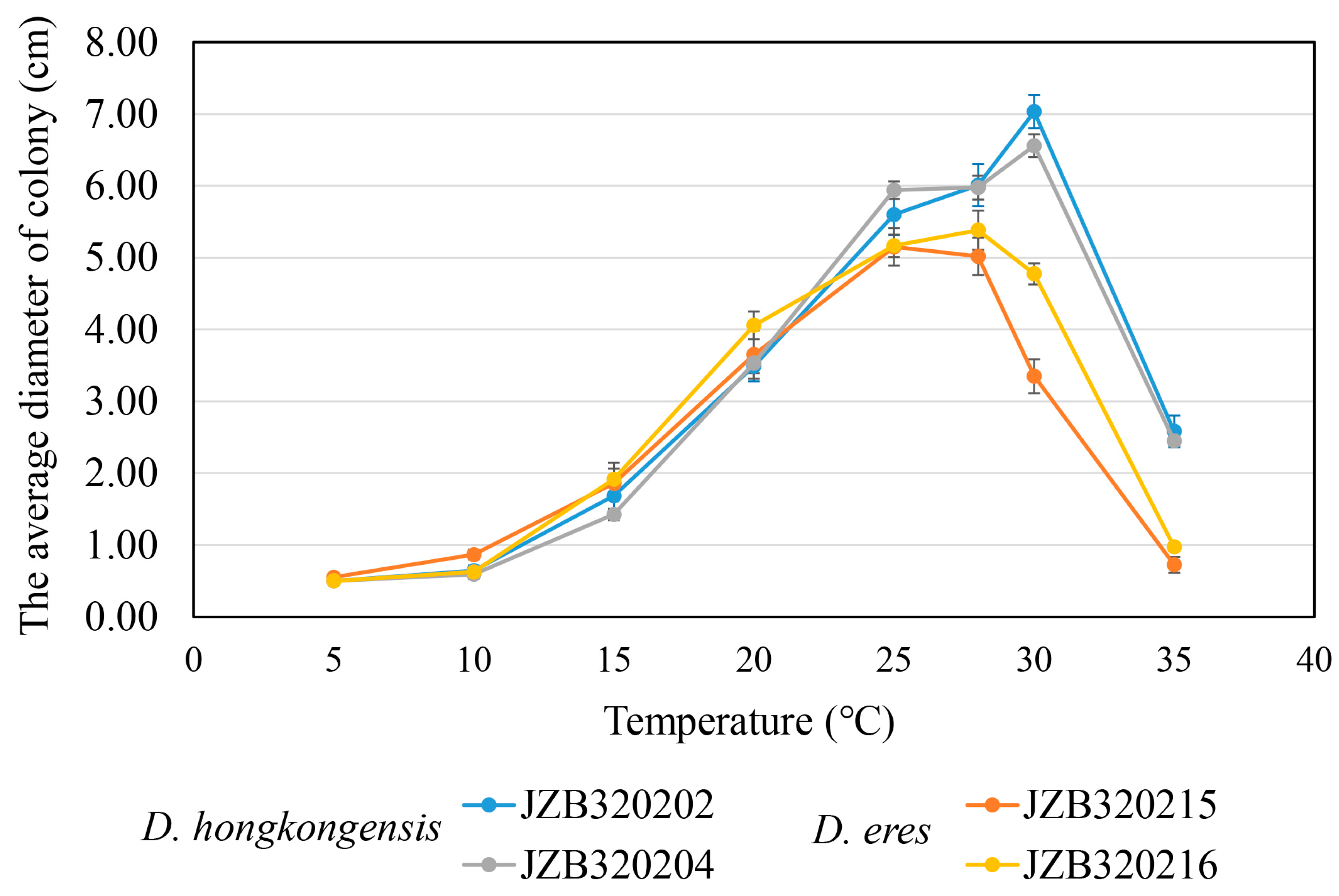
3.5. Temperature Growth Studies
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chethana, K.W.T.; Jayawardene, R.S.; Zhang, W.; Zhou, Y.Y.; Liu, M.; Hyde, K.D.; Li, X.H.; Wang, J.; Zhang, K.C.; Yan, J.Y. Molecular characterization and pathogenicity of fungal taxa associated with cherry leaf spot disease. Mycosphere 2019, 10, 490–530. [Google Scholar] [CrossRef]
- Kumar, A.; Sharma, M.K.; Wani, T.F.; Sharma, A.; Nyorak, G. Varietal Wealth of Prunus Species. In Prunus-Recent Advances; IntechOpen: Rijeka, Croatia, 2021. [Google Scholar]
- Zhang, Q.J.; Gu, D.J.; Yu, K.H.; Zhou, Z.H. A model system for off season cherry production in northern China. Acta Hortic. 2018, 1208, 221–226. [Google Scholar] [CrossRef]
- Zhang, K.; Yan, G.; Zhang, X.; Wang, J.; Duan, X. Sweet cherry growing in China. Acta Hortic. 2019, 1235, 133–140. [Google Scholar] [CrossRef]
- Chen, H.; Zhang, K.C.; Zhou, J.Z.; Zhang, Y.Y.; Ding, C.X.; Zhang, W. Global cherry production status and suggestions for cherry industry development of Beijing. Deciduous Fruits 2020, 52, 27–30. [Google Scholar]
- Uyemoto, J.K.; Ogawa, J.M.; Jaffee, B.A. Common Names of Plant Diseases: Diseases of Sweet Cherry (Prunus avium L.) and Sour Cherry (P. cerasus L.); The American Phyto-Pathological Society: St. Paul, MN, USA, 2018. [Google Scholar]
- Cooke, T.; Persley, D.; House, S. Diseases of Fruit Crops in Australia; Csiro Publishing: Collingwood, Australia, 2009. [Google Scholar]
- Gramaje, D.; Agustí-Brisach, C.; Pérez-Sierra, A.; Morale-jo, E.; Olmo, D.; Mostert, L.; Damm, U.; Armengol, J. Fungal trunk pathogens associated with wood decay of almond trees on Mallorca (Spain). Pers. Mol. Phylogeny Evol. 2012, 28, 1–13. [Google Scholar] [CrossRef]
- Espargham, N.; Mohammadi, H.; Gramaje, D. Survey of trunk disease pathogens within citrus trees in Iran. Plants 2020, 9, 754. [Google Scholar] [CrossRef]
- Sohrabi, M.; Mohammadi, H.; León, M.; Armengol, J.; Banihashemi, Z. Fungal pathogens associated with branch and trunk cankers of nut crops in Iran. Eur. J. Plant Pathol. 2020, 157, 327–351. [Google Scholar] [CrossRef]
- Guarnaccia, V.; Kraus, C.; Markakis, E.; Alves, A.; Armengol, J.; Eichmeier, A.; Compant, S.; Gramaje, D. Fungal trunk diseases of fruit trees in Europe: Pathogens, spread and future directions. Phytopathol. Mediterr. 2022, 61, 563–599. [Google Scholar] [CrossRef]
- Slippers, B.; Wingfield, M.J. Botryosphaeriaceae as endophytes and latent pathogens of woody plants: Diversity, ecology and impact. Fungal Biol. Rev. 2007, 21, 90–106. [Google Scholar] [CrossRef]
- Senanayake, I.C.; Rathnayaka, A.R.; Marasinghe, D.S.; Calabon, M.S.; Gentekaki, E.; Lee, H.B.; Hurdeal, V.G.; Pem, D.; Dissanayake, L.S.; Wijesinghe, S.N.; et al. Morphological approaches in studying fungi: Collection, examination, isolation, sporulation and preservation. Mycosphere 2020, 11, 2678–2754. [Google Scholar] [CrossRef]
- Abeywickrama, P.D.; Zhang, W.; Li, X.; Jayawardena, R.S.; Hyde, K.D.; Yan, J. Campylocarpon fasciculare (Nectriaceae, Sordariomycetes); Novel Emergence of Black-Foot Causing Pathogen on Young Grapevines in China. Pathogens 2021, 10, 1555. [Google Scholar] [CrossRef] [PubMed]
- Thesiya, M.R.; Rakholiya, K.B.; Lokesh, R. Isolation, Cultural and Morphological Characterization of Phomopsis vexans (Sacc. and Syd.) Harter, causing Stem Blight and Fruit Rot of Brinjal. Int. J. Curr. Microbiol. App. Sci. 2020, 9, 2851–2859. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. In PCR Protocols; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; Volume 38, pp. 315–322. [Google Scholar]
- Alves, A.; Crous, P.W.; Correia, A.; Phillips, A.J.L. Morphological and molecular data reveal cryptic species in Lasiodiplodia theobromae. Fungal Diver. 2008, 28, 1–13. [Google Scholar]
- Carbone, I.; Kohn, L.M. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999, 93, 553–556. [Google Scholar] [CrossRef]
- Glass, N.L.U.O.; Donaldson, G.C. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microbiol. 1995, 61, 1323–1330. [Google Scholar] [CrossRef] [PubMed]
- Woudenberg, J.H.C.; Aveskamp, M.M.; de Gruyter, J.; Spiers, A.G.; Crous, P.W. Multiple Didymella teleomorphs are linked to the Phoma clematidina morphotype. Persoonia 2009, 22, 56–62. [Google Scholar] [CrossRef] [PubMed]
- Norphanphoun, C.; Gentekaki, E.; Hongsanan, S.; Jayawardena, R.; Senanayake, I.C.; Manawasinghe, I.S.; Abeywickrama, P.D.; Bhunjun, C.S.; Hyde, K.D. Diaporthe: Formalizing the species-group concept. Mycosphere 2022, 13, 752–819. [Google Scholar] [CrossRef]
- Hongsanan, S.; Norphanphoun, C.; Senanayake, I.C.; Jayawardena, R.S.; Manawasinghe, I.S.; Abeywickrama, P.D.; Khuna, S.; Suwannarach, N.; Senwanna, C.; Monkai, J.; et al. Annotated notes on Diaporthe species. Mycosphere 2023, 14, 918–1189. [Google Scholar]
- Guindon, S.; Gascuel, O. A simple, fast and accurate method to estimate large phylogenies by maximum-likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the Gateway Computing Environments Workshop (GCE), New Orleans, LA, USA, 14 November 2010; pp. 1–8. [Google Scholar]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. Mrbayes 3.2: Efficient Bayesian phylogenetic inference and model selection across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed]
- Udayanga, D.; Castlebury, L.A.; Rossman, A.Y.; Chukeatirote, E.; Hyde, K.D. Insights into the genus Diaporthe: Phylogenetic species delimitation in the D. eres species complex. Fungal Divers. 2014, 67, 203–229. [Google Scholar] [CrossRef]
- Abeywickrama, P.D.; Camporesi, E.; Jayawardena, R.S.; Hyde, K.D.; Yan, J.; Zhang, W.; Li, X. Novel and surprising host associations of Diaporthe (Diaporthaceae, Diaporthales) species from Italy. Chiang Mai J. Sci. 2022, 49, 223–247. [Google Scholar] [CrossRef]
- Cline, E.T.; Farr, D.F. Synopsis of fungi listed as regulated plant pests by the USDA Animal and Plant Health Inspection Service: Notes on nomenclature, disease, plant hosts, and geographic distribution. Plant Health Prog. 2006, 7, 1. Available online: http://www.plantmanagementnetwork.org/php/ (accessed on 15 May 2023). [CrossRef]
- Gomes, R.; Glienke, C.; Videira, S.; Lombard, L.; Groenewald, J.Z.; Crous, P.W. Diaporthe: A genus of endophytic, saprobic and plant pathogenic fungi. Pers. Mol. Phylogeny Evol. 2013, 31, 1–41. [Google Scholar] [CrossRef]
- Dissanayake, A.J.; Phillips, A.J.L.; Hyde, K.D.; Yan, J.Y.; Li, X.H. The current status of species in Diaporthe. Mycosphere 2017, 8, 1106–1156. [Google Scholar] [CrossRef]
- Erper, I.; Turkkan, M.; Ozean, M.; Luongo, L.; Belisario, A. Characterization of Diaporthe hongkongensis species causing stem-end rot on kiwifruit in turkey. J. Plant Pathol. 2017, 99, 779–782. [Google Scholar]
- Du, Y.; Wang, X.; Guo, Y.; Xiao, F.; Peng, Y.; Hong, N.; Wang, G. Biological and molecular characterization of seven Diaporthe species associated with kiwifruit shoot blight and leaf spot in China. Phytopathol. Mediterr. 2021, 60, 177–198. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhang, Z.B.; Huang, Y.T.; Wang, F.X.; Hu, W.H.; Dai, L.Y.; Zhong, J.; Liu, Y.; Zhu, J.Z. First report of Diaporthe hongkongensis causing fruit rot on peach (Prunus persica) in China. Plant Dis. 2021, 105, 2017. [Google Scholar] [CrossRef]
- Liao, Y.C.Z.; Sun, J.W.; Li, D.W.; Nong, M.L.; Zhu, L.H. First report of top blight of Cunninghamia lanceolata caused by Diaporthe unshiuensis and Diaporthe hongkongensis in China. Plant Dis. 2023, 107, 962. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.S.; Crous, P.W.; Bai, Q.; Fu, M.; Yang, M.M.; Wang, X.H.; Du, Y.M.; Hong, N.; Xu, W.X.; Wang, G.P. High diversity of Diaporthe species associated with pear shoot canker in China. Pers. Mol. Phylogeny Evol. 2020, 45, 132–162. [Google Scholar] [CrossRef] [PubMed]
- Udayanga, D.; Liu, X.; Crous, P.W.; McKenzie, E.H.C.; Chukeatirote, E.; Hyde, K.D. A multi-locus phylogenetic evaluation of Diaporthe (Phomopsis). Fungal Divers. 2012, 56, 157–171. [Google Scholar] [CrossRef]
- Abeywickrama, P.D.; Wanasinghe, D.N.; Karunarathna, S.C.; Jayawardena, R.S.; Hyde, K.D.; Zhang, W.; Li, X.; Yan, J. A new host report of Diaporthe manihotia (Diaporthales, Ascomycota) from Camellia sp. in Yunnan province, China. Asian J. Mycol. 2020, 3, 473–489. [Google Scholar] [CrossRef]
- Manawasinghe, I.S.; Dissanayake, A.J.; Li, X.; Liu, M.; Wanasinghe, D.N.; Xu, J.; Zhao, W.; Zhang, W.; Zhou, Y.; Hyde, K.D. High Genetic Diversity and Species Complexity of Diaporthe Associated with Grapevine Dieback in China. Front. Microbiol. 2019, 10, 1936. [Google Scholar] [CrossRef]
- Havenga, M.; Gatsi, G.M.; Halleen, F.; Spies, C.F.J.; van der Merwe, R.; Mostert, L. Canker and Wood Rot Pathogens Present in Young Apple Trees and Propagation Material in the Western Cape of South Africa. Plant Dis. 2019, 103, 3129–3141. [Google Scholar] [CrossRef] [PubMed]
- Hilario, S.; Amaral, I.A.; Goncalves, M.; Lopes, A.; Santos, L.; Alves, A. Diaporthe species associated with twig blight and dieback of Vaccinium corymbosum in Portugal, with description of four new species. Mycologia 2020, 112, 293–308. [Google Scholar] [CrossRef] [PubMed]
- Ando, Y. Diaporthe toxicodendri sp. nov., a causal fungus of the canker disease on Toxicodendron vernicifluum in Japan. Mycosphere 2017, 8, 1157–1167. [Google Scholar] [CrossRef]
- van Rensburg, J.C.J.; Lamprecht, S.C.; Groenewald, J.Z.; Castlebury, L.A.; Crous, P.W. Characterisation of Phomopsis spp. associated with die-back of rooibos (Aspalathus linearis) in South Africa. Stud. Mycol. 2006, 55, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Liu, F.; Cai, L. Unravelling Diaporthe species associated with Camellia. Syst. Biodivers. 2016, 14, 102–117. [Google Scholar] [CrossRef]
- Huang, F.; Udayanga, D.; Wang, X.; Hou, X.; Mei, X.; Fu, Y.; Hyde, K.D.; Li, H. Endophytic Diaporthe associated with Citrus: A phylogenetic reassessment with seven new species from China. Fungal Biol. 2015, 119, 331–347. [Google Scholar] [CrossRef] [PubMed]
- Santos, L.; Phillips, A.J.L.; Crous, P.; Alves, A. Diaporthe species on Rosaceae with descriptions of D. pyracanthae sp. nov. and D. malorum sp. nov. Mycosphere 2017, 8, 485–511. [Google Scholar] [CrossRef]
- Wang, C.X.; Li, B.H.; Dong, X.L.; Li, G.F. First Report of Stem Canker on Cherry Caused by Phomopsis perniciosa in Shandong Peninsula, Eastern China. Plant Dis. 2011, 95, 1316. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.Z.; Zhang, Q.; Yang, P.; Niu, Y.; Niu, W. First Report of Gummosis Disease of Sweet Cherry Caused by Botryosphaeria dothidea in China. Plant Dis. 2019, 103, 3283. [Google Scholar] [CrossRef]
- Dong, X.L.; Li, B.H.; Chen, L.M.; Zhang, F.X. Valsa mali, an important fungus pathogen causes cherry gummosis. In Proceedings of the 10th International Congress of Plant Pathology, Beijing, China, 25–30 August 2013; Chinese Society of Plant Pathology: Beijing, China, 2013. p16.024. [Google Scholar]
- Xu, L.; Wang, J.W.; Chen, X.; Wei, H.R.; Zong, X.J.; Liu, Q.Z. Identification and pathogenicity detection of the cherry gummosis pathogen. Acta Phytopathol. Sin. 2015, 45, 350–355. [Google Scholar]
- León, M.; Berbegal, M.; Rodríguez-Reina, J.M.; Elena, G.; Abad-Campos, P.; Ramón-Albalat, A.; Olmo, D.; Vicent, A.; Luque, J.; Miarnau, X.; et al. Identification and Characterization of Diaporthe spp. Associated with Twig Cankers and Shoot Blight of Almonds in Spain. Agronomy 2020, 10, 1062. [Google Scholar] [CrossRef]


| Gene Region | Primer Pairs | Sequence (5′—3′) | Reference |
|---|---|---|---|
| ITS | ITS1 ITS4 | TCCGTAGGTGAACCTGCGG TCCTCCGCTTATTGATATGC | [16] |
| Tef1-α | EF1-688F EF1-1251R EF1-728F EF1-986R | CGGTCACTTGATCTACAAGTGC CCTCGAACTCACCAGTACCG CATCGAGAAGTTCGAGAAGG TACTTGAAGGAACCCTTACC | [17,18] |
| tub-2 | Bt2a Bt2b TUB2Fd TUB4Rd | GGTAACCAAATCGGTGCTGCTTTC ACCCTCAGTGTAGTGACCCTTGGC GTBCACCTYCARACCGGYCARTG CCRGAYTGRCCRAARACRAAGTTGTC | [19,20] |
| Cal | CAL-228F CAL-737R | GAGTTCAAGGAGGCCTTCTCCC CATCTTTCTGGCCATCATGG | [18] |
| Species | Culture No. | Origin | GenBank Number | |||
|---|---|---|---|---|---|---|
| ITS | Cal | tef1-α | tub-2 | |||
| D. acuta | PSCG 046 | China | MK626958 | MK691124 | MK654803 | MK691224 |
| D. acuta | PSCG 047 * | China | MK626957 | MK691125 | MK654802 | MK691225 |
| D. alleghaniensis | CBS 495.72 = ATCC 24097 * | Canada | KC343007 | KC343249 | KC343733 | KC343975 |
| D. apiculatum | CGMCC 3.17533 | China | KP267896 | - | KP267970 | KP293476 |
| D. arecae | CBS 161.64 * | India | KC343032 | KC343274 | KC343758 | KC344000 |
| D. arecae | CBS 535.75 | Suriname | KC343033 | KC343275 | KC343759 | KC344001 |
| D. australiana | BRIP 66145 * | Australia | MN708222 | - | MN696522 | MN696530 |
| D. australiana | BRIP 66147 | Australia | MN708224 | - | MN696523 | MN696532 |
| D. cercidis | CFCC 52565* | China | MH121500 | MH121424 | MH121542 | MH121582 |
| D. cercidis | CFCC 52566 | China | MH121501 | MH121425 | MH121543 | MH121583 |
| D. charlesworthii | BRIP 54884 m * | Australia | KJ197288 | - | KJ197250 | KJ197268 |
| D. eres D. eres | AR5193 * CBS 101742 | Germany Netherlands | KJ210529 KC343073 | KJ434999 KC343315 | KJ210550 KC343799 | KJ420799 KC344041 |
| D. eres (=D. biguttusis) | CGMCC 3.17081 * | China | KF576282 | - | KF576257 | KF576306 |
| D. eres (=D. castaneae-mollisimae) | DNP 128 * | China | JF957786 | JX197430 | JX275401 | JX275438 |
| D. eres (=D. castaneae-mollisimae) | DNP 129 | China | JQ619886 | JX197431 | JX275402 | JX275439 |
| D. eres (=D. cotoneastri) | DP0667 | - | KC843328 | KC843155 | KC843121 | KC843229 |
| D. eres (=D. ellipicola) | CGMCC 3.17084 * | - | KF576270 | - | KF576245 | KF576294 |
| D. eres (=D. nobilis) | CBS 200.39 | - | KC343151 | KC343393 | KC343877 | KC344119 |
| D. eres (=D. nobilis) | CBS 587.79 | - | KC343153 | KC343395 | KC343879 | KC344121 |
| D. eres | JZB320206 | Guizhou, China | OM980309 | OQ473424 | OQ513364 | ON152804 |
| D. eres | JZB320207 | Guizhou, China | OM980310 | OQ473425 | OQ513365 | ON152805 |
| D. eres | JZB320208 | Guizhou, China | OM980311 | OQ473426 | OQ513366 | ON152806 |
| D. eres | JZB320209 | Guizhou, China | OM980312 | OQ473427 | - | ON152807 |
| D. eres | JZB320210 | Guizhou, China | OM980313 | OQ473428 | - | ON152808 |
| D. eres | JZB320211 | Beijing, China | OM980314 | OQ473429 | OQ513367 | ON152809 |
| D. eres | JZB320212 | Beijing, China | OM980315 | OQ473430 | OQ513368 | ON152810 |
| D. eres | JZB320213 | Beijing, China | OM980316 | OQ473431 | OQ513369 | ON152811 |
| D. eres | JZB320214 | Beijing, China | OM980317 | OQ473432 | OQ513370 | ON152812 |
| D. eres | JZB320215 | Beijing, China | OM980318 | OQ473433 | OQ513371 | ON152813 |
| D. eres | JZB320216 | Beijing, China | OM980319 | OQ473434 | OQ513372 | ON152814 |
| D. eres | JZB320217 | Shandong, China | OM980320 | OQ473435 | OQ513373 | ON152815 |
| D. eres | JZB320218 | Shandong, China | OM980321 | OQ473436 | OQ513374 | ON152816 |
| D. eres | JZB320219 | Shandong, China | OM980322 | OQ473437 | OQ513375 | ON152817 |
| D. fusicola | CGMCC 3.17087 * | China | KF576281 | KF576233 | KF576256 | KF576305 |
| D. fusicola | CGMCC 3.17088 | China | KF576263 | KF576221 | KF576238 | KF576287 |
| D. hongkongensis | CBS 115448 * | China | KC343119 | KC343361 | KC343845 | KC344087 |
| D. hongkongensis | ZJUD74 | China | KJ490609 | - | KJ490488 | KJ490430 |
| D. hongkongensis (=D. lithocarpus) | CGMCC 3.15175 * | - | KC153104 | KF576235 | KC153095 | KF576311 |
| D. hongkongensis (=D. lithocarpus) | CGMCC 3.17098 | - | KF576276 | KF576228 | KF576251 | KF576300 |
| D. hongkongensis | JZB320202 | Guizhou, China | OM980305 | OQ473420 | OQ513360 | OL845879 |
| D. hongkongensis | JZB320203 | Guizhou, China | OM980306 | OQ473421 | OQ513361 | OL845880 |
| D. hongkongensis | JZB320204 | Guizhou, China | OM980307 | OQ473422 | OQ513362 | OL845881 |
| D. hongkongensis | JZB320205 | Guizhou, China | OM980308 | OQ473423 | OQ513363 | OL845882 |
| D. longispora | CBS 194.36 | - | MH855769 | KC343377 | KC343861 | KC344103 |
| D. padina | CFCC 52590 * | China | MH121525 | MH121443 | MH121567 | MH121604 |
| D. padina | PSCG 160 | - | MK626892 | MK691172 | MK654851 | MK691261 |
| D. penetriteum | CGMCC 3.17532 | China | KP267879 | - | KP267953 | KP293459 |
| D. sennicola | CFCC 51634 * | China | KY203722 | - | KY228883 | KY228889 |
| D. vaccinii | CBS 160.32 = IFO 32646 * | USA | KC343228 | KC343470 | KC343954 | KC344196 |
| D. viniferae | JZB320071 * | Guangxi, China | MK341551 | MK500119 | MK500107 | MK500112 |
| D. viniferae | JZB320072 | Guangxi, China | MK341552 | MK500120 | MK500108 | MK500113 |
| Diaporthella corylina | CBS 121124 * | China | KC343004 | KC343246 | KC343730 | KC343972 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, P.; Abeywickrama, P.D.; Ji, S.; Zhou, Y.; Li, X.; Zhang, W.; Yan, J. Molecular Identification and Pathogenicity of Diaporthe eres and D. hongkongensis (Diaporthales, Ascomycota) Associated with Cherry Trunk Diseases in China. Microorganisms 2023, 11, 2400. https://doi.org/10.3390/microorganisms11102400
Chen P, Abeywickrama PD, Ji S, Zhou Y, Li X, Zhang W, Yan J. Molecular Identification and Pathogenicity of Diaporthe eres and D. hongkongensis (Diaporthales, Ascomycota) Associated with Cherry Trunk Diseases in China. Microorganisms. 2023; 11(10):2400. https://doi.org/10.3390/microorganisms11102400
Chicago/Turabian StyleChen, Pengzhao, Pranami D. Abeywickrama, Shuxian Ji, Yueyan Zhou, Xinghong Li, Wei Zhang, and Jiye Yan. 2023. "Molecular Identification and Pathogenicity of Diaporthe eres and D. hongkongensis (Diaporthales, Ascomycota) Associated with Cherry Trunk Diseases in China" Microorganisms 11, no. 10: 2400. https://doi.org/10.3390/microorganisms11102400
APA StyleChen, P., Abeywickrama, P. D., Ji, S., Zhou, Y., Li, X., Zhang, W., & Yan, J. (2023). Molecular Identification and Pathogenicity of Diaporthe eres and D. hongkongensis (Diaporthales, Ascomycota) Associated with Cherry Trunk Diseases in China. Microorganisms, 11(10), 2400. https://doi.org/10.3390/microorganisms11102400









