Molecular Epidemiology of Group B Streptococcus Colonization in Egyptian Women
Abstract
1. Introduction
2. Materials and Methods
2.1. MLST Types
2.2. Pili and GBS Surface Protein Genes
3. Results
3.1. MLST Types
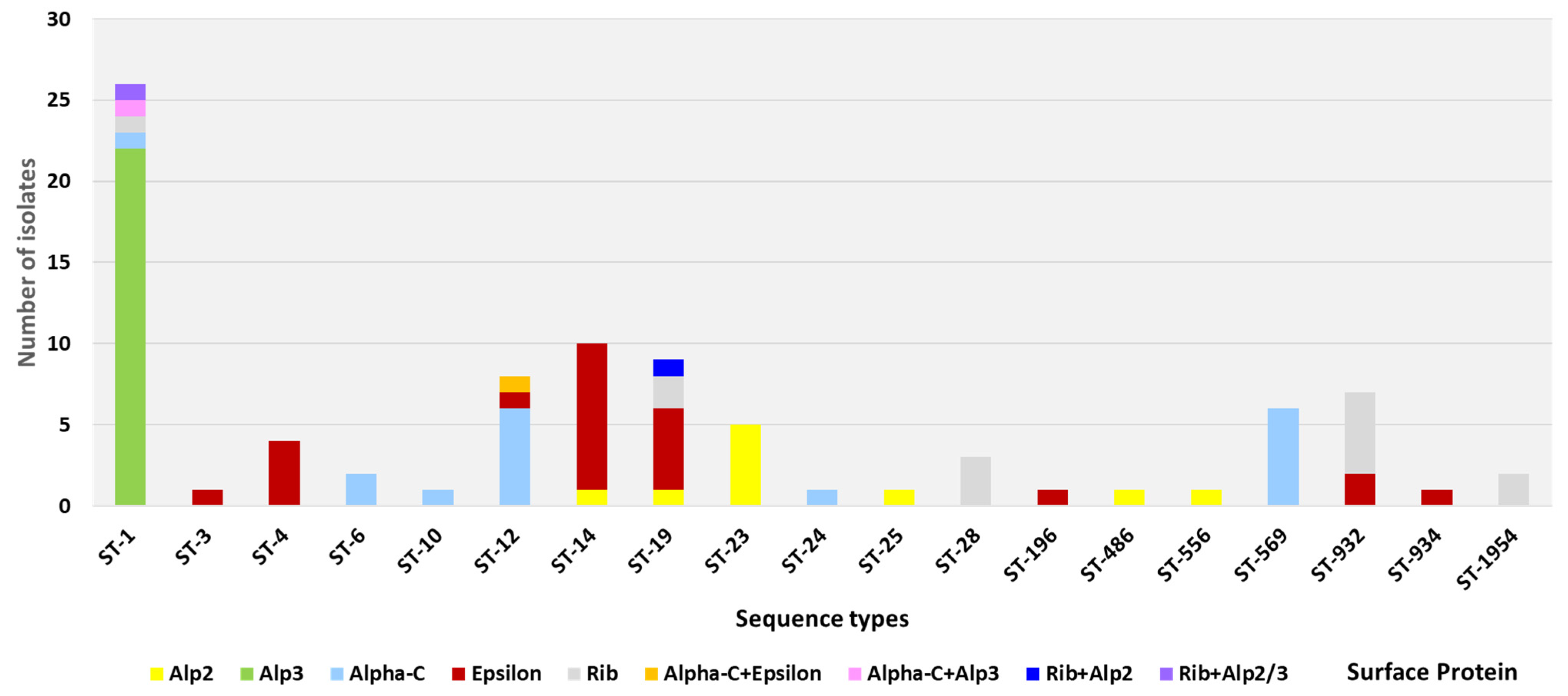
3.2. Pili and GBS Surface Protein Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Brokaw, A.; Furuta, A.; Dacanay, M.; Rajagopal, L.; Adams Waldorf, K.M. Bacterial and Host Determinants of Group B Streptococcal Vaginal Colonization and Ascending Infection in Pregnancy. Front. Cell. Infect. Microbiol. 2021, 11, 720789. [Google Scholar] [CrossRef] [PubMed]
- Verani, J.R.; McGee, L.; Schrag, S.J. Prevention of perinatal group B streptococcal disease—Revised guidelines from CDC, 2010. MMWR. Recomm. Rep. Morb. Mortal. Wkly. Report. Recomm. Rep. 2010, 59, 1–36. [Google Scholar]
- World Health Organization. Group B Streptococcus Vaccine: Full Value of Vaccine Assessment; World Health Organization: Geneva, Switzerland, 2021. [Google Scholar]
- Furfaro, L.L.; Chang, B.J.; Payne, M.S. Perinatal Streptococcus agalactiae Epidemiology and Surveillance Targets. Clin. Microbiol. Rev. 2018, 31, e00049-18. [Google Scholar] [CrossRef] [PubMed]
- Jones, N.; Bohnsack, J.F.; Takahashi, S.; Oliver, K.A.; Chan, M.S.; Kunst, F.; Glaser, P.; Rusniok, C.; Crook, D.W.; Harding, R.M.; et al. Multilocus sequence typing system for group B streptococcus. J. Clin. Microbiol. 2003, 41, 2530–2536. [Google Scholar] [CrossRef]
- Da Cunha, V.; Davies, M.R.; Douarre, P.E.; Rosinski-Chupin, I.; Margarit, I.; Spinali, S.; Perkins, T.; Lechat, P.; Dmytruk, N.; Sauvage, E.; et al. Streptococcus agalactiae clones infecting humans were selected and fixed through the extensive use of tetracycline. Nat. Commun. 2014, 5, 4544. [Google Scholar] [CrossRef]
- Gori, A.; Harrison, O.B.; Mlia, E.; Nishihara, Y.; Chan, J.M.; Msefula, J.; Mallewa, M.; Dube, Q.; Swarthout, T.D.; Nobbs, A.H.; et al. Pan-GWAS of Streptococcus agalactiae Highlights Lineage-Specific Genes Associated with Virulence and Niche Adaptation. mBio 2020, 11. [Google Scholar] [CrossRef]
- Meehan, M.; Eogan, M.; McCallion, N.; Cunney, R.; Bray, J.E.; Jolley, K.A.; Unitt, A.; Maiden, M.C.J.; Harrison, O.B.; Drew, R.J. Genomic epidemiology of group B streptococci spanning 10 years in an Irish maternity hospital, 2008–2017. J. Infect. 2021, 83, 37–45. [Google Scholar] [CrossRef]
- Springman, A.C.; Lacher, D.W.; Wu, G.; Milton, N.; Whittam, T.S.; Davies, H.D.; Manning, S.D. Selection, recombination, and virulence gene diversity among group B streptococcal genotypes. J. Bacteriol. 2009, 191, 5419–5427. [Google Scholar] [CrossRef]
- Tazi, A.; Disson, O.; Bellais, S.; Bouaboud, A.; Dmytruk, N.; Dramsi, S.; Mistou, M.Y.; Khun, H.; Mechler, C.; Tardieux, I.; et al. The surface protein HvgA mediates group B streptococcus hypervirulence and meningeal tropism in neonates. J. Exp. Med. 2010, 207, 2313–2322. [Google Scholar] [CrossRef]
- Brochet, M.; Couvé, E.; Bercion, R.; Sire, J.M.; Glaser, P. Population structure of human isolates of Streptococcus agalactiae from Dakar and Bangui. J. Clin. Microbiol. 2009, 47, 800–803. [Google Scholar] [CrossRef]
- Huber, C.A.; McOdimba, F.; Pflueger, V.; Daubenberger, C.A.; Revathi, G. Characterization of invasive and colonizing isolates of Streptococcus agalactiae in East African adults. J. Clin. Microbiol. 2011, 49, 3652–3655. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Capan, M.; Mombo-Ngoma, G.; Akerey-Diop, D.; Basra, A.; Würbel, H.; Lendamba, W.; Auer-Hackenberg, L.; Mackanga, R.; Melser, J.; Belard, S.; et al. Epidemiology and management of group B streptococcal colonization during pregnancy in Africa. Wien. Klin. Wochenschr. 2012, 124 (Suppl. S3), 14–16. [Google Scholar] [CrossRef] [PubMed]
- Russell, L.B.; Kim, S.Y.; Cosgriff, B.; Pentakota, S.R.; Schrag, S.J.; Sobanjo-Ter Meulen, A.; Verani, J.R.; Sinha, A. Cost-effectiveness of maternal GBS immunization in low-income sub-Saharan Africa. Vaccine 2017, 35, 6905–6914. [Google Scholar] [CrossRef] [PubMed]
- Seale, A.C.; Baker, C.J.; Berkley, J.A.; Madhi, S.A.; Ordi, J.; Saha, S.K.; Schrag, S.J.; Sobanjo-Ter Meulen, A.; Vekemans, J. Vaccines for maternal immunization against Group B Streptococcus disease: WHO perspectives on case ascertainment and case definitions. Vaccine 2019, 37, 4877–4885. [Google Scholar] [CrossRef]
- Le Doare, K.; Kampmann, B.; Vekemans, J.; Heath, P.T.; Goldblatt, D.; Nahm, M.H.; Baker, C.; Edwards, M.S.; Kwatra, G.; Andrews, N.; et al. Serocorrelates of protection against infant group B streptococcus disease. Lancet Infect. Dis. 2019, 19, e162–e171. [Google Scholar] [CrossRef]
- Absalon, J.; Segall, N.; Block, S.L.; Center, K.J.; Scully, I.L.; Giardina, P.C.; Peterson, J.; Watson, W.J.; Gruber, W.C.; Jansen, K.U.; et al. Safety and immunogenicity of a novel hexavalent group B streptococcus conjugate vaccine in healthy, non-pregnant adults: A phase 1/2, randomised, placebo-controlled, observer-blinded, dose-escalation trial. Lancet Infect. Dis. 2021, 21, 263–274. [Google Scholar] [CrossRef]
- Buurman, E.T.; Timofeyeva, Y.; Gu, J.; Kim, J.H.; Kodali, S.; Liu, Y.; Mininni, T.; Moghazeh, S.; Pavliakova, D.; Singer, C.; et al. A Novel Hexavalent Capsular Polysaccharide Conjugate Vaccine (GBS6) for the Prevention of Neonatal Group B Streptococcal Infections by Maternal Immunization. J. Infect. Dis. 2019, 220, 105–115. [Google Scholar] [CrossRef]
- Vekemans, J.; Crofts, J.; Baker, C.J.; Goldblatt, D.; Heath, P.T.; Madhi, S.A.; Le Doare, K.; Andrews, N.; Pollard, A.J.; Saha, S.K.; et al. The role of immune correlates of protection on the pathway to licensure, policy decision and use of group B Streptococcus vaccines for maternal immunization: Considerations from World Health Organization consultations. Vaccine 2019, 37, 3190–3198. [Google Scholar] [CrossRef]
- Project JUNO: A Global Genomic Survey of Streptococcus Agalactiae [Website]. Hinxton: Wellcome Genome Campus. 2021. Available online: https://www.gbsgen.net/ (accessed on 30 December 2021).
- Shabayek, S.; Abdalla, S.; Abouzeid, A.M. Serotype and surface protein gene distribution of colonizing group B streptococcus in women in Egypt. Epidemiol. Infect. 2014, 142, 208–210. [Google Scholar] [CrossRef]
- Jolley, K.A.; Bray, J.E.; Maiden, M.C.J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018, 3, 124. [Google Scholar] [CrossRef]
- Luan, S.L.; Granlund, M.; Sellin, M.; Lagergård, T.; Spratt, B.G.; Norgren, M. Multilocus sequence typing of Swedish invasive group B streptococcus isolates indicates a neonatally associated genetic lineage and capsule switching. J. Clin. Microbiol. 2005, 43, 3727–3733. [Google Scholar] [CrossRef] [PubMed]
- McGee, L.; Chochua, S.; Li, Z.; Mathis, S.; Rivers, J.; Metcalf, B.; Ryan, A.; Alden, N.; Farley, M.M.; Harrison, L.H.; et al. Multistate, Population-Based Distributions of Candidate Vaccine Targets, Clonal Complexes, and Resistance Features of Invasive Group B Streptococci Within the United States, 2015–2017. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2021, 72, 1004–1013. [Google Scholar] [CrossRef]
- Bertels, F.; Silander, O.K.; Pachkov, M.; Rainey, P.B.; van Nimwegen, E. Automated reconstruction of whole-genome phylogenies from short-sequence reads. Mol. Biol. Evol. 2014, 31, 1077–1088. [Google Scholar] [CrossRef]
- Shabayek, S.; Spellerberg, B. Group B Streptococcal Colonization, Molecular Characteristics, and Epidemiology. Front. Microbiol. 2018, 9, 437. [Google Scholar] [CrossRef] [PubMed]
- Rosini, R.; Rinaudo, C.D.; Soriani, M.; Lauer, P.; Mora, M.; Maione, D.; Taddei, A.; Santi, I.; Ghezzo, C.; Brettoni, C.; et al. Identification of novel genomic islands coding for antigenic pilus-like structures in Streptococcus agalactiae. Mol. Microbiol. 2006, 61, 126–141. [Google Scholar] [CrossRef] [PubMed]
- Margarit, I.; Rinaudo, C.D.; Galeotti, C.L.; Maione, D.; Ghezzo, C.; Buttazzoni, E.; Rosini, R.; Runci, Y.; Mora, M.; Buccato, S.; et al. Preventing bacterial infections with pilus-based vaccines: The group B streptococcus paradigm. J. Infect. Dis. 2009, 199, 108–115. [Google Scholar] [CrossRef]
- Springman, A.C.; Lacher, D.W.; Waymire, E.A.; Wengert, S.L.; Singh, P.; Zadoks, R.N.; Davies, H.D.; Manning, S.D. Pilus distribution among lineages of group b streptococcus: An evolutionary and clinical perspective. BMC Microbiol. 2014, 14, 159. [Google Scholar] [CrossRef]
- Sheen, T.R.; Jimenez, A.; Wang, N.Y.; Banerjee, A.; van Sorge, N.M.; Doran, K.S. Serine-rich repeat proteins and pili promote Streptococcus agalactiae colonization of the vaginal tract. J. Bacteriol. 2011, 193, 6834–6842. [Google Scholar] [CrossRef]
- Wang, N.Y.; Patras, K.A.; Seo, H.S.; Cavaco, C.K.; Rösler, B.; Neely, M.N.; Sullam, P.M.; Doran, K.S. Group B streptococcal serine-rich repeat proteins promote interaction with fibrinogen and vaginal colonization. J. Infect. Dis. 2014, 210, 982–991. [Google Scholar] [CrossRef]
- Santi, I.; Maione, D.; Galeotti, C.L.; Grandi, G.; Telford, J.L.; Soriani, M. BibA induces opsonizing antibodies conferring in vivo protection against group B Streptococcus. J. Infect. Dis. 2009, 200, 564–570. [Google Scholar] [CrossRef]
- Santi, I.; Scarselli, M.; Mariani, M.; Pezzicoli, A.; Masignani, V.; Taddei, A.; Grandi, G.; Telford, J.L.; Soriani, M. BibA: A novel immunogenic bacterial adhesin contributing to group B Streptococcus survival in human blood. Mol. Microbiol. 2007, 63, 754–767. [Google Scholar] [CrossRef] [PubMed]
- Brochet, M.; Couvé, E.; Zouine, M.; Vallaeys, T.; Rusniok, C.; Lamy, M.C.; Buchrieser, C.; Trieu-Cuot, P.; Kunst, F.; Poyart, C.; et al. Genomic diversity and evolution within the species Streptococcus agalactiae. Microbes Infect. 2006, 8, 1227–1243. [Google Scholar] [CrossRef] [PubMed]
- Kolter, J.; Henneke, P. Codevelopment of Microbiota and Innate Immunity and the Risk for Group B Streptococcal Disease. Front. Immunol. 2017, 8, 1497. [Google Scholar] [CrossRef] [PubMed]
- Bianchi-Jassir, F.; Paul, P.; To, K.N.; Carreras-Abad, C.; Seale, A.C.; Jauneikaite, E.; Madhi, S.A.; Russell, N.J.; Hall, J.; Madrid, L.; et al. Systematic review of Group B Streptococcal capsular types, sequence types and surface proteins as potential vaccine candidates. Vaccine 2020, 38, 6682–6694. [Google Scholar] [CrossRef] [PubMed]
- Manning, S.D.; Lewis, M.A.; Springman, A.C.; Lehotzky, E.; Whittam, T.S.; Davies, H.D. Genotypic diversity and serotype distribution of group B streptococcus isolated from women before and after delivery. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2008, 46, 1829–1837. [Google Scholar] [CrossRef] [PubMed]
- Bergal, A.; Loucif, L.; Benouareth, D.E.; Bentorki, A.A.; Abat, C.; Rolain, J.M. Molecular epidemiology and distribution of serotypes, genotypes, and antibiotic resistance genes of Streptococcus agalactiae clinical isolates from Guelma, Algeria and Marseille, France. Eur. J. Clin. Microbiol. Infect. Dis. Off. Publ. Eur. Soc. Clin. Microbiol. 2015, 34, 2339–2348. [Google Scholar] [CrossRef]
- Flores, A.R.; Galloway-Peña, J.; Sahasrabhojane, P.; Saldaña, M.; Yao, H.; Su, X.; Ajami, N.J.; Holder, M.E.; Petrosino, J.F.; Thompson, E.; et al. Sequence type 1 group B Streptococcus, an emerging cause of invasive disease in adults, evolves by small genetic changes. Proc. Natl. Acad. Sci. USA 2015, 112, 6431–6436. [Google Scholar] [CrossRef]
- Hirai, N.; Kasahara, K.; Nakano, R.; Ogawa, Y.; Suzuki, Y.; Ogawa, M.; Hishiya, N.; Nakano, A.; Ichimura, S.; Yano, H.; et al. Clinical characteristics and molecular epidemiology of invasive Streptococcus agalactiae infections between 2007 and 2016 in Nara, Japan. PLoS ONE 2020, 15, e0240590. [Google Scholar] [CrossRef]
- Salloum, M.; van der Mee-Marquet, N.; Domelier, A.S.; Arnault, L.; Quentin, R. Molecular characterization and prophage DNA contents of Streptococcus agalactiae strains isolated from adult skin and osteoarticular infections. J. Clin. Microbiol. 2010, 48, 1261–1269. [Google Scholar] [CrossRef]
- Teatero, S.; Ferrieri, P.; Martin, I.; Demczuk, W.; McGeer, A.; Fittipaldi, N. Serotype Distribution, Population Structure, and Antimicrobial Resistance of Group B Streptococcus Strains Recovered from Colonized Pregnant Women. J. Clin. Microbiol. 2017, 55, 412–422. [Google Scholar] [CrossRef]
- Furfaro, L.L.; Chang, B.J.; Kahler, C.M.; Payne, M.S. Genomic characterisation of perinatal Western Australian Streptococcus agalactiae isolates. PLoS ONE 2019, 14, e0223256. [Google Scholar] [CrossRef] [PubMed]
- Bohnsack, J.F.; Whiting, A.; Gottschalk, M.; Dunn, D.M.; Weiss, R.; Azimi, P.H.; Philips, J.B., 3rd; Weisman, L.E.; Rhoads, G.G.; Lin, F.Y. Population structure of invasive and colonizing strains of Streptococcus agalactiae from neonates of six U.S. Academic Centers from 1995 to 1999. J. Clin. Microbiol. 2008, 46, 1285–1291. [Google Scholar] [CrossRef] [PubMed]
- Nagano, N.; Koide, S.; Hayashi, W.; Taniguchi, Y.; Tanaka, H.; Maeyama, Y.; Suzuki, M.; Kimura, K.; Arakawa, Y.; Nagano, Y. Population-level transition of capsular polysaccharide types among sequence type 1 group B Streptococcus isolates with reduced penicillin susceptibility during a long-term hospital epidemic. Int. J. Antimicrob. Agents 2019, 53, 203–210. [Google Scholar] [CrossRef] [PubMed]
- Björnsdóttir, E.S.; Martins, E.R.; Erlendsdóttir, H.; Haraldsson, G.; Melo-Cristino, J.; Kristinsson, K.G.; Ramirez, M. Changing epidemiology of group B streptococcal infections among adults in Iceland: 1975–2014. Clin. Microbiol. Infect. Off. Publ. Eur. Soc. Clin. Microbiol. Infect. Dis. 2016, 22, 379.e9–379.e16. [Google Scholar] [CrossRef] [PubMed]
- De Francesco, M.A.; Caracciolo, S.; Gargiulo, F.; Manca, N. Phenotypes, genotypes, serotypes and molecular epidemiology of erythromycin-resistant Streptococcus agalactiae in Italy. Eur. J. Clin. Microbiol. Infect. Dis. Off. Publ. Eur. Soc. Clin. Microbiol. 2012, 31, 1741–1747. [Google Scholar] [CrossRef]
- Eickel, V.; Kahl, B.; Reinisch, B.; Dübbers, A.; Küster, P.; Brandt, C.; Spellerberg, B. Emergence of respiratory Streptococcus agalactiae isolates in cystic fibrosis patients. PLoS ONE 2009, 4, e4650. [Google Scholar] [CrossRef]
- Gherardi, G.; Imperi, M.; Baldassarri, L.; Pataracchia, M.; Alfarone, G.; Recchia, S.; Orefici, G.; Dicuonzo, G.; Creti, R. Molecular epidemiology and distribution of serotypes, surface proteins, and antibiotic resistance among group B streptococci in Italy. J. Clin. Microbiol. 2007, 45, 2909–2916. [Google Scholar] [CrossRef][Green Version]
- Neemuchwala, A.; Teatero, S.; Liang, L.; Martin, I.; Demzcuk, W.; McGeer, A.; Fittipaldi, N. Genetic Diversity and Antimicrobial Drug Resistance of Serotype VI Group B Streptococcus, Canada. Emerg. Infect. Dis. 2018, 24, 1941–1942. [Google Scholar] [CrossRef]
- Creti, R.; Fabretti, F.; Orefici, G.; von Hunolstein, C. Multiplex PCR assay for direct identification of group B streptococcal alpha-protein-like protein genes. J. Clin. Microbiol. 2004, 42, 1326–1329. [Google Scholar] [CrossRef][Green Version]
- Dhanoa, A.; Karunakaran, R.; Puthucheary, S.D. Serotype distribution and antibiotic susceptibility of group B streptococci in pregnant women. Epidemiol. Infect. 2010, 138, 979–981. [Google Scholar] [CrossRef]
- Eskandarian, N.; Ismail, Z.; Neela, V.; van Belkum, A.; Desa, M.N.; Amin Nordin, S. Antimicrobial susceptibility profiles, serotype distribution and virulence determinants among invasive, non-invasive and colonizing Streptococcus agalactiae (group B streptococcus) from Malaysian patients. Eur. J. Clin. Microbiol. Infect. Dis. Off. Publ. Eur. Soc. Clin. Microbiol. 2015, 34, 579–584. [Google Scholar] [CrossRef] [PubMed]
- Lachenauer, C.S.; Kasper, D.L.; Shimada, J.; Ichiman, Y.; Ohtsuka, H.; Kaku, M.; Paoletti, L.C.; Ferrieri, P.; Madoff, L.C. Serotypes VI and VIII predominate among group B streptococci isolated from pregnant Japanese women. J. Infect. Dis. 1999, 179, 1030–1033. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.C.; Chen, C.J.; Chiang, K.H.; Yen, T.Y.; Ho, C.M.; Hwang, K.P.; Su, B.H.; Lin, H.C.; Li, T.C.; Lu, J.J. Clonal dissemination of invasive and colonizing clonal complex 1 of serotype VI group B Streptococcus in central Taiwan. J. Microbiol. Immunol. Infect. Wei Mian Yu Gan Ran Za Zhi 2016, 49, 902–909. [Google Scholar] [CrossRef]
- Lo, C.W.; Liu, H.C.; Lee, C.C.; Lin, C.L.; Chen, C.L.; Jeng, M.J.; Chiu, C.H. Serotype distribution and clinical correlation of Streptococcus agalactiae causing invasive disease in infants and children in Taiwan. J. Microbiol. Immunol. Infect. Wei Mian Yu Gan Ran Za Zhi 2019, 52, 578–584. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.H.; Lu, C.C.; Chiu, C.H.; Wang, M.H.; Yang, T.H.; Chu, C. Genetically diverse serotypes III and VI substitute major clonal disseminated serotypes Ib and V as prevalent serotypes of Streptococcus agalactiae from 2007 to 2012. J. Microbiol. Immunol. Infect. Wei Mian Yu Gan Ran Za Zhi 2016, 49, 672–678. [Google Scholar] [CrossRef] [PubMed]
- Morozumi, M.; Wajima, T.; Takata, M.; Iwata, S.; Ubukata, K. Molecular Characteristics of Group B Streptococci Isolated from Adults with Invasive Infections in Japan. J. Clin. Microbiol. 2016, 54, 2695–2700. [Google Scholar] [CrossRef]
- Mazzuoli, M.V.; Daunesse, M.; Varet, H.; Rosinski-Chupin, I.; Legendre, R.; Sismeiro, O.; Gominet, M.; Kaminski, P.A.; Glaser, P.; Chica, C.; et al. The CovR regulatory network drives the evolution of Group B Streptococcus virulence. PLoS Genet. 2021, 17, e1009761. [Google Scholar] [CrossRef]
- Shabayek, S.; Spellerberg, B. Acid Stress Response Mechanisms of Group B Streptococci. Front. Cell. Infect. Microbiol. 2017, 7, 395. [Google Scholar] [CrossRef]
- Santi, I.; Grifantini, R.; Jiang, S.M.; Brettoni, C.; Grandi, G.; Wessels, M.R.; Soriani, M. CsrRS regulates group B Streptococcus virulence gene expression in response to environmental pH: A new perspective on vaccine development. J. Bacteriol. 2009, 191, 5387–5397. [Google Scholar] [CrossRef]
- Maeland, J.A.; Afset, J.E.; Lyng, R.V.; Radtke, A. Survey of immunological features of the alpha-like proteins of Streptococcus agalactiae. Clin. Vaccine Immunol. CVI 2015, 22, 153–159. [Google Scholar] [CrossRef]
- Almeida, A.; Rosinski-Chupin, I.; Plainvert, C.; Douarre, P.E.; Borrego, M.J.; Poyart, C.; Glaser, P. Parallel Evolution of Group B Streptococcus Hypervirulent Clonal Complex 17 Unveils New Pathoadaptive Mutations. mSystems 2017, 2, e00074-17. [Google Scholar] [CrossRef] [PubMed]
- Teatero, S.; Athey, T.B.; Van Caeseele, P.; Horsman, G.; Alexander, D.C.; Melano, R.G.; Li, A.; Flores, A.R.; Shelburne, S.A., 3rd; McGeer, A.; et al. Emergence of Serotype IV Group B Streptococcus Adult Invasive Disease in Manitoba and Saskatchewan, Canada, Is Driven by Clonal Sequence Type 459 Strains. J. Clin. Microbiol. 2015, 53, 2919–2926. [Google Scholar] [CrossRef] [PubMed]
- Campisi, E.; Rinaudo, C.D.; Donati, C.; Barucco, M.; Torricelli, G.; Edwards, M.S.; Baker, C.J.; Margarit, I.; Rosini, R. Serotype IV Streptococcus agalactiae ST-452 has arisen from large genomic recombination events between CC23 and the hypervirulent CC17 lineages. Sci. Rep. 2016, 6, 29799. [Google Scholar] [CrossRef] [PubMed]
- Ali, M.M.; Woldeamanuel, Y.; Asrat, D.; Fenta, D.A.; Beall, B.; Schrag, S.; McGee, L. Features of Streptococcus agalactiae strains recovered from pregnant women and newborns attending different hospitals in Ethiopia. BMC Infect. Dis. 2020, 20, 848. [Google Scholar] [CrossRef]
- Seale, A.C.; Koech, A.C.; Sheppard, A.E.; Barsosio, H.C.; Langat, J.; Anyango, E.; Mwakio, S.; Mwarumba, S.; Morpeth, S.C.; Anampiu, K.; et al. Maternal colonization with Streptococcus agalactiae and associated stillbirth and neonatal disease in coastal Kenya. Nat. Microbiol. 2016, 1, 16067. [Google Scholar] [CrossRef]
- Perme, T.; Golparian, D.; Bombek Ihan, M.; Rojnik, A.; Lučovnik, M.; Kornhauser Cerar, L.; Fister, P.; Lozar Krivec, J.; Grosek, Š.; Ihan, A.; et al. Genomic and phenotypic characterisation of invasive neonatal and colonising group B Streptococcus isolates from Slovenia, 2001–2018. BMC Infect. Dis. 2020, 20, 958. [Google Scholar] [CrossRef]
- Gizachew, M.; Tiruneh, M.; Moges, F.; Adefris, M.; Tigabu, Z.; Tessema, B. Molecular characterization of Streptococcus agalactiae isolated from pregnant women and newborns at the University of Gondar Comprehensive Specialized Hospital, Northwest Ethiopia. BMC Infect. Dis. 2020, 20, 35. [Google Scholar] [CrossRef]

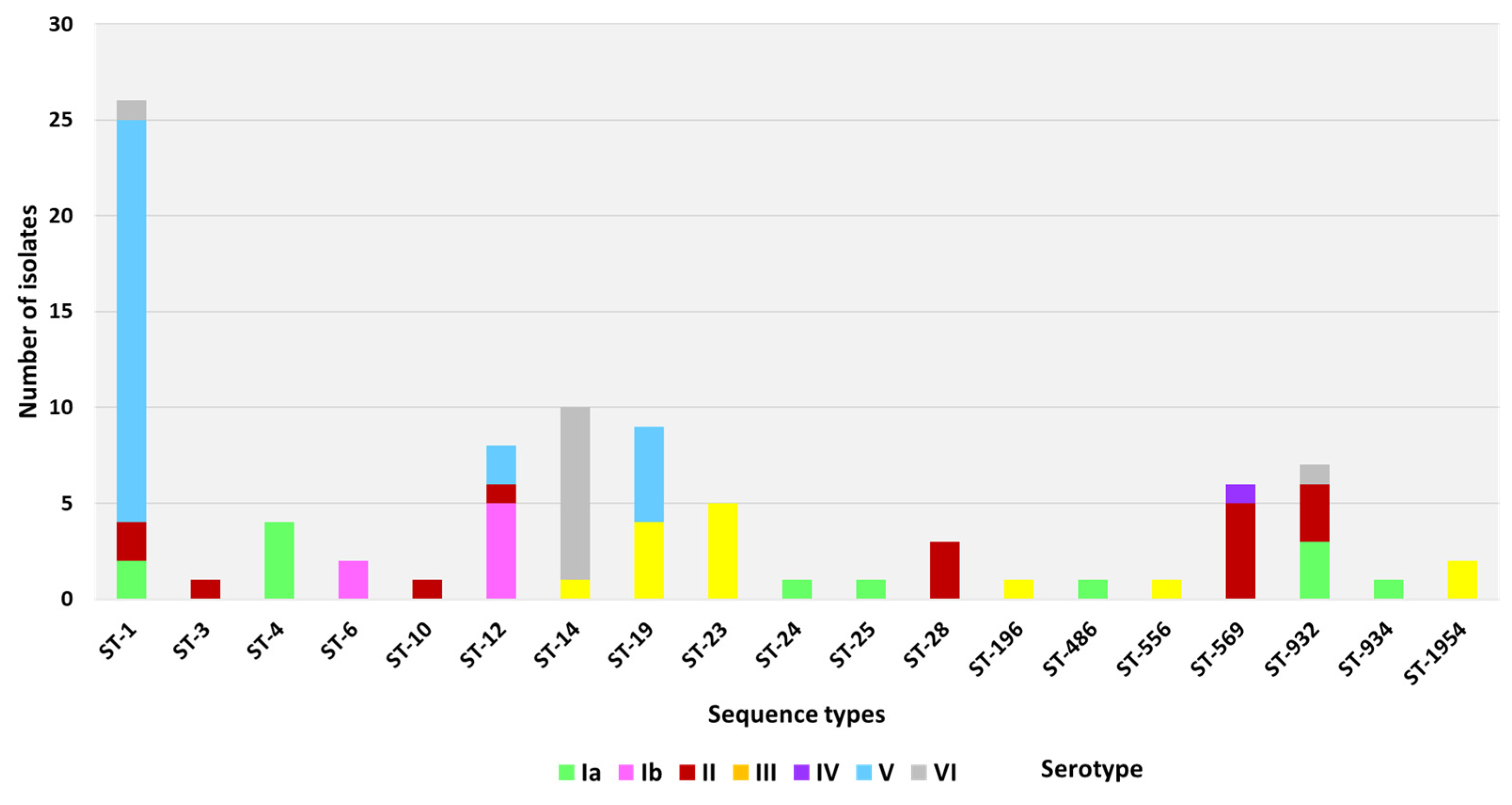

| Clonal Complex/Sequence Type (Number) | Serotypes | Pilus Islands | Srr Proteins | HvgA | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ia | Ib | II | III | IV | V | VI | 1+2A | 1+2B | 2A | Srr1 | Srr2 | ||
| CC1 | |||||||||||||
| ST-1 (26) | 2 | 2 | 21 | 1 | 24 | 2 | 25 | ||||||
| ST-14 (10) | 1 | 9 | 10 | 10 | |||||||||
| CC4 | |||||||||||||
| ST-3 (1) | 1 | 1 | 1 | ||||||||||
| ST-4 (4) | 4 | 4 | 4 | ||||||||||
| ST-932 (7) | 3 | 3 | 1 | 1 | 6 | 6 | |||||||
| CC10 | |||||||||||||
| ST-10 (1) | 1 | 1 | 1 | ||||||||||
| ST-12 (8) | 5 | 1 | 2 | 8 | 8 | ||||||||
| CC17 | |||||||||||||
| ST-1954 (2) | 2 | 2 | 2 | 2 | |||||||||
| CC19 | |||||||||||||
| ST-19 (9) | 4 | 5 | 9 | 9 | |||||||||
| ST-28 (3) | 3 | 3 | 1 | ||||||||||
| CC23 | |||||||||||||
| ST-23 (5) | 5 | 5 | 5 | ||||||||||
| ST-25 (1) | 1 | 1 | 1 | ||||||||||
| ST-556 (1) | 1 | 1 | 1 | ||||||||||
| Singletons | |||||||||||||
| ST-6 (2) | 2 | 2 | 2 | ||||||||||
| ST-24 (1) | 1 | 1 | 1 | ||||||||||
| ST-196 (1) | 1 | 1 | 1 | ||||||||||
| ST-486 (1) | 1 | 1 | 1 | ||||||||||
| ST-569 (6) | 5 | 1 | 6 | 6 | |||||||||
| ST-934 (1) | 1 | 1 | 1 | 1 | |||||||||
| Total | 13 | 7 | 16 | 14 | 1 | 28 | 11 | 61 | 26 | 3 | 84 | 2 | 3 |
| Clonal Complex/Sequence Type (Number) | Alpha-Like Surface Proteins | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Alp2 | Alp3 | Alpha-C | Epsilon | Rib | Alpha-C + Epsilon | Alpha-C + Alp3 | Rib + Alp2 | Rib + Alp2/3 | |
| CC1 | |||||||||
| ST-1 (26) | 22 | 1 | 1 | 1 | 1 | ||||
| ST-14 (10) | 1 | 9 | |||||||
| CC4 | |||||||||
| ST-3 (1) | 1 | ||||||||
| ST-4 (4) | 4 | ||||||||
| ST-932 (7) | 2 | 5 | |||||||
| CC10 | |||||||||
| ST-10 (1) | 1 | ||||||||
| ST-12 (8) | 6 | 1 | 1 | ||||||
| CC17 | |||||||||
| ST-1954 (2) | 2 | ||||||||
| CC19 | |||||||||
| ST-19 (9) | 1 | 5 | 2 | 1 | |||||
| ST-28 (3) | 3 | ||||||||
| CC23 | |||||||||
| ST-23 (5) | 5 | ||||||||
| ST-25 (1) | 1 | ||||||||
| ST-556 (1) | 1 | ||||||||
| Singletons | |||||||||
| ST-6 (2) | 2 | ||||||||
| ST-24 (1) | 1 | ||||||||
| ST-196 (1) | 1 | ||||||||
| ST-486 (1) | 1 | ||||||||
| ST-569 (6) | 6 | ||||||||
| ST-934 (1) | 1 | ||||||||
| Total | 10 | 22 | 17 | 24 | 13 | 1 | 1 | 1 | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shabayek, S.; Vogel, V.; Jamrozy, D.; Bentley, S.D.; Spellerberg, B. Molecular Epidemiology of Group B Streptococcus Colonization in Egyptian Women. Microorganisms 2023, 11, 38. https://doi.org/10.3390/microorganisms11010038
Shabayek S, Vogel V, Jamrozy D, Bentley SD, Spellerberg B. Molecular Epidemiology of Group B Streptococcus Colonization in Egyptian Women. Microorganisms. 2023; 11(1):38. https://doi.org/10.3390/microorganisms11010038
Chicago/Turabian StyleShabayek, Sarah, Verena Vogel, Dorota Jamrozy, Stephen D. Bentley, and Barbara Spellerberg. 2023. "Molecular Epidemiology of Group B Streptococcus Colonization in Egyptian Women" Microorganisms 11, no. 1: 38. https://doi.org/10.3390/microorganisms11010038
APA StyleShabayek, S., Vogel, V., Jamrozy, D., Bentley, S. D., & Spellerberg, B. (2023). Molecular Epidemiology of Group B Streptococcus Colonization in Egyptian Women. Microorganisms, 11(1), 38. https://doi.org/10.3390/microorganisms11010038








