Preclinical Safety Assessment of Bacillus subtilis BS50 for Probiotic and Food Applications
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacillus subtilis BS50 Isolation
2.2. Genome Sequencing
2.3. BS50 Taxonomic Classification via Multilocus Sequence Typing
2.4. BLASTn Screen for Known Bacillus Toxins
2.5. BLASTx Screen for Known Bacillus Toxins
2.6. In Silico PCR Amplification of BS50 for Bacillus Toxins
2.7. Secondary Metabolite Screen via AntiSMASH
2.8. Secreted Protein via SignalP 6.0 Analysis
2.9. Virulence Factor Screen via VFDB
2.10. Antimicrobial Resistance Gene and Mobile Genetic Element Screen
2.11. Antibiotic Minimum Inhibitory Concentration (MIC) Evaluation of BS50
2.12. Blood Hemolysis Assay
2.13. Caco-2 Cell Viability Assay
2.14. Caco-2 Cell Transepithelial Electrical Resistance (TEER) Assay
3. Results
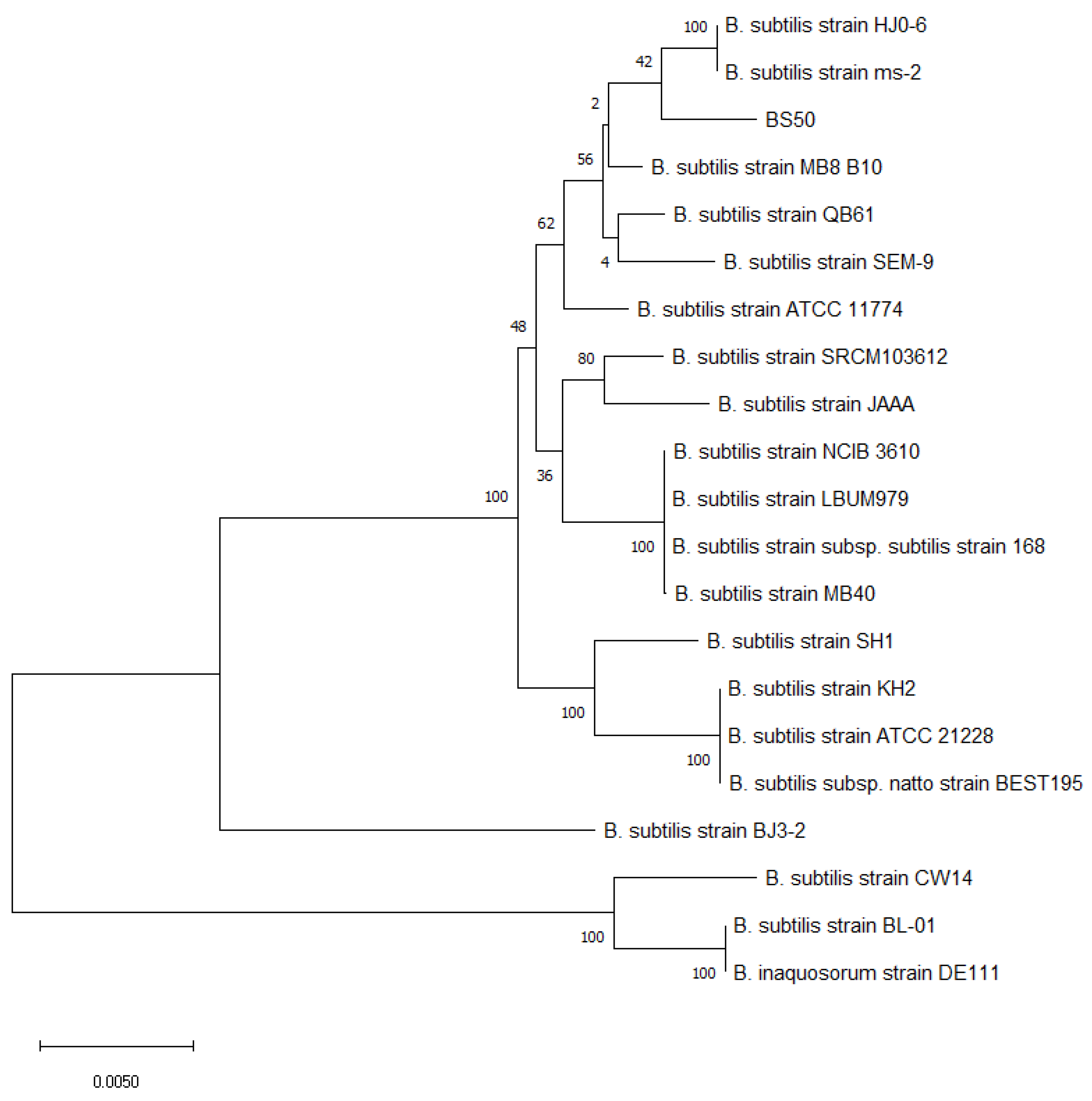
3.1. Taxonomic Classification of BS50
3.2. BLASTn Screen for Known Bacillus Toxins
3.3. BLASTx Screen for Known Bacillus Toxins
3.4. In Silico PCR Amplification of BS50 for Bacillus Toxins
3.5. Secondary Metabolite Screen via AntiSMASH
3.6. Secreted Protein via SignalP 6.0 Analysis
3.7. Virulence Factor Screen via VFDB
3.8. Antibiotic Resistance Gene Analysis
3.9. Insertion Sequences and Mobile Genetic Element Analysis
3.10. MIC Evaluation of BS50 against Eight Antibiotics
3.11. Blood Hemolysis Assay
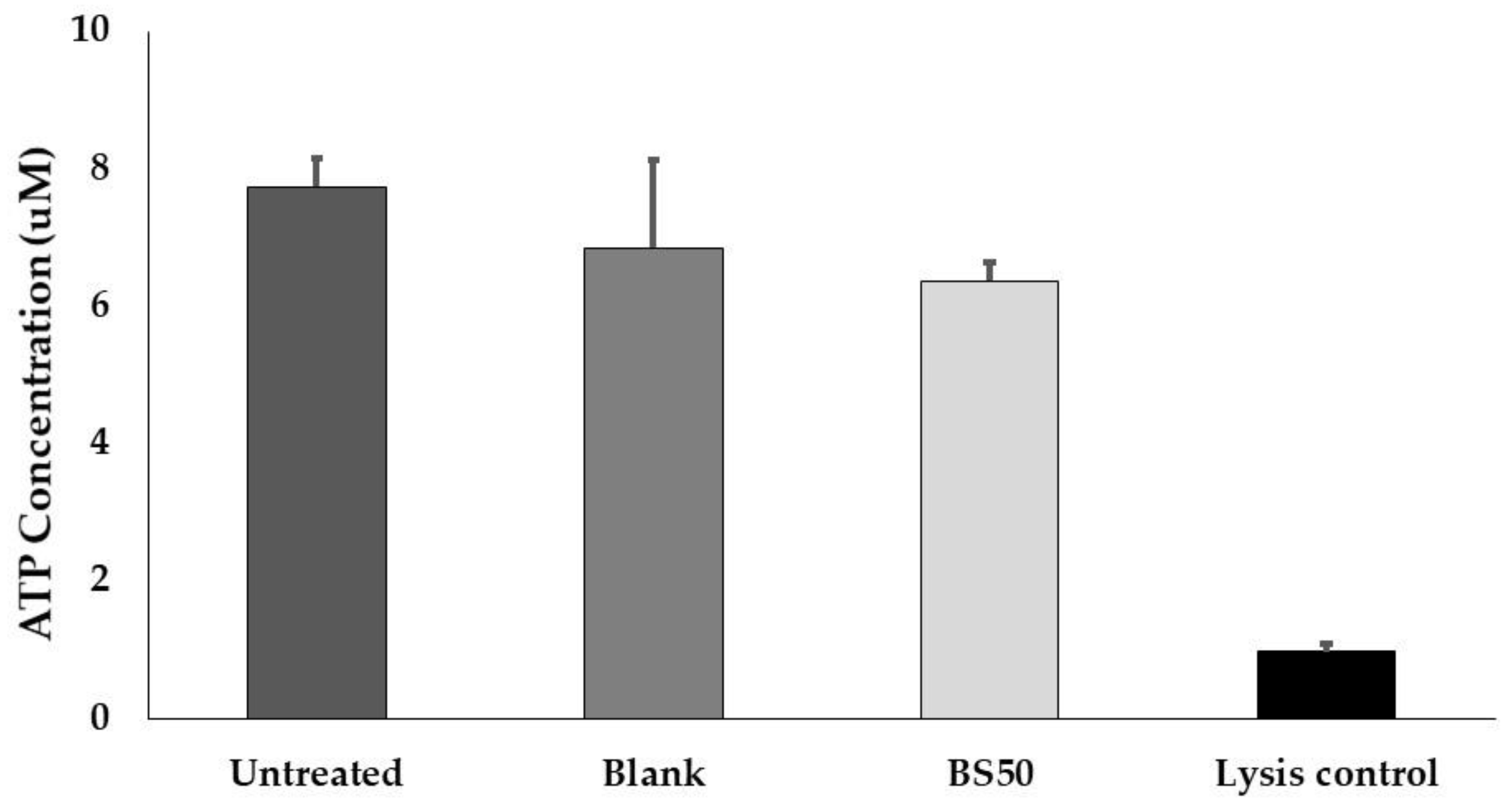
3.12. Caco-2 Cell Viability Assay
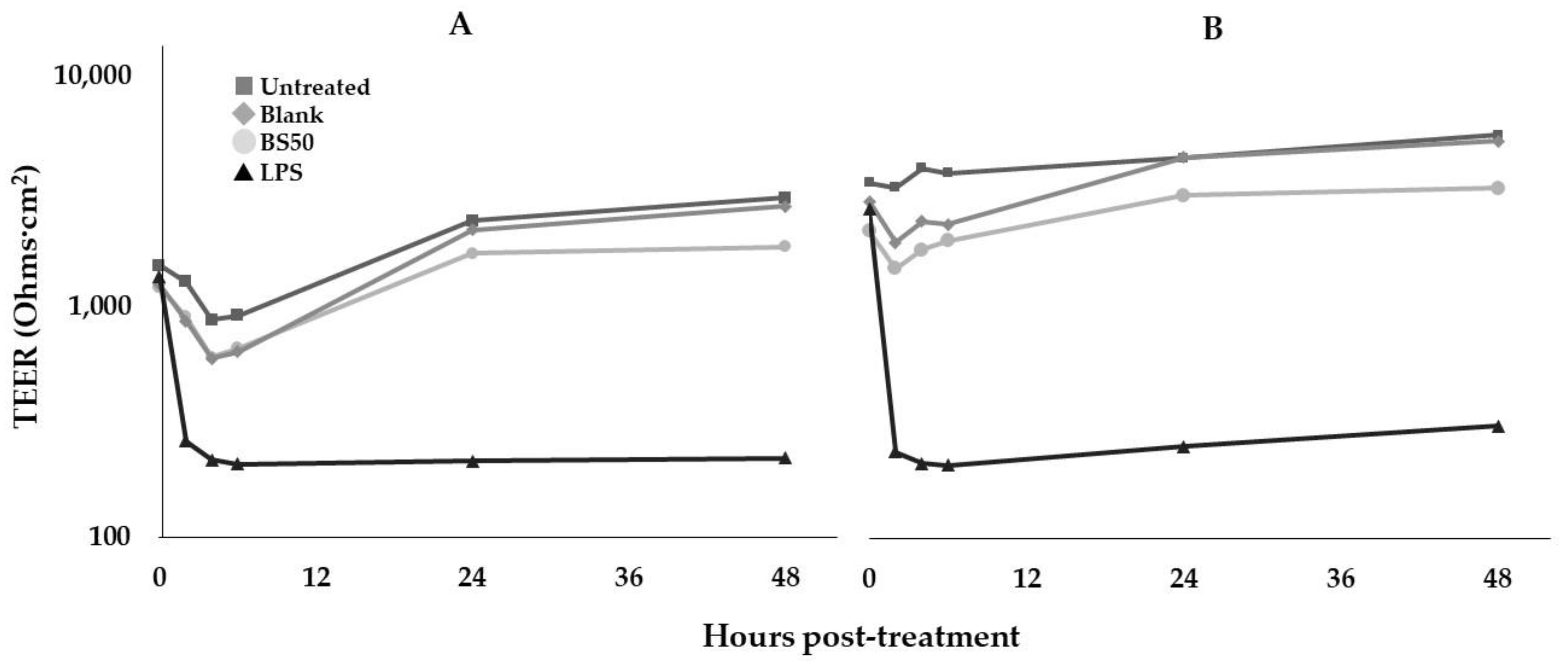
3.13. Caco-2 Cell TEER Assay
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Harwood, C.R. Bacillus subtilis and its relatives: Molecular biological and industrial workhorses. Trends Biotechnol. 1992, 10, 247–256. [Google Scholar] [CrossRef]
- Su, Y.; Liu, C.; Fang, H.; Zhang, D. Bacillus subtilis: A universal cell factory for industry, agriculture, biomaterials and medicine. Microb. Cell Fact. 2020, 19, 173. [Google Scholar] [CrossRef]
- Hill, C.; Guarner, F.; Reid, G.; Gibson, G.R.; Merenstein, D.J.; Pot, B.; Morelli, L.; Canani, R.B.; Flint, H.J.; Salminen, S.; et al. Expert consensus document: The international scientific association for probiotics and prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 506–514. [Google Scholar] [CrossRef] [PubMed]
- Patterson, E.; Griffin, S.M.; Ibarra, A.; Ellsiepen, E.; Hellhammer, J. Lacticaseibacillus paracasei Lpc-37® improves psychological and physiological markers of stress and anxiety in healthy adults: A randomized, double-blind, placebo-controlled and parallel clinical trial (the Sisu study). Neurobiol. Stress 2020, 13, 100277. [Google Scholar] [CrossRef] [PubMed]
- Venkataraman, R.; Madempudi, R.S.; Neelamraju, J.; Ahire, J.J.; Vinay, H.R.; Lal, A.; Thomas, G.; Stephen, S. Effect of Multi-strain Probiotic Formulation on Students Facing Examination Stress: A Double-Blind, Placebo-Controlled Study. Probiotics Antimicrob. Proteins 2021, 13, 12–18. [Google Scholar] [CrossRef]
- Wauters, L.; Van Oudenhove, L.; Accarie, A.; Geboers, K.; Geysen, H.; Toth, J.; Luypaerts, A.; Verbeke, K.; Smokvina, T.; Raes, J.; et al. Lactobacillus rhamnosus CNCM I-3690 decreases subjective academic stress in healthy adults: A randomized placebo-controlled trial. Gut Microbes 2022, 14, 2031695. [Google Scholar] [CrossRef]
- Plaza-Diaz, J.; Ruiz-Ojeda, F.J.; Gil-Campos, M.; Gil, A. Mechanisms of Action of Probiotics. Adv. Nutr. 2019, 10, S49–S66. [Google Scholar] [CrossRef]
- Sanders, M.E.; Merenstein, D.J.; Reid, G.; Gibson, G.R.; Rastall, R.A. Probiotics and prebiotics in intestinal health and disease: From biology to the clinic. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 605–616. [Google Scholar] [CrossRef]
- Cutting, S.M. Bacillus probiotics. Food Microbiol. 2011, 28, 214–220. [Google Scholar] [CrossRef]
- Shurtleff, W.; Aoyagi, A. History of Natto and Its Relatives. Soyinfo Center 2012. [Google Scholar]
- Jeon, H.L.; Lee, N.K.; Yang, S.J.; Kim, W.S.; Paik, H.D. Probiotic characterization of Bacillus subtilis P223 isolated from kimchi. Food Sci. Biotechnol. 2017, 26, 1641–1648. [Google Scholar] [CrossRef] [PubMed]
- Kotb, E. Purification and partial characterization of serine fibrinolytic enzyme from Bacillus megaterium KSK-07 isolated from kishk, a traditional Egyptian fermented food. Appl. Biochem. Microbiol. 2015, 51, 34–43. [Google Scholar] [CrossRef]
- Chantawannakul, P.; Oncharoen, A.; Klanbut, K.; Chukeatirote, E.; Lumyong, S. Characterization of proteases of Bacillus subtilis strain 38 isolated from traditionally fermented soybean in Northern Thailand. Sci. Asia 2002, 28, 241. [Google Scholar] [CrossRef]
- Inatsu, Y.; Nakamura, N.; Yuriko, Y.; Fushimi, T.; Watanasiritum, L.; Kawamoto, S. Characterization of Bacillus subtilis strains in Thua nao, a traditional fermented soybean food in northern Thailand. Lett. Appl. Microbiol. 2006, 43, 237–242. [Google Scholar] [CrossRef]
- Lee, N.K.; Kim, W.S.; Paik, H.D. Bacillus strains as human probiotics: Characterization, safety, microbiome, and probiotic carrier. Food Sci. Biotechnol. 2019, 28, 1297–1305. [Google Scholar] [CrossRef]
- Sorokulova, I.B.; Pinchuk, I.V.; Denayrolles, M.; Osipova, I.G.; Huang, J.M.; Cutting, S.M.; Urdaci, M.C. The safety of two Bacillus probiotic strains for human use. Dig. Dis. Sci. 2008, 53, 954–963. [Google Scholar] [CrossRef]
- Spears, J.L.; Kramer, R.; Nikiforov, A.I.; Rihner, M.O.; Lambert, E.A. Safety Assessment of Bacillus subtilis MB40 for Use in Foods and Dietary Supplements. Nutrients 2021, 13, 733. [Google Scholar] [CrossRef]
- Tompkins, T.A.; Hagen, K.E.; Wallace, T.D.; Fillion-Forté, V. Safety evaluation of two bacterial strains used in asian probiotic products. Can. J. Microbiol. 2008, 54, 391–400. [Google Scholar] [CrossRef]
- Hong, H.A.; Huang, J.M.; Khaneja, R.; Hiep, L.V.; Urdaci, M.C.; Cutting, S.M. The safety of Bacillus subtilis and Bacillus indicus as food probiotics. J. Appl. Microbiol. 2008, 105, 510–520. [Google Scholar] [CrossRef]
- Li, A.; Jiang, X.; Wang, Y.; Zhang, L.; Zhang, H.; Mehmood, K.; Li, Z.; Waqas, M.; Li, J. The impact of Bacillus subtilis 18 isolated from Tibetan yaks on growth performance and gut microbial community in mice. Microb. Pathog. 2019, 128, 153–161. [Google Scholar] [CrossRef]
- Dolin, B.J. Effects of a propietary Bacillus coagulans preparation on symptoms of diarrhea-predominant irritable bowel syndrome. Methods Find. Exp. Clin. Pharmacol. 2009, 31, 655. [Google Scholar] [CrossRef] [PubMed]
- Hun, L. Original Research: Bacillus coagulans Significantly Improved Abdominal Pain and Bloating in Patients with IBS. Postgrad. Med. 2009, 121, 119–124. [Google Scholar] [CrossRef] [PubMed]
- Majeed, M.; Nagabhushanam, K.; Natarajan, S.; Sivakumar, A.; Ali, F.; Pande, A.; Majeed, S.; Karri, S.K. Bacillus coagulans MTCC 5856 supplementation in the management of diarrhea predominant Irritable Bowel Syndrome: A double blind randomized placebo controlled pilot clinical study. Nutr. J. 2016, 15, 21. [Google Scholar] [CrossRef]
- Madempudi, R.S.; Ahire, J.J.; Neelamraju, J.; Tripathi, A.; Nanal, S. Randomized clinical trial: The effect of probiotic Bacillus coagulans Unique IS2 vs. placebo on the symptoms management of irritable bowel syndrome in adults. Sci. Rep. 2019, 9, 12210. [Google Scholar] [CrossRef]
- Gupta, A.K.; Maity, C. Efficacy and safety of Bacillus coagulans LBSC in irritable bowel syndrome: A prospective, interventional, randomized, double-blind, placebo-controlled clinical study [CONSORT Compliant]. Medicine 2021, 100, e23641. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Sudha, K.; Bennur, S.; Dhanasekar, K. A prospective, randomized, open-label, placebo-controlled comparative study of Bacillus coagulans GBI-30,6086 with digestive enzymes in improving indigestion in geriatric population. J. Fam. Med. Prim. Care 2020, 9, 1108. [Google Scholar] [CrossRef]
- Wauters, L.; Slaets, H.; De Paepe, K.; Ceulemans, M.; Wetzels, S.; Geboers, K.; Toth, J.; Thys, W.; Dybajlo, R.; Walgraeve, D.; et al. Efficacy and safety of spore-forming probiotics in the treatment of functional dyspepsia: A pilot randomised, double-blind, placebo-controlled trial. Lancet Gastroenterol. Hepatol. 2021, 6, 784–792. [Google Scholar] [CrossRef]
- Tompkins, T.; Xu, X.; Ahmarani, J. A comprehensive review of post-market clinical studies performed in adults with an Asian probiotic formulation. Benef. Microbes 2010, 1, 93–106. [Google Scholar] [CrossRef]
- Zeng, J.; Wang, C.T.; Zhang, F.S.; Qi, F.; Wang, S.F.; Ma, S.; Wu, T.J.; Tian, H.; Tian, Z.T.; Zhang, S.L.; et al. Effect of probiotics on the incidence of ventilator-associated pneumonia in critically ill patients: A randomized controlled multicenter trial. Intensive Care Med. 2016, 42, 1018–1028. [Google Scholar] [CrossRef]
- Maity, C.; Gupta, A.K. A prospective, interventional, randomized, double-blind, placebo-controlled clinical study to evaluate the efficacy and safety of Bacillus coagulans LBSC in the treatment of acute diarrhea with abdominal discomfort. Eur. J. Clin. Pharmacol. 2019, 75, 21–31. [Google Scholar] [CrossRef]
- Stecker, R.A.; Moon, J.M.; Russo, T.J.; Ratliff, K.M.; Mumford, P.W.; Jäger, R.; Purpura, M.; Kerksick, C.M. Bacillus coagulans GBI-30, 6086 improves amino acid absorption from milk protein. Nutr. Metab. 2020, 17, 93. [Google Scholar] [CrossRef] [PubMed]
- Tarik, M.; Ramakrishnan, L.; Bhatia, N.; Goswami, R.; Kandasamy, D.; Roy, A.; Chandran, D.S.; Singh, A.; Upadhyay, A.D.; Kalaivani, M.; et al. The effect of Bacillus coagulans Unique IS-2 supplementation on plasma amino acid levels and muscle strength in resistance trained males consuming whey protein: A double-blind, placebo-controlled study. Eur. J. Nutr. 2022. [Google Scholar] [CrossRef] [PubMed]
- Hanifi, A.; Culpepper, T.; Mai, V.; Anand, A.; Ford, A.L.; Ukhanova, M.; Christman, M.; Tompkins, T.A.; Dahl, W.J. Evaluation of Bacillus subtilis R0179 on gastrointestinal viability and general wellness: A randomised, double-blind, placebo-controlled trial in healthy adults. Benef. Microbes 2015, 6, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Hatanaka, M.; Yamamoto, K.; Suzuki, N.; Iio, S.; Takara, T.; Morita, H.; Takimoto, T.; Nakamura, T. Effect of Bacillus subtilis C-3102 on loose stools in healthy volunteers. Benef. Microbes 2018, 9, 357–365. [Google Scholar] [CrossRef] [PubMed]
- Penet, C.; Kramer, R.; Little, R.; Spears, J.L.; Parker, J.; Iyer, J.K.; Guthrie, N.; Evans, M. A Randomized, Double-blind, Placebo-controlled, Parallel Study Evaluating the Efficacy of Bacillus subtilis MB40 to Reduce Abdominal Discomfort, Gas, and Bloating. Altern. Ther. Health Med. 2021, 27, 146–157. [Google Scholar]
- Hatanaka, M.; Kanzato, H.; Tsuda, R.; Nadaoka, I.; Yasue, M.; Hoshino, T.; Iio, S.; Takara, T. Safety evaluation of the excessive intake of Bacillus subtilis C-3102 in healthy Japanese adults: A randomized, placebo-controlled, double-blind, parallel-group, comparison trial. Toxicol. Rep. 2020, 7, 46–58. [Google Scholar] [CrossRef]
- Lefevre, M.; Racedo, S.M.; Ripert, G.; Housez, B.; Cazaubiel, M.; Maudet, C.; Jüsten, P.; Marteau, P.; Urdaci, M.C. Probiotic strain Bacillus subtilis CU1 stimulates immune system of elderly during common infectious disease period: A randomized, double-blind placebo-controlled study. Immun. Ageing 2015, 12, 24. [Google Scholar] [CrossRef]
- Horosheva, T.V.; Vodyanoy, V.; Sorokulova, I. Efficacy of Bacillus probiotics in prevention of antibiotic-associated diarrhoea: A randomized, double-blind, placebo-controlled clinical trial. JMM Case Rep. 2014, 1, e004036. [Google Scholar] [CrossRef]
- Lefevre, M.; Racedo, S.M.; Denayrolles, M.; Ripert, G.; Desfougères, T.; Lobach, A.R.; Simon, R.; Pélerin, F.; Jüsten, P.; Urdaci, M.C. Safety assessment of Bacillus subtilis CU1 for use as a probiotic in humans. Regul. Toxicol. Pharmacol. 2017, 83, 54–65. [Google Scholar] [CrossRef]
- Kalman, D.S.; Schwartz, H.I.; Alvarez, P.; Feldman, S.; Pezzullo, J.C.; Krieger, D.R. A prospective, randomized, double-blind, placebo-controlled parallel-group dual site trial to evaluate the effects of a Bacillus coagulans-based product on functional intestinal gas symptoms. BMC Gastroenterol. 2009, 9, 85. [Google Scholar] [CrossRef]
- Freedman, K.E.; Hill, J.L.; Wei, Y.; Vazquez, A.R.; Grubb, D.S.; Trotter, R.E.; Wrigley, S.D.; Johnson, S.A.; Foster, M.T.; Weir, T.L. Examining the gastrointestinal and immunomodulatory effects of the novel probiotic Bacillus subtilis de111. Int. J. Mol. Sci. 2021, 22, 2453. [Google Scholar] [CrossRef]
- Paytuví-Gallart, A.; Sanseverino, W.; Winger, A.M. Daily intake of probiotic strain Bacillus subtilis DE111 supports a healthy microbiome in children attending day-care. Benef. Microbes 2020, 11, 611–620. [Google Scholar] [CrossRef]
- Toohey, J.C.; Townsend, J.R.; Johnson, S.B.; Toy, A.M.; Vantrease, W.C.; Bender, D.; Crimi, C.C.; Stowers, K.L.; Ruiz, M.D.; VanDusseldorp, T.A.; et al. Effects of Probiotic (Bacillus subtilis) Supplementation During Offseason Resistance Training in Female Division I Athletes. J. Strength Cond. Res. 2020, 34, 3173–3181. [Google Scholar] [CrossRef] [PubMed]
- Townsend, J.R.; Bender, D.; Vantrease, W.C.; Sapp, P.A.; Toy, A.M.; Woods, C.A.; Johnson, K.D. Effects of probiotic (Bacillus subtilis de111) supplementation on immune function, hormonal status, and physical performance in division i baseball players. Sports 2018, 6, 70. [Google Scholar] [CrossRef]
- Townsend, J.R.; Vantrease, W.C.; Jones, M.D.; Sapp, P.A.; Johnson, K.D.; Beuning, C.N.; Haase, A.A.; Boot, C.M. Plasma amino acid response to whey protein ingestion following 28 days of probiotic (Bacillus subtilis de111) supplementation in active men and women. J. Funct. Morphol. Kinesiol. 2021, 6, 1. [Google Scholar] [CrossRef] [PubMed]
- Trotter, R.E.; Vazquez, A.R.; Grubb, D.S.; Freedman, K.E.; Grabos, L.E.; Jones, S.; Gentile, C.L.; Melby, C.L.; Johnson, S.A.; Weir, T.L. Bacillus subtilis DE111 intake may improve blood lipids and endothelial function in healthy adults. Benef. Microbes 2020, 11, 621–630. [Google Scholar] [CrossRef] [PubMed]
- Colom, J.; Freitas, D.; Simon, A.; Brodkorb, A.; Buckley, M.; Deaton, J.; Deaton, J. Presence and Germination of the Probiotic Bacillus subtilis DE111® in the Human Small Intestinal Tract: A Randomized, Crossover, Double- Blind, and Placebo-Controlled Study. Front. Microbiol. 2021, 12, 2189. [Google Scholar] [CrossRef] [PubMed]
- Cuentas, A.M.; Deaton, J.; Khan, S.; Davidson, J.; Ardita, C. The Effect of Bacillus subtilis DE111 on the Daily Bowel Movement Profile for People with Occasional Gastrointestinal Irregularity. J. Probiotics Health 2017, 5, 4. [Google Scholar] [CrossRef]
- FDA GRN. GRAS Notification 905. Available online: https://www.fda.gov/media/139501/download (accessed on 18 April 2022).
- FDA GRN. GRAS Notification 831. Available online: https://www.fda.gov/media/132389/download (accessed on 18 April 2022).
- FDA GRN. GRAS Notification 955. Available online: https://www.fda.gov/media/146438/download (accessed on 18 April 2022).
- FDA GRN. GRAS Notification 969. Available online: https://fda.report/media/150352/GRAS-Notice-GRN-969-Bacillus-subtilis.pdf (accessed on 18 April 2022).
- FDA GRN. GRAS Notification 956. Available online: https://www.fda.gov/media/146998/download (accessed on 18 April 2022).
- Koutsoumanis, K.; Allende, A.; Alvarez-Ordóñez, A.; Bolton, D.; Bover-Cid, S.; Chemaly, M.; Davies, R.; De Cesare, A.; Hilbert, F.; Lindqvist, R.; et al. Update of the list of QPS-recommended biological agents intentionally added to food or feed as notified to EFSA 14: Suitability of taxonomic units notified to EFSA until March 2021. EFSA J. 2021, 19, e06689. [Google Scholar] [CrossRef]
- Damgaard, P.H.; Granum, P.E.; Bresciani, J.; Torregrossa, M.V.; Eilenberg, J.; Valentino, L. Characterization of Bacillus thuringiensis isolated from infections in burn wounds. FEMS Immunol. Med. Microbiol. 1997, 18, 47–53. [Google Scholar] [CrossRef]
- Hernandez, E.; Ramisse, F.; Ducoureau, J.P.; Cruel, T.; Cavallo, J.D. Bacillus thuringiensis subsp. konkukian (Serotype H34) superinfection: Case report and experimental evidence of pathogenicity in immunosuppressed mice. J. Clin. Microbiol. 1998, 36, 2138–2139. [Google Scholar] [CrossRef]
- Little, S.F.; Ivins, B.E. Molecular pathogenesis of Bacillus anthracis infection. Microbes Infect. 1999, 1, 131–139. [Google Scholar] [CrossRef]
- Kotiranta, A.; Lounatmaa, K.; Haapasalo, M. Epidemiology and pathogenesis of Bacillus cereus infections. Microbes Infect. 2000, 2, 189–198. [Google Scholar] [CrossRef]
- Raymond, B.; Johnston, P.R.; Nielsen-LeRoux, C.; Lereclus, D.; Crickmore, N. Bacillus thuringiensis: An impotent pathogen? Trends Microbiol. 2010, 18, 189–194. [Google Scholar] [CrossRef] [PubMed]
- Dietrich, R.; Jessberger, N.; Ehling-Schulz, M.; Märtlbauer, E.; Granum, P.E. The Food Poisoning Toxins of Bacillus cereus. Toxins 2021, 13, 98. [Google Scholar] [CrossRef] [PubMed]
- Schoeni, J.L.; Lee Wong, A.C. Bacillus cereus Food Poisoning and Its Toxins. J. Food Prot. 2005, 68, 636–648. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, S.; Chattopadhyay, M.K.; Grossart, H.P. The multifaceted roles of antibiotics and antibiotic resistance in nature. Front. Microbiol. 2013, 4, 47. [Google Scholar] [CrossRef] [PubMed]
- Alekshun, M.N.; Levy, S.B. Molecular Mechanisms of Antibacterial Multidrug Resistance. Cell 2007, 128, 1037–1050. [Google Scholar] [CrossRef]
- van Reenen, C.A.; Dicks, L.M.T. Horizontal gene transfer amongst probiotic lactic acid bacteria and other intestinal microbiota: What are the possibilities? A review. Arch. Microbiol. 2011, 193, 157–168. [Google Scholar] [CrossRef]
- Santagati, M.; Campanile, F.; Stefani, S. Genomic diversification of enterococci in hosts: The role of the mobilome. Front. Microbiol. 2012, 3, 95. [Google Scholar] [CrossRef]
- Blair, J.M.A.; Webber, M.A.; Baylay, A.J.; Ogbolu, D.O.; Piddock, L.J.V. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 2015, 13, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Wozniak, R.A.F.; Waldor, M.K. Integrative and conjugative elements: Mosaic mobile genetic elements enabling dynamic lateral gene flow. Nat. Rev. Microbiol. 2010, 8, 552–563. [Google Scholar] [CrossRef] [PubMed]
- Kolmogorov, M.; Yuan, J.; Lin, Y.; Pevzner, P.A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 2019, 37, 540–546. [Google Scholar] [CrossRef] [PubMed]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Burkholder, P.R.; Giles, N.H. Induced biochemical mutations in Bacillus subtilis. Am. J. Bot. 1947, 34, 345–348. [Google Scholar] [CrossRef]
- Zeigler, D.R.; Prágai, Z.; Rodriguez, S.; Chevreux, B.; Muffler, A.; Albert, T.; Bai, R.; Wyss, M.; Perkins, J.B. The origins of 168, W23, and other Bacillus subtilis legacy strains. J. Bacteriol. 2008, 190, 6983–6995. [Google Scholar] [CrossRef]
- Kubo, Y.; Rooney, A.P.; Tsukakoshi, Y.; Nakagawa, R.; Hasegawa, H.; Kimura, K. Phylogenetic analysis of Bacillus subtilis strains applicable to natto (fermented soybean) production. Appl. Environ. Microbiol. 2011, 77, 6463–6469. [Google Scholar] [CrossRef]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2018, 20, 1160–1166. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Tamura, K.; Nei, M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 1993, 10, 512–526. [Google Scholar] [CrossRef] [PubMed]
- Mei, X.; Xu, K.; Yang, L.; Yuan, Z.; Mahillon, J.; Hu, X. The genetic diversity of cereulide biosynthesis gene cluster indicates a composite transposon Tnces in emetic Bacillus weihenstephanensis. BMC Microbiol. 2014, 14, 149. [Google Scholar] [CrossRef]
- Ehling-Schulz, M.; Fricker, M.; Grallert, H.; Rieck, P.; Wagner, M.; Scherer, S. Cereulide synthetase gene cluster from emetic Bacillus cereus: Structure and location on a mega virulence plasmid related to Bacillus anthracis toxin plasmid pXO1. BMC Microbiol. 2006, 6, 20. [Google Scholar] [CrossRef]
- San Millán, R.M.; Martínez-Ballesteros, I.; Rementeria, A.; Garaizar, J.; Bikandi, J. Online exercise for the design and simulation of PCR and PCR-RFLP experiments. BMC Res. Notes 2013, 6, 2–5. [Google Scholar] [CrossRef] [PubMed]
- Agata, N.; Ohta, M.; Mori, M.; Isobe, M. A novel dodecadepsipeptide, cereulide, is an emetic toxin of Bacillus cereus. FEMS Microbiol. Lett. 1995, 129, 17–19. [Google Scholar] [CrossRef] [PubMed]
- Asano, S.I.; Nukumizu, Y.; Bando, H.; Iizuka, T.; Yamamoto, T. Cloning of novel enterotoxin genes from Bacillus cereus and Bacillus thuringiensis. Appl. Environ. Microbiol. 1997, 63, 1054–1057. [Google Scholar] [CrossRef]
- Mäntynen, V.; Lindström, K. A rapid PCR-based DNA test for enterotoxic Bacillus cereus. Appl. Environ. Microbiol. 1998, 64, 1634–1639. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Kloosterman, A.M.; Charlop-Powers, Z.; van Wezel, G.P.; Medema, M.H.; Weber, T. antiSMASH 6.0: Improving cluster detection and comparison capabilities. Nucleic Acids Res. 2021, 49, W29–W35. [Google Scholar] [CrossRef]
- Wattam, A.R.; Abraham, D.; Dalay, O.; Disz, T.L.; Driscoll, T.; Gabbard, J.L.; Gillespie, J.J.; Gough, R.; Hix, D.; Kenyon, R.; et al. PATRIC, the bacterial bioinformatics database and analysis resource. Nucleic Acids Res. 2014, 42, 581–591. [Google Scholar] [CrossRef]
- Brettin, T.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Olsen, G.J.; Olson, R.; Overbeek, R.; Parrello, B.; Pusch, G.D.; et al. RASTtk: A modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci. Rep. 2015, 5, 8365. [Google Scholar] [CrossRef]
- Teufel, F.; Almagro Armenteros, J.J.; Johansen, A.R.; Gíslason, M.H.; Pihl, S.I.; Tsirigos, K.D.; Winther, O.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat. Biotechnol. 2022. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Zheng, D.; Jin, Q.; Chen, L.; Yang, J. VFDB 2019: A comparative pathogenomic platform with an interactive web interface. Nucleic Acids Res. 2019, 47, D687–D692. [Google Scholar] [CrossRef]
- McArthur, A.G.; Waglechner, N.; Nizam, F.; Yan, A.; Azad, M.A.; Baylay, A.J.; Bhullar, K.; Canova, M.J.; De Pascale, G.; Ejim, L.; et al. The comprehensive antibiotic resistance database. Antimicrob. Agents Chemother. 2013, 57, 3348–3357. [Google Scholar] [CrossRef]
- Alcock, B.P.; Raphenya, A.R.; Lau, T.T.Y.; Tsang, K.K.; Bouchard, M.; Edalatmand, A.; Huynh, W.; Nguyen, A.L.V.; Cheng, A.A.; Liu, S.; et al. CARD 2020: Antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 2020, 48, D517–D525. [Google Scholar] [CrossRef] [PubMed]
- Leplae, R.; Lima-Mendez, G.; Toussaint, A. ACLAME: A CLAssification of mobile genetic elements, update 2010. Nucleic Acids Res. 2009, 38, 57–61. [Google Scholar] [CrossRef]
- Siguier, P.; Perochon, J.; Lestrade, L.; Mahillon, J.; Chandler, M. ISfinder: The reference centre for bacterial insertion sequences. Nucleic Acids Res. 2006, 34, 32–36. [Google Scholar] [CrossRef] [PubMed]
- Ben Maamar, S.; Glawe, A.J.; Brown, T.K.; Hellgeth, N.; Hu, J.; Wang, J.P.; Huttenhower, C.; Hartmann, E.M. Mobilizable antibiotic resistance genes are present in dust microbial communities. PLoS Pathog. 2020, 16, e1008211. [Google Scholar] [CrossRef]
- Weinstein, M.P.; Patel, J.B.; Burnhman, C.-A.; ZImmer, B.L. M07 Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2018; p. 91. [Google Scholar]
- Rychen, G.; Aquilina, G.; Azimonti, G.; Bampidis, V.; Bastos, M.D.L.; Bories, G.; Chesson, A.; Cocconcelli, P.S.; Flachowsky, G.; Gropp, J.; et al. Guidance on the characterisation of microorganisms used as feed additives or as production organisms. EFSA J. 2018, 16, e05206. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA) European Food Safety Authority, EFSA Panel on Additives and Products or Substances used in Animal Feed (FEEDAP). Scientific opinion Guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance. EFSA J. 2012, 10, 2740. [Google Scholar] [CrossRef]
- Konstantinidis, K.T.; Tiedje, J.M. Genomic insights that advance the species definition for prokaryotes. Proc. Natl. Acad. Sci. USA 2005, 102, 2567–2572. [Google Scholar] [CrossRef]
- Richter, M.; Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl. Acad. Sci. USA 2009, 106, 19126–19131. [Google Scholar] [CrossRef]
- Chaucheyras-Durand, F.; Durand, H. Probiotics in Animal Nutrition and Health. Benef. Microbes 2016, 1, 3–9. [Google Scholar] [CrossRef] [PubMed]
- From, C.; Pukall, R.; Schumann, P.; Hormazábal, V.; Granum, P.E. Toxin-producing ability among Bacillus spp. outside the Bacillus cereus group. Appl. Environ. Microbiol. 2005, 71, 1178–1183. [Google Scholar] [CrossRef] [PubMed]
- Salkinoja-Salonen, M.S.; Vuorio, R.; Andersson, M.A.; Kämpfer, P.; Andersson, M.C.; Honkanen-Buzalski, T.; Scoging, A.C. Toxigenic strains of Bacillus licheniformis related to food poisoning. Appl. Environ. Microbiol. 1999, 65, 4637–4645. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.M.W.; Sutherland, A.D.; Aidoo, K.E.; Logan, N.A. Heat-stable toxin production by strains of Bacillus cereus, Bacillus firmus, Bacillus megaterium, Bacillus simplex and Bacillus licheniformis. FEMS Microbiol. Lett. 2005, 242, 313–317. [Google Scholar] [CrossRef][Green Version]
- Harwood, C.R.; Mouillon, J.M.; Pohl, S.; Arnau, J. Secondary metabolite production and the safety of industrially important members of the Bacillus subtilis group. FEMS Microbiol. Rev. 2018, 42, 721–738. [Google Scholar] [CrossRef]
- Raza, W.; Hussain, Q.; Shen, Q. Production, regulation and transportation of bacillibactin in Bacillus subtilis. J. Chem. Soc. Pak. 2012, 34, 996–1002. [Google Scholar]
- May, J.J.; Wendrich, T.M.; Marahiel, M.A. The dhb Operon of Bacillus subtilis Encodes the Biosynthetic Template for the Catecholic Siderophore 2,3-Dihydroxybenzoate-Glycine-Threonine Trimeric Ester Bacillibactin. J. Biol. Chem. 2001, 276, 7209–7217. [Google Scholar] [CrossRef]
- Kaur, D.; Yantiri, F.; Rajagopalan, S.; Kumar, J.; Mo, J.Q.; Boonplueang, R.; Viswanath, V.; Jacobs, R.; Yang, L.; Beal, M.F.; et al. Genetic or Pharmacological Iron Chelation Prevents MPTP-Induced Neurotoxicity In Vivo. Neuron 2003, 37, 899–909. [Google Scholar] [CrossRef]
- Piewngam, P.; Zheng, Y.; Nguyen, T.H.; Dickey, S.W.; Joo, H.-S.; Villaruz, A.E.; Glose, K.A.; Fisher, E.L.; Hunt, R.L.; Li, B.; et al. Pathogen elimination by probiotic Bacillus via signalling interference. Nature 2018, 562, 532–537. [Google Scholar] [CrossRef]
- Kenig, M.; Abraham, E.P. Antimicrobial Activities and Antagonists of Bacilysin and Anticapsin. J. Gen. Microbiol. 1976, 94, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Nannan, C.; Vu, H.Q.; Gillis, A.; Caulier, S.; Nguyen, T.T.T.; Mahillon, J. Bacilysin within the Bacillus subtilis group: Gene prevalence versus antagonistic activity against Gram-negative foodborne pathogens. J. Biotechnol. 2021, 327, 28–35. [Google Scholar] [CrossRef] [PubMed]
- Steinborn, G.; Hajirezaei, M.R.; Hofemeister, J. bac genes for recombinant bacilysin and anticapsin production in Bacillus host strains. Arch. Microbiol. 2005, 183, 71–79. [Google Scholar] [CrossRef]
- Chen, X.H.; Vater, J.; Piel, J.; Franke, P.; Scholz, R.; Schneider, K.; Koumoutsi, A.; Hitzeroth, G.; Grammel, N.; Strittmatter, A.W.; et al. Structural and functional characterization of three polyketide synthase gene clusters in Bacillus amyloliquefaciens FZB 42. J. Bacteriol. 2006, 188, 4024–4036. [Google Scholar] [CrossRef]
- Patel, P.S.; Huang, S.; Fisher, S.; Pirnik, D.; Aklonis, C.; Dean, L.; Meyers, E.; Fernandes, P.; Mayerl, F. Bacillaene, a novel inhibitor of procaryotic protein synthesis produced by Bacillus subtilis: Production, taxonomy, isolation, physico-chemical characterization and biological activity. J. Antibiot. 1995, 48, 997–1003. [Google Scholar] [CrossRef]
- Li, H.; Han, X.; Dong, Y.; Xu, S.; Chen, C.; Feng, Y.; Cui, Q.; Li, W. Bacillaenes: Decomposition Trigger Point and Biofilm Enhancement in Bacillus. ACS Omega 2021, 6, 1093–1098. [Google Scholar] [CrossRef] [PubMed]
- Butcher, R.A.; Schroeder, F.C.; Fischbach, M.A.; Straight, P.D.; Kolter, R.; Walsh, C.T.; Clardy, J. The identification of bacillaene, the product of the PksX megacomplex in Bacillus subtilis. Proc. Natl. Acad. Sci. USA 2007, 104, 1506–1509. [Google Scholar] [CrossRef]
- Sharma, S.; Bhatnagar, R.; Gaur, D. Bacillus anthracis Poly-γ-D-Glutamate Capsule Inhibits Opsonic Phagocytosis by Impeding Complement Activation. Front. Immunol. 2020, 11, 462. [Google Scholar] [CrossRef]
- Kocianova, S.; Vuong, C.; Yao, Y.; Voyich, J.M.; Fischer, E.R.; DeLeo, F.R.; Otto, M. Key role of poly-γ-DL-glutamic acid in immune evasion and virulence of Staphylococcus epidermidis. J. Clin. Investig. 2005, 115, 688–694. [Google Scholar] [CrossRef]
- Ijadi Bajestani, M.; Mousavi, S.M.; Mousavi, S.B.; Jafari, A.; Shojaosadati, S.A. Purification of extra cellular poly-γ-glutamic acid as an antibacterial agent using anion exchange chromatography. Int. J. Biol. Macromol. 2018, 113, 142–149. [Google Scholar] [CrossRef]
- Stein, T.; Düsterhus, S.; Stroh, A.; Entian, K.D. Subtilosin Production by Two Bacillus subtilis Subspecies and Variance of the sbo-alb Cluster. Appl. Environ. Microbiol. 2004, 70, 2349–2353. [Google Scholar] [CrossRef]
- Zheng, G.; Slavik, M.F. Isolation, partial purification and characterization of a bacteriocin produced by a newly isolated Bacillus subtilis strain. Lett. Appl. Microbiol. 1999, 28, 363–367. [Google Scholar] [CrossRef] [PubMed]
- Bolocan, A.S.; Pennone, V.; O’Connor, P.M.; Coffey, A.; Nicolau, A.I.; McAuliffe, O.; Jordan, K. Inhibition of Listeria monocytogenes biofilms by bacteriocin-producing bacteria isolated from mushroom substrate. J. Appl. Microbiol. 2017, 122, 279–293. [Google Scholar] [CrossRef]
- Bhat, A.R.; Irorere, V.U.; Bartlett, T.; Hill, D.; Kedia, G.; Morris, M.R.; Charalampopoulos, D.; Radecka, I. Bacillus subtilis natto: A non-toxic source of poly-γ-glutamic acid that could be used as a cryoprotectant for probiotic bacteria. AMB Express 2013, 3, 36. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.-E.; Rhee, J.-H.; Park, C.; Sung, M.-H.; Lee, I.-H. Distribution of poly-gamma-glutamate producers in Korean fermented foods, Cheongkukjang, Doenjang, and Kochujang. Food Sci. Biotechnol. 2005, 14, 704–708. [Google Scholar]
- Laxminarayan, R.; Duse, A.; Wattal, C.; Zaidi, A.K.M.; Wertheim, H.F.L.; Sumpradit, N.; Vlieghe, E.; Hara, G.L.; Gould, I.M.; Goossens, H.; et al. Antibiotic resistance—The need for global solutions. Lancet Infect. Dis. 2013, 13, 1057–1098. [Google Scholar] [CrossRef]
- Davies, J. Origins and evolution of antibiotic resistance. Microbiologia 1996, 12, 9–16. [Google Scholar] [CrossRef]
- Aarts, H.; Margolles, A. Antibiotic resistance genes in food and gut (non pathogenic) bacteria. Bad genes in good bugs. Front. Microbiol. 2014, 5, 2014–2015. [Google Scholar] [CrossRef]
- Agersø, Y.; Bjerre, K.; Brockmann, E.; Johansen, E.; Nielsen, B.; Siezen, R.; Stuer-Lauridsen, B.; Wels, M.; Zeidan, A.A. Putative antibiotic resistance genes present in extant Bacillus licheniformis and Bacillus paralicheniformis strains are probably intrinsic and part of the ancient resistome. PLoS ONE 2019, 14, e0210363. [Google Scholar] [CrossRef]
- Deng, F.; Chen, Y.; Sun, T.; Wu, Y.; Su, Y.; Liu, C.; Zhou, J.; Deng, Y.; Wen, J. Antimicrobial resistance, virulence characteristics and genotypes of Bacillus spp. from probiotic products of diverse origins. Food Res. Int. 2021, 139, 109949. [Google Scholar] [CrossRef]
- Yelin, I.; Flett, K.B.; Merakou, C.; Mehrotra, P.; Stam, J.; Snesrud, E.; Hinkle, M.; Lesho, E.; McGann, P.; McAdam, A.J.; et al. Genomic and epidemiological evidence of bacterial transmission from probiotic capsule to blood in ICU patients. Nat. Med. 2019, 25, 1728–1732. [Google Scholar] [CrossRef] [PubMed]
| Gene | Organism | Accession | Max Score | % Coverage | E-Value | % Identity |
|---|---|---|---|---|---|---|
| gatA | B. subtilis | 938748 | 2405 | 100% | 0 | 98% |
| metG | B. cereus | 61578313 | 911 | 95% | 0 | 71% |
| HblA | B. licheniformis | KM514479.1 | No significant similarity | |||
| HblA | B. cereus | KF681259.1 | 35.6 | 12% | 0.021 | 82% |
| HblC | B. cereus | JQ039142.1 | No significant similarity | |||
| HblD | B. cereus | JQ039158.1 | No significant similarity | |||
| NheA,B,C | B. cereus | DQ885236.1 | 424 | 22% | 9 × 10−118 | 70% |
| NheA,B,C | B. mycoides | DQ153260.1 | 82.8 | 3% | 0.002 | 68% |
| NheA | B. cereus | FN825684.1 | No significant similarity | |||
| NheA,B,C | B. thuringiensis | EU925144.1 | No significant similarity | |||
| entFM | B. cereus | AY789084.1 | 59 | 14% | 9 × 10−09 | 75% |
| cytK | B. mycoides | AY871809.1 | No significant similarity | |||
| cytK | B. licheniformis | KM657965.1 | No significant similarity | |||
| cytK | B. cereus | DQ019311.1 | 37.4 | 1% | 0.044 | 92% |
| HlyII | B. thuringiensis | 564444080 | No significant similarity | |||
| cesHPTABCD | B. cereus | NC_010924.1 | 109 | 50% | 1 × 10−22 | 79% |
| Protein | Organism | Accession | Max Score | E-Value | % Identity |
|---|---|---|---|---|---|
| GatA | B. subtilis | NP_388550.1 | 879 | 0 | 100% |
| MetG | B. cereus | WP_079994147.1 | 946 | 0 | 74.16% |
| cytK | B. mycoides | AAW56196.1 | No significant similarity found | ||
| EntFM | B. cereus | AAX14641.1 | 121 | 4 × 10−29 | 52.21% |
| cytK | B. cereus | AAY84864.1 | No significant similarity found | ||
| NheA | B. mycoides | AAZ82480.1 | No significant similarity found | ||
| NheB | B. mycoides | AAZ82481.1 | No significant similarity found | ||
| NheC | B. mycoides | AAZ82482.1 | No significant similarity found | ||
| NheA | B. cereus | ABI52601.1 | No significant similarity found | ||
| NheB | B. cereus | ABI52602.1 | No significant similarity found | ||
| NheC | B. cereus | ABI52603.1 | No significant similarity found | ||
| NheA | B. cereus | CBL95107.1 | No significant similarity found | ||
| NheA, partial | B. thuringiensis | ACM18211.1 | No significant similarity found | ||
| NheB | B. thuringiensis | ACM18212.1 | No significant similarity found | ||
| NheC, partial | B. thuringiensis | ACM18213.1 | No significant similarity found | ||
| HblD | B. cereus | AFN08801.1 | No significant similarity found | ||
| HblC | B. cereus | AFN08807.1 | No significant similarity found | ||
| HblA | B. cereus | AII31101.1 | No significant similarity found | ||
| HblD | B. licheniformis | AIR07774.1 | No significant similarity found | ||
| HblA | B. licheniformis | AIR07775.1 | No significant similarity found | ||
| cytK | B. licheniformis | AIS75096.1 | No significant similarity found | ||
| CesA | B. cereus | WP_002081542.1 | 1250 | 0 | 34.42% |
| CesB | B. cereus | WP_000953496.1 | 776 | 0 | 36.32% |
| CesC | B. cereus | WP_000590108.1 | 144 | 6 × 10−38 | 31.51% |
| CesD | B. cereus | WP_001008264.1 | No significant similarity found | ||
| CesH | B. cereus | WP_000291846.1 | 53 | 2 × 10−07 | 22.05% |
| CesP | B. cereus | WP_000680399.1 | 129 | 3 × 10−33 | 31.16% |
| CesT | B. cereus | WP_000764755.1 | 116 | 4 × 10−29 | 30.22% |
| Cluster Type | Most Similar Cluster | % Identity |
|---|---|---|
| NRPS (Non-ribosomal peptide synthases) | Surfactin | 78% |
| NRPS | Fengycin | 100% |
| NRPS | Bacillibactin | 100% |
| Other | Bacilysin | 100% |
| Polyketide + NRP | Bacillaene | 100% |
| RiPP: Thiopeptide | Subtilosin A | 100% |
| RiPP: Thiopeptide | Subtilomycin | 100% |
| CDPS | N/A | N/A |
| Terpene | N/A | N/A |
| T3PKS | N/A | N/A |
| Gene | Category | Organism | Accession | % Ident | % Coverage | E |
|---|---|---|---|---|---|---|
| non-ribosomal peptide synthetase, DhbF | Bacillibactin; Nutritional/ Metabolic factor | B. sub 168 | NP_391076 | 99.1 | 99 | 0 |
| 2,3-dihydroxybenzoate adenylase DhbE | Bacillibactin; Nutritional/ Metabolic factor | B. sub 168 | NP_389723 | 99.4 | 100 | 0 |
| isochorismate synthase DhbC | Bacillibactin; Nutritional/ Metabolic factor | B. sub 168 | NP_391078 | 98.5 | 100 | 0 |
| isochorismatase, DhbB | Bacillibactin; Nutritional/ Metabolic factor | B. sub 168 | NP_391471 | 99.7 | 100 | 0 |
| 2,3-dihydroxybenzoate-2,3- dehydrogenase, DhbA | Bacillibactin; Nutritional/ Metabolic factor | B. sub 168 | NP_391079 | 99.2 | 100 | 0 |
| gamma-glutamyltranspeptidase, required for polyglutamate anchoring to peptidoglycan | Capsule; Immune modulation | B. sub 168 | NP_391469 | 98.9 | 100 | 0 |
| CapB, involved in Poly-gamma-glutamate synthesis | Capsule; Immune modulation | B. sub 168 | NP_391077 | 99.3 | 100 | 0 |
| CapA, required for Poly-gamma-glutamate transport | Capsule; Immune modulation | B. sub 168 | NP_391080 | 99.2 | 100 | 0 |
| CapC, involved in Poly-gamma-glutamate synthesis | Capsule; Immune modulation | B. sub 168 | NP_390062 | 100 | 100 | 0 |
| endopeptidase Clp ATP-binding chain C | ClpC; Stress survival | B. sub 168 | NP_391470 | 98.7 | 100 | 0 |
| (tufA) elongation factor Tu | EF-Tu; Adherence | Lm EGD-e | NP_463763 | 72.6 | 89 | 0 |
| (hlyIII) putative membrane hydrolase | Hemolysin III; Exotoxin | Franc. | WP_013922406 | 74.7 | 99 | 1.39 × 10−142 |
| ARO Term (Gene) | AMR Gene Family | Drug Class | % Identity | % Length | RGI Criteria |
|---|---|---|---|---|---|
| ykkD | small multidrug resistance (SMR) antibiotic efflux pump | aminoglycoside antibiotic, tetracycline antibiotic, phenicol antibiotic | 100 | 101.9 | Strict |
| lmrB | ATP-binding cassette (ABC) antibiotic efflux pump | lincosamide antibiotic | 96.7 | 100.42 | Strict |
| ykkC | small multidrug resistance (SMR) antibiotic efflux pump | aminoglycoside antibiotic, tetracycline antibiotic, phenicol antibiotic | 100 | 100 | Perfect |
| tet(45) | major facilitator superfamily (MFS) antibiotic efflux pump | tetracycline antibiotic | 75.8 | 100 | Strict |
| mphK | macrolide phosphotransferase (MPH) | macrolide antibiotic | 97.7 | 100 | Strict |
| blt | major facilitator superfamily (MFS) antibiotic efflux pump | fluoroquinolone antibiotic, acridine dye | 99.8 | 98.5 | Strict |
| Bacillus subtilis pgsA with mutation conferring resistance to daptomycin | daptomycin resistant pgsA | peptide antibiotic | 99.7 | 90.53 | Strict |
| Bacillus subtilis mprF | defensin resistant mprF | peptide antibiotic | 99.7 | 76.87 | Strict |
| vmlR | ABC-F ATP-binding cassette ribosomal protection protein | macrolide antibiotic, lincosamide antibiotic, streptogramin antibiotic, tetracycline antibiotic, oxazolidinone antibiotic, phenicol antibiotic, pleuromutilin antibiotic | 98.5 | 75.5 | Strict |
| aadK | ANT(6) | aminoglycoside antibiotic | 97.8 | 63.03 | Strict |
| bmr | major facilitator superfamily (MFS) antibiotic efflux pump | fluoroquinolone antibiotic, nucleoside antibiotic, acridine dye, phenicol antibiotic | 100 | 47.3 | Strict |
| tmrB | tunicamycin resistance protein | nucleoside antibiotic | 97.6 | 42.13 | Strict |
| aadK | ANT(6) | aminoglycoside antibiotic | 97.2 | 39.44 | Strict |
| vmlR | ABC-F ATP-binding cassette ribosomal protection protein | macrolide antibiotic, lincosamide antibiotic, streptogramin antibiotic, tetracycline antibiotic, oxazolidinone antibiotic, phenicol antibiotic, pleuromutilin antibiotic | 96.4 | 27.24 | Strict |
| tmrB | tunicamycin resistance protein | nucleoside antibiotic | 100 | 26.9 | Strict |
| Bacillus subtilis mprF | defensin resistant mprF | peptide antibiotic | 100 | 16.36 | Strict |
| Antibiotics | Type | MIC (µg/mL) | EFSA MIC (µg/mL) Resistance Threshold |
|---|---|---|---|
| Chloramphenicol | Phenicol | 2 | 8 |
| Clindamycin | Macrolides, lincosamides | 0.5 | 4 |
| Erythromycin | Macrolides, lincosamides | <0.0625 | 4 |
| Gentamicin | Aminoglycosides | 0.5 | 4 |
| Kanamycin | Aminoglycoside | 2 | 8 |
| Streptomycin | Aminoglycoside | 125 | 8 |
| Oxytetracycline | Tetracycline | 8 | 8 |
| Vancomycin | Glycopeptide | 0.25 | 4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brutscher, L.M.; Borgmeier, C.; Garvey, S.M.; Spears, J.L. Preclinical Safety Assessment of Bacillus subtilis BS50 for Probiotic and Food Applications. Microorganisms 2022, 10, 1038. https://doi.org/10.3390/microorganisms10051038
Brutscher LM, Borgmeier C, Garvey SM, Spears JL. Preclinical Safety Assessment of Bacillus subtilis BS50 for Probiotic and Food Applications. Microorganisms. 2022; 10(5):1038. https://doi.org/10.3390/microorganisms10051038
Chicago/Turabian StyleBrutscher, Laura M., Claudia Borgmeier, Sean M. Garvey, and Jessica L. Spears. 2022. "Preclinical Safety Assessment of Bacillus subtilis BS50 for Probiotic and Food Applications" Microorganisms 10, no. 5: 1038. https://doi.org/10.3390/microorganisms10051038
APA StyleBrutscher, L. M., Borgmeier, C., Garvey, S. M., & Spears, J. L. (2022). Preclinical Safety Assessment of Bacillus subtilis BS50 for Probiotic and Food Applications. Microorganisms, 10(5), 1038. https://doi.org/10.3390/microorganisms10051038






