Comparison Study of Four Extraction Methods Combined with PCR and LAMP for Feline Tritrichomonas foetus Detection in Fecal Samples
Abstract
:1. Introduction
2. Results
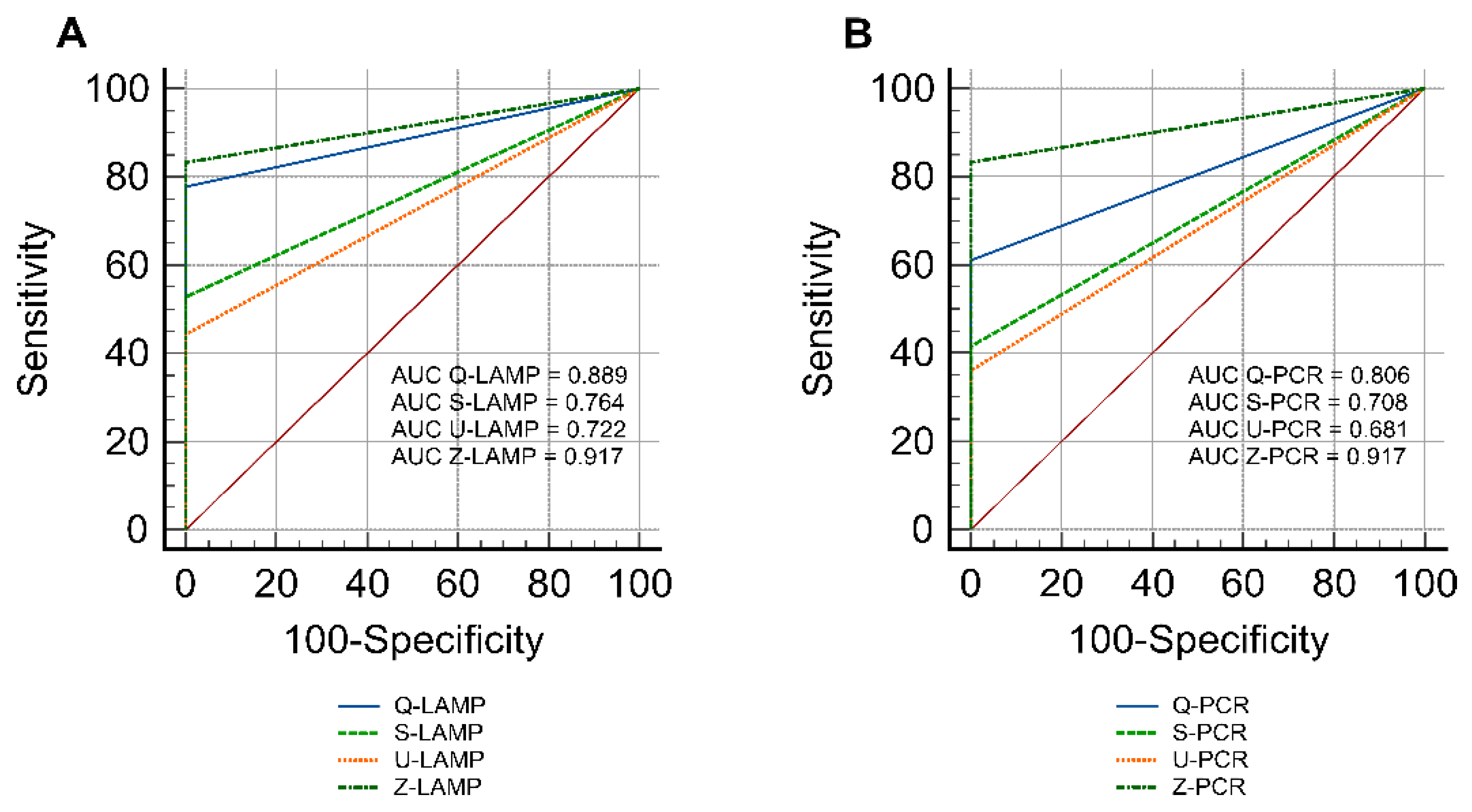
Influence of DNA Extraction Methods on the Performance of Molecular Assays
3. Discussion
4. Material and Methods
4.1. T. foetus Cells
4.2. Feline Fecal Samples
4.3. Spiking of Fecal Samples with T. foetus Trophozoites
4.4. DNA Extraction
4.5. Molecular Analysis
4.5.1. Positive Controls
4.5.2. PCR
4.5.3. LAMP
4.5.4. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Burgener, I.; Frey, C.; Kook, P.; Gottstein, B. Tritrichomonas foetus: A new intestinal parasite in Swiss cats. Companion Anim. Pract. 2010, 20, 113. [Google Scholar]
- Raab, O.; Greenwood, S.; Vanderstichel, R.; Gelens, H. A cross-sectional study of Tritrichomonas foetus infection in feral and shelter cats in Prince Edward Island, Canada. Can. Vet. J. 2016, 57, 265. [Google Scholar]
- Gookin, J.L.; Hanrahan, K.; Levy, M.G. The conundrum of feline trichomonosis: The more we learn the ‘trickier’it gets. J. Feline Med. Surg. 2017, 19, 261–274. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheraghi, B.; Vafaei, R.; Nassiri, S. The first report of clinical case of intestinal trichomoniasis caused by Tritrichomonas foetus in a cat with chronic diarrhea in Iran. Iran. J. Vet. Res. 2021, 22, 248. [Google Scholar]
- Yildiz, K.; Sursal, N. The first report of Tritrichomonas foetus in cats from Turkey. Isr. J. Vet. Med. 2019, 74, 127–133. [Google Scholar]
- Stockdale, H.D.; Givens, M.D.; Dykstra, C.C.; Blagburn, B.L. Tritrichomonas foetus infections in surveyed pet cats. Vet. Parasitol. 2009, 160, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Dąbrowska, J.; Karamon, J.; Kochanowski, M.; Sroka, J.; Skrzypek, K.; Zdybel, J.; Różycki, M.; Jabłoński, A.; Cencek, T. Tritrichomonas foetus: A study of prevalence in animal hosts in Poland. Pathogens 2020, 9, 203. [Google Scholar] [CrossRef] [Green Version]
- Kuehner, K.A.; Marks, S.L.; Kass, P.H.; Sauter-Louis, C.; Grahn, R.A.; Barutzki, D.; Hartmann, K. Tritrichomonas foetus infection in purebred cats in Germany: Prevalence of clinical signs and the role of co-infection with other enteroparasites. J. Feline Med. Surg. 2011, 13, 251–258. [Google Scholar] [CrossRef]
- Gookin, J.L.; Stebbins, M.E.; Hunt, E.; Burlone, K.; Fulton, M.; Hochel, R.; Talaat, M.; Poore, M.; Levy, M.G. Prevalence of and risk factors for feline Tritrichomonas foetus and Giardia infection. J. Clin. Microbiol. 2004, 42, 2707–2710. [Google Scholar] [CrossRef] [Green Version]
- Hosein, A.; Kruth, S.A.; Pearl, D.L.; Richardson, D.; Maggs, J.C.; Peach, H.A.; Peregrine, A.S. Isolation of Tritrichomonas foetus from cats sampled at a cat clinic, cat shows and a humane society in southern Ontario. J. Feline Med. Surg. 2013, 15, 706–711. [Google Scholar] [CrossRef]
- Levy, M.G.; Gookin, J.L.; Poore, M.; Birkenheuer, A.J.; Dykstra, M.J.; Litaker, R.W. Tritrichomonas foetus and not Pentatrichomonas hominis is the etiologic agent of feline trichomonal diarrhea. J. Parasitol. 2003, 89, 99–104. [Google Scholar] [CrossRef]
- Dąbrowska, J.; Karamon, J.; Kochanowski, M.; Sroka, J.; Cencek, T. Unexpected cross-reaction with Honigbergiella-like DNA in a PCR for detection of bovine Tritrichomonas foetus. Pathogens 2021, 10, 441. [Google Scholar] [CrossRef] [PubMed]
- Leski, T.A.; Taitt, C.R.; Swaray, A.G.; Bangura, U.; Reynolds, N.D.; Holtz, A.; Yasuda, C.; Lahai, J.; Lamin, J.M.; Baio, V. Use of real-time multiplex PCR, malaria rapid diagnostic test and microscopy to investigate the prevalence of Plasmodium species among febrile hospital patients in Sierra Leone. Malar. J. 2020, 19, 1–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mori, Y.; Notomi, T. Loop-mediated isothermal amplification (LAMP): A rapid, accurate, and cost-effective diagnostic method for infectious diseases. J. Infect. Chemother. 2009, 15, 62–69. [Google Scholar] [CrossRef] [PubMed]
- Soroka, M.; Wasowicz, B.; Rymaszewska, A. Loop-Mediated Isothermal Amplification (LAMP): The Better Sibling of PCR? Cells 2021, 10, 1931. [Google Scholar] [CrossRef]
- Gookin, J.L.; Birkenheuer, A.J.; Breitschwerdt, E.B.; Levy, M.G. Single-tube nested PCR for detection of Tritrichomonas foetus in feline feces. J. Clin. Microbiol. 2002, 40, 4126–4130. [Google Scholar] [CrossRef] [Green Version]
- Hawash, Y. DNA extraction from protozoan oocysts/cysts in feces for diagnostic PCR. Korean J. Parasitol. 2014, 52, 263. [Google Scholar] [CrossRef]
- Paulos, S.; Mateo, M.; De Lucio, A.; Hernández-De Mingo, M.; Bailo, B.; Saugar, J.M.; Cardona, G.A.; Fuentes, I.; Mateo, M.; Carmena, D. Evaluation of five commercial methods for the extraction and purification of DNA from human faecal samples for downstream molecular detection of the enteric protozoan parasites Cryptosporidium spp., Giardia duodenalis, and Entamoeba spp. J. Microbiol. Methods 2016, 127, 68–73. [Google Scholar] [CrossRef]
- Le Govic, Y.; Guyot, K.; Certad, G.; Deschildre, A.; Novo, R.; Mary, C.; Sendid, B.; Viscogliosi, E.; Favennec, L.; Dei-Cas, E. Assessment of microscopic and molecular tools for the diagnosis and follow-up of cryptosporidiosis in patients at risk. Eur. J. Clin. Microbiol. Infect. Dis. 2016, 35, 137–148. [Google Scholar] [CrossRef]
- Yoshikawa, H.; Dogruman-Ai, F.; Turk, S.; Kustimur, S.; Balaban, N.; Sultan, N. Evaluation of DNA extraction kits for molecular diagnosis of human Blastocystis subtypes from fecal samples. Parasitol. Res. 2011, 109, 1045–1050. [Google Scholar] [CrossRef]
- Subrungruang, I.; Mungthin, M.; Chavalitshewinkoon-Petmitr, P.; Rangsin, R.; Naaglor, T.; Leelayoova, S. Evaluation of DNA extraction and PCR methods for detection of Enterocytozoon bienuesi in stool specimens. J. Clin. Microbiol. 2004, 42, 3490–3494. [Google Scholar] [CrossRef] [Green Version]
- Karakavuk, M.; Aldemir, D.; Mercier, A.; Atalay Şahar, E.; Can, H.; Murat, J.-B.; Döndüren, Ö.; Can, Ş.; Özdemir, H.G.; Değirmenci Döşkaya, A. Prevalence of toxoplasmosis and genetic characterization of Toxoplasma gondii strains isolated in wild birds of prey and their relation with previously isolated strains from Turkey. PLoS ONE 2018, 13, e0196159. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stauffer, S.H.; Birkenheuer, A.J.; Levy, M.G.; Marr, H.; Gookin, J.L. Evaluation of four DNA extraction methods for the detection of Tritrichomonas foetus in feline stool specimens by polymerase chain reaction. J. Vet. Diagn. 2008, 20, 639–641. [Google Scholar] [CrossRef] [Green Version]
- Felleisen, R. Comparative sequence analysis of 5· 8S rRNA genes and internal transcribed spacer (ITS) regions of trichomonadid protozoa. Parasitology 1997, 115, 111–119. [Google Scholar] [CrossRef]
- Dąbrowska, J.; Karamon, J.; Kochanowski, M.; Gottstein, B.; Cencek, T.; Frey, C.F.; Müller, N. Development and comparative evaluation of different LAMP and PCR assays for coprological diagnosis of feline tritrichomonosis. Vet. Parasitol. 2019, 273, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Maksimov, P.; Schares, G.; Press, S.; Fröhlich, A.; Basso, W.; Herzig, M.; Conraths, F.J. Comparison of different commercial DNA extraction kits and PCR protocols for the detection of Echinococcus multilocularis eggs in faecal samples from foxes. Vet. Parasitol. 2017, 237, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Frey, C.F.; Müller, N.; Stäuber, N.; Marreros, N.; Hofmann, L.; Hentrich, B.; Hirsbrunner, G. Simplicimonas-like DNA in vaginal swabs of cows and heifers cross-reacting in the real-time PCR for T. foetus. Vet. Parasitol. 2017, 237, 30–36. [Google Scholar] [CrossRef] [PubMed]
| Suspensions of T. foetus Cells | Z (n = 6) | U (n = 6) | Q (n = 6) | S (n = 6) | ||||
|---|---|---|---|---|---|---|---|---|
| PCR | LAMP | PCR | LAMP | PCR | LAMP | PCR | LAMP | |
| 10,000 | 100% | 100% | 100% | 100% | 100% | 100% | 100% | 100% |
| 1000 | 100% | 100% | 100% | 100% | 100% | 100% | 100% | 100% |
| 100 | 100% | 100% | 16.6% | 33.3% | 100% | 100% | 50% | 100% |
| 10 | 100% | 100% | n.d | n.d | 33.3% | 100% | n.d | 16.6% |
| 1 | 100% | 100% | n.d | n.d | 33.3% | 66.6% | n.d | n.d |
| 0.1 | n.d | n.d | n.d | n.d | n.d | n.d | n.d | n.d |
| Full Name of the Kit | Manufacturers Details | Kit Name Abbreviation | Recommended Sample Starting Amount | Extraction Method | Elution Volume (µL) |
|---|---|---|---|---|---|
| QIAamp® DNA Stool Mini Kit | Qiagen Inc., Valencia, CA, USA | Q | 180–220 mg | Manual | 200 |
| UltraClean Fecal DNA Kit (50 preps) | MO BIO, San Diego, CA, USA | U | 250 mg | Manual | 50 |
| ZR Fecal DNA MiniPrep (50 preps) | Zymo Research, Irvine, CA, USA | Z | 150 mg | Manual | 100 |
| Sherlock AX/100 isolations | A&A Biotechnology, Gdynia, Poland | S | 10–20 mg | Manual | 350 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dąbrowska, J.; Karamon, J.; Kochanowski, M.; Sroka, J.; Zdybel, J.; Cencek, T. Comparison Study of Four Extraction Methods Combined with PCR and LAMP for Feline Tritrichomonas foetus Detection in Fecal Samples. Pathogens 2022, 11, 604. https://doi.org/10.3390/pathogens11050604
Dąbrowska J, Karamon J, Kochanowski M, Sroka J, Zdybel J, Cencek T. Comparison Study of Four Extraction Methods Combined with PCR and LAMP for Feline Tritrichomonas foetus Detection in Fecal Samples. Pathogens. 2022; 11(5):604. https://doi.org/10.3390/pathogens11050604
Chicago/Turabian StyleDąbrowska, Joanna, Jacek Karamon, Maciej Kochanowski, Jacek Sroka, Jolanta Zdybel, and Tomasz Cencek. 2022. "Comparison Study of Four Extraction Methods Combined with PCR and LAMP for Feline Tritrichomonas foetus Detection in Fecal Samples" Pathogens 11, no. 5: 604. https://doi.org/10.3390/pathogens11050604
APA StyleDąbrowska, J., Karamon, J., Kochanowski, M., Sroka, J., Zdybel, J., & Cencek, T. (2022). Comparison Study of Four Extraction Methods Combined with PCR and LAMP for Feline Tritrichomonas foetus Detection in Fecal Samples. Pathogens, 11(5), 604. https://doi.org/10.3390/pathogens11050604







