Diversity and Phylogenetic Analyses of Bacterial Symbionts in Three Whitefly Species from Southeast Europe
Abstract
1. Introduction
2. Materials and Methods
2.1. Whitefly Populations
2.2. Screening and Sequencing of Secondary Bacterial Symbionts and Molecular Identification of B. tabaci Species
2.3. Phylogenetic Analyses
2.4. Nucleotide Sequence Accession Numbers
3. Results
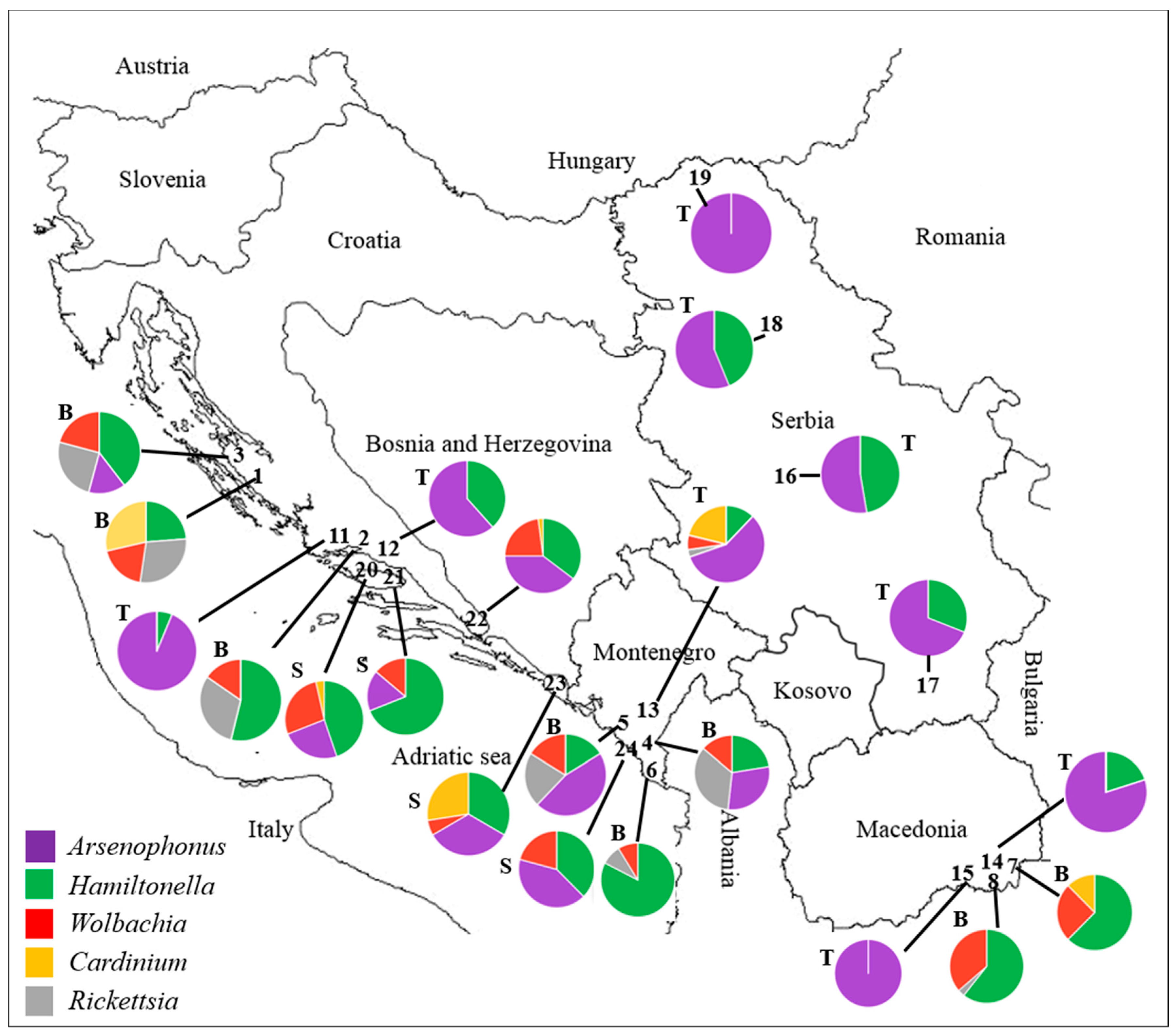
3.1. Whitefly Infection with Secondary Bacterial Symbionts
3.1.1. Bemisia tabaci Infection with Secondary Bacterial Symbionts
3.1.2. Trialeurodes vaporariorum Infection with Secondary Bacterial Symbionts
3.1.3. Siphoninus phillyreae Infection with Secondary Bacterial Symbionts
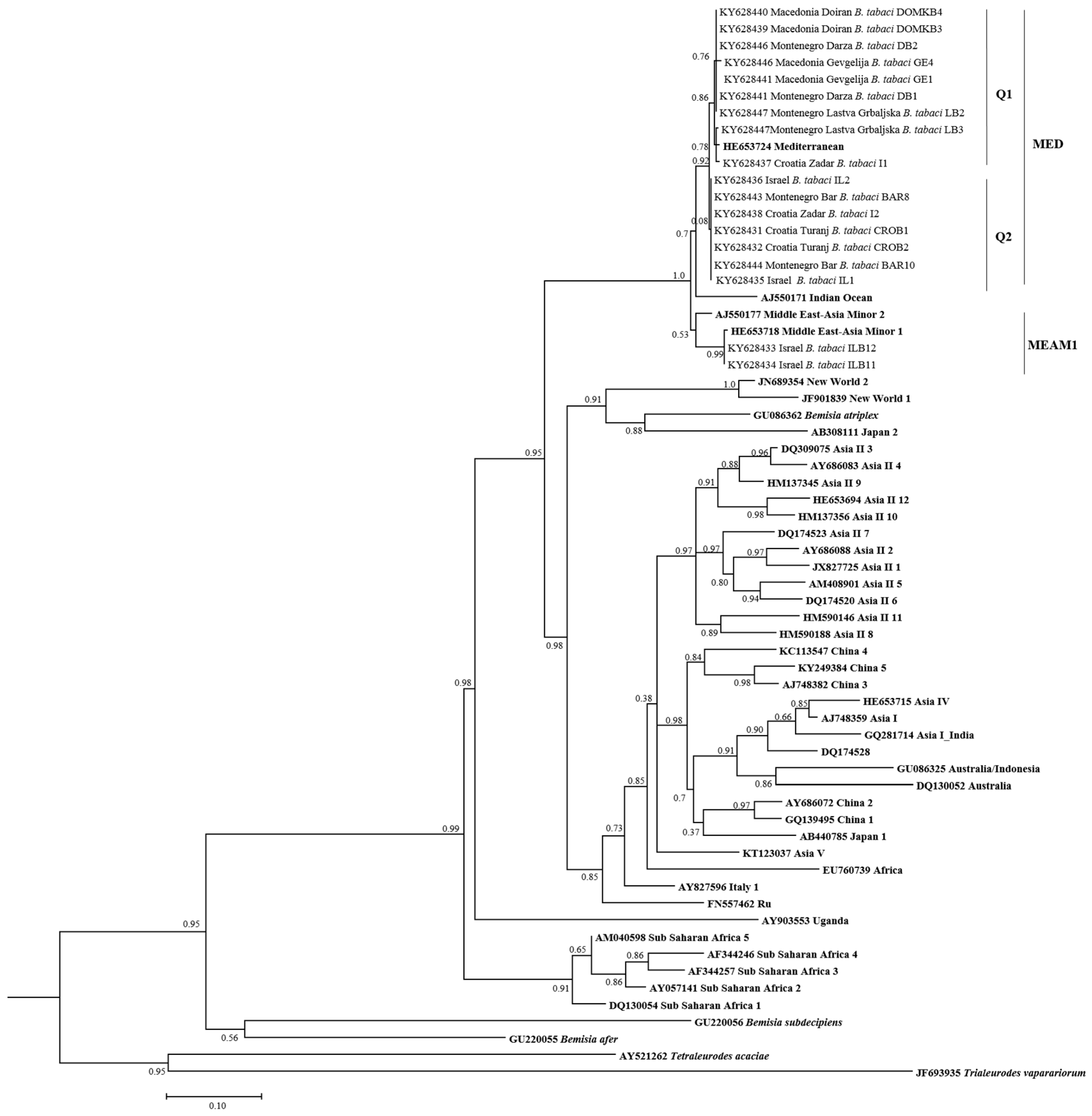
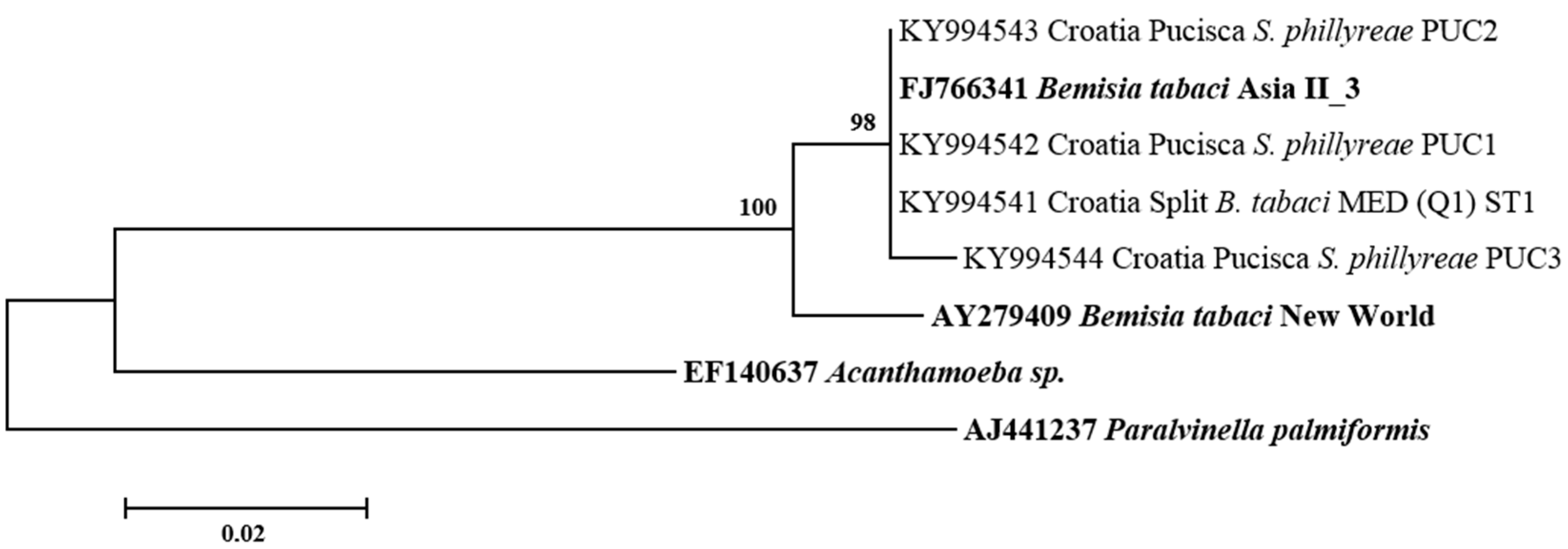
3.2. Phylogenetic Relationships of B. tabaci Sequences
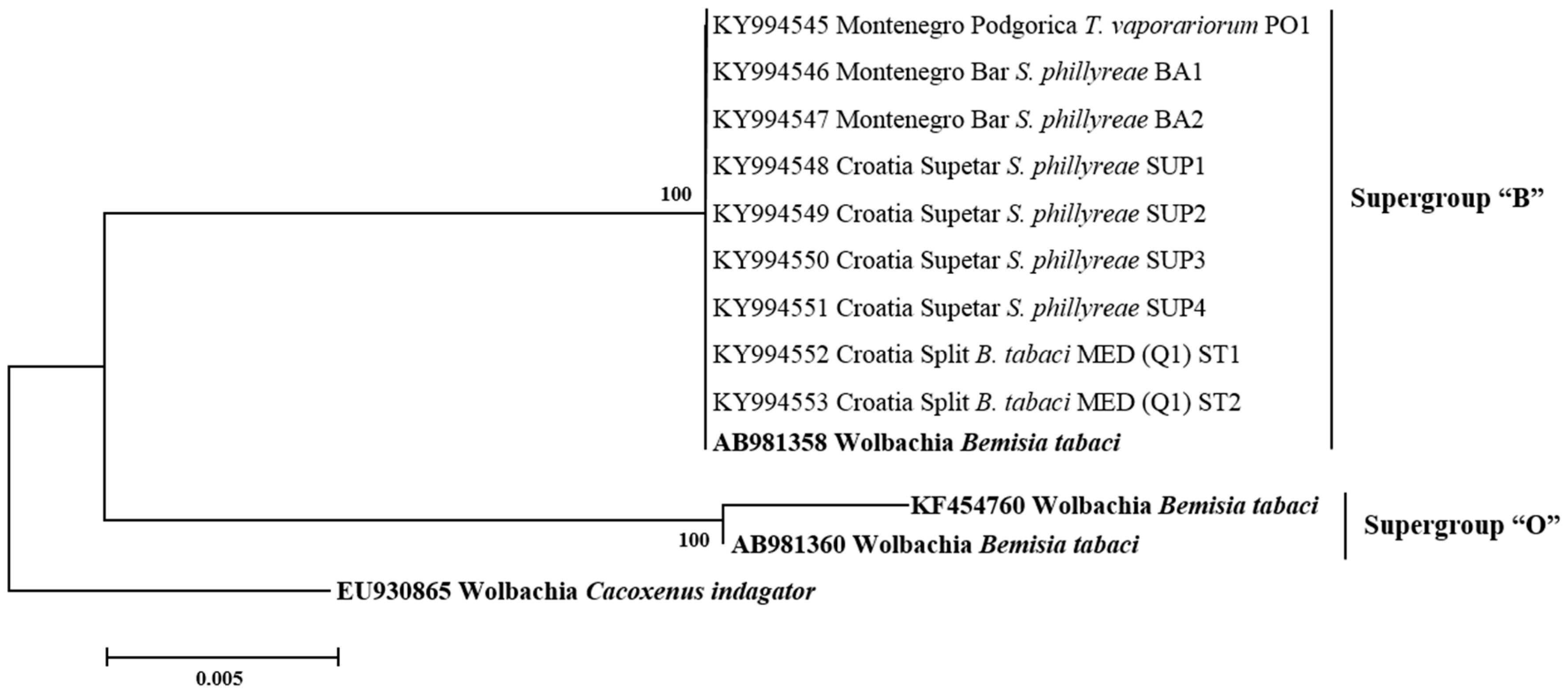
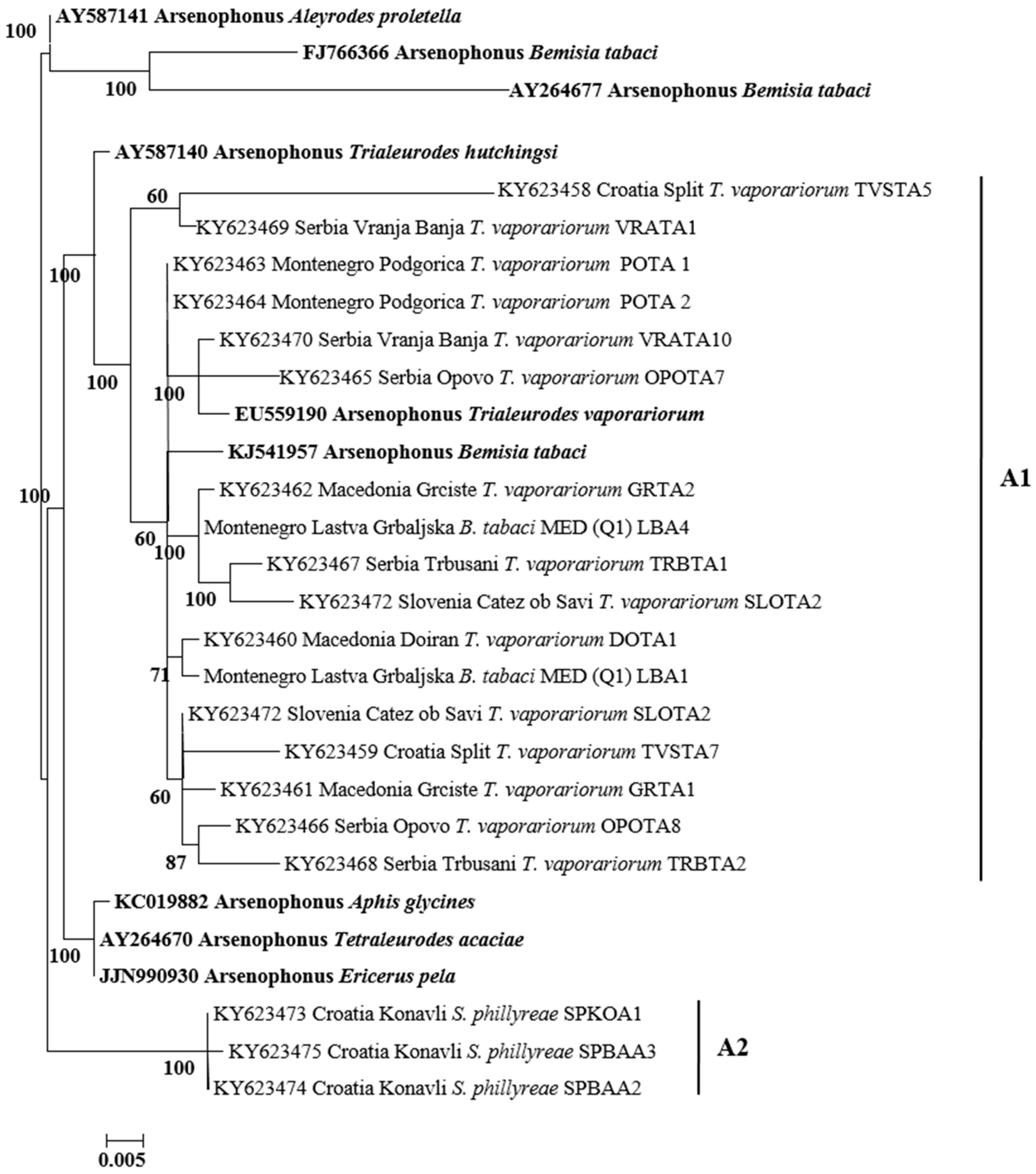
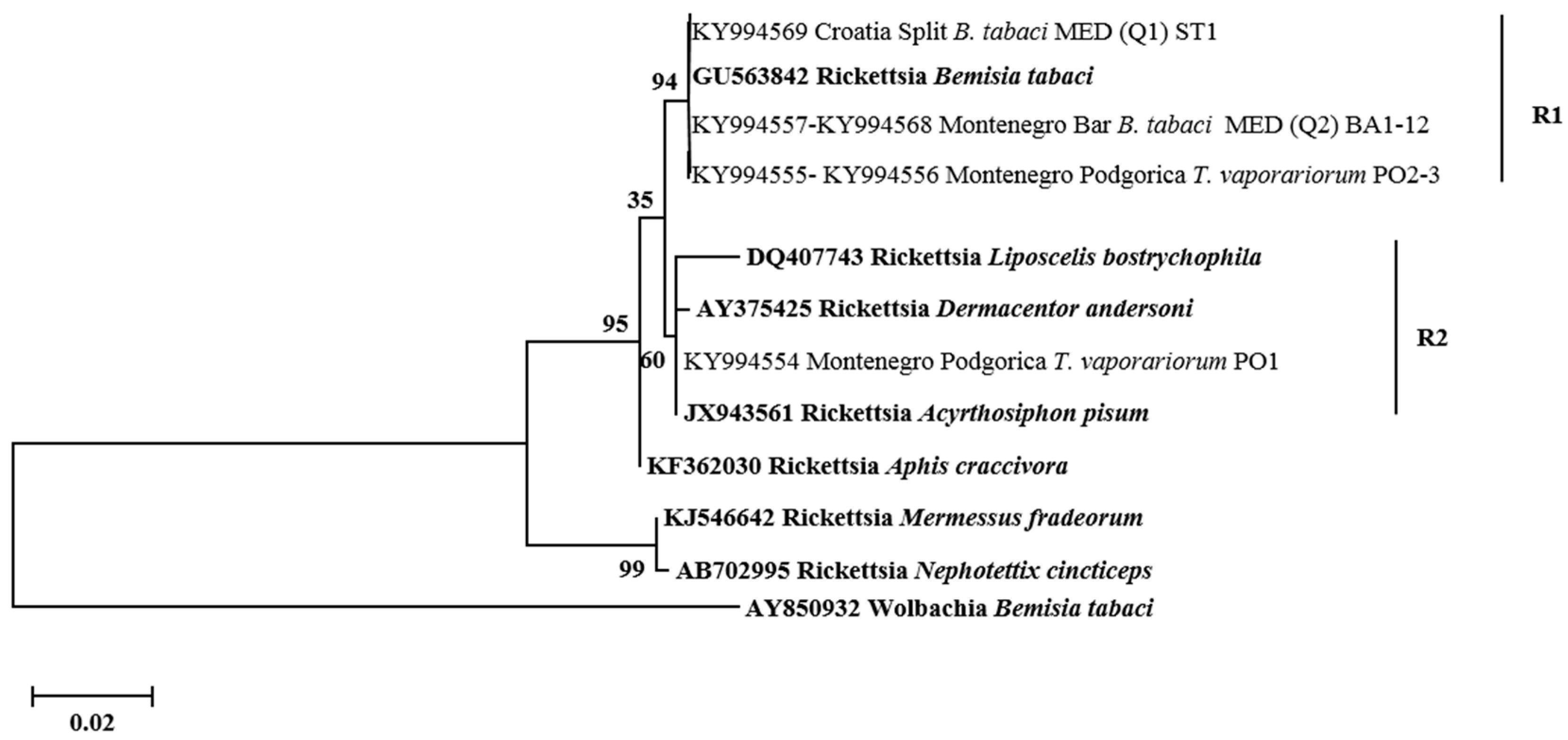
3.3. Phylogenetic Analysis of Secondary Symbionts in Whiteflies
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Moran, N.A.; McCutcheon, J.P.; Nakabachi, A. Genomics and evolution of heritable bacterial symbionts. Annu. Rev. Genet. 2008, 42, 165–190. [Google Scholar] [CrossRef] [PubMed]
- Dale, C.; Moran, N.A. Molecular interactions between bacterial symbionts and their hosts. Cell 2006, 126, 453–465. [Google Scholar] [CrossRef] [PubMed]
- Brownlie, J.C.; Johnson, K.N. Symbiont-mediated protection in insect hosts. Trends Microbiol. 2009, 17, 348–354. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Contreras, M.; Vlisidou, I. The diversity of insect-bacteria interactions and its applications for disease control. Biotechnol. Genet. Eng. Rev. 2008, 25, 203–244. [Google Scholar] [CrossRef]
- Kikuchi, Y.; Hosokawa, T.; Nikoh, N.; Meng, X.Y.; Kamagata, Y.; Fukatsu, T. Host-symbiont co-speciation and reductive genome evolution in gut symbiotic bacteria of acanthosomatid stinkbugs. BMC Biol. 2009, 7, 2. [Google Scholar] [CrossRef] [PubMed]
- Feldhaar, H. Bacterial symbionts as mediators of ecologically important traits of insect hosts. Ecol. Entomol. 2011, 36, 533–543. [Google Scholar] [CrossRef]
- Dunbar, H.E.; Wilson, A.C.C.; Ferguson, N.R.; Moran, N.A. Aphid thermal tolerance is governed by a point mutation in bacterial symbionts. PLoS Biol. 2007, 5. [Google Scholar] [CrossRef] [PubMed]
- Tsuchida, T.; Koga, R.; Matsumoto, S.; Fukatsu, T. Interspecific symbiont transfection confers a novel ecological trait to the recipient insect. Biol. Lett. 2011, 7, 245–248. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, J.; Vavre, F. Bacterial symbionts in insects or the story of communities affecting communities. Phil. Trans. R. Soc. B Biol. Sci. 2011, 366, 1389–1400. [Google Scholar] [CrossRef] [PubMed]
- De Barro, P.J.; Liu, S.S.; Boykin, L.M.; Dinsdale, A.B. Bemisia tabaci: A statement of species status. Annu. Rev. Entomol. 2011, 56, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Ash Whitefly, Siphoninus Phillyreae (Haliday) (Insecta: Aleyrodidae: Aleyrodinae). Available online: http://www.doacs. state.fl.us/pi/enpp/ento/entcirc/ent337.pdf (accessed on 18 May 2012).
- Brown, J.K. The bemisia tabaci complex: Genetic and phenotypic variation and relevance to tylcv-vector interactions. In Tomato Yellow Leaf Curl Virus Disease (Managment, Molecular Biology, Breeding for Resistance); Czosnek, H., Ed.; Springer: Houten, The Netherlands, 2007; pp. 25–57. [Google Scholar]
- CABI. Improving Lives by Solving Problems in Agriculture and the Environment. Available online: http://www.cabi.org (accessed on 1 September 2017).
- Jones, D.R. Plant viruses transmitted by whiteflies. Eur. J. Plant Pathol. 2003, 109, 195–219. [Google Scholar] [CrossRef]
- Hu, J.; De Barro, P.; Zhao, H.; Wang, J.; Nardi, F.; Liu, S.-S. An extensive field survey combined with a phylogenetic analysis reveals rapid and widespread invasion of two alien whiteflies in china. PLoS ONE 2011, 6. [Google Scholar] [CrossRef] [PubMed]
- Gao, R.R.; Zhang, W.P.; Wu, H.T.; Zhang, R.M.; Zhou, H.X.; Pan, H.P.; Zhang, Y.J.; Brown, J.K.; Chu, D. Population structure of the greenhouse whitefly, trialeurodes vaporariorum (westwood), an invasive species from the americas, 60 years after invading china. Int. J. Mol. Sci. 2014, 15, 13514–13528. [Google Scholar] [CrossRef] [PubMed]
- Sparks, T.C.; Nauen, R. Irac: Mode of action classification and insecticide resistance management. Pestic. Biochem. Physiol. 2015, 121, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Kapantaidaki, D.E.; Sadikoglou, E.; Tsakireli, D.; Kampanis, V.; Stavrakaki, M.; Schorn, C.; Ilias, A.; Riga, M.; Tsiamis, G.; Nauen, R.; et al. Insecticide resistance in trialeurodes vaporariorum populations and novel diagnostics for kdr mutations. Pest. Manag. Sci. 2017. [Google Scholar] [CrossRef] [PubMed]
- Dinsdale, A.; Cook, L.; Riginos, C.; Buckley, Y.; Barro, P.D. Refined global analysis of bemisia tabaci (hemiptera: Sternorrhyncha: Aleyrodoidea: Aleyrodidae) mitochondrial cytochrome oxidase 1 to identify species level genetic boundaries. Ann. Entomol. Soc. Am. 2010, 103, 196–208. [Google Scholar] [CrossRef]
- Firdaus, S.; Vosman, B.; Hidayati, N.; Jaya Supena, E.D.; Visser, R.G.F.; van Heusden, A.W. The bemisia tabaci species complex: Additions from different parts of the world. Insect Sci. 2013, 20, 723–733. [Google Scholar] [CrossRef] [PubMed]
- Chowda-Reddy, R.; Kirankumar, M.; Seal, S.E.; Muniyappa, V.; Valand, G.B.; Govindappa, M.; Colvin, J. Bemisia tabaci phylogenetic groups in india and the relative transmission efficacy of tomato leaf curl bangalore virus by an indigenous and an exotic population. J. Integr. Agric. 2012, 11, 235–248. [Google Scholar] [CrossRef]
- Hu, J.; Jiang, Z.L.; Nardi, F.; Liu, Y.Y.; Luo, X.R.; Li, H.X.; Zhang, Z.K. Members of bemisia tabaci (hemiptera: Aleyrodidae) cryptic species and the status of two invasive alien species in the Yunnan province (China). J. Insect Sci. Online 2014, 14. [Google Scholar] [CrossRef] [PubMed]
- Alemandri, V.; De Barro, P.; Bejerman, N.; Arguello Caro, E.B.; Dumon, A.D.; Mattio, M.F.; Rodriguez, S.M.; Truoli, G. Species within the bemisia tabaci (hemiptera: Aleyrodidae) complex in soybean and bean crops in argentina. J. Econ. Entomol. 2012, 105, 48–53. [Google Scholar] [CrossRef] [PubMed]
- Ueda, S.; Kitamura, T.; Kijima, K.; Honda, K.I.; Kanmiya, K. Distribution and molecular characterization of distinct asian populations of bemisia tabaci (hemiptera: Aleyrodidae) in Japan. J. Appl. Entomol. 2009, 133, 355–366. [Google Scholar] [CrossRef]
- Parrella, G.; Scassillo, L.; Giorgini, M. Evidence for a new genetic variant in the bemisia tabaci species complex and the prevalence of the biotype q in Southern Italy. J. Pest Sci. 2012, 85, 227–238. [Google Scholar] [CrossRef]
- Esterhuizen, L.L.; Mabasa, K.G.; van Heerden, S.W.; Czosnek, H.; Brown, J.K.; van Heerden, H.; Rey, M.E.C. Genetic identification of members of the bemisia tabaci cryptic species complex from south africa reveals native and introduced haplotypes. J. Appl. Entomol. 2013, 137, 122–135. [Google Scholar] [CrossRef]
- Hu, J.; Zhang, X.; Jiang, Z.; Zhang, F.; Liu, Y.; Li, Z.; Zhang, Z. New putative cryptic species detection and genetic network analysis of bemisia tabaci (hempitera: Aleyrodidae) in China based on mitochondrial coi sequences. Mitochondrial DNA Part A DNA Mapp. Seq. Anal. 2017, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Tsagkarakou, A.; Mouton, L.; Kristoffersen, J.B.; Dokianakis, E.; Grispou, M.; Bourtzis, K. Population genetic structure and secondary endosymbionts of q bemisia tabaci (hemiptera: Aleyrodidae) from greece. Bull. Entomol. Res. 2012, 102, 353–365. [Google Scholar] [CrossRef] [PubMed]
- Gauthier, N.; Clouet, C.; Perrakis, A.; Kapantaidaki, D.; Peterschmitt, M.; Tsagkarakou, A. Genetic structure of bemisia tabaci med populations from home-range countries, inferred by nuclear and cytoplasmic markers: Impact on the distribution of the insecticide resistance genes. Pest Manag. Sci. 2014, 70, 1477–1491. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Hasegawa, D.K.; Kaur, N.; Kliot, A.; Pinheiro, P.V.; Luan, J.; Stensmyr, M.C.; Zheng, Y.; Liu, W.; Sun, H.; et al. The draft genome of whitefly bemisia tabaci meam1, a global crop pest, provides novel insights into virus transmission, host adaptation, and insecticide resistance. BMC Biol. 2016, 14, 110. [Google Scholar] [CrossRef] [PubMed]
- Boykin, L.M.; Shatters, R.G., Jr.; Rosell, R.C.; McKenzie, C.L.; Bagnall, R.A.; De Barro, P.; Frohlich, D.R. Global relationships of bemisia tabaci (hemiptera: Aleyrodidae) revealed using bayesian analysis of mitochondrial coi DNA sequences. Mol. Phylogenet. Evol. 2007, 44, 1306–1319. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.; Wan, F.-H.; Tao, Y.-L.; Liu, G.-X.; Fan, Z.-X.; Bi, Y.-P. Genetic differentiation of bemisia tabaci (gennadius) (hemiptera: Aleyrodidae) biotype q based on mitochondrial DNA markers. Insect Sci. 2008, 15, 115–123. [Google Scholar] [CrossRef]
- Gueguen, G.; Vavre, F.; Gnankine, O.; Peterschmitt, M.; Charif, D.; Chiel, E.; Gottlieb, Y.; Ghanim, M.; Zchori-Fein, E.; Fleury, F. Endosymbiont metacommunities, mtdna diversity and the evolution of the bemisia tabaci (hemiptera: Aleyrodidae) species complex. Mol. Ecol. 2010, 19, 4365–4376. [Google Scholar] [CrossRef] [PubMed]
- Sloan, D.B.; Moran, N.A. Endosymbiotic bacteria as a source of carotenoids in whiteflies. Biol. Lett. 2012, 8, 986–989. [Google Scholar] [CrossRef] [PubMed]
- Zchori-Fein, E.; Brown, J.K. Diversity of prokaryotes associated with bemisia tabaci (gennadius) (hemiptera: Aleyrodidae). Ann. Entomol. Soc. Am. 2002, 95, 711–718. [Google Scholar] [CrossRef]
- Gottlieb, Y.; Ghanim, M.; Chiel, E.; Gerling, D.; Portnoy, V.; Steinberg, S.; Tzuri, G.; Horowitz, A.R.; Belausov, E.; Mozes-Daube, N.; et al. Identification and localization of a rickettsia sp. In bemisia tabaci (homoptera: Aleyrodidae). Appl. Environ. Microbiol. 2006, 72, 3646–3652. [Google Scholar] [CrossRef] [PubMed]
- Chiel, E.; Gottlieb, Y.; Zchori-Fein, E.; Mozes Daube, N.; Katzir, N.; Inbar, M. Biotype-dependent secondary symbiont communities in sympatric populations of bemisia tabaci. Bull. Entomol. Res. 2007, 97, 407. [Google Scholar] [CrossRef] [PubMed]
- Gottlieb, Y.; Ghanim, M.; Gueguen, G.; Kontsedalov, S.; Vavre, F.; Fleury, F.; Zchori-Fein, E. Inherited intracellular ecosystem: Symbiotic bacteria share bacteriocytes in whiteflies. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2008, 22, 2591–2599. [Google Scholar] [CrossRef] [PubMed]
- Skaljac, M.; Zanic, K.; Ban, S.G.; Kontsedalov, S.; Ghanim, M. Co-infection and localization of secondary symbionts in two whitefly species. BMC Microbiol. 2010, 10, 142. [Google Scholar] [CrossRef] [PubMed]
- Marubayashi, J.M.; Kliot, A.; Yuki, V.A.; Rezende, J.A.M.; Krause-Sakate, R.; Pavan, M.A.; Ghanim, M. Diversity and localization of bacterial endosymbionts from whitefly species collected in brazil. PLoS ONE 2014, 9, e108363. [Google Scholar] [CrossRef] [PubMed]
- Skaljac, M.; Zanic, K.; Hrncic, S.; Radonjic, S.; Perovic, T.; Ghanim, M. Diversity and localization of bacterial symbionts in three whitefly species (hemiptera: Aleyrodidae) from the east coast of the adriatic sea. Bull. Entomol. Res. 2013, 103, 48–59. [Google Scholar] [CrossRef] [PubMed]
- Gherna, R.L.; Werren, J.H.; Weisburg, W.; Cote, R.; Woese, C.R.; Mandelco, L.; Brenner, D.J. Arsenophonus nasoniae gen. Nov., sp. Nov., the causative agent of the son-killer trait in the parasitic wasp nasonia vitripennis. Int. J. Syst. Evol. Microbiol. 1991, 41, 563–565. [Google Scholar] [CrossRef]
- Zchori-Fein, E.; Perlman, S.J. Distribution of the bacterial symbiont cardinium in arthropods. Mol. Ecol. 2004, 13, 2009–2016. [Google Scholar] [CrossRef] [PubMed]
- Werren, J.H.; Baldo, L.; Clark, M.E. Wolbachia: Master manipulators of invertebrate biology. Nat. Rev. Microbiol. 2008, 6, 741–751. [Google Scholar] [CrossRef] [PubMed]
- Brumin, M.; Kontsedalov, S.; Ghanim, M. Rickettsia influences thermotolerance in the whitefly bemisia tabaci b biotype. Insect Sci. 2011, 18, 57–66. [Google Scholar] [CrossRef]
- Himler, A.G.; Adachi-Hagimori, T.; Bergen, J.E.; Kozuch, A.; Kelly, S.E.; Tabashnik, B.E. Rapid spread of a bacterial symbiont in an invasive whitefly is driven by fitness benefits and female bias. Science 2011, 332, 254–256. [Google Scholar] [CrossRef] [PubMed]
- Kontsedalov, S.; Zchori-Fein, E.; Chiel, E.; Gottlieb, Y.; Inbar, M.; Ghanim, M. The presence of rickettsia is associated with increased susceptibility of bemisia tabaci (homoptera: Aleyrodidae) to insecticides. Pest. Manag. Sci. 2008, 64, 789–792. [Google Scholar] [CrossRef] [PubMed]
- Mahadav, A.; Gerling, D.; Gottlieb, Y.; Czosnek, H.; Ghanim, M. Parasitization by the wasp eretmocerus mundus induces transcription of genes related to immune response and symbiotic bacteria proliferation in the whitefly bemisia tabaci. BMC Genom. 2008, 9, 342. [Google Scholar] [CrossRef] [PubMed]
- Allen, J.M.; Reed, D.L.; Perotti, M.A.; Braig, H.R. Evolutionary relationships of “candidatus riesia spp.,” endosymbiotic enterobacteriaceae living within hematophagous primate lice. Appl. Environ. Microbiol. 2007, 73, 1659–1664. [Google Scholar] [CrossRef] [PubMed]
- Perotti, M.A.; Allen, J.M.; Reed, D.L.; Braig, H.R. Host-symbiont interactions of the primary endosymbiont of human head and body lice. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2007, 21, 1058–1066. [Google Scholar] [CrossRef] [PubMed]
- Rana, V.S.; Singh, S.T.; Priya, N.G.; Kumar, J.; Rajagopal, R. Arsenophonus groel interacts with clcuv and is localized in midgut and salivary gland of whitefly b. Tabaci. PLoS ONE 2012, 7, e42168. [Google Scholar] [CrossRef] [PubMed]
- Gottlieb, Y.; Zchori-Fein, E.; Mozes-Daube, N.; Kontsedalov, S.; Skaljac, M.; Brumin, M.; Sobol, I.; Czosnek, H.; Vavre, F.; Fleury, F.; et al. The transmission efficiency of tomato yellow leaf curl virus by the whitefly bemisia tabaci is correlated with the presence of a specific symbiotic bacterium species. J. Virol. 2010, 84, 9310–9317. [Google Scholar] [CrossRef] [PubMed]
- Oliver, K.M.; Russell, J.A.; Moran, N.A.; Hunter, M.S. Facultative bacterial symbionts in aphids confer resistance to parasitic wasps. Proc. Natl. Acad. Sci. USA 2003, 100, 1803–1807. [Google Scholar] [CrossRef] [PubMed]
- Russell, J.A.; Moran, N.A. Horizontal transfer of bacterial symbionts: Heritability and fitness effects in a novel aphid host. Appl. Environ. Microbiol. 2005, 71, 7987–7994. [Google Scholar] [CrossRef] [PubMed]
- Rao, Q.; Wang, S.; Su, Y.-L.; Bing, X.-L.; Liu, S.-S.; Wang, X.-W. Draft genome sequence of “candidatus hamiltonella defensa,” an endosymbiont of the whitefly bemisia tabaci. J. Bacteriol. 2012, 194. [Google Scholar] [CrossRef] [PubMed]
- Su, Q.; Xie, W.; Wang, S.; Wu, Q.; Liu, B.; Fang, Y.; Xu, B.; Zhang, Y. The endosymbiont hamiltonella increases the growth rate of its host bemisia tabaci during periods of nutritional stress. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Thao, M.L.; Baumann, L.; Hess, J.M.; Falk, B.W.; Ng, J.C.; Gullan, P.J.; Baumann, P. Phylogenetic evidence for two new insect-associated chlamydia of the family simkaniaceae. Curr. Microbiol. 2003, 47, 46–50. [Google Scholar] [CrossRef] [PubMed]
- Everett, K.D.E.; Thao, M.; Horn, M.; Dyszynski, G.E.; Baumann, P. Novel chlamydiae in whiteflies and scale insects: Endosymbionts “candidatus fritschea bemisiae” strain falk and “candidatus fritschea eriococci” strain elm. Int. J. Syst. Evol. Microbiol. 2005, 55, 1581–1587. [Google Scholar] [CrossRef] [PubMed]
- De Barro, P.J.; Scott, K.D.; Graham, G.C.; Lange, C.L.; Schutze, M.K. Isolation and characterization of microsatellite loci in bemisia tabaci. Mol. Ecol. Notes 2003, 3, 40–43. [Google Scholar] [CrossRef]
- Frohlich, D.R.; Torres-Jerez, I.I.; Bedford, I.D.; Markham, P.G.; Brown, J.K. A phylogeographical analysis of the bemisia tabaci species complex based on mitochondrial DNA markers. Mol. Ecol. 1999, 8, 1683–1691. [Google Scholar] [CrossRef] [PubMed]
- Thao, M.L.; Baumann, P. Evidence for multiple acquisition of arsenophonus by whitefly species (sternorrhyncha: Aleyrodidae). Curr. Microbiol. 2004, 48, 140–144. [Google Scholar] [CrossRef] [PubMed]
- Weeks, A.; Breeuwer, J. A new bacterium from the cytophaga-flavobacterium—bacteroides phylum that causes sex-ratio distortion. In Insect Symbiosis; CRC Press: Boca Raton, FL, USA, 2003; pp. 165–176. [Google Scholar]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. Mafft multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the cipres science gateway for inference of large phylogenetic trees. In Gateway Computing Environments Workshop (GCE), 2010; IEEE: New Orleans, LA, USA, 2010; pp. 1–8. [Google Scholar]
- Ronquist, F.; Huelsenbeck, J.P. Mrbayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. Mega6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Glowska, E.; Dragun-Damian, A.; Dabert, M.; Gerth, M. New wolbachia supergroups detected in quill mites (acari: Syringophilidae). Infect. Genet. Evol. 2015, 30, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Ghanim, M.; Kontsedalov, S. Susceptibility to insecticides in the q biotype of bemisia tabaci is correlated with bacterial symbiont densities. Pest. Manag. Sci. 2009, 65, 939–942. [Google Scholar] [CrossRef] [PubMed]
- Degnan, P.H.; Yu, Y.; Sisneros, N.; Wing, R.A.; Moran, N.A. Hamiltonella defensa, genome evolution of protective bacterial endosymbiont from pathogenic ancestors. Proc. Natl. Acad. Sci. USA 2009, 106, 9063–9068. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.T.; Priya, N.G.; Kumar, J.; Rana, V.S.; Ellango, R.; Joshi, A. Diversity and phylogenetic analysis of endosymbiotic bacteria from field caught bemisia tabaci from different locations of north india based on 16s rdna library screening. Infect. Genet. Evol. 2012, 12, 411–419. [Google Scholar] [CrossRef] [PubMed]
- Xie, W.; Meng, Q.S.; Wu, Q.J.; Wang, S.L.; Yang, X.; Yang, N.N.; Li, R.M.; Jiao, X.G.; Pan, H.P.; Liu, B.M.; et al. Pyrosequencing the bemisia tabaci transcriptome reveals a highly diverse bacterial community and a robust system for insecticide resistance. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Caspi-Fluger, A.; Inbar, M.; Mozes-Daube, N.; Katzir, N.; Portnoy, V.; Belausov, E.; Hunter, M.S.; Zchori-Fein, E. Horizontal transmission of the insect symbiont rickettsia is plant-mediated. Proc. R. Soc. B Biol. Sci. 2012, 279, 1791–1796. [Google Scholar] [CrossRef] [PubMed]
- Gonella, E.; Pajoro, M.; Marzorati, M.; Crotti, E.; Mandrioli, M.; Pontini, M.; Bulgari, D.; Negri, I.; Sacchi, L.; Chouaia, B.; et al. Plant-mediated interspecific horizontal transmission of an intracellular symbiont in insects. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Chiel, E.; Zchori-Fein, E.; Inbar, M.; Gottlieb, Y.; Adachi-Hagimori, T.; Kelly, S.E.; Asplen, M.K.; Hunter, M.S. Almost there: Transmission routes of bacterial symbionts between trophic levels. PLoS ONE 2009, 4. [Google Scholar] [CrossRef] [PubMed]
- Terraz, G.; Gueguen, G.; Arno, J.; Fleury, F.; Mouton, L. Nuclear and cytoplasmic differentiation among mediterranean populations of bemisia tabaci: Testing the biological relevance of cytotypes. Pest. Manag. Sci. 2014, 70, 1503–1513. [Google Scholar] [CrossRef] [PubMed]
- Parrella, G.; Nappo, A.G.; Manco, E.; Greco, B.; Giorgini, M. Invasion of the q2 mitochondrial variant of mediterranean bemisia tabaci in southern italy: Possible role of bacterial endosymbionts. Pest. Manag. Sci. 2014, 70, 1514–1523. [Google Scholar] [CrossRef] [PubMed]
- Bing, X.; Ruan, Y.; Rao, Q.; Wang, X.; Liu, S. Diversity of secondary endosymbionts among different putative species of the whitefly bemisia tabaci. Insect Sci. 2013, 20, 194–206. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.; Gao, C.S.; De Barro, P.; Zhang, Y.J.; Wan, F.H.; Khan, I.A. Further insights into the strange role of bacterial endosymbionts in whitefly, bemisia tabaci: Comparison of secondary symbionts from biotypes b and q in china. Bull. Entomol. Res. 2011, 101, 477–486. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Bouvaine, S.; Maruthi, M. Prevalence and genetic diversity of endosymbiotic bacteria infecting cassava whiteflies in africa. BMC Microbiol. 2015, 15. [Google Scholar] [CrossRef] [PubMed]
- Kapantaidaki, D.E.; Ovcarenko, I.; Fytrou, N.; Knott, K.E.; Bourtzis, K.; Tsagkarakou, A. Low levels of mitochondrial DNA and symbiont diversity in the worldwide agricultural pest, the greenhouse whitefly trialeurodes vaporariorum (hemiptera: Aleyrodidae). J. Hered. 2014, 106, 80–92. [Google Scholar] [CrossRef] [PubMed]
- Correa, C.C.; Ballard, J. Wolbachia associations with insects: Winning or losing against a master manipulator. Front. Ecol. Evol. 2016, 3. [Google Scholar] [CrossRef]
- Vautrin, E.; Vavre, F. Interactions between vertically transmitted symbionts: Coperation or conflict? Trends Microbiol. 2009, 17, 95–99. [Google Scholar] [CrossRef] [PubMed]
- Sintupachee, S.; Milne, J.R.; Poonchaisri, S.; Baimai, V.; Kittayapong, P. Closely related wolbachia strains within the pumpkin arthropod community and the potential for horizontal transmission via the plant. Microb. Ecol. 2006, 51, 294–301. [Google Scholar] [CrossRef] [PubMed]
- Mouton, L.; Thierry, M.; Henri, H.; Baudin, R.; Gnankine, O.; Reynaud, B.; Zchori-Fein, E.; Becker, N.; Fleury, F.; Delatte, H. Evidence of diversity and recombination in arsenophonus symbionts of the bemisia tabaci species complex. BMC Microbiol. 2012, 12 (Suppl. 1). [Google Scholar] [CrossRef] [PubMed]
- Vavre, F.; Fleury, F.; Lepetit, D.; Fouillet, P.; Bouletreau, M. Phylogenetic evidence for horizontal transmission of wolbachia in host-parasitoid associations. Mol. Biol. Evol. 1999, 16, 1711–1723. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.Z.; De Barro, P.J.; Ren, S.X.; Greeff, J.M.; Qiu, B.L. Evidence for horizontal transmission of secondary endosymbionts in the bemisia tabaci cryptic species complex. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Tajebe, L.S.; Guastella, D.; Cavalieri, V.; Kelly, S.E.; Hunter, M.S.; Lund, O.S.; Legg, J.P.; Rapisarda, C. Diversity of symbiotic bacteria associated with bemisia tabaci (homoptera: Aleyrodidae) in cassava mosaic disease pandemic areas of tanzania. Ann. Appl. Biol. 2015, 166, 297–310. [Google Scholar] [CrossRef]
- Ghosh, S.; Bouvaine, S.; Richardson, S.C.W.; Ghanim, M.; Maruthi, M.N. Fitness costs associated with infections of secondary endosymbionts in the cassava whitefly species bemisia tabaci. J. Pest Sci. 2017. [Google Scholar] [CrossRef]


| Primer Name | Sequence (5′→3′) | Annealing (°C)/Size (bp) | Gene | Reference |
|---|---|---|---|---|
| Bem 23 F Bem 23 R | CGGAGCTTGCGCCTTAGTC CGGCTTTATCATAGCTCTCGT | 55/MEAM1 = 200; MED = 400 | Microsatellite | [59] |
| C1-J-2195 L2-N-3014 | TTGATTTTT TGGTCATCCAGAAGT TCCAATGCACTAATCTGCCATATTA | 51/850 | Cytochrome oxidase I (mtCOI) | [60] |
| Por-F Por-R | TGCAAGTCGAGCGGCATCAT AAAGTTCCCGCCTTATGCGT | 59/1000 | Portiera 16S rDNA | [35] |
| Rb F Rb R | GCTCAGAACGAACGCTATC GAAGGAAAGCATCTCTGC | 59/900 | Rickettsia 16S rDNA | [36] |
| 92 F Hb R | TGAGTAAAGTCTGGGAATCTGG AGTTCAAGACCGCAACCTC | 62/700 | Hamiltonella 16S rDNA | [35] |
| Ars23S-1 Ars23S-2 | CGTTTGATGAATTCATAGTCAAA GGTCCTCCAGTTAGTGTTACCCAAC | 59/600 | Arsenophonus 23S rDNA | [61] |
| Wol16S F Wol16S R | CGG GGGAAAAATTTATTGCT AGCTGTAATACAGAAAGTAAA | 55/650 | Wolbachia 16S rDNA | [38] |
| CFB F CFB R | GCGGTGTAAAATGAGCGTG ACCTMTTCTTAACTCAAGCCT | 59/500 | Cardinium 16S rDNA | [62] |
| Population Number, Location and Species | Host Plant | n | Infection of Bacterial Symbiont (%) | Reference | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| H | A | R | W | C | ||||||
| 1 | Croatia/Turanj | MED (Q2) | Cucumis sativus | 20 | 25 | - | 30 | 20 | 30 | [39] |
| 2 | Croatia/Split | MED (Q1) | Euphorbia pulcherrima | 20 | 35 | - | 20 | 10 | - | This study |
| 3 | Croatia/Zadar ¶ | MED (Q1 and Q2) | Hibiscus sp. | 20 | 95 | 35 | 60 | 50 | - | [41] |
| 4 | Montenegro/Bar | MED (Q2) | Dipladenia sanderi | 20 | 65 | 85 | 100 | 40 | - | |
| 5 | Montenegro/Lastva Grbaljska | MED (Q1) | Cucumis sativus | 20 | 30 | 85 | 40 | 30 | - | This study |
| 6 | Montenegro/Darza | MED (Q1) | Cucumis melo | 20 | 95 | - | 10 | 10 | - | |
| 7 | Macedonia/Doiran | MED (Q1) | Lycopersicon esculentum | 20 | 100 | - | - | 40 | 20 | |
| 8 | Macedonia/Gevgelija | MED (Q1) | Cucumis melo | 20 | 100 | - | 5 | 60 | - | |
| 9 | Israel | MED (Q2) | Gossypium hirsutum | 20 | - | 100 | 75 | 20 | - | [39] |
| 10 | Israel | MEAM1 | Gossypium hirsutum | 20 | 100 | - | 70 | - | - | |
| 11 | Croatia/Split | T. vaporariorum | Sonchus oleraceus | 20 | 5 | 75 | - | - | - | |
| 12 | Croatia/Split§ | T. vaporariorum | Euphorbia pulcherrima | 10 | 50 | 80 | - | - | - | [41]; This study |
| 13 | Montenegro/Podgorica | T. vaporariorum | Sonchus oleraceus | 20 | 20 | 95 | 5 | 10 | 35 | |
| 14 | Macedonia/Doiran | T. vaporariorum | Lycopersicon esculentum | 20 | 25 | 100 | - | - | - | This study |
| 15 | Macedonia/Grciste | T. vaporariorum | Lycopersicon esculentum | 20 | - | 100 | - | - | - | |
| 16 | Serbia/Trbusani | T. vaporariorum | Cucurbita pepo | 20 | 90 | 100 | - | - | - | |
| 17 | Serbia/Vranjska Banja | T. vaporariorum | Gerbera sp. | 20 | 45 | 100 | - | - | - | |
| 18 | Serbia/Opovo | T. vaporariorum | Lycopersicon esculentum | 20 | 70 | 90 | - | - | - | |
| 19 | Serbia/Zorka Subotica | T. vaporariorum | Chrysanthemum sp. | 20 | - | 95 | - | - | - | |
| 20 | Croatia/Brac-Supetar | S. phillyreae | Punica granatum | 20 | 65 | 35 | - | 40 | 5 | [41] |
| 21 | Croatia/Brac-Pucisca | S. phillyreae | Punica granatum | 20 | 100 | 25 | - | 20 | - | |
| 22 | Croatia/Opuzen | S. phillyreae | Punica granatum | 20 | 85 | 95 | - | 55 | 5 | |
| 23 | Croatia/Ljuta | S. phillyreae | Punica granatum | 20 | 85 | 85 | - | 15 | 70 | |
| 24 | Montenegro/Bar | S. phillyreae | Punica granatum | 20 | 90 | 100 | - | 50 | - | |
| Population Number, Location, and Species | Infection Frequencies of Bacterial Symbiont Combination (%) | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| R | H | A | W | C | RH | RA | HA | HW | HC | AW | AC | WC | RHA | RHW | RAW | RWC | HAW | HAC | HWC | AWC | RHWC | RHAW | HAWC | No Infection | ||
| 1 | Croatia/Turanj MED (Q2) | 10 | 10 | - | 5 | 15 | 15 | - | - | - | - | - | - | 5 | - | - | - | 5 | - | - | - | - | 5 | - | - | 35 |
| 2 | Croatia/Split MED (Q1) | 20 | 35 | - | 10 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 35 |
| 3 | Croatia/Zadar¶ MED (Q1 and Q2) | - | 20 | - | - | - | 15 | - | 15 | 5 | - | - | - | - | - | 25 | 5 | - | - | - | - | - | - | 15 | - | - |
| 4 | Montenegro/Bar MED (Q2) | 5 | - | - | - | - | 5 | 20 | - | - | - | - | - | - | 30 | 5 | 10 | - | - | - | - | - | - | 25 | - | - |
| 5 | Montenegro/L. Grbaljska MED (Q1) | - | - | 35 | 5 | - | - | 15 | - | - | - | 5 | - | - | 10 | - | - | - | 5 | - | - | - | - | 15 | - | 10 |
| 6 | Montenegro/Darza MED (Q1) | - | 75 | - | - | - | 10 | - | - | 10 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 5 |
| 7 | Macedonia/Doiran MED (Q1) | - | 55 | - | - | - | - | - | - | 25 | 5 | - | - | - | - | - | - | - | - | - | 15 | - | - | - | - | - |
| 8 | Macedonia/Gevgelija MED (Q1) | - | 40 | - | - | - | - | - | - | 55 | - | - | - | - | - | 5 | - | - | - | - | - | - | - | - | - | - |
| 9 | Israel MED (Q2) | - | - | 25 | - | - | - | 55 | - | - | - | - | - | - | - | - | 20 | - | - | - | - | - | - | - | - | - |
| 10 | Israel MEAM1 | - | 30 | - | - | - | 70 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| 11 | Croatia/Split§ T. vaporariorum | - | - | 70 | - | - | - | - | 5 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 25 |
| 12 | Croatia§/Split T. vaporariorum | - | - | 30 | - | - | - | - | 50 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 20 |
| 13 | Montenegro/Podgorica T. vaporariorum | - | - | 35 | 5 | - | - | - | 20 | - | - | 5 | 35 | - | - | - | - | - | - | - | - | - | - | - | - | - |
| 14 | Macedonia/Doiran T. vaporariorum | - | - | 75 | - | - | - | - | 25 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| 15 | Macedonia/Grciste T. vaporariorum | - | - | 100 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| 16 | Serbia/Trbusani T. vaporariorum | - | - | 10 | - | - | - | - | 90 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| 17 | Serbia/Vranjska Banja T. vaporariorum | - | - | 55 | - | - | - | - | 45 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| 18 | Serbia/Opovo T. vaporariorum | - | 5 | 25 | - | - | - | - | 65 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 5 |
| 19 | Serbia/Z. Subotica T. vaporariorum | - | - | 95 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 5 |
| 20 | Croatia/Brac-Supetar S. phillyreae | - | 25 | - | 10 | - | - | - | 20 | 10 | - | 5 | - | 5 | - | - | - | - | 10 | - | - | - | - | - | - | 15 |
| 21 | Croatia/Brac-Pucisca S. phillyreae | - | 60 | - | - | - | - | - | 20 | 15 | - | - | - | - | - | - | - | - | 5 | - | - | - | - | - | - | - |
| 22 | Croatia/Opuzen S. phillyreae | - | 5 | 5 | - | - | - | - | 35 | - | - | 10 | - | - | - | - | - | - | 40 | - | - | - | - | - | 5 | - |
| 23 | Croatia/Ljuta S. phillyreae | - | 10 | 5 | - | - | - | - | 15 | - | - | - | 5 | - | - | - | - | - | - | 50 | - | 5 | - | - | 10 | - |
| 24 | Montenegro/Bar S. phillyreae | - | - | 5 | - | - | - | - | 45 | - | - | 5 | - | - | - | - | - | - | 45 | - | - | - | - | - | - | - |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Skaljac, M.; Kanakala, S.; Zanic, K.; Puizina, J.; Pleic, I.L.; Ghanim, M. Diversity and Phylogenetic Analyses of Bacterial Symbionts in Three Whitefly Species from Southeast Europe. Insects 2017, 8, 113. https://doi.org/10.3390/insects8040113
Skaljac M, Kanakala S, Zanic K, Puizina J, Pleic IL, Ghanim M. Diversity and Phylogenetic Analyses of Bacterial Symbionts in Three Whitefly Species from Southeast Europe. Insects. 2017; 8(4):113. https://doi.org/10.3390/insects8040113
Chicago/Turabian StyleSkaljac, Marisa, Surapathrudu Kanakala, Katja Zanic, Jasna Puizina, Ivana Lepen Pleic, and Murad Ghanim. 2017. "Diversity and Phylogenetic Analyses of Bacterial Symbionts in Three Whitefly Species from Southeast Europe" Insects 8, no. 4: 113. https://doi.org/10.3390/insects8040113
APA StyleSkaljac, M., Kanakala, S., Zanic, K., Puizina, J., Pleic, I. L., & Ghanim, M. (2017). Diversity and Phylogenetic Analyses of Bacterial Symbionts in Three Whitefly Species from Southeast Europe. Insects, 8(4), 113. https://doi.org/10.3390/insects8040113







