Comparison of Model Predictions and Laboratory Observations of Transgene Frequencies in Continuously-Breeding Mosquito Populations
Abstract
:1. Introduction
2. Materials and Methods
2.1. Modelling
2.2. Laboratory Experiments
2.2.1. Mosquito Strains
2.2.2. Study Protocol
2.3. Statistical Analysis
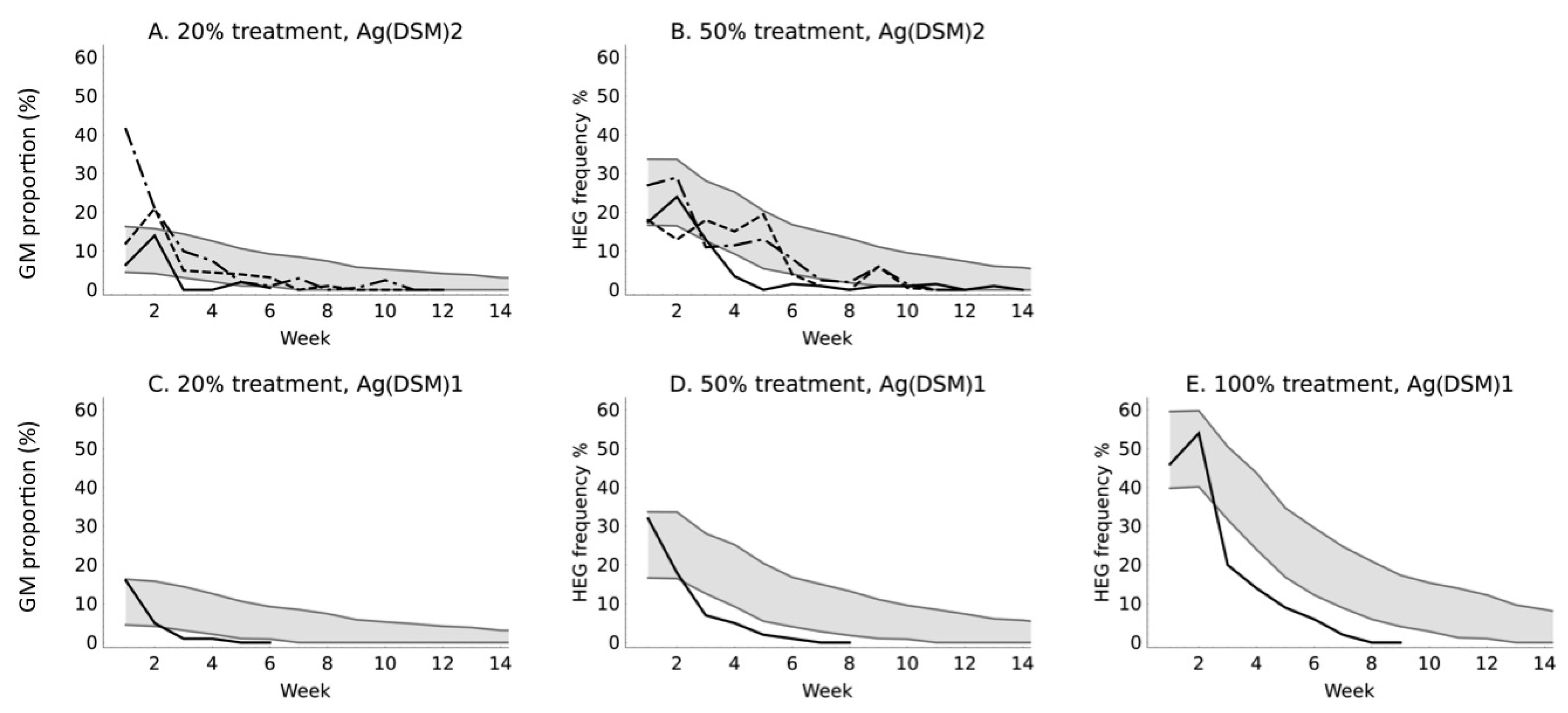
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- EFSA Panel on Genetically Modified Organisms (GMO). Guidance on the environmental risk assessment of genetically modified animals. EFSA J. 2013. [Google Scholar] [CrossRef]
- WHO-TDR/FNIH. Guidance Framework for Testing of Genetically Modified Mosquitoes; WHO: Geneva, Switzerland, 2014; p. 159. [Google Scholar]
- Harris, A.F.; McKemey, A.R.; Nimmo, D.; Curtis, Z.; Black, I.; Morgan, S.A.; Oviedo, M.N.; Lacroix, R.; Naish, N.; Morrison, N.I.; et al. Successful suppression of a field mosquito population by sustained release of engineered male mosquitoes. Nat. Biotechnol. 2012, 30, 828–830. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, D.O.; McKemey, A.R.; Garziera, L.; Lacroix, R.; Donnelly, C.A.; Alphey, L.; Malavasi, A.; Capurro, M.L. Suppression of a Field Population of Aedes aegypti in Brazil by Sustained Release of Transgenic Male Mosquitoes. PLoS Negl. Trop. Dis. 2015, 9, e0003864. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, K.W.; Robert, M.A.; Gould, F.; Lloyd, A.L. Feasible Introgression of an Anti-pathogen Transgene into an Urban Mosquito Population without Using Gene-Drive. PLoS Negl. Trop. Dis. 2014, 8, e2827. [Google Scholar] [CrossRef] [PubMed]
- Burt, A. Heritable strategies for controlling insect vectors of disease. Philos. Trans. R. Soc. B Biol. Sci. 2014. [Google Scholar] [CrossRef]
- Alphey, L. Genetic Control of Mosquitoes. Annu. Rev. Entomol. 2013, 59, 205–224. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, A.A.; Montgomery, B.L.; Popovici, J.; Iturbe-Ormaetxe, I.; Johnson, P.H.; Muzzi, F.; Greenfield, M.; Durkan, M.; Leong, Y.S.; Dong, Y.; et al. Successful establishment of Wolbachia in Aedes populations to suppress dengue transmission. Nature 2011, 476, 454–457. [Google Scholar] [CrossRef] [PubMed]
- Burt, A. Site-specific selfish genes as tools for the control and genetic engineering of natural populations. Proc. R. Soc. B Biol. Sci. 2003, 270, 921–928. [Google Scholar] [CrossRef] [PubMed]
- Bernardini, F.; Galizi, R.; Menichelli, M.; Papathanos, P.A.; Dritsou, V.; Marois, E.; Crisanti, A.; Windbichler, N. Site-specific genetic engineering of the Anopheles gambiae Y chromosome. Proc. Natl. Acad. Sci. USA 2014, 111, 7600–7605. [Google Scholar] [CrossRef] [PubMed]
- Hammond, A.; Galizi, R.; Kyrou, K.; Simoni, A.; Siniscalchi, C.; Katsanos, D.; Gribble, M.; Baker, D.; Marois, E.; Russell, S.; et al. A CRISPR-Cas9 gene drive system targeting female reproduction in the malaria mosquito vector Anopheles gambiae. Nat. Biotechnol. 2016, 34, 78–83. [Google Scholar] [CrossRef] [PubMed]
- Gantz, V.M.; Jasinskiene, N.; Tatarenkova, O.; Fazekas, A.; Macias, V.M.; Bier, E.; James, A.A. Highly efficient Cas9-mediated gene drive for population modification of the malaria vector mosquito Anopheles stephensi. Proc. Natl. Acad. Sci. USA 2015, 112, E6736–E6743. [Google Scholar] [CrossRef] [PubMed]
- Windbichler, N.; Papathanos, P.A.; Crisanti, A. Targeting the X chromosome during spermatogenesis induces Y chromosome transmission ratio distortion and early dominant embryo lethality in Anopheles gambiae. PLoS Genet. 2008, 4, e1000291. [Google Scholar] [CrossRef] [PubMed]
- Alphey, L.; Andreasen, M. Dominant lethality and insect population control. Mol. Biochem. Parasitol. 2002, 121, 173–178. [Google Scholar] [CrossRef]
- Irvin, N.; Hoddle, M.S.; O’Brochta, D.A.; Carey, B.; Atkinson, P.W. Assessing fitness costs for transgenic Aedes aegypti expressing the GFP marker and transposase genes. Proc. Natl. Acad. Sci. USA 2004, 101, 891–896. [Google Scholar] [CrossRef] [PubMed]
- Harvey-Samuel, T.; Ant, T.; Gong, H.; Morrison, N.I.; Alphey, L. Population-level effects of fitness costs associated with repressible female-lethal transgene insertions in two pest insects. Evol. Appl. 2014, 7, 597–606. [Google Scholar] [CrossRef] [PubMed]
- Catteruccia, F.; Godfray, H.C.J.; Crisanti, A. Impact of genetic manipulation on the fitness of Anopheles stephensi mosquitoes. Science 2003, 299, 1225–1227. [Google Scholar] [CrossRef] [PubMed]
- Marrelli, M.T.; Li, C.; Rasgon, J.L.; Jacobs-Lorena, M. Transgenic malaria-resistant mosquitoes have a fitness advantage when feeding on Plasmodium-infected blood. Proc. Natl. Acad. Sci. USA 2007, 104, 5580–5583. [Google Scholar] [CrossRef] [PubMed]
- Collins, F.H.; Mendez, M.A.; Rasmussen, M.O.; Mehaffey, P.C.; Besansky, N.J.; Finnerty, V. A ribosomal RNA gene probe differentiates member species of the Anopheles gambiae complex. Am. J. Trop. Med. Hyg. 1987, 37, 37–41. [Google Scholar] [PubMed]
- Wilkins, E.E.; Howell, P.I.; Benedict, M.Q. X and Y chromosome inheritance and mixtures of rDNA intergenic spacer regions in Anopheles gambiae. Insect Mol. Biol. 2007, 16, 735–741. [Google Scholar] [CrossRef] [PubMed]
- Galizi, R.; Doyle, L.A.; Menichelli, M.; Bernardini, F.; Deredec, A.; Burt, A.; Stoddard, B.L.; Windbichler, N.; Crisanti, A. A synthetic sex ratio distortion system for the control of the human malaria mosquito. Nat. Commun. 2014. [Google Scholar] [CrossRef] [PubMed]
- Klein, T.A.; Windbichler, N.; Deredec, A.; Burt, A.; Benedict, M.Q. Infertility resulting from transgenic I-PpoI male Anopheles gambiae in large cage trials. Pathog. Glob. Health 2012, 106, 20–31. [Google Scholar] [CrossRef] [PubMed]
- Benedict, M.Q.; Hood-Nowotny, R.C.; Howell, P.I.; Wilkins, E.E. Methylparaben in Anopheles gambiae s.l. sugar meals increases longevity and malaria oocyst abundance but is not a preferred diet. J. Insect Physiol. 2009, 55, 197–204. [Google Scholar] [CrossRef] [PubMed]
- Toni, T.; Welch, D.; Strelkowa, N.; Ipsen, A.; Stumpf, M.P.H. Approximate Bayesian computation scheme for parameter inference and model selection in dynamical systems. J. R. Soc. Interface 2009, 6, 187–202. [Google Scholar] [CrossRef] [PubMed]
- Csilléry, K.; Blum, M.G.B.; Gaggiotti, O.E.; François, O. Approximate Bayesian Computation (ABC) in practice. Trends Ecol. Evol. 2010, 25, 410–418. [Google Scholar] [CrossRef] [PubMed]
- Malaria Research and Reference Reagent Resource Center (MR4). Available online: https://www.beiresources.org/Catalog/livingMosquitoes/MRA-112.aspx (accessed on 6 April 2016).
- Damiens, D.; Benedict, M.Q.; Wille, M.; Gilles, J.R.L. An Inexpensive and Effective Larval Diet for Anopheles arabiensis (Diptera: Culicidae): Eat Like a Horse, a Bird, or a Fish? J. Med. Entomol. 2012, 49, 1001–1111. [Google Scholar] [CrossRef] [PubMed]
- Valerio, L.; Collins, C.M.; Lees, R.S.; Benedict, M.Q. Benchmarking vector arthropod culture: an example using the African malaria mosquito, Anopheles gambiae (Diptera: Culicidae). Malar. J. 2016. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, J.; Bates, D.; DebRoy, S.; Sarkar, D. NLME: Linear and Nonlinear Mixed Effect Models; R Foundation for Statistical Computing: Vienna, Austria, 2015; Available online: https://cran.r-project.org/web/packages/nlme/index.html (accessed on 6 April 2016).
- R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013; Available online: http://www.R-project.org/ (accessed on 6 April 2016).
- Facchinelli, L.; Valerio, L.; Lees, R.S.; Oliva, C.F.; Persampieri, T.; Collins, C.M.; Crisanti, A.; Spaccapelo, R.; Benedict, M.Q. Stimulating Anopheles gambiae swarms: application for behavioural and fitness studies. Malar. J. 2015, 14, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Facchinelli, L.; University of Perugia, Perugia, Italy. Unpublished data. 2014.
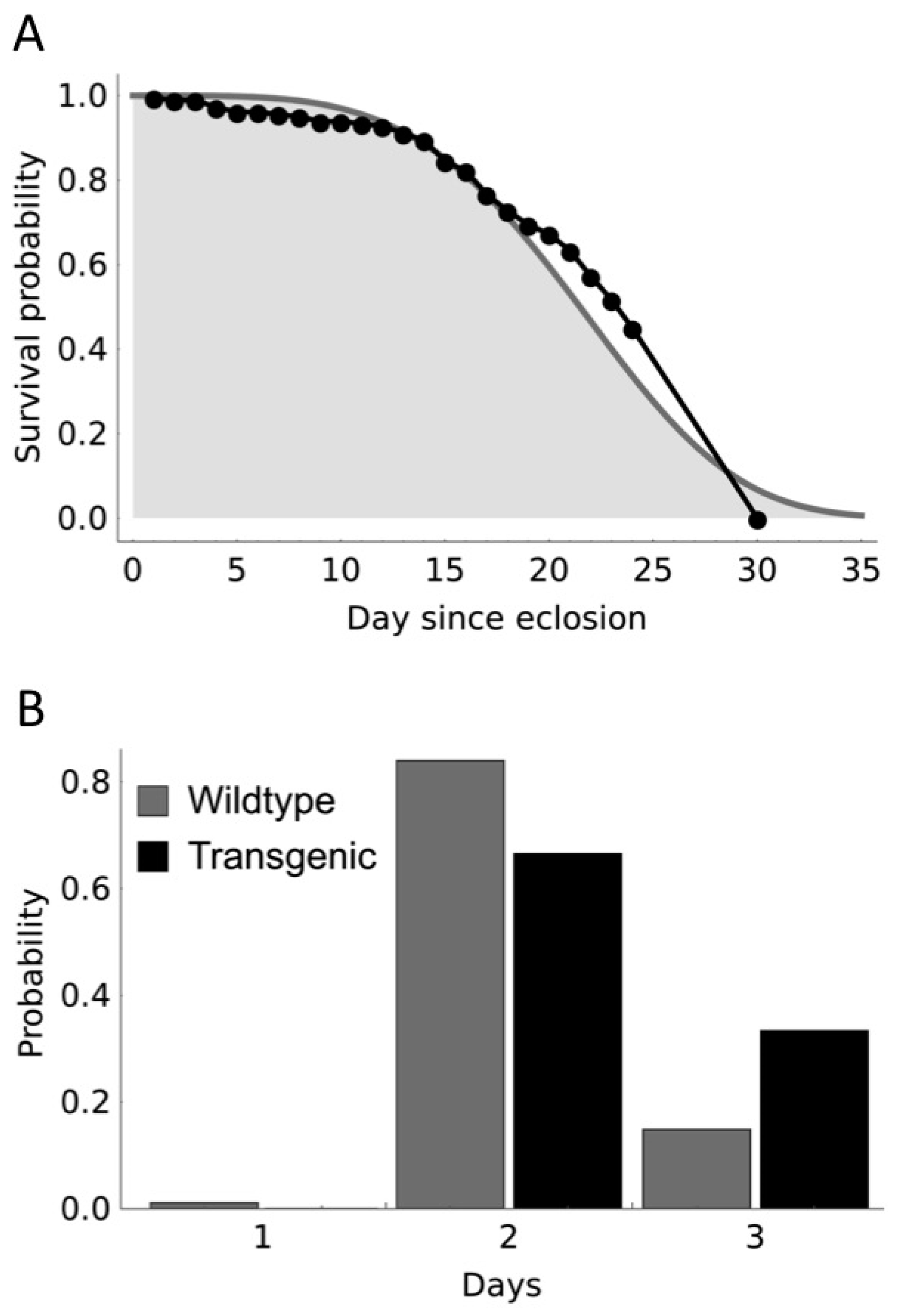
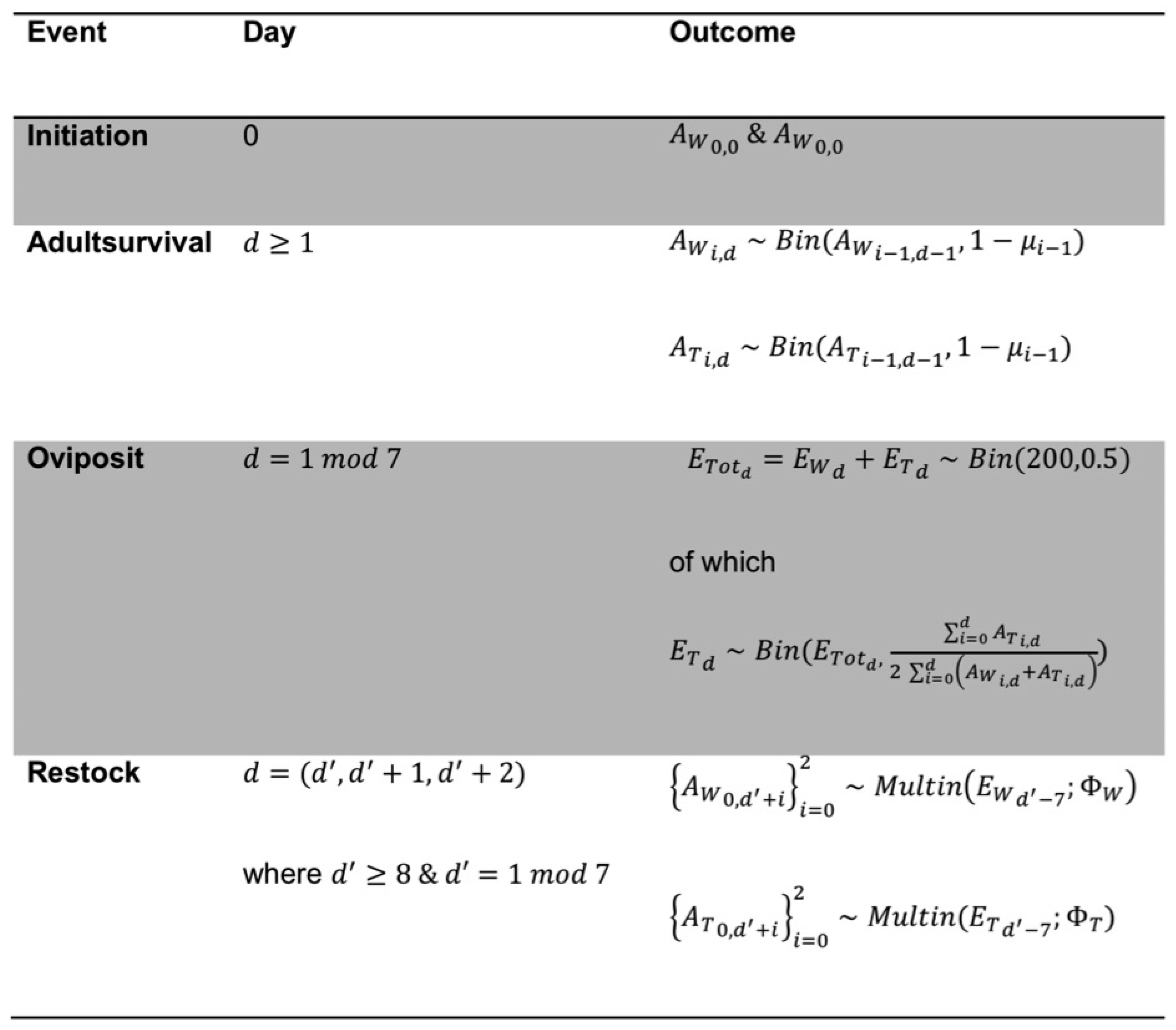
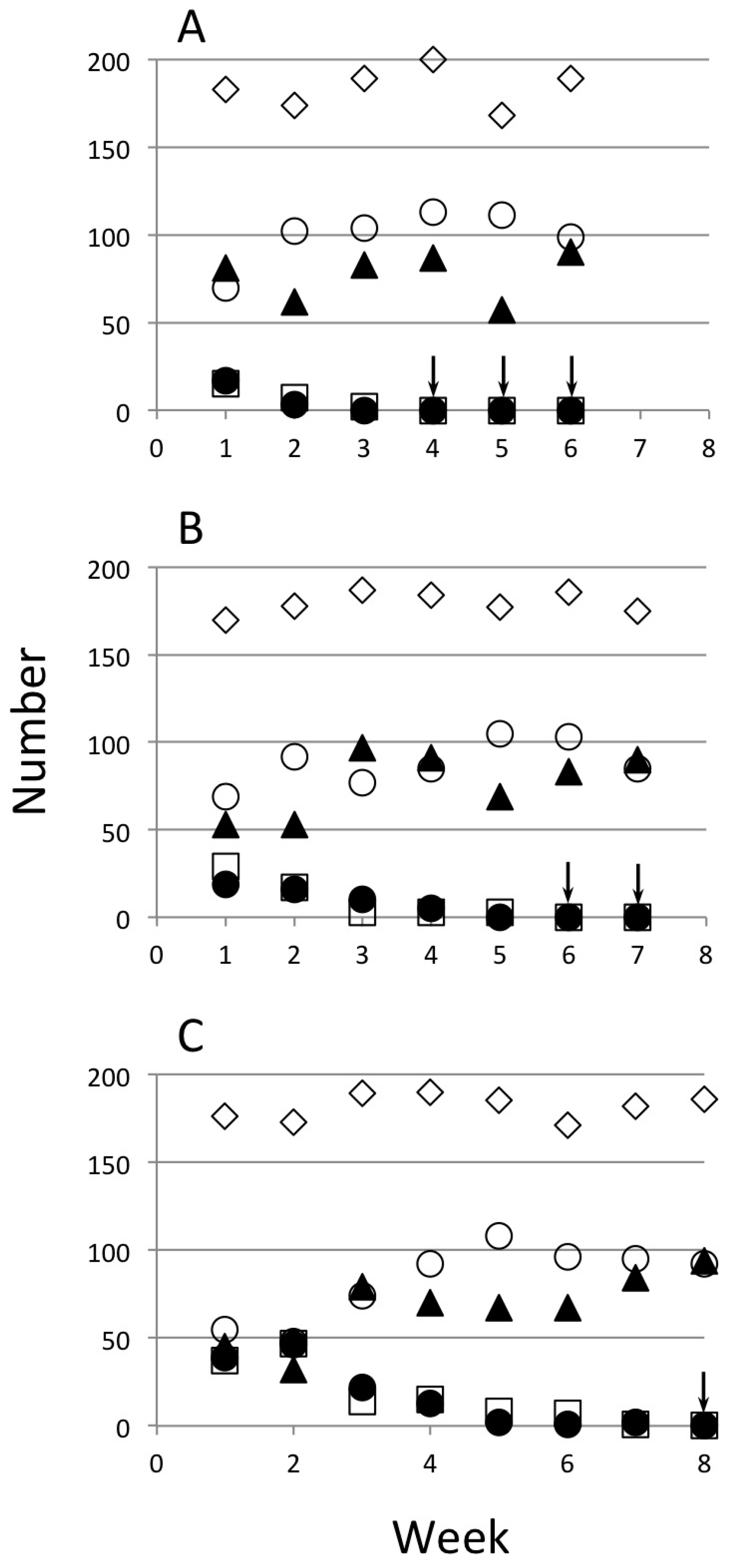
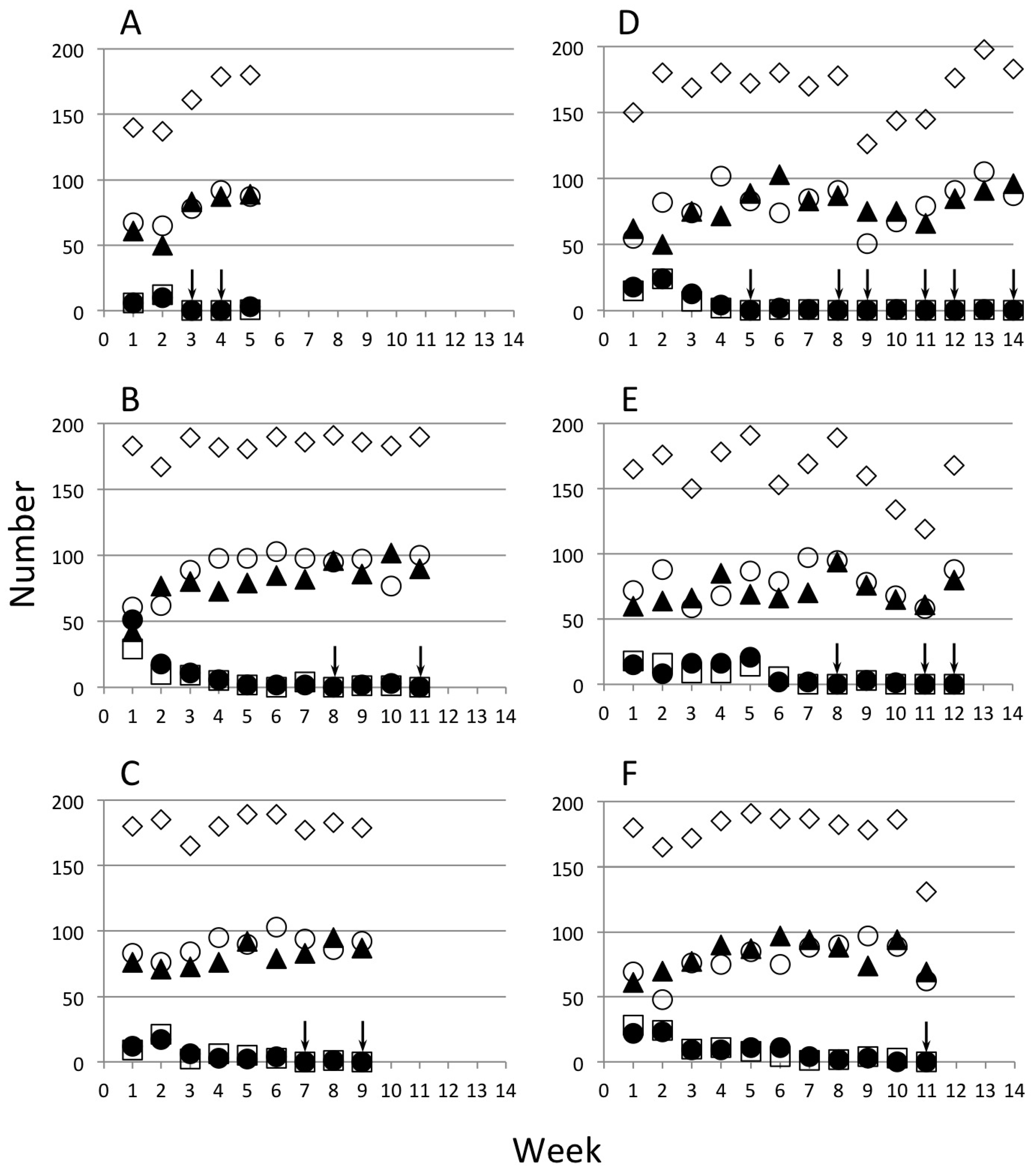
| Strain | Initial Release Level | L Ratio | d.f. | p |
|---|---|---|---|---|
| Ag(DSM)1 | 100% | 34.47 | 5,6 | <0.001 |
| 50% | 21.07 | 6,7 | <0.001 | |
| 20% | 4.62 | 6,7 | <0.05 | |
| Ag(DSM)2 | 50% | 32.93 | 6,7 | <0.001 |
| 20% | 9.39 | 6,7 | <0.01 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Valerio, L.; North, A.; Collins, C.M.; Mumford, J.D.; Facchinelli, L.; Spaccapelo, R.; Benedict, M.Q. Comparison of Model Predictions and Laboratory Observations of Transgene Frequencies in Continuously-Breeding Mosquito Populations. Insects 2016, 7, 47. https://doi.org/10.3390/insects7040047
Valerio L, North A, Collins CM, Mumford JD, Facchinelli L, Spaccapelo R, Benedict MQ. Comparison of Model Predictions and Laboratory Observations of Transgene Frequencies in Continuously-Breeding Mosquito Populations. Insects. 2016; 7(4):47. https://doi.org/10.3390/insects7040047
Chicago/Turabian StyleValerio, Laura, Ace North, C. Matilda Collins, John D. Mumford, Luca Facchinelli, Roberta Spaccapelo, and Mark Q. Benedict. 2016. "Comparison of Model Predictions and Laboratory Observations of Transgene Frequencies in Continuously-Breeding Mosquito Populations" Insects 7, no. 4: 47. https://doi.org/10.3390/insects7040047
APA StyleValerio, L., North, A., Collins, C. M., Mumford, J. D., Facchinelli, L., Spaccapelo, R., & Benedict, M. Q. (2016). Comparison of Model Predictions and Laboratory Observations of Transgene Frequencies in Continuously-Breeding Mosquito Populations. Insects, 7(4), 47. https://doi.org/10.3390/insects7040047




_Collins.png)


