Abstract
Since the original description of Helicosporidium parasiticum in 1921, members of the genus Helicosporidium have been reported to infect a wide variety of invertebrates, but their characterization has remained dependent on occasional reports of infection. Recently, several new Helicosporidium isolates have been successfully maintained in axenic cultures. The ability to produce large quantity of biological material has led to very significant advances in the understanding of Helicosporidium biology and its interactions with insect hosts. In particular, the unique infectious process has been well documented; the highly characteristic cyst and its included filamentous cell have been shown to play a central role during host infection and have been the focus of detailed morphological and developmental studies. In addition, phylogenetic analyses inferred from a multitude of molecular sequences have demonstrated that Helicosporidium are highly specialized non-photosynthetic algae (Chlorophyta: Trebouxiophyceae), and represent the first described entomopathogenic algae. This review provides an overview of (i) the morphology of Helicosporidium cell types, (ii) the Helicosporidium life cycle, including the entire infectious sequence and its impact on insect hosts, (iii) the phylogenetic analyses that have prompted the taxonomic classification of Helicosporidium as green algae, and (iv) the documented host range for this novel group of entomopathogens.
1. Introduction
The genus Helicosporidium was first described by Keilin in 1921 [1]. The original specimen was isolated in England from the ceratopogonid larvae Dasyhelea obscura (Diptera), and named Helicosporidium parasiticum. In 1931, the genus and species names were validated and placed in a newly created order, Helicosporidia, in an attempt to classify these organisms [2]. The second report of Helicosporidium parasiticum in an insect host occurred in 1970, and interestingly, this pathogen was reported in a different insect host order (Lepidoptera) isolated from a different continent (Argentina) [3]. Albeit rare, Helicosporidium infections have since been reported from a variety of invertebrate hosts isolated from diverse geographic locations. In insect hosts, this pathogen has predominantly been found to infect larvae. As noted throughout the vast majority of reports, the unique morphological features of Helicosporidium facilitate identification, although it was recognized for a long time as a major obstacle to a precise taxonomic classification, especially during early descriptions [1,3].
2. The Cyst, Which Includes a Filamentous Cell, Is the Characteristic and Diagnostic Feature for the Genus Helicosporidium
Virtually all reports of Helicosporidium infection in invertebrate hosts rely on the observation of unique and characteristic four-cell structures [1,3,4,5,6,7,8,9,10,11,12,13]. These structures have been alternatively termed spores [1,3,4,5,6,7,9] or cysts [8,10,11,12,13]. Although the term spore was used during the original description of Helicosporidium parasiticum [1], and can still be found in occasional modern reports [9], most current studies, including this review, refer to the diagnostic feature of Helicosporidium as cyst. The Helicosporidium cyst is a barrel-shape structure that contains a core of three superposed ovoid cells surrounded by a single elongated, filamentous cell. These four cells are enclosed in a pellicle. The original H. parasiticum description featured elaborate drawings and microphotographs of the cyst and the filamentous cell [1]. Since then, electron microscopy photographs of the characteristic cyst have routinely been provided to support the identification of Helicosporidium in insect and other invertebrates. Recent reports of Helicosporidium sp. in various Coleopteran hosts, including the great European spruce bark beetle Dendroctonus micans (Coleoptera: Curculionidae), the predator beetle Rhizophagus grandis (Coleoptera: Rhizophaginae), and the weevil Cyrtobagous salviniae (Coleoptera: Curculionidae), all incorporated transmission electron micrographs (TEM) depicting cysts with the peripheral filamentous cell surrounding the three ovoid cells [10,11,12]. Cell measurements [7,8,10,12] indicated that the cysts are rather small, and range from 3 to 6 μm, although some of this variation might be due to differences in preparation methods [7]. Inside the cysts, the filamentous cell typically wraps around the core of ovoid cells three or four times, and can be distinguished on the narrow outer surface of the cysts (Figure 1).
In addition to TEM pictures, a recent study presented scanning electron microscopes pictures of cysts that were purified using Ludox gradient centrifugation, as well as light microscopy and SEM pictures of the filamentous cell being liberated from the cyst and separated from the remaining three ovoid cells [8]. This process is known as dehiscence (Figure 2). It has been observed both in vitro, by applying pressure to the microscope slide [8], and in vivo, in the gut lumen of susceptible hosts [14], and in the host hemolymph, after a series of desiccation events [1]. Significantly, dehisced cysts were used to single out the filamentous cells and highlight the presence of barbs [8] (Figure 2). Purified filamentous cells range in length from 37 to 62 μm [1,4,8,10,14]. It remains unclear if these ultrastructural differences can be related to the hosts from which these Helicosporidium were isolated.
The dividing stage of Helicosporidium corresponds to vegetative cells. These cells are characterized by the mitotic production of multi-cellular structures containing two, four or eight cells within a pellicle [14,15,16]. Similar to the cyst, the structure can rupture and release daughter cells from an empty pellicle. This mode of reproduction was termed autosporulation in reference to a similar process described for related taxa [16]. Meiosis has yet to be reported. Vegetative growth have been successfully obtained in vitro using cell-free artificial media, demonstrating that Helicosporidium can replicate without living host cells [8]. In heterologous, lepidopteran hosts, only a minority (15%–21%) of four-cell vegetative cells from a dipteran isolate developed into the typical cysts [16], and it is unknown if cyst morphogenesis is more efficient in natural hosts. Cyst differentiation has yet to be observed in in vitro cultures, suggesting that host–derived stimuli are required for this process [16]. Rather than developing into cysts, Helicosporidium cells allowed to undergo repetitive autosporulation cycles in artificial media were observed to form palmelloid colonies, which correspond to the agglutination of potentially abnormal vegetative cells [16]. Another notable difference between in vitro and in vivo vegetative development of Helicosporidium cells involves cell concentrations, as in vitro cultures obtained in artificial media failed to match the cell density observed in the host hemolymph [16].
Following the development of axenic cultures [8], three strains have been deposited the American Type Culture Collection (ATCC). These strains, labeled Sj-1, Dm-1, and Cs-1, refer to isolates, respectively collected from the black fly Simulium jonesi, the bark beetle Dendroctonus micans, and the weevil Cyrtobagous salviniae [8,10,12]. All three strains have been prudently named Helicosporidium sp., without a full species name, since it remains unclear if the genus is comprised of only one species, and if all Helicosporidium isolates are Helicosporidium parasiticum Keilin [1]. Comparative analyses between isolates are rare and have been complicated by difficulties in locating the original specimen. Hence, the vast majority of new Helicosporidium reports have been based on the limited description of the diagnostic cysts and filamentous cells, with little attempts to further the identification past the genus level, leading to reports of Helicosporidium sp. [6,7,8,9,10,11,12,13]. Polymorphic traits associated with cell morphology have been identified, but it is unclear if they are taxonomically significant [8,10,17]. Potentially, in vitro cultures of Helicosporidium may provide a basis to develop comparative analyses between isolates, and identify differences justifying the description of several Helicosporidium species. Comparative analyses of in vitro cultures of Helicosporidium Sj-1 and related taxa have already been performed [18]. In addition, axenic cultures will facilitate the characterization of the different cell phenotypes at the molecular level. In particular, the compositions of the Helicosporidium pellicle, or the vegetative cell wall, remain largely unknown, even if these structures play an important role in pathogenicity. Although molecular data has been generated in the form of expressed sequence tags (ESTs), they have been mainly used for phylogenetic analyses and not for molecular characterization of Helicosporidium cells [19]. To date, in vitro cultures have mostly proved crucial to obtain molecular sequences that precisely established the taxonomic affinities of Helicosporidium spp., and to investigate the transitions between cell types in order to fully understand the Helicosporidium life cycle.
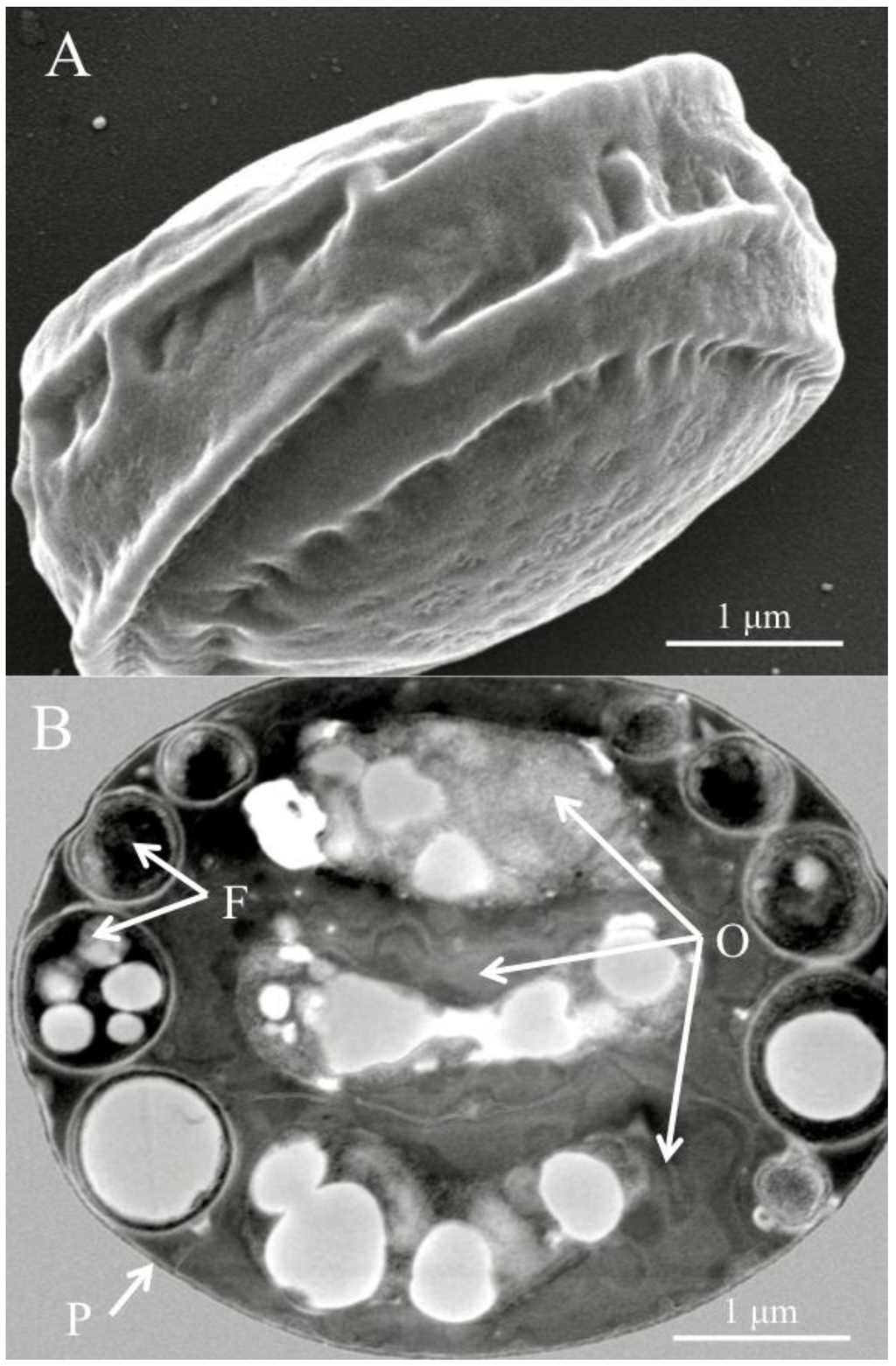
Figure 1.
(A) Scanning electron micrograph of a diagnostic Helicosporidium cyst. (B) Transmission electron micrograph (cross section) detailing the core of three stacked ovoid cells (O) and the filamentous cell (F) contained within a pellicle (P).
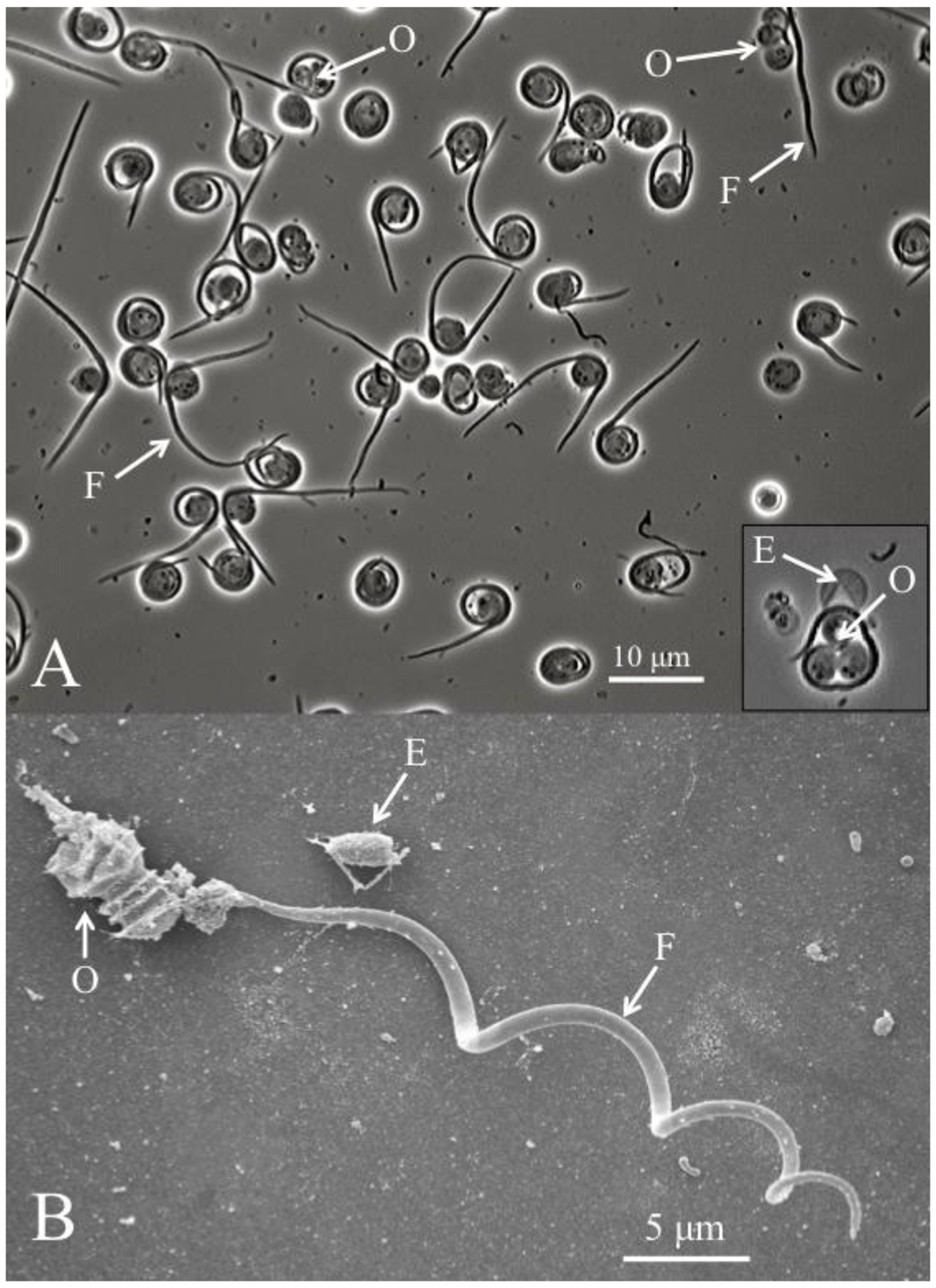
Figure 2.
(A) Helicosporidium cyst dehiscence observed under light microscopy, showing groups of three ovoid cells (O), diagnostic filamentous cells (F) and empty pellicles (E). (B) Scanning electron micrograph detailing the filamentous cell and its barbs (pointing away from the core of ovoid cells).
3. The Cyst, and the Included Filamentous Cell, Initiates the Helicosporidium Infectious Cycle in Susceptible Insect Hosts
The pathogenicity process of Helicosporidium spp. has been inferred principally from two long‑term studies that detailed the infection sequence in heterologous lepidopteran hosts maintained in laboratories [8,14,15,16,20]. A Helicosporidium originally isolated from larvae and adults of Carpophilus mutilatus (Coleoptera, Nitidulidae) was tracked in the navel orangeworm Paramyelois transitella [14,15]. Later, Helicosporidium Sj-1 was used to obtain infections in common laboratory insects such as the corn earworm Helicoverpa zea, the tobacco hornworm Manduca sexta, or the beet armyworm Spodoptera exigua [8,16,20]. Both studies established that cysts are the infective propagules and that infection occurs per os, when cysts are ingested by susceptible hosts. During insect bioassays, significant infection rates have routinely be obtained after per os challenge, even though a significant portion of intact cysts may be retrieved in the animal’s feces [20]. Gut dissections clearly demonstrated that cyst dehiscence was induced in the gut lumen [8,14,20]. The impact of insect gut on cysts was confirmed in vitro, as cysts incubated in midgut fluids were induced to rupture and release the filamentous cells [8]. Ovoid cells contained in the cysts were observed to lyse in the midgut [8,20]. In vivo cyst dehiscence is illustrated in Figure 3, complementing additional micrographs of insect gut lumen that demonstrated the infection process [8,20].
An initial investigation suggested that cysts might bind to the insect peritrophic membrane before dehiscence, and therefore be in a position where the released filamentous cells penetrate the peritrophic matrix immediately after dehiscence [8]. Although cyst dehiscence was initially described in the host hemocoel [1], it is now well established that filamentous cells are not found freely circulating in the hemolymph, but mediate the transition from midgut tissues to the host hemolymph, where Helicosporidium vegetative cells can be observed 24 h or 48 h post infection [8,14,16,20]. Detailed microscopic observations revealed that the released filamentous cells pass through the midgut epithelium, reach the hemocoel, and may create significant damage to the peritrophic membrane so that remaining cysts and cells gain access to the underlying tissues, and eventually, to the hemocoel [20]. Even the smallest described filamentous cells (37 μm) were estimated to be long enough to puncture the peritrophic membrane and the ectoperitrophic space, and gain access to the midgut cells [8]. The orientation of the filamentous cells barbs was found to be highly consistent, suggesting that the host‑pathogen interactions may be highly regulated at the cellular and molecular level. The barbs were first observed to be oriented towards the gut lumen as the filamentous cells initiated contact with the matrix, but then switched orientation and pointed towards the hemocoel as the filamentous cells gain ingress in the hemolymph [20].
The transition from invasive filamentous cells to replicating vegetative cells was elucidated recently under in vitro conditions [16]. In a manner similar to the more commonly observed autosporulation process, filamentous cells swell and undergo two division cycles within the original cell wall, releasing four bean-shaped daughter cells. These elongated cells divide again to produce spherical vegetative cells [16]. It is assumed that a similar differentiation takes place in vivo. Although uncommon, elongated cells were observed in both gut and hemolymph preparations of infected P. transitella [14]. Remnants of filamentous cells and bean-shaped cells were also observed being phagocytosed in infected S. exigua hemocytes [16]. The vegetative cells are responsible for hemolymph colonization. Since no filamentous cells are observed in the hemolymph, it is hypothesized that the transition occurs either in the host phagocytes for the filamentous cells that reach the hemolymph, or during the ingress from the gut tissue to the hemocoel, once the peritrophic matrix has been disrupted [20].
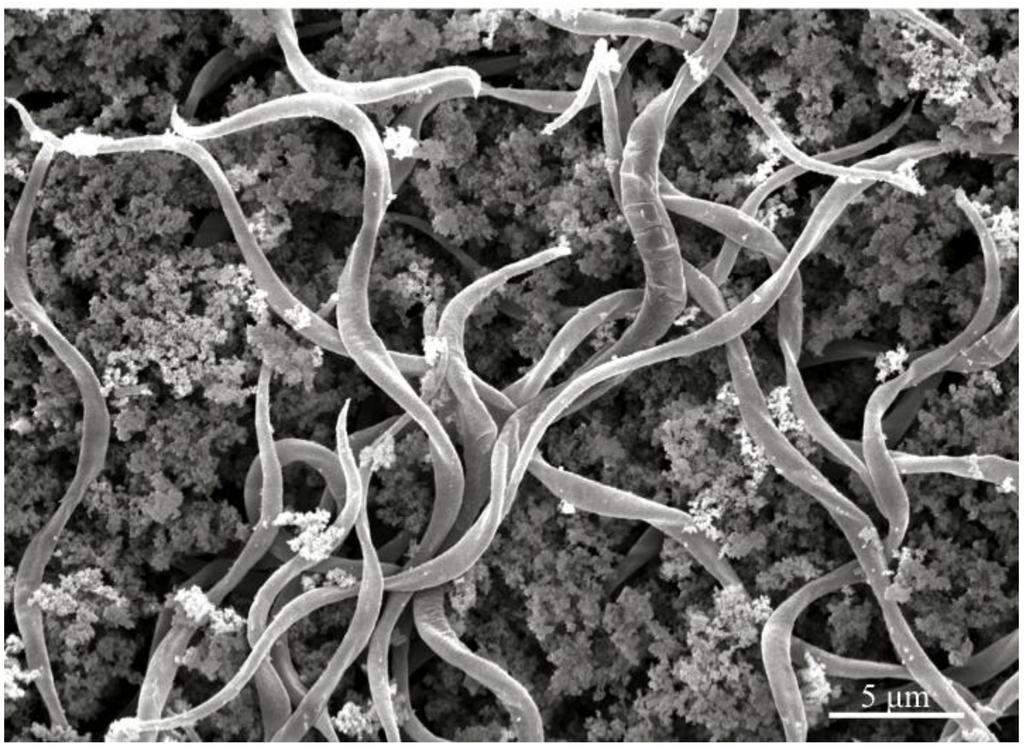
Figure 3.
Scanning electron micrograph of the gut content of insect hosts challenged with Helicosporidium cysts, demonstrating that filamentous cells are released in the gut lumen.
The vegetative cells are not recognized by the defense system of susceptible hosts, and replicate freely in the hemocoel, or in the hemocytes. They eventually fill the hemocoel, and have been showed to reach very high concentrations [16,20]. The Helicosporidium life cycle culminates as a portion of the vegetative cells differentiate into cysts [1,8,14,15,16,20,21]. This process has been observed both in the hemolymph and in hemocytes [20], and is impacted by both host age and pathogen dosage [21]. Although cyst production has yet to occur in artificial media, the vegetative cells obtained in vitro were shown to maintain their competence to differentiate into cysts when injected into host hemolymph, suggesting that a host-derived stimulus is required for cyst morphogenesis [16]. Direct injection of Helicosporidium cells in host hemolymph also resulted in 100% infection rates, which contrasted with the 50% rates reported for per os challenges [22]. This observation not only confirmed that cells are undetected by the host defense system, but also suggested that mechanisms of resistance against Helicosporidium infection are related to the ingress of the pathogen through the midgut [22].
High cell concentration and cyst morphogenesis in the hemolymph has variable impact on the insect host. There is no consistent or characteristic symptom associated with Helicosporidium infection in insects. Noticeable infection symptoms, such as change in hemolymph color and decreased host weight gain [23], or decreased host mobility [24], have occasionally been reported. Host death may be considered as the ultimate symptom, as suggested by the original Helicosporidium description [1]. However, several studies also indicated that the morphology [11,13] and the development (pupation) of infected insects [9,17,22,25] appeared unaffected. The lack of characteristic symptoms for Helicosporidium infection is consistent with the variable infectivity and mortality rates reported for this pathogen. Infection rates are dependent on both hosts and strains. A Helicosporidium sp. isolate tested on various mosquito species resulted in infection rates ranging from 0 to 93% [17]. In heterologous lepidopteran hosts, Helicosporidium Cs-1 produced an infection rate of 50% [22]. Bioassays demonstrated that infection rates are dose dependent [21,23]. Mortality rates have been equally variable, and can also be modulated by the host age and the concentration of cysts used to challenge potential hosts [21,26]. Significant mortality rates have been reported in several mosquito hosts, spurring substantial investigations on the potential of Helicosporidium as a biological control agent against mosquito larvae [9,17,24,25,26,27]. Overall, the variability in pathogenicity and virulence observed in different studies may also be correlated to the subsampled diversity within the genus Helicosporidium. The small number of infection bioassays that attempted to compare several Helicosporidium isolates all highlighted the fact that distinct isolates can be readily distinguished, based on differences in infectivity or virulence towards a single host [23,26]. These observations support the existence of several Helicosporidium species.
Lastly, although the Helicosporidium life cycle, from cyst ingestion to cyst morphogenesis in the hemolymph has been clearly established, no study has unambiguously described how Helicosporidium cysts exit the hemolymph of infected insects and are transmitted to subsequent hosts. The original H. parasiticum description indicated that infected larvae might become so impaired that the exoskeleton ruptures and the entire hemocoel content (including the infective cysts) is released into the environment [1]. This release has also been reported in mosquito larvae [26]. In pathosystems where the hosts appeared mostly unaffected and infection was trans-stadially maintained, vertical transmission of Helicosporidium was investigated. In mosquito hosts, infection of ovarian tissues was not observed [25], but vertical transmission was reported in noctuids [22]. Studies that described infections in both insects of a prey/predator system (the bark beetle Dendroctonus micans and its predator Rhizophagus grandis) suggested that Helicosporidium cysts may be directly transmitted by feeding when insects consume an infected prey, or potentially cannibalize an infected conspecies [10,11,13].
4. Phylogenetic and Phylogenomic Analyses Revealed that Helicosporidium spp. Are Non‑Photosynthetic Green Algae
The taxonomic classification for the genus Helicosporidium was historically a matter of debate, with these organisms alternatively proposed as Protozoa [1,2] or lower Fungi [3], based on morphological characteristics. However, the phylogenetic affinity of Helicosporidium has been unequivocally established by molecular-based analyses. The first sequenced gene fragments from Helicosporidium Sj-1 corresponded to rDNA, actin and β-tubulin. Phylogenetic reconstructions inferred from these genes revealed that members of the genus Helicosporidium are green algae (Chlorophyta) [28]. The SSU-rDNA gene fragment was also used to demonstrate that the genus Helicosporidium belongs to the chlorophyte class Trebouxiophyceae, and is closely related to the non‑photosynthetic algae of the genus Prototheca [28]. A set of 69 ribosomal protein sequences from the Sj-1 strain provided strong support, not only for the reclassification of Helicosporidium as green algae, but also for the specific relationship with Prototheca spp. [19]. Following this strongly supported, molecular-based taxonomic classification, similarities between Helicosporidium cells and trebouxiophyte green algae have also been highlighted at the cellular level, principally in regards to the vegetative cell multiplication and the autosporulation process [8,16].
The identification of Helicosporidium spp. as non-photosynthetic green algae prompted several studies aimed at characterizing chloroplast-like genetic material, since achlorophytic algae and plants have been shown to retain vestigial chloroplasts, or plastids. Amplification and analysis of plastid 16S rDNA gene fragments suggested that Helicosporidium cells had retained such an organelle [29]. Sequencing of a cluster of protein encoding genes demonstrated that the Helicosporidium plastid DNA was potentially transcribed [30], and a subset of expressed sequences tags annotated as homologs to nucleus-encoded, plastid-targeted proteins offered additional indirect evidence for a plastid and its included genome [31]. Although the organelle itself has yet to be observed in ultra-thin sections of Helicosporidium cells, the presence of a vestigial chloroplast has been thoroughly demonstrated by the sequencing of the entire plastid genome [32]. This genome lacks genes involved in photosynthesis and is so reduced that it is among the smallest known plastid genome. Overall, the plastid genome, and the phylogenetic inferred from its genes, have confirmed that Helicosporidium spp. are highly specialized non-photosynthetic trebouxiophyte green algae. Additional phylogenomic evidence was most recently provided by the analysis of the complete mitochondrial genome for Helicosporidium Sj-1 [33].
Trebouxiophyte algae include Coccomyxa spp., which are known pathogens of invertebrates such as mussels, scallops, geoducks, and starfishes [34,35,36]. However, Coccomyxa spp. are photosynthetic and have never been reported in insect hosts. The non-photosynthetic Helicosporidium spp. remain the only described achlorophytic entomopathogenic algae, and recent phylogenetic analyses indicated that the genera Helicosporidium and Coccomyxa are not closely related [37]. Virtually all phylogeny reconstructions have associated Helicosporidium with Prototheca, which harbors emerging pathogens of vertebrates, including isolates infecting humans, pets, and farm animals [38]. Both Prototheca and Helicosporidium are non photosynthetic, and since most phylogenetic analyses have depicted Helicosporidium and Prototheca as members of a monophyletic clade, it is hypothesized that the loss of photosynthesis represents a synapomorphic character for these algae. Only selected species of Prototheca are known to be pathogenic, and although there is similarities in cell division, pathogenic isolates of P. wickerhamii or P. zopfii have never been associated with a cyst stage, or filamentous cells, similar to Helicosporidium. To date, these unique cellular structures are thought to be specific to the genus Helicosporidium and, potentially, to its infectious process.
In addition to the studies aimed at precisely positioning Helicosporidium within a taxonomic framework, independent phylogenetic analyses focused on the genus Prototheca have incorporated the sequences generated from Helicosporidium spp. and confirmed the close relationship between the two genera [39,40]. Both Prototheca and Helicosporidium have appeared in a monophyletic clade that included the photosynthetic species Auxenochlorella protothecoides, and that has been referred to as the Auxenochlorella-Helicosporidium-Prototheca (AHP) clade [39]. In taxa-rich phylogenetic reconstructions focused on these genera, the species-level relationships have consistently been obscured by very poor resolution, and therefore the precise taxonomic position of Helicosporidium within the Trebouxiophyceae remains unclear. Lineages within the class Trebouxiophyceae are currently poorly resolved [41], and in the case of the AHP clade, it has been further complicated by the proposals for two novel Prototheca species, which have yet to appear in large-scale taxonomic classifications [42,43]. A general consensus might be emerging, as the inclusion of Helicosporidium has consistently led to phylogenetic trees that depicted Prototheca as a paraphyletic genus. Recent studies indicated that Helicosporidium is a sister taxon to a strongly supported clade that includes P. zopfii, P. ulmea, P. moriformis, P. stagnora, and the newly proposed P. blaschkeae [37,40]. Other species, such as P. wickerhamii and P. cutis, were not identified as members of this Prototheca sensu stricto clade, and therefore several authors have suggested that they should be re-assigned to alternative genera [39,40]. Although they have yet to include comparative analyses with Helicosporidium, alternative cladistic approaches, based on nutritional requirements [40], or biochemical profiles [38,39], have confirmed the strong heterogeneity of the genus Prototheca. To date, the closest relative to Helicosporidium spp. remain unidentified. A refined and potentially revised Prototheca taxonomy may be necessary to precisely establish species-level relationships between non‑photosynthetic trebouxiophyte algae, and determine if the acquisition of pathogenicity occurred once or multiple times in these algae.
Attempts to obtain strongly supported phylogenetic trees featuring both Helicosporidium and Prototheca have led to the generation of sequence data from multiple Helicosporidium isolates, and revealed an unsuspected genetic diversity. Sequences from plastid 16S rDNA genes [29], nuclear 18S rDNA and β-tubulin genes [36] contained polymorphic loci, suggesting that distinct Helicosporidium isolates might be differentiated at a molecular level. In phylogenetic trees, all Helicosporidium isolates were grouped in a strongly monophyletic clade, but analyses indicated that isolates collected from coleopteran hosts (strain Dm-1 and Cs-1) were more closely related to each other than they were to the dipteran isolate (strain Sj-1) [36]. Combined with the previously noted differences in relation to morphological structures, and infectivity and virulence, these polymorphic characters may serve as a basis to develop comprehensive studies aimed at distinguishing more than one Helicosporidium species. A refined understanding of the Helicosporidium biodiversity may help establishing whether the currently observed host range for these pathogens is reflective of one or more species.
5. The Helicosporidium Host Range Includes a Wide Variety of Insects and Other Invertebrates
Insect infections with Helicosporidium have been reported in three orders: Lepidoptera, Coleoptera and Diptera (Table 1). Overall, 23 species (11 families and 16 genera) of insects are known natural hosts for these pathogenic algae. Coleopteran and dipteran infections are most common. Only two instances of lepidopteran infections have been reported. A hepialid larva was first identified as a Helicosporidium host [3]. Later, Helicosporidium parasiticum was catalogued as a pathogen of the light brown apple moth (Epiphyas postvittana) in New Zealand [44]. In laboratory settings, lepidopteran larvae have been used extensively as heterologous hosts, clearly demonstrating that these insects are susceptible to infections [8,14,15,16,17,20,21,22,23,24].

Table 1.
Reported insect host record for Helicosporidium spp.
| Original insect hosts | Location | Heterologous insect hosts 1 | Reference |
|---|---|---|---|
| Diptera | |||
| Ceratopogonidae | |||
| Dasyhelea obscura | England | nr | [1] |
| Rhiphidae | |||
| Mycetobia pallipes | England | nr | [1] |
| Culicidae | |||
| Culex territans | USA | nr | [45] |
| Culex nigripalpus | USA | Diptera (15) Coleoptera (3) Lepidoptera (2) | [26] |
| Culex quinquefasciatus | Thailand | nr | [24] |
| Culex pipiens | Egypt | Diptera (4) Lepidoptera (1) | [9] |
| Aedes aegypti | Thailand | Diptera (3) Lepidoptera (1) | [24] |
| Sciaridae | |||
| Ctenosciara hyalipennis | Germany | nr | [46] |
| Simuliidae | |||
| Simulium jonesi | USA | Diptera (4) Coleoptera (1) Lepidoptera (4) | [8] |
| Coleoptera | |||
| Nitidulidae | |||
| Carpophilus dimidiatus | USA | nr | [47] |
| Carpophilus freeman | USA | nr | [47] |
| Carpophilus hemipterus | USA | nr | [47] |
| Carpophilus mutilatus | USA, Mexico | Diptera (1) Coleoptera (9) Lepidoptera (5) | [47,48] |
| Carpophilus pilosellus | USA | nr | [47] |
| Conotelus stenoides | USA | nr | [47] |
| Stelidota geminata | USA | nr | [47] |
| Urophorus humeralis | Mexico | nr | [47] |
| Scarabaeidae | |||
| Orycetes monoceros | Tanzania | nr | [49] |
| Curculionidae | |||
| Cyrtobagous salviniae | South Africa | Diptera (3) Coleoptera (1) Lepidoptera (2) | [12] |
| Dendroctonus micans | Turkey | nr | [10] |
| Rhizophaginae | |||
| Rhizophagus grandis | Turkey | nr | [11,13] |
| Lepidoptera | |||
| Hepialidae | |||
| Hepialis pallens | Argentina | nr | [3] |
| Tortricidae | |||
| Epiphyas postvittana | New Zealand | nr | [44] |
1 “nr” denotes that no heterologous hosts have been reported. When available, the numbers of heterologous host species is indicated in parenthesis.
Similar to the two infections reported in Lepidoptera, most coleopteran and dipteran accounts have corresponded to episodic observations (Table 1). However, notable exceptions exist. In Diptera, Helicosporidium infections have been recurrently reported in mosquitoes, especially in the genus Culex, which has been found as a host on four different occasions on various continents (North America, Asia, Africa), suggesting that Helicosporidium may be ubiquitous and chronic in mosquito populations [9,24,26,45]. Bioassays demonstrated that mosquito Helicosporidium isolates were infectious to additional mosquito species [9,26]. A large survey of nitidulid beetles pathogens suggested that Helicosporidium might be widespread in Coleoptera, as infections were reported in over 50% of the examined species [47]. A tentative identification in the rhinoceros beetle [49], combined with the recent detection of Helicosporidium in two members of the Curculionidae family [10,12] supports the possibility that coleopterans serve as major hosts for these pathogens. The first study aimed at investigating the abundance and importance of Helicosporidium on natural insect populations was recently performed on the coleopteran host Dendroctonus micans, and results indicated that the pathogen is widely distributed and can be detected over a period of three years in different geographic locales, with estimated infection rates reaching 71% [50]. A similar study focused on the occurrence of Helicosporidium in a second coleopteran host, the predator beetle Rhizophagus grandis (a predator of D. micans), although it was limited to laboratory-reared insect [51]. These two studies suggest that Helicosporidium investigations may move from isolated and occasional reports of infection in a few insects to a broader understanding of the abundance and impact of these organisms in insect populations.
In addition to the observation that Helicosporidium can be detected in Lepidoptera, Coleoptera and Diptera, one of the most striking and recurrent characteristics of these insect pathogens is arguably their ability to be horizontally transferred irrespectively of the host order from which they have been isolated. Isolates detected in dipteran hosts can readily infect coleopterans or lepidopterans [8,9,16,17,20,21,22,23]. Isolates collected in Coleoptera have showed similar broad host range, and were used to infect Lepidoptera or Diptera taxa [12,14,15,22,23,48]. The most comprehensive study of Helicosporidium host range involved an isolate collected from the nitidulid beetle Carpophilus mutilatus that was transmitted to nine other coleopteran species, five lepidopteran species and one dipteran species (Table 1) [48]. This isolate also failed to infect orthopteran and hymenopteran insects [48], providing the basis for, and supporting, the current understanding that the Helicosporidium host range is restricted to three insect orders: Coleoptera, Lepidoptera and Diptera, as indicated in Table 1. This study was also instrumental is demonstrating that the Helicosporidium host range includes non-insect invertebrates, as the coleopteran isolate was shown to be infectious to three species of mites. Mites and collembolans have been reported as potential natural hosts for Helicosporidium, as early as during the first description of Helicosporidium parasiticum [1,5,6,17,46]. In addition, Helicosporidium spp. were detected in trematodes [7] and cladocerans [4]. Overall, pathosystems involving non-insect hosts remain largely uncharacterized.
Finally, in agreement with the observation that the cuticle of heavily infected insect hosts may rupture and release Helicosporidium cysts and vegetative cells in the environment [1,52], several reports have indicated that Helicosporidium can be isolated [17,25,53], or at least detected [37], from lentic water samples such as ponds or ditches. Pathogenicity of these isolates was evaluated in the corn earworm Heliothis zea, or mosquitoes [17,25,53], suggesting that pathogenic isolates can be uncovered independently of host infections. When tested on mosquitoes, a Helicosporidium isolate collected from field water samples was compared to the Thailand mosquito isolate (from Aedes aegyptii), and was shown to be not as effective or as lethal [17]. This result prompted the proposal for distinct species of Helicosporidium, and was reminiscent of other comparative studies that indicated that pathogenicity to Culex pipiens quinquefasciatus was variable [26]. Although a Helicosporidium isolate from a coleopteran host was reportedly infectious to Cx. p. quinquefasciatus [48], this mosquito species appeared non-susceptible to an isolate collected from Cx. nigripalpus [26]. Although rare, the noted differences in host susceptibility and Helicosporidium virulence suggest that the list of natural hosts presented in Table 1 may be reflective of the combined range of several isolates that have yet to be formally distinguished and taxonomically separated.
As noted previously [37], the ability to consistently detect Helicosporidium in selected environments [17,25,37,53] or insect populations [50,51] will likely further the current understanding on these organisms by not only providing robust ecological data to estimate their impact on host populations but also expanding the current knowledge of Helicosporidium biodiversity and the identification of polymorphic characters that may be informative for the distinction of several Helicosporidium species.
6. Conclusion
The protists Helicosporidium spp. represent the first described entomopathogenic non‑photosynthetic algae. They are characterized by unique and diagnostic cell phenotypes that primarily include the infective cyst, a four-cell structure that is ingested by susceptible hosts and releases a filamentous cell that mediates host ingress and infection. Susceptible hosts include insects belonging to the Diptera, Coleoptera and Lepidoptera orders, as well as non-insect invertebrates such as mites and collembolans. Techniques associated with maintaining Helicosporidium spp. in axenic cultures in vitro and evaluating pathogenicity in insect bioassays have been well established. Despite the recent advances in understanding Helicosporidium biology, several challenges remain. One of these challenges involves characterizing the molecular basis of Helicosporidium-host interactions, and identifying the host factors potentially responsible for both cyst dehiscence in the gut, and cyst morphogenesis in the host hemocoel. Comprehensive investigations on the molecular composition of Helicosporidium cell phenotypes may shed light on the role of the filamentous cell barbs during ingress, or the ability of the vegetative cell wall to evade host detection systems. The filamentous cell is the most diagnostic feature of Helicosporidium, but also represents an extremely unique structure in the green algae phylum. Genomic analyses may be aimed at identifying the genes associated with the development of these cells and determining if these genes share a green algal origin. A second challenge includes developing a more global estimation of the occurrence and abundance of Helicosporidium in insect populations. Increasing the number of isolates available for comparative analyses will undoubtedly lead to an improved understanding of Helicosporidium diversity, refined trebouxiophyte phylogenies, and as noted throughput this review, may lead to the distinction of multiple Helicosporidium species.
Acknowledgments
The author thanks Robert Smith for critical comments on earlier drafts of the manuscript, and Drion Boucias, Verena-Ulrike Lietze and Michael Botts for providing micrographs.
References
- Keilin, D. On the like-history of Helicosporidium parasiticum, n.g., n.sp., a new type of protist parasitic in the larva of Dasyhelea obscura Winn. (Diptera, Ceratopogonidae) and in some other arthropods. Parasitology 1921, 13, 97–113. [Google Scholar]
- Kudo, R.R. Handbook of Protozoology; Thomas: Springfield, IL, USA, 1931; pp. 643–679. [Google Scholar]
- Weiser, J. Helicosporidium parasiticum Keilin infection in the caterpillar of a Hepialid moth in Argentina. J. Protozool. 1970, 17, 436–440. [Google Scholar]
- Sayre, R.M.; Clark, T.B. Daphnia magna (Cladocera: Chydoroidea) A new host of a Helicosporidium sp. (Protozoa: Helicosporidia). J. Invertebr. Pathol. 1978, 31, 260–261. [Google Scholar]
- Purrini, K. Studies on some Amoebae (Amoebida) and Helicosporidium parasiticum (Helicosporida) infecting moss-mites (Oribatei, Acarina) in forest soil samples. Arch. Protistenkd 1981, 124, 303–311. [Google Scholar] [CrossRef]
- Purrini, K. Light and electron microscope studies on Helicosporidium sp. parasitizing Oribatid mites (Oribatei, Acarina) and Collembola (Apterygota, Insecta) in forest soils. J. Invertebr. Pathol. 1984, 44, 18–27. [Google Scholar] [CrossRef]
- Pekkarinen, M. Bucephalid trematode sporocysts in brackish-water Mytilus edulis, new host of a Helicosporidium sp. (Protozoa: Helicosporida). J. Invertebr. Pathol. 1993, 61, 214–216. [Google Scholar]
- Boucias, D.G.; Becnel, J.J.; White, S.E.; Bott, M. In vivo and in vitro development of the protist Helicosporidium sp. J. Eukaryot. Microbiol. 2001, 48, 460–470. [Google Scholar]
- Seif, A.I.; Rifaat, M.M.A. Laboratory evaluation of a Helicosporidium sp. (Protozoa: Helicosporida) as an agent for the microbial control of mosquitoes. J. Egypt. Soc. Parasitol. 2001, 31, 21–35. [Google Scholar]
- Yaman, M.; Radek, R. Helicosporidium infection of the great European spruce bark beetle, Dendroctonus micans (Coleoptera: Scolytidae). Eur. J. Protistol. 2005, 41, 203–207. [Google Scholar]
- Yaman, M.; Radek, R. Infection of the predator beetle Rhizophagus grandis Gyll. (Coleoptera, Rhizophagidae) with the insect pathogenic alga Helicosporidium sp. (Chlorophyta: Trebouxiophyceae). Biol. Contr. 2007, 41, 384–388. [Google Scholar]
- White, S.E.; Tipping, P.W.; Becnel, J.J. First isolation of a Helicosporidium sp. (Chlorophyta: Trebouxiophyceae) from the biological control agent Cyrtobagous salviniae (Coleoptera: Curculionidae). Biol. Contr. 2007, 40, 243–245. [Google Scholar]
- Yaman, M.; Radek, R.; Aydin, C.; Tosun, O.; Ertuk, O. First record of the insect pathogenic alga Helicosporidium sp. (Chlorophyta: Trebouxiophyceae) infection in larvae and pupae of Rhizophagus grandis Gyll. (Coleoptera, Rhizophaginae) from Turkey. J. Invertebr. Pathol. 2009, 102, 182–184. [Google Scholar]
- Kellen, W.R.; Lindegren, J.E. Life cycle of Helicosporidium parasiticum in the navel orangeworm, Paramyelois transitella. J. Invertebr. Pathol. 1974, 23, 202–208. [Google Scholar] [CrossRef]
- Lindegren, J.E.; Hoffmann, D.F. Ultrastructure of some developmental stages of Helicosporidium sp. in the navel orangeworm, Paramyelois transitella. J. Invertebr. Pathol. 1976, 27, 105–113. [Google Scholar]
- Bläske-Lietze, V.-U.; Shapiro, A.M.; Denton, J.S.; Botts, M.; Becnel, B.; Boucias, D.G. Development of the insect pathogenic alga Helicosporidium. J. Eukaryot. Microbiol. 2006, 53, 165–176. [Google Scholar]
- Avery, S.W.; Undeen, A.H. Some characteristics of a new isolate of Helicosporidium and its effect upon mosquitoes. J. Invertebr. Pathol. 1987, 49, 246–251. [Google Scholar] [CrossRef]
- Hamana, K.; Aizaki, T.; Arai, E.; Saito, A.; Uchikata, K.; Ohnishi, H. Distribution of norspermidine as a cellular polyamine within micro green algae including non-photosynthetic achlorophyllous Polytoma, Polytomella, Prototheca and Helicosporidium. J. Gen. Appl. Microbiol. 2004, 50, 289–295. [Google Scholar]
- De Koning, A.P.; Tartar, A.; Boucias, D.G.; Keeling, P.J. Expressed Sequence Tag (EST) survey of the highly adapted green algal parasite, Helicosporidium. Protist 2005, 156, 181–190. [Google Scholar] [CrossRef]
- Bläske-Lietze, V.-U.; Boucias, D.G. Pathogenesis of Helicosporidium sp. (Chlorophyta: Trebouxiophyceae) in susceptible noctuid larvae. J. Invertebr. Pathol. 2005, 90, 161–168. [Google Scholar]
- Denton, J.S.S.; Lietze, V.-U.; Boucias, D.G. Host age and pathogen dosage impact cyst morphogenesis in the invertebrate pathogenic alga Helicosporidium sp. (Chlorophyta: Trebouxiophyceae). J. Invertebr. Pathol. 2009, 102, 36–39. [Google Scholar]
- Bläske, V.-U.; Boucias, D.G. Influence of Helicosporidium spp. (Chlorophyta: Trebouxiophyceae) infection on the development and survival of three noctuid species. Environ. Entomol. 2004, 33, 54–61. [Google Scholar]
- Conklin, T.; Bläske-Lietze, V.-U.; Becnel, J.J.; Boucias, D.G. Infectivity of two isolates of Helicosporidium spp. (Chlorophyta; Trebouxiophyceae) in heterologous host insects. Fla. Entomol. 2005, 88, 431–440. [Google Scholar]
- Hembree, S.C. Preliminary report of some mosquito pathogens from Thailand. Mosquito News 1979, 39, 575–582. [Google Scholar]
- Kim, S.S.; Avery, S.W. Effects of Helicosporidium sp. infection on larval mortality, adult longevity, and fecundity of Culex Salinarius Coq. Kor. J. Entomol. 1986, 16, 153–156. [Google Scholar]
- Fukuda, T.; Lindegren, J.E.; Chapman, H.C. Helicosporidium sp. a new parasite of mosquitoes. Mosquito News 1976, 36, 514–517. [Google Scholar]
- Hembree, S.C. Evaluation of the microbial control potential of a Helicosporidium sp (Protozoa: Helicosporida) from Aedes aegypti and Culex quinquefasciatus from Thailand. Mosquito News 1981, 41, 770–783. [Google Scholar]
- Tartar, A.; Boucias, D.G.; Adams, B.J.; Becnel, J.J. Phylogenetic analysis identifies the invertebrate pathogen Helicosporidium sp. as a green alga (Chlorophyta). Int. J. Syst. Evol. Microbiol. 2002, 52, 273–279. [Google Scholar]
- Tartar, A.; Boucias, D.G.; Becnel, J.J.; Adams, B.J. Comparison of plastid 16S rDNA (rrn16) genes from Helicosporidium spp.: Evidence supporting the reclassification of Helicosporidia as green algae (Chlorophyta). Int. J. Syst. Evol. Microbiol. 2003, 53, 1719–1723. [Google Scholar]
- Tartar, A.; Boucias, D.G. The non-photosynthetic, pathogenic green alga Helicosporidium sp. has retained a modified, functional plastid genome. FEMS Microbiol. Lett. 2004, 233, 153–157. [Google Scholar]
- De Koning, A.P.; Keeling, P.J. Nucleus-encoded genes for plastid-targeted proteins in Helicosporidium: functional diversity of a cryptic plastid in a parasitic alga. Eukaryot. Cell 2004, 3, 1198–1205. [Google Scholar] [CrossRef]
- De Koning, A.P.; Keeling, P.J. The complete plastid genome sequence of the parasitic green alga Helicosporidium sp. is highly reduced and structured. BMC Biol. 2006, 4, 12. [Google Scholar] [CrossRef]
- Pombert, J.-F.; Keeling, P.J. The mitochondrial genome of the entomoparasitic green alga Helicosporidium. PLoS One 2010, 5, e8954. [Google Scholar] [CrossRef]
- Rodríguez, F.; Feist, S.W.; Guillou, L.; Harkestad, L.S.; Bateman, K.; Renault, T.; Mortensen, S. Phylogenetic and morphological characterization of the green algae infesting blue mussel Mytilus edulis in the North and South Atlantic oceans. Dis. Aquat. Org. 2008, 81, 231–240. [Google Scholar] [CrossRef]
- Crespo, C.; Rodriguez, H.; Segade, P.; Iglesias, R.; Garcia-Estevez, J.M. Coccomyxa sp. (Chlorophyta: Chlorococcales), a new pathogen in mussels (Mytilus galloprovincialis) of Vigo estuary (Galicia, NW Spain). J. Invertebr. Pathol. 2009, 102, 214–219. [Google Scholar]
- Vázquez, N.; Rodríguez, F.; Ituarte, C.; Klaich, J.; Cremonte, F. Host-parasite relationship of the geoduck Panopea abbreviata and the green alga Coccomyxa parasitica in the Argentinean Patagonian coast. J. Invertebr. Pathol. 2010, 105, 254–260. [Google Scholar] [CrossRef]
- Mancera, N.; Douma, L.G.; James, S.; Liu, S.; Van, A.; Boucias, D.G.; Tartar, A. Detection of Helicosporidium spp. in metagenomic DNA. J. Invertebr. Pathol. 2012, 111, 13–19. [Google Scholar] [CrossRef]
- Von Bergen, M.; Eidner, A.; Schmidt, F.; Murugaiyan, J.; Wirth, H.; Binder, H.; Maier, T.; Roesler, U. Identification of harmless and pathogenic algae of the genus Prototheca by MALDI-MS. Proteonomics Clin. Appl. 2009, 3, 774–784. [Google Scholar] [CrossRef]
- Ueno, R.; Hanagata, N.; Urano, N.; Suzuki, M. Molecular phylogeny and phenotypic variation in the heterotrophic green algal genus Prototheca (Trebouxiophyecae, Chlorophyta). J. Phycol. 2005, 41, 1268–1280. [Google Scholar] [CrossRef]
- Roesler, U.; Möller, A.; Hansel, A.; Baumann, D.; Truyen, U. Diversity within the current algal species Prototheca zopfii: A proposal for two Prototheca zopfii genotypes and description of a novel species Prototheca blaschkeae sp. nov. Int. J. Syst. Evol. Microbiol. 2006, 56, 1419–1425. [Google Scholar] [CrossRef]
- Leliaert, F.; Smith, D.R.; Moreau, H.; Herron, M.; Verbruggen, H.; Delwiche, C.F.; DeClerck, O. Phylogeny and molecular evolution of the green algae. Crit. Rev. Plant Sci. 2012, 31, 1–46. [Google Scholar] [CrossRef]
- Satoh, K.; Ooe, K.; Nagayama, H.; Makimura, K. Prototheca cutis sp. nov., a newly discovered pathogen of protothecosis isolated from inflamed human skin. Int. J. Syst. Evol. Microbiol. 2010, 60, 1236–1240. [Google Scholar]
- Marques, S.; Silva, E.; Kraft, C.; Carvalheira, J.; Videira, A.; Huss, V.A.; Thompson, G. Bovine mastitis associated with Prototheca blaschkeae. J. Clin. Microbiol. 2008, 46, 1941–1945. [Google Scholar] [CrossRef]
- Glare, T.; O’Callaghan, M.; Wigley, P.J. Checklist of naturally occurring entomopathogenic microbes and nematodes in New Zealand. New Zeal. J. Zool. 1993, 20, 95–120. [Google Scholar]
- Chapman, H.C.; Woodward, D.B.; Petersen, J.J. Pathogens and parasites in Louisiana Culicidae and Chaoboridae. Proc. N. J. Mosq. Exterm. Assoc. 1967, 54, 54–60. [Google Scholar]
- Purrini, K. On the incidence and distribution of parasites of soil fauna of mixed coniferous forests, mixed leaf forests, and pure beech forests in Lower Saxony, West Germany. In Soil Biology as Related to Land Use Practices, Proceedings of the VII International Colloquium of Soil Zoology, Syracuse, NY, USA, 29 July–3 August 1979; Dindal, D.L., Ed.; 1980; pp. 561–582. [Google Scholar]
- Lindegren, J.E.; Okumura, G.T. Pathogens from economically important nitidulid beetles. USDA ARS 1973, W-9, 1–7. [Google Scholar]
- Kellen, W.R.; Lindegren, J.E. New host records for Helicosporidium parasiticum. J. Invertebr. Pathol. 1973, 22, 296–297. [Google Scholar] [CrossRef]
- Purrini, K. On disease agents of insect pests of wild palms and forests in Tanzania. J. Appl. Entomol. 1985, 99, 237–240. [Google Scholar]
- Yaman, M. First results on distribution and occurrence of the insect pathogenic alga Helicosporidium sp. (Chlorophyta: Trebouxiophyceae) in the populations of the great spruce bark beetle, Dendroctonus micans (Kugelann) (Coleoptera: Curculionidae, Scolytinae). N. West. J. Zool 2008, 4, 99–107. [Google Scholar]
- Yaman, M.; Tosun, O.; Aydin, C.; Erturk, O. Distribution and occurrence of the insect pathogenic alga Helicosporidium sp. (Chlorophyta: Trebouxiophyceae) in the predator beetle Rhizophagus grandis Gyll. (Coleoptera: Rhizophagidae)-rearing laboratories. Folia Microbiol. 2011, 56, 44–48. [Google Scholar]
- Fukuda, T. Helicosporidium (Protozoa). In Biological Control of Mosquitoes; Chapman, H.C., Ed.; Bulletin #6 of the American Control of Mosquito Association; American Control of Mosquito Association: Mount Laurel, NJ, USA, 1985; pp. 59–61. [Google Scholar]
- Avery, S.W.; Undeen, A.H. The isolation of microsporidia and other pathogens from concentrated ditch water. J. Am. Mosq. Contr. Assoc. 1987, 3, 54–58. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).