Insights into the Maternal Ancestry of Côte d’Ivoire Honeybees Using the Intergenic Region COI-COII
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area
2.2. Collection of Honeybees
2.3. DNA Extraction, Sequencing, and DraI Test
2.4. Data Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ruttner, F. Biogeography and Taxonomy of Honeybees; Springer: New York, NY, USA, 1988. [Google Scholar]
- Chávez-Galarza, J.; Garnery, L.; Henriques, D.; Neves, C.J.; Loucif-Ayad, W.; Jonhston, J.S.; Pinto, M.A. Mitochondrial DNA variation of Apis mellifera iberiensis: Further insights from a large-scale study using sequence data of the tRNAleu-cox2 intergenic region. Apidologie 2017, 48, 533–544. [Google Scholar] [CrossRef]
- Franck, P.; Garnery, L.; Loiseau, A.; Oldroyd, B.P.; Hepburn, H.R.; Solignac, M.; Cornuet, J.M. Genetic diversity of the honeybee in Africa: Microsatellite and mitochondrial data. Heredity 2001, 86, 420–430. [Google Scholar] [CrossRef]
- Garnery, L.; Cornuet, J.M.; Solignac, M. Evolutionary history of the honey bee Apis mellifera inferred from mitochondrial DNA analysis. Mol. Ecol. 1992, 1, 145–154. [Google Scholar] [CrossRef]
- Clarke, K.E.; Rinderer, T.E.; Franck, P.; Quezada-Euán, J.J.G.; Oldroyd, B.P. The Africanization of honey bees (Apis mellifera L.) of the Yucatan: A study of a massive hybridization event across time. Evolution 2002, 56, 1462–1474. [Google Scholar]
- Jensen, A.B.; Palmer, K.A.; Boomsma, J.J.; Pedersen, B.V. Varying degrees of Apis mellifera ligustica introgression in protected populations of the black honey bee, Apis mellifera mellifera, in northwest Europe. Mol. Ecol. 2005, 14, 93–106. [Google Scholar] [CrossRef]
- Franck, P.; Garnery, L.; Solignac, M.; Cornuet, J.M. Molecular confirrmation of a fourth lineage in honeybees from Middle-East. Apidologie 2000, 31, 167–180. [Google Scholar] [CrossRef]
- Cánovas, F.; De La Rúa, P.; Serrano, J.; Galián, J. Geographical patterns of mitochondrial DNA variation in Apis mellifera iberiensis (Hymenoptera: Apidae). J. Zool. Syst. Evol. Res. 2008, 46, 24–30. [Google Scholar] [CrossRef]
- De La Rúa, P.; Galián, J.; Pedersen, B.V.; Serrano, J. Molecular characterization and population structure of Apis mellifera from Madeira and the Azores. Apidologie 2006, 38, 699–708. [Google Scholar] [CrossRef]
- Palmer, M.R.; Smith, D.R.; Kaftanoglu, O. Turkish honeybees: Genetic variation and evidence for a fourth lineage of Apis mellifera mt DNA. J. Hered. 2000, 91, 42–46. [Google Scholar] [CrossRef]
- Whitfield, C.W.; Behura, S.K.; Stewart, I.; Berlocher, H.; Clark, A.G.; Johnston, J.S.; Sheppard, W.S.; Smith, D.R.; Suarez, A.V.; Weaver, D.; et al. Thrice out of Africa: Ancient and recent expansions of the honey bee, Apis mellifera. Science 2006, 314, 642–645. [Google Scholar] [CrossRef]
- Dietemann, V.; Pirk, C.; Crewe, R. Is there a need for conservation of honeybees in Africa? Apidologie 2009, 40, 285–295. [Google Scholar] [CrossRef]
- Radloff, S.E.; Hepburn, H.R.; Fuchs, S. Ecological and morphological differentiation of the Apis mellifera linnaeus honeybees (Hymenoptera: Apidae) of West Africa. Afr. Entomol. 1998, 6, 17–23. [Google Scholar]
- Cockerell, T.D.A. New rocky mountain bees, and other notes. Can. Entomol. 1906, 38, 160–166. [Google Scholar] [CrossRef]
- Ruttner, F. Les races d’abeille de l’Afrique. In Proceedings of the XXV Congrès International d’Apiculture, Grenoble, France, 4 September 1976; pp. 347–367. [Google Scholar]
- Maa, T.C. An inquiry into the systematics of the tribus Apidini or honeybees (Hymenoptera). Treubia 1953, 21, 525–640. [Google Scholar]
- Baldensperger, P.J. Variétés d’abeilles en Afrique du nord. In Proceedings of the 5th Congrès International d’Entomologie, Paris, France, 18 July 1932. [Google Scholar]
- Latreille, P.A. Notice des espèces d’abeilles vivant en grande société, ou abeilles proprement dites, et description d’espèces nouvelles. Ann. Mus. Natl. Hist. Nat. 1804, 5, 161–178. [Google Scholar]
- Lepeletier, A. Histoire Naturelle des Insectes. Suites à Buffon; Hymenopte Áres: Roret, Paris, 1836. [Google Scholar]
- Eschscholtz, J.F. Entomographien; Reimer: Berlin, Germany, 1822. [Google Scholar]
- Smith, F.G. The races of honeybees in Africa. Bee World 1961, 42, 255–260. [Google Scholar] [CrossRef]
- Meixner, M.D.; Leta, M.A.; Koeniger, N.; Fuchs, S. The honey bees of Ethiopia represent a new subspecies of Apis mellifera—Apis mellifera simensis n. ssp. Apidologie 2011, 42, 425–437. [Google Scholar] [CrossRef]
- Cornuet, J.M.; Garnery, L. Mitochondrial DNA variability in honey bees and its phylogeographic implications. Apidologie 1991, 22, 627–642. [Google Scholar] [CrossRef]
- Nikolenko, A.G.; Poskryakov, A.V. Polymorphism of locus COI-COII of mitochondrial DNA in the honeybee Apis mellifera L. from the southern Ural Region. Russ. J. Genet. 2002, 38, 364–368. [Google Scholar] [CrossRef]
- Tan, K.; Warrit, N.; Smith, D.R. Mitochondrial DNA diversity of the Chinese Apis cerana. Apidologie 2007, 38, 238–246. [Google Scholar] [CrossRef]
- Achou, M.; Loucif-Ayad, W.; Legout, H.; Hmidan, H.; Alburaki, M.; Garnery, L. An insightful molecular analysis reveals foreign honeybees among Algerian honeybee populations (Apis mellifera L.). J. Data Min. Genomics Proteomics 2015, 6, 166. [Google Scholar] [CrossRef]
- Arias, M.C.; Sheppard, W.S. Molecular phylogenetics of honey bee subspecies (Apis mellifera L.) inferred from mitochondrial DNA sequence. Mol. Phylogenet. Evol. 1996, 5, 557–566. [Google Scholar] [CrossRef]
- De la Rúa, P.J.; Serrano, J.; Galián, J. Mitochondrial DNA variability in the Canary Islands honeybees (Apis mellifera L.). Mol. Ecol. 1998, 7, 1543–1547. [Google Scholar] [CrossRef] [PubMed]
- Alburaki, M.; Moulin, S.; Legout, H.; Alburaki, A.; Garnery, L. Mitochondrial structure of Eastern honeybee populations from Syria, Lebanon and Iraq. Apidologie 2011, 42, 628. [Google Scholar] [CrossRef]
- Alburaki, M.; Bertrand, B.; Legout, H.; Moulin, S.; Alburaki, A.; Sheppard, W.S.; Garnery, L. A fifth major genetic group among honeybees revealed in Syria. BMC Genet. 2013, 14, 117. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D. Molecular cloning: A laboratory manual (3-volume set). Immunology 2001, 49, 895–909. [Google Scholar]
- Kumar, S.; Nei, M.; Dudley, J.; Tamura, K. MEGA: A biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 2008, 9, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.M.; Proter, J.B.; Martin, D.M.A. Jalview version 2-a multiple sequence alignement editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef]
- Szalanski, A.L.; Magnus, M.R. Mitochondrial DNA characterization of Africanized honey bee (Apis mellifera L.) populations from the USA. J. Apic. Res. 2010, 49, 177–185. [Google Scholar] [CrossRef]
- El-Niweiri, M.A.; Moritz, R.F. Mitochondrial discrimination of honeybees (Apis mellifera) of Sudan. Apidologie 2008, 39, 566–573. [Google Scholar] [CrossRef]
- Fletcher, D.J. The African bee, Apis mellifera adansonii, in Africa. Annu. Rev. Entomol. 1978, 23, 151–171. [Google Scholar] [CrossRef]
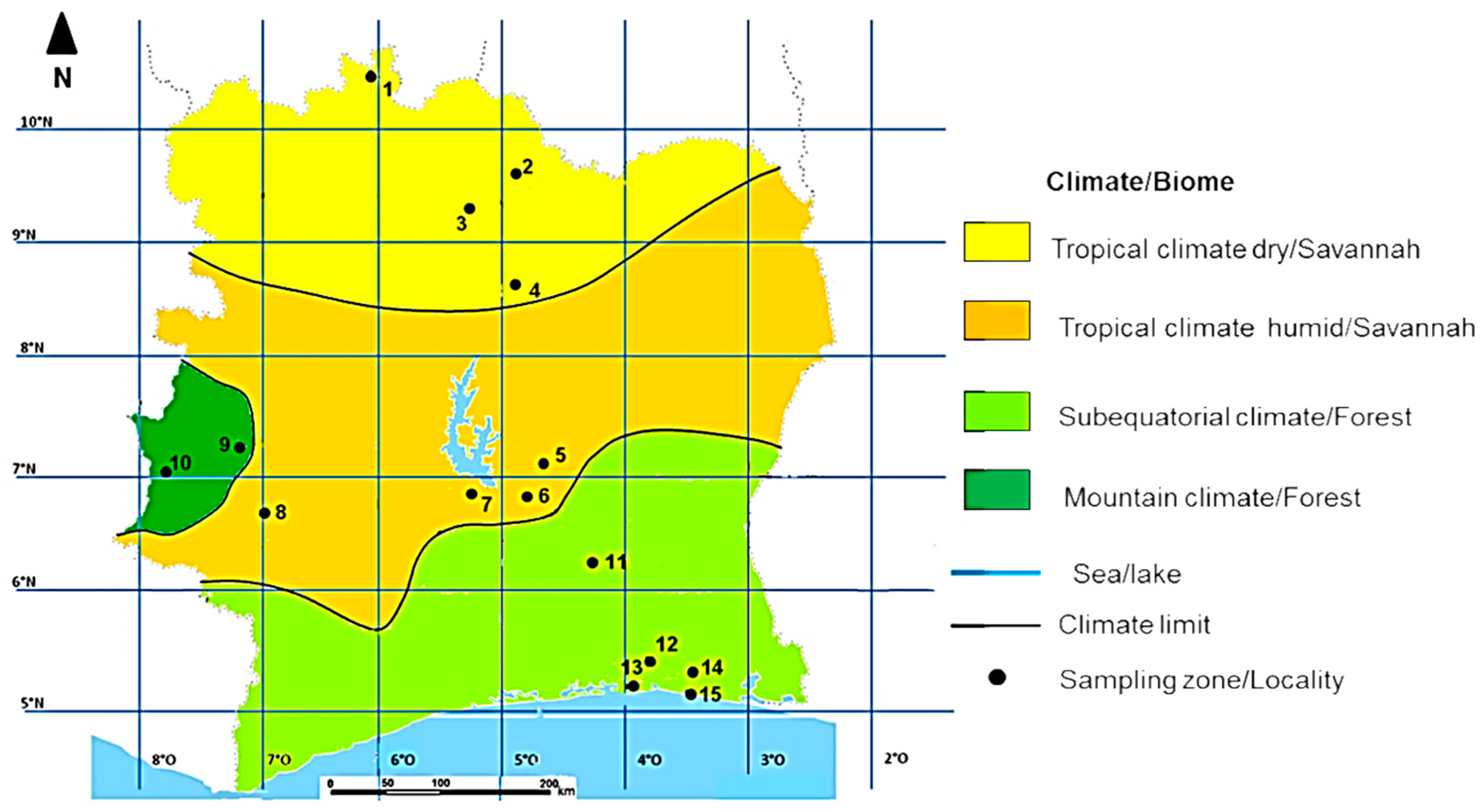
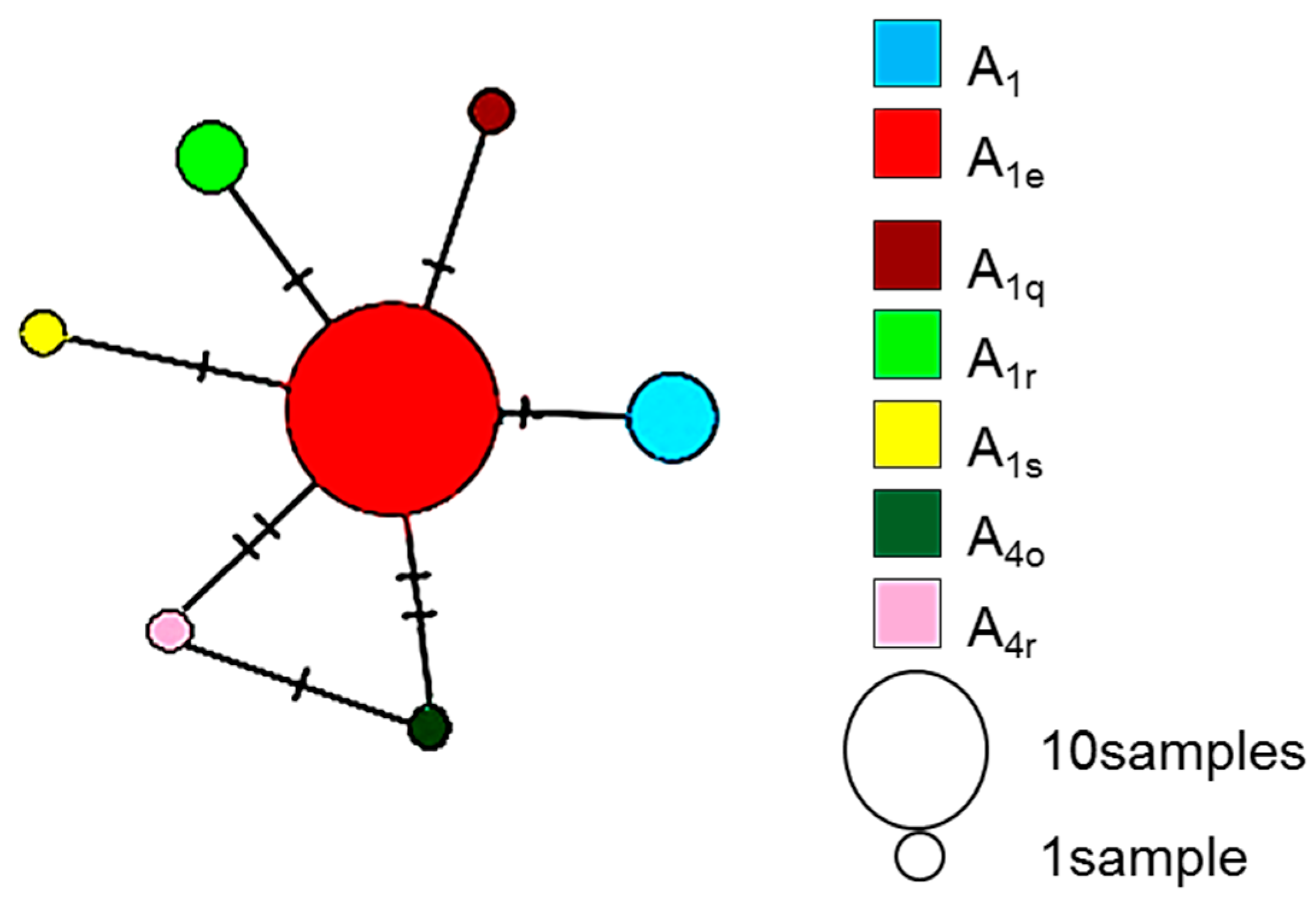

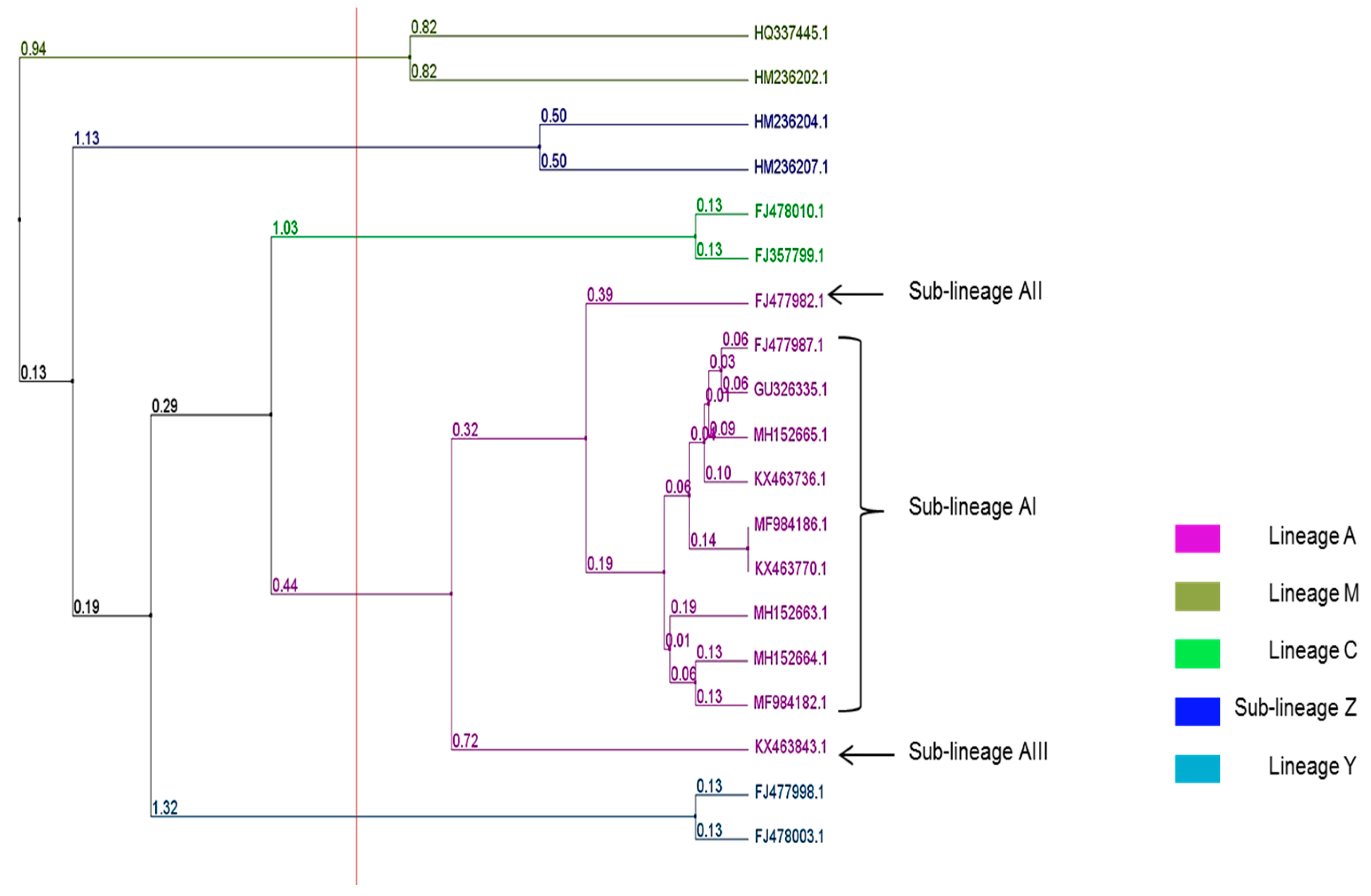
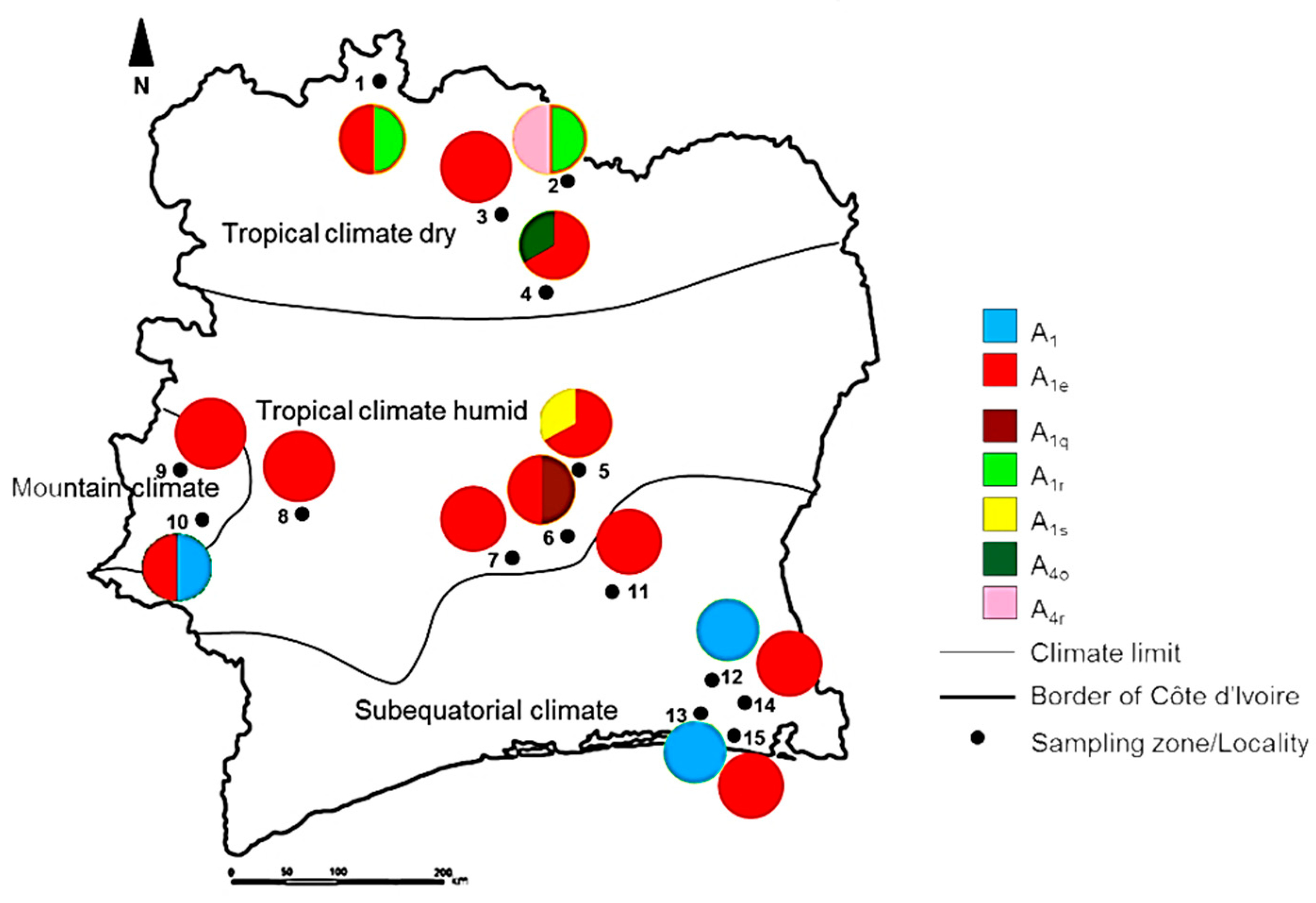
| Climate Type | Total Number of Colonies (N) | Sampling Localities (Map Code) | Number of Colonies per Sampling Location | Latitude | Longitude | Altitude | Mean Annual Temperature (°C) | Mean Relative Humidity (%) | Mean Annual Precipitation (mm) | Biome Type |
|---|---|---|---|---|---|---|---|---|---|---|
| Tropical dry climate | 10 | Tengrela (1) | 2 | 10°2′ N | 6°42′ W | 382 | 27.6 | 60.7 | 841 | Savannah |
| Ferkéssedougou (2) | 2 | 9°6′ N | 5°19′ W | 370 | 27 | 68.0 | 951 | |||
| Korhogo (3) | 3 | 9°36′ N | 5°63′ W | 342 | 27 | 65.4 | 951 | |||
| Tafiére (4) | 3 | 8°4′ N | 5°17′ W | 385 | 26.6 | 65.0 | 1100 | |||
| Tropical humid climate | 10 | Didievi (5) | 3 | 7°4′ N | 4°3′ W | 116 | 27.2 | 74.1 | 1090.6 | |
| Yamoussoukrou (6) | 4 | 6°54′ N | 5°21′ W | 213 | 26 | 76.5 | 1118 | |||
| Bouaflé (7) | 2 | 6°59′ N | 5°45′ W | 205 | 26.6 | 75.8 | 1242 | |||
| Duekoué (8) | 1 | 6°45′ N | 7°21′ W | 234 | 25.4 | 78.2 | 1572 | |||
| Mountain climate | 3 | Man (9) | 1 | 7°23′ N | 7°31′ W | 339 | 25 | 78 | 1930 | Forest |
| Danané (10) | 2 | 7°16′ N | 8°09′ W | 354 | 24.9 | 77.2 | 1930 | |||
| Subequatorial climate | 10 | Dimbokro (11) | 4 | 6°39′ N | 4°42′ W | 92 | 27 | 75 | 1176 | |
| Alepé (12) | 1 | 5°3′ N | 3°36′ W | 89 | 26.5 | 82.7 | 1544 | |||
| Dabou (13) | 2 | 5°2′ N | 4°23′ W | 20 | 26.4 | 83.5 | 1743 | |||
| Abobo (14) | 1 | 5°25′ N | 4°01′ W | 108 | 26.5 | 82.7 | 1544 | |||
| Bingerville (15) | 2 | 5°21′ N | 4°53′ W | 59 | 26.5 | 90.4 | 1230 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Coulibaly, K.A.S.; Majeed, M.Z.; Chen, C.; YEO, K.; Shi, W.; Ma, C.-S. Insights into the Maternal Ancestry of Côte d’Ivoire Honeybees Using the Intergenic Region COI-COII. Insects 2019, 10, 90. https://doi.org/10.3390/insects10040090
Coulibaly KAS, Majeed MZ, Chen C, YEO K, Shi W, Ma C-S. Insights into the Maternal Ancestry of Côte d’Ivoire Honeybees Using the Intergenic Region COI-COII. Insects. 2019; 10(4):90. https://doi.org/10.3390/insects10040090
Chicago/Turabian StyleCoulibaly, Krouholé Abdoul Salam, Muhammad Zeeshan Majeed, Chao Chen, Kolo YEO, Wei Shi, and Chun-Sen Ma. 2019. "Insights into the Maternal Ancestry of Côte d’Ivoire Honeybees Using the Intergenic Region COI-COII" Insects 10, no. 4: 90. https://doi.org/10.3390/insects10040090
APA StyleCoulibaly, K. A. S., Majeed, M. Z., Chen, C., YEO, K., Shi, W., & Ma, C.-S. (2019). Insights into the Maternal Ancestry of Côte d’Ivoire Honeybees Using the Intergenic Region COI-COII. Insects, 10(4), 90. https://doi.org/10.3390/insects10040090





