Automated Lung Cancer Segmentation in Tissue Micro Array Analysis Histopathological Images Using a Prototype of Computer-Assisted Diagnosis
Abstract
1. Introduction
- The study aimed to develop a computer-aided diagnostic system to detect lung cancer early by segmenting tumor and non-tumor tissue on Tissue Micro Array Analysis (TMA) histopathological images.
- Histopathological medical imaging is the “gold standard” in the early detection of most diseases, including lung cancer.
- The study compares three image enhancement techniques with histogram equalization showing significant improvement in final segmentation of the CAD system.
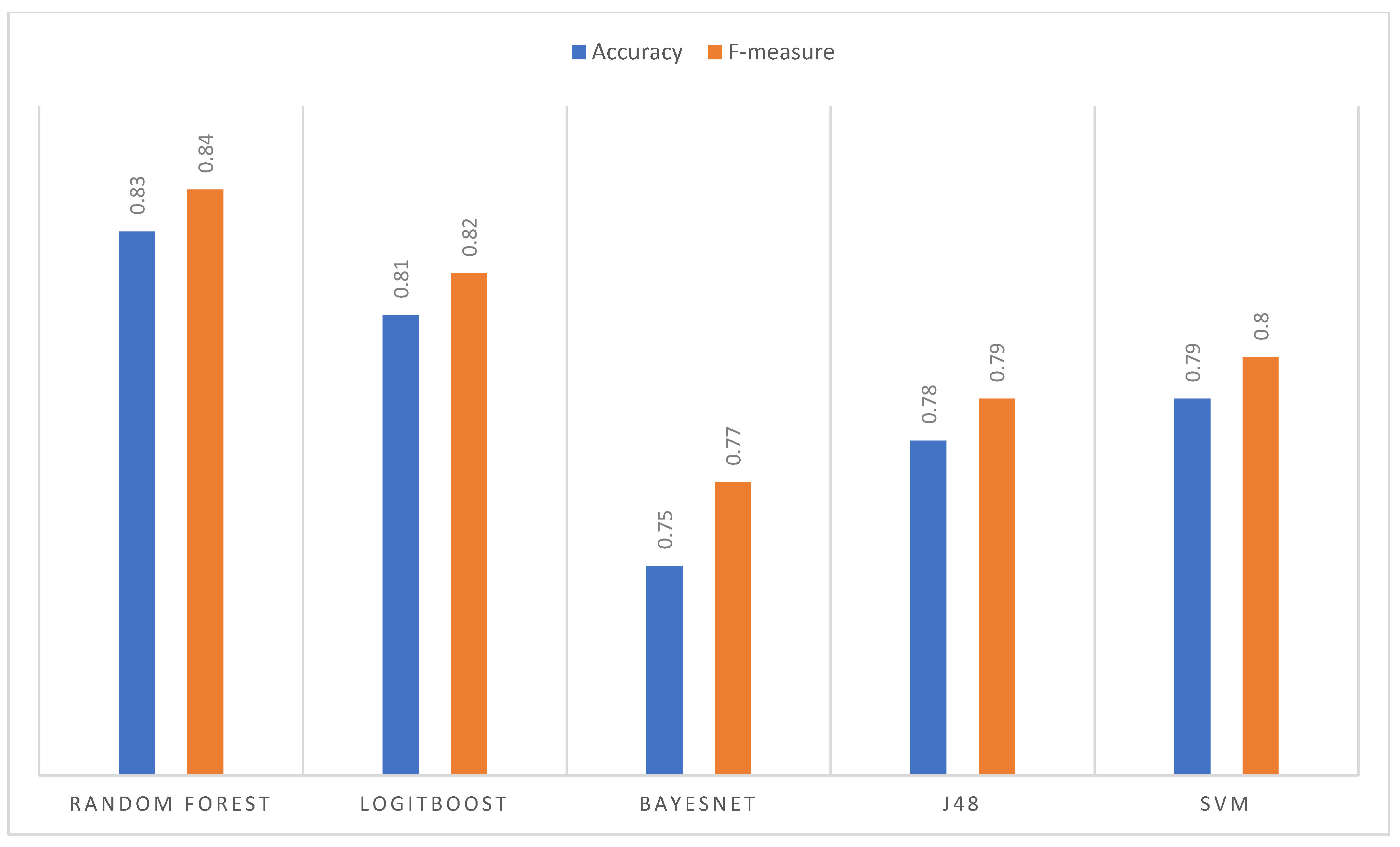
- The classification algorithm that presented the best performance for the three data sets of the three image improvement transformations was Random Forest.
Review of Literature
2. Materials
2.1. The Data Set
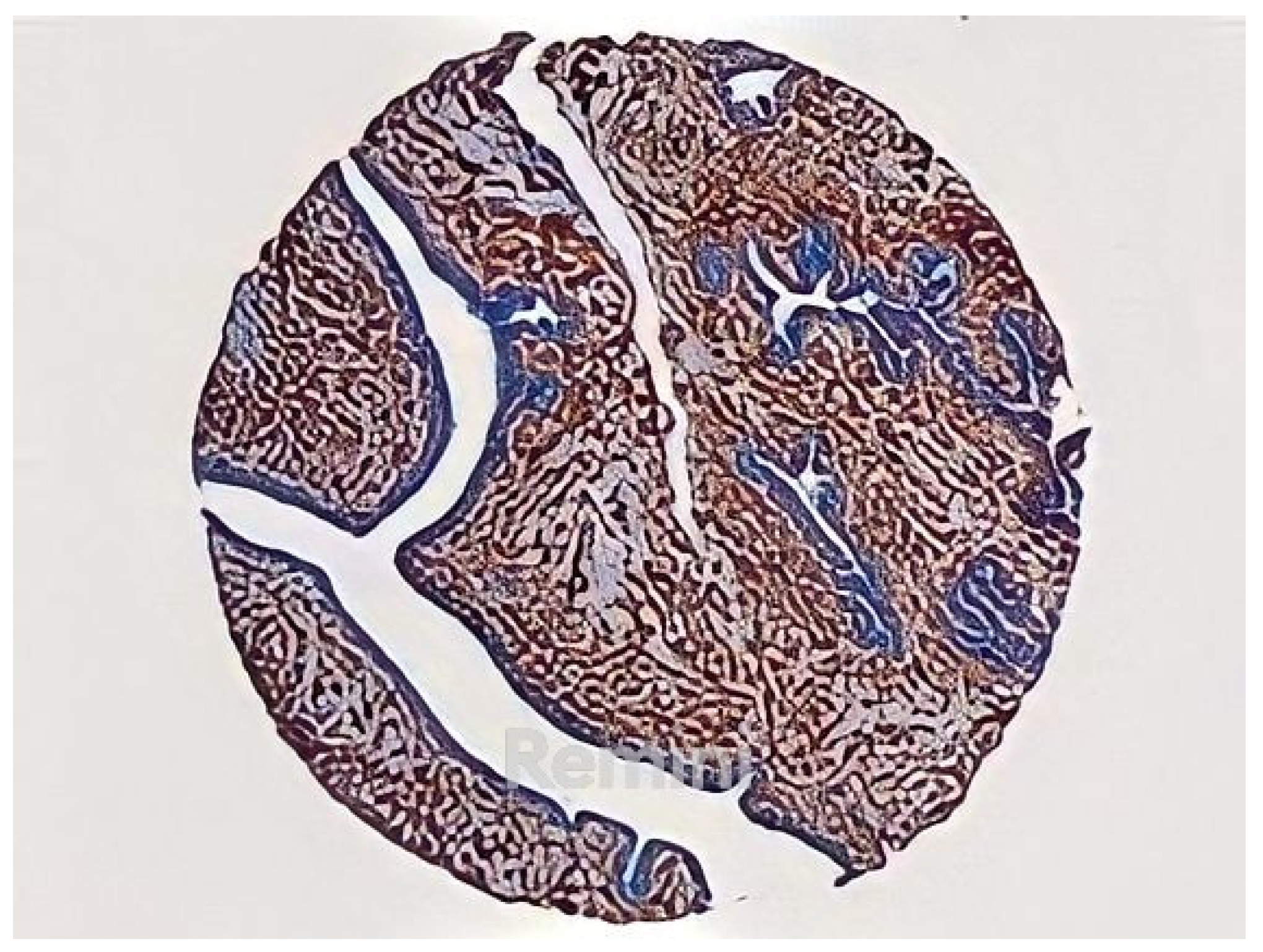
2.1.1. TMA Images
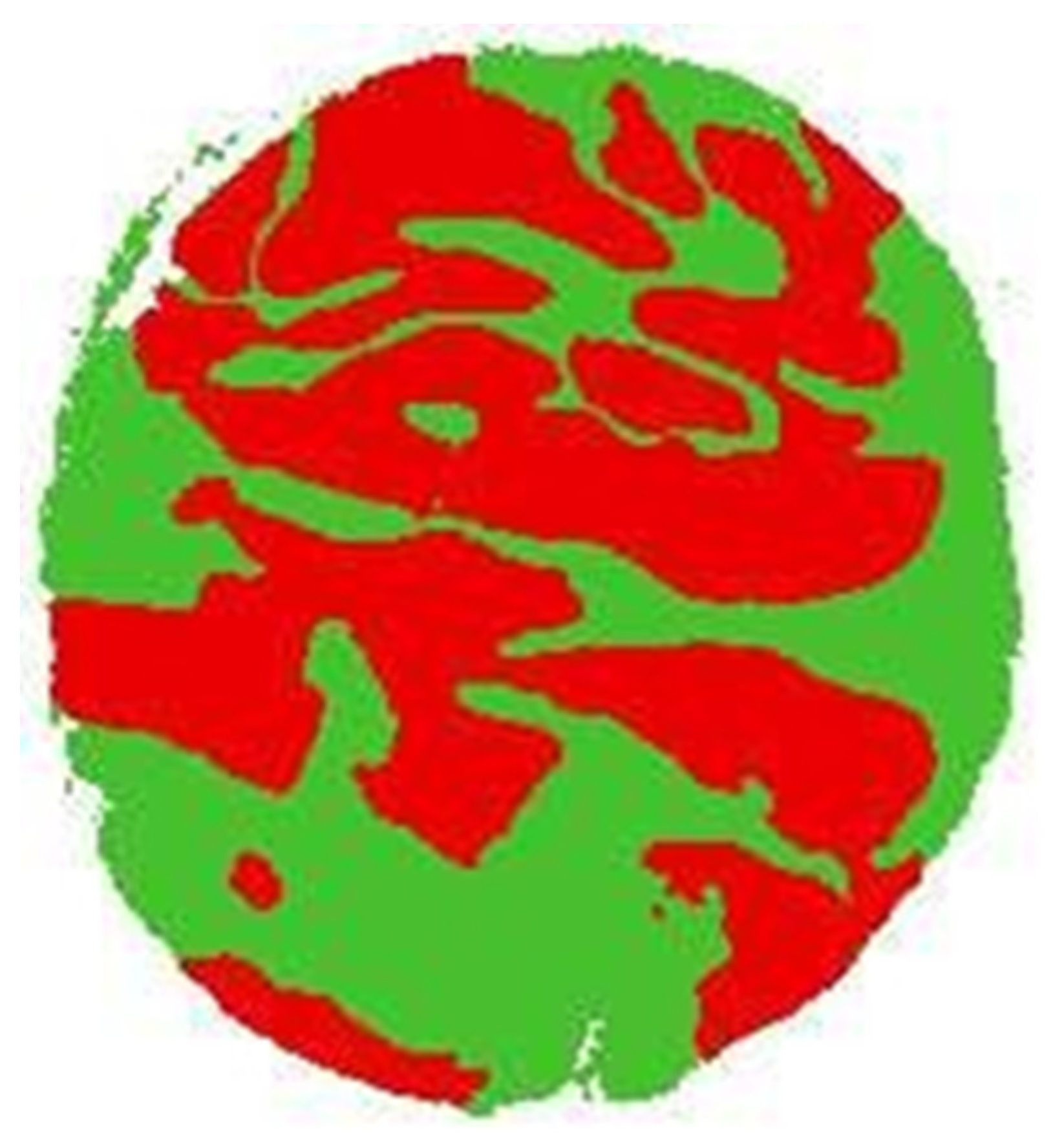
2.1.2. Labeled TMA Images
2.2. The Software
KNIME Analytics Platform
- KNIME Image Processing Extension: KNIME has an image processing plugin that contains about 100 nodes to deal with various types of images (2D and 3D) and videos, in addition to applying common methods such as pre-processing segmentation, feature extraction, tracking, and classification. Currently, this extension can be used with various image processing tools such as BioFormats, SCFIO, ImageJ, Omero, and SciJava [19].
2.3. Proposed Method
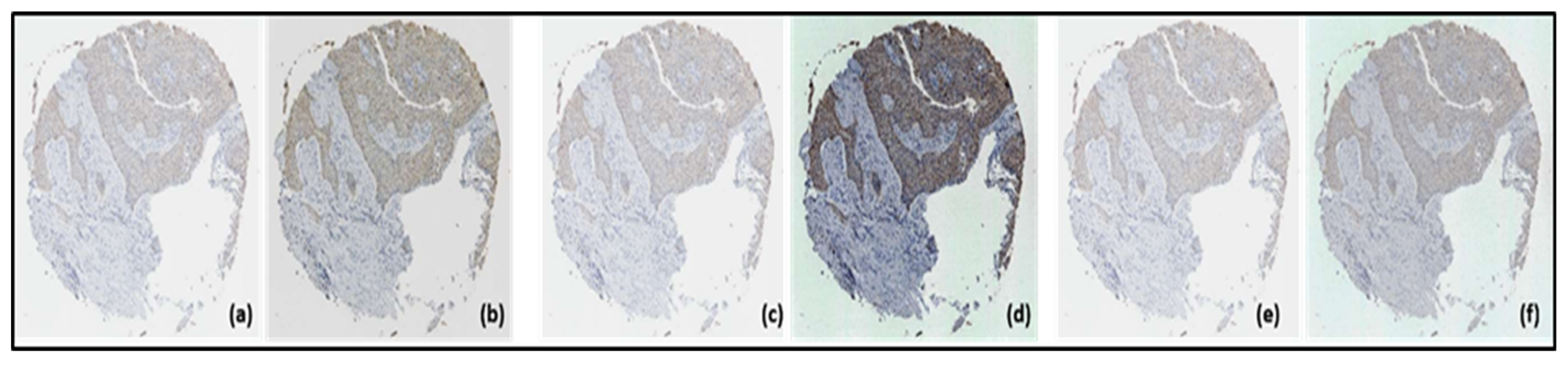
- Pre-processing: The TMA images were initially very large, and it was found that they had a very high computational load without providing many benefits against a possible reduction in their dimension. Then, evaluating the available hardware resources, it was decided to scale them to 25%, leaving each image with a resolution of approximately 1500 × 1500 pixels. The best one was selected for the segmentation phase. These techniques were applied separately, producing three new sets of TMA images, one for each transformation.
- Image normalization: Normalizing an image changes the range of pixel intensity values in order to obtain greater consistency within the data set. In this case, the TMA images were normalized by applying the spatial contrast enhancement method.
- Histogram equalization: Histogram equalization is a nonlinear transformation technique that modifies the intensity value of pixels by distributing it over the entire spectrum, which produces a more constant histogram.
- Histogram matching: When an image’s histogram matches the specified histogram, it is known as histogram matching [18]. When a mathematical function or another image’s histogram is matched to an input image’s own histogram, the operation is called histogram matching. It is very useful for comparing and contrasting images. A valid comparison can be made if the histograms of the two photos in question are similar.
- SLIC super pixel construction: Groups of pixels that share features like color, brightness, or texture are known as super pixels. In addition to offering an easy starting point from which to compute image characteristics, they also considerably reduce the complexity of subsequent image processing operations. Histopathological pictures have shown it to be effective [20].
- Experimentation: The SLIC method was applied to the original (untransformed), normalized, and histogram equalization TMA images using ImageJ with the jSLIC plugin [20], which corresponds to a computationally more efficient variation of SLIC but retains the original method. To begin with, jSLIC is basically governed by two parameters, which are:
- ○
- Start grid size: the initial average size of super pixels (30 by default)
- ○
- Regularization: degree of deformation of the estimated super pixels. The range is from 0 for very elastic super pixels to 1 for almost square super pixels (0.20 by default).
- Feature extraction: This stage consisted of obtaining a representation of the image (in the form of data) that would help discriminate between the different classes (tumor, non-tumor, and fundus). According to Jurmeister, et al. [21] the performance of a CAD system depends more on the extraction and selection of features than on the classification method.
- First-order statistics: The variance and other pixel neighborhood associations are not taken into account when computing statistics for first-order texture measures. A technique for texture analysis based on histograms examines how an image’s intensity concentrations—represented visually as a histogram—change over time. Moments such as mean, variance, spread, root mean square or mean energy, entropy, skewness, and kurtosis are among the most common characteristics of a distribution. A texture measure is a change in gray levels near a pixel, according to Fernández-Carrobles et al. [22].
- Tamura Characteristics: In Tamura et al. [23], six textural features that take advantage of human visual perception are introduced: roughness, roughness, contrast, directionality, linearity, and regularity. Fernández-Carrobles et al. [22] demonstrated the effectiveness of these visual characteristics, helping classifiers achieve success rates of up to 97% on histopathological images. The contrast and directionality features were selected as they were considered “more relevant” to obtain a better representation of the TMA images.
- Extracted features: For the problem addressed, it was determined that the texture characteristics were the most appropriate: 23 texture features were selected, divided into 17 first-order statistics and 6 Tamura-type features.
- Decomposition of RGB space: Because the original TMA images were in RGB space, it was necessary to decompose them into their three channels R (red), G (green), and B (blue), as a requirement for the calculation of the different types of features. In the end, a total of 69 features were obtained for each super pixel. For this, KNIME was used with the Splitter node to separate the image into channels and the Image Segment Features node to extract the selected features from each channel.
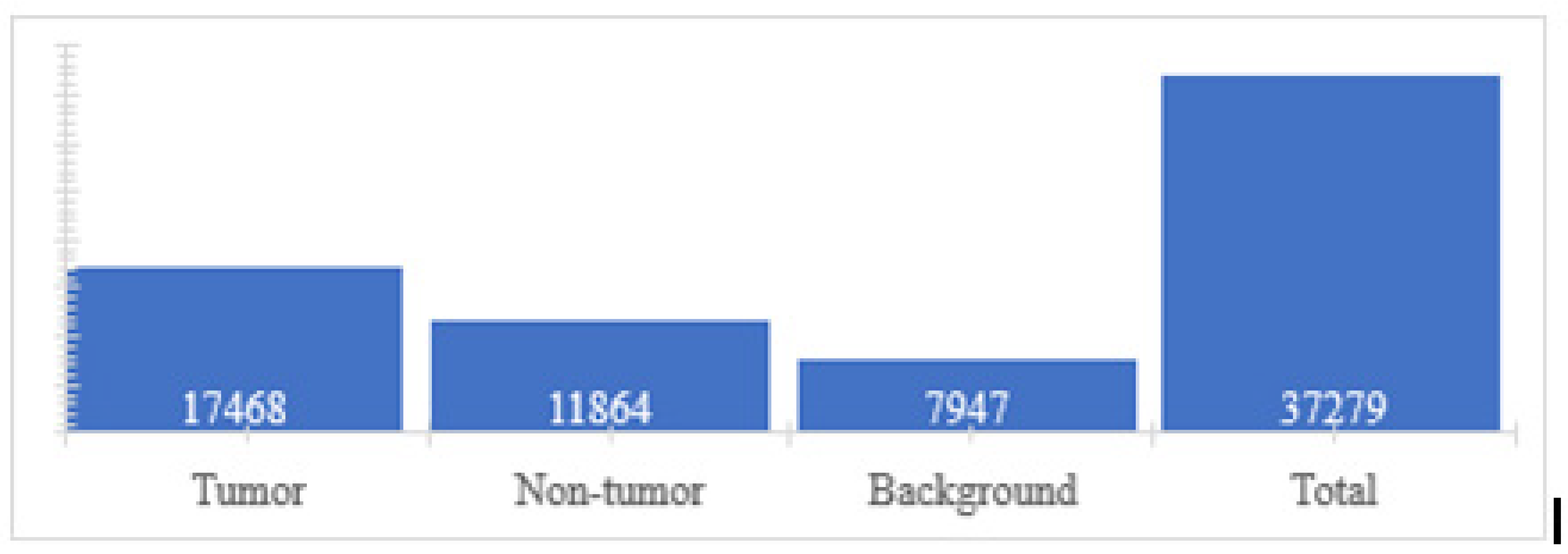
- Class assignment: In this step, the class variable (tumor, non-tumor, or background) was created and assigned to each feature vector (super pixel). For this, super pixel images were used, together with label images. The procedure consisted of taking the super pixel image by traversing each super pixel and interposing it over its corresponding area within the label image. This, in turn, has information on the class to which each pixel belongs, for which the number of pixels belonging to each class is counted through a voting system, and finally assigned to the super pixel the class that has the most votes. In other words, the class of the majority region to which the super pixel belongs is assigned.
- Segmentation: This stage consisted of the construction and comparison of five supervised classification models, of which the best was chosen and then used in the segmentation of the TMA images. First, five algorithms were selected from among the most relevant as evidenced in state of the art; then, once the models were trained for comparison, their performance was evaluated, and finally, the best segmentation algorithm was applied.
2.3.1. Classification Algorithms
- Random Forest: Breiman [24] introduced this method, which is currently one of the best and most often used. Using Random Forests is a mixture of prediction trees where each tree is dependent on the values of a random vector that is tested individually and with the same distribution for each one. A significant change from bagging is the construction and averaging of a large number of uncorrelated trees.
- Support Vector Machines (SVM): In huge data sets, the effectiveness of Sequential Minimal Optimization stands out. Training a support vector classifier is made easier by using Platt’s sequential minimum optimization method [25]. With this approach, no values are left out, and the nominal properties are converted from decimal to binary. All default attributes are also normalized. (In such a scenario, the output coefficients are based on the normalized data, not the original data, which is critical for evaluating the classifier).
- LogitBoost: It is the Weka implementation of the additive logistic regression algorithm introduced [26]. It can handle a wide range of issues and perform classification using a regression approach similar to the base learner. This algorithm was chosen over other boosting algorithms because of its superior performance in this case.
2.3.2. Evaluation Metrics
- F-measure (F1-score): If you are asked to sort items into a class, the accuracy (also known as positive predictive value (PPV)) of a class is equal to one divided by the total number of positives. Classified as positive are those things.
- Accuracy: There are many ways to quantify accuracy in binary classification tests, and precision is one of the most commonly utilized. In other words, the precision of a test measures the proportion of real outcomes (including real positives and real negatives) to the total number of instances investigated.
2.4. Pre-Processing
3. Results
3.1. SLIC Super Pixels
3.2. Extracted Features
3.3. Classification
3.3.1. Results in Normalized Images
3.3.2. Histogram Equalization Imaging Results
3.3.3. Image Results of Histogram Matching
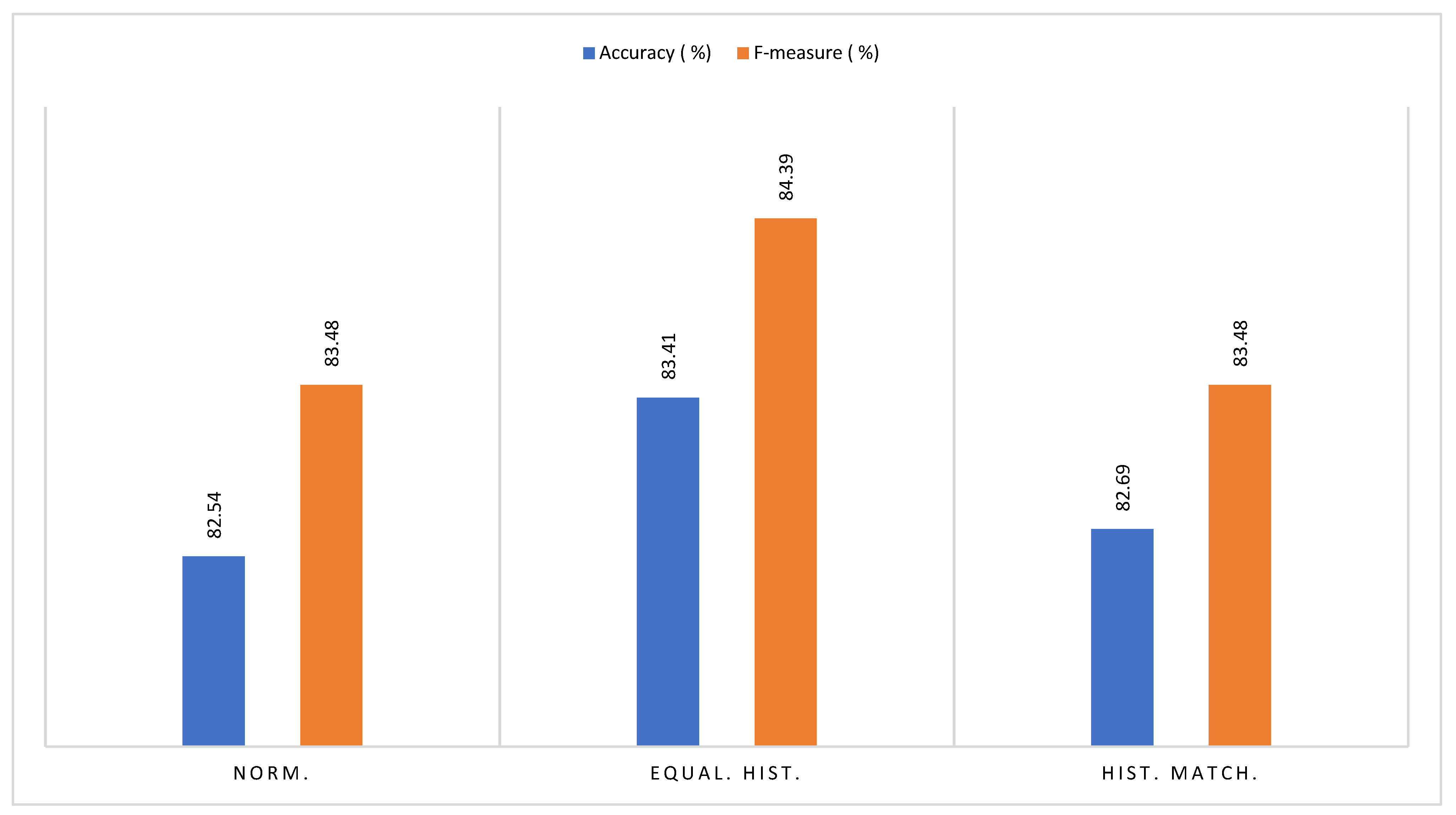
3.3.4. Comparative Analysis
3.4. Segmented Images
4. Discussion
5. Conclusions
Future Work
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Walser, T.; Cui, X.; Yanagawa, J.; Lee, J.M.; Heinrich, E.; Lee, G.; Sharma, S.; Dubinett, S.M. Smoking and Lung Cancer: The Role of Inflammation. Proc. Am. Thorac. Soc. 2008, 5, 811–815. [Google Scholar] [CrossRef]
- Habib, O.S.; Al-Asadi, J.; Hussien, O.G. Lung cancer in Basrah, Iraq during 2005-2012. Saudi Med. J. 2016, 37, 1214–1219. [Google Scholar] [CrossRef]
- Mustafa, M.; Azizi, A.J.; Iiizam, E.; Nazirah, A.; Sharifa, S.; Abbas, S. Lung Cancer: Risk Factors, Management, And Prognosis. IOSR J. Dent. Med. Sci. 2016, 15, 94–101. [Google Scholar] [CrossRef]
- Ruano-Ravina, A.; Figueiras, A.; Barros-Dios, J. Lung cancer and related risk factors: An update of the literature. Public Health 2003, 117, 149–156. [Google Scholar] [CrossRef]
- Purandare, N.C.; Rangarajan, V. Imaging of lung cancer: Implications on staging and management. Indian J. Radiol. Imaging 2015, 25, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Traverso, A.; Torres, E.L.; Fantacci, M.E.; Cerello, P. Computer-aided detection systems to improve lung cancer early diagnosis: State-of-the-art and challenges. J. Phys. Conf. Ser. 2017, 841, 12013. [Google Scholar] [CrossRef]
- Camp, R.L.; Neumeister, V.; Rimm, D.L. A Decade of Tissue Microarrays: Progress in the Discovery and Validation of Cancer Biomarkers. J. Clin. Oncol. 2008, 26, 5630–5637. [Google Scholar] [CrossRef] [PubMed]
- Voduc, D.; Kenney, C.; Nielsen, T.O. Tissue Microarrays in Clinical Oncology. Semin. Radiat. Oncol. 2008, 18, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Terán, M.D.; Brock, M.V. Staging lymph node metastases from lung cancer in the mediastinum. J. Thorac. Dis. 2014, 6, 230–236. [Google Scholar] [CrossRef] [PubMed]
- Gurcan, M.N.; Boucheron, L.E.; Can, A.; Madabhushi, A.; Rajpoot, N.M.; Yener, B. Histopathological Image Analysis: A Review. IEEE Rev. Biomed. Eng. 2009, 2, 147–171. [Google Scholar] [CrossRef]
- Das, A.; Nair, M.S.; Peter, S.D. Computer-Aided Histopathological Image Analysis Techniques for Automated Nuclear Atypia Scoring of Breast Cancer: A Review. J. Digit. Imaging 2020, 33, 1091–1121. [Google Scholar] [CrossRef]
- He, L.; Long, L.R.; Antani, S.; Thoma, G.R. Histology image analysis for carcinoma detection and grading. Comput. Methods Programs Biomed. 2012, 107, 538–556. [Google Scholar] [CrossRef]
- Hamilton, P.W.; Bankhead, P.; Wang, Y.; Hutchinson, R.; Kieran, D.; McArt, D.G.; James, J.; Salto-Tellez, M. Digital pathology and image analysis in tissue biomarker research. Methods 2014, 70, 59–73. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, W.; Sisk, A.; Ye, H.; Wallace, W.D.; Speier, W.; Arnold, C.W. A multi-resolution model for histopathology image classification and localization with multiple instance learning. Comput. Biol. Med. 2021, 131, 104253. [Google Scholar] [CrossRef]
- Yu, K.-H.; Lee, T.-L.M.; Yen, M.-H.; Kou, S.C.; Rosen, B.; Chiang, J.-H.; Kohane, I.S. Reproducible Machine Learning Methods for Lung Cancer Detection Using Computed Tomography Images: Algorithm Development and Validation. J. Med. Internet Res. 2020, 22, e16709. [Google Scholar] [CrossRef] [PubMed]
- Mejia, T.M.; Perez, M.G.; Andaluz, V.H.; Conci, A. Automatic Segmentation and Analysis of Thermograms Using Texture Descriptors for Breast Cancer Detection. In Proceedings of the 2015 Asia-Pacific Conference on Computer Aided System Engineering, Quito, Ecuador, 14–16 July 2015; IEEE: New York, NY, USA, 2015; pp. 24–29. [Google Scholar] [CrossRef]
- Marinelli, R.J.; Montgomery, K.; Liu, C.L.; Shah, N.H.; Prapong, W.; Nitzberg, M.; Zachariah, Z.K.; Sherlock, G.; Natkunam, Y.; West, R.B.; et al. The Stanford Tissue Microarray Database. Nucleic Acids Res. 2007, 36, D871–D877. [Google Scholar] [CrossRef]
- Berthold, M.R.; Cebron, N.; Dill, F.; Gabriel, T.R.; Kötter, T.; Meinl, T.; Ohl, P.; Sieb, C.; Thiel, K.; Wiswedel, B. KNIME: The Konstanz information miner. First publ. In Data Analysis, Machine Learning and Applications, Proceedings of the 31st Annual Conference of the Gesellschaft für Klassifikation e.V., Freiburg im Breisgau, Germany, 7–9 March 2007; Springer: Berlin/Heidelberg, Germany, 2008; Volume 5, p. 5. [Google Scholar] [CrossRef]
- Sydow, D.; Wichmann, M.; Rodríguez-Guerra, J.; Goldmann, D.; Landrum, G.; Volkamer, A. TeachOpenCADD-KNIME: A Teaching Platform for Computer-Aided Drug Design Using KNIME Workflows. J. Chem. Inf. Model. 2019, 59, 4083–4086. [Google Scholar] [CrossRef]
- Fouad, S.; Randell, D.; Galton, A.; Mehanna, H.; Landini, G. Epithelium and Stroma Identification in Histopathological Images Using Unsupervised and Semi-Supervised Superpixel-Based Segmentation. J. Imaging 2017, 3, 61. [Google Scholar] [CrossRef]
- Jurmeister, P.; Vollbrecht, C.; Jöhrens, K.; Aust, D.; Behnke, A.; Stenzinger, A.; Penzel, R.; Endris, V.; Schirmacher, P.; Fisseler-Eckhoff, A.; et al. Status quo of ALK testing in lung cancer: Results of an EQA scheme based on in-situ hybridization, immunohistochemistry, and RNA/DNA sequencing. Virchows Arch. 2021, 479, 247–255. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Carrobles, M.M.; Tadeo, I.; Bueno, G.; Noguera, R.; Déniz, O.; Salido, J.; García-Rojo, M. TMA Vessel Segmentation Based on Color and Morphological Features: Application to Angiogenesis Research. Sci. World J. 2013, 2013, 263190. [Google Scholar] [CrossRef]
- El Mehdi, E.A.; Hassan, S. An Effective Content Based image Retrieval Utilizing Color Features and Tamura Features. In Proceedings of the 4th International Conference on Big Data and Internet of Things, Rabat, Morocco, 23–24 October 2019; Association for Computing Machinery: New York, NY, USA, 2019; pp. 1–6. [Google Scholar] [CrossRef]
- Fawagreh, K.; Gaber, M.M.; Elyan, E. Random forests: From early developments to recent advancements. Syst. Sci. Control Eng. Open Access J. 2014, 2, 602–609. [Google Scholar] [CrossRef]
- Furey, T.S.; Cristianini, N.; Duffy, N.; Bednarski, D.W.; Schummer, M.; Haussler, D. Support vector machine classification and validation of cancer tissue samples using microarray expression data. Bioinformatics 2000, 16, 906–914. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Delgado, M.; Cernadas, E.; Barro, S.; Amorim, D. Do we Need Hundreds of Classifiers to Solve Real World Classification Problems? J. Mach. Learn. Res. 2014, 15, 3133–3181. [Google Scholar]
- Qadri, S.F.; Shen, L.; Ahmad, M.; Qadri, S.; Zareen, S.S.; Akbar, M.A. SVseg: Stacked Sparse Autoencoder-Based Patch Classification Modeling for Vertebrae Segmentation. Mathematics 2022, 10, 796. [Google Scholar] [CrossRef]
- Gertych, A.; Swiderska-Chadaj, Z.; Ma, Z.; Ing, N.; Markiewicz, T.; Cierniak, S.; Salemi, H.; Guzman, S.; Walts, A.E.; Knudsen, B.S. Convolutional neural networks can accurately distinguish four histologic growth patterns of lung adenocarcinoma in digital slides. Sci. Rep. 2019, 9, 1483. [Google Scholar] [CrossRef] [PubMed]
- Adhyam, M.; Gupta, A.K. A Review on the Clinical Utility of PSA in Cancer Prostate. Indian J. Surg. Oncol. 2012, 3, 120–129. [Google Scholar] [CrossRef]
- Kumar, Y.; Gupta, S.; Singla, R.; Hu, Y.-C. A Systematic Review of Artificial Intelligence Techniques in Cancer Prediction and Diagnosis. Arch. Comput. Methods Eng. 2021, 29, 2043–2070. [Google Scholar] [CrossRef] [PubMed]








| Positive (P) | Negative (N) | |
|---|---|---|
| Actual True | TP | FN |
| Actual False | FP | TN |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Althubaity, D.D.; Alotaibi, F.F.; Osman, A.M.A.; Al-khadher, M.A.; Abdalla, Y.H.A.; Alwesabi, S.A.; Abdulrahman, E.E.H.; Alhemairy, M.A. Automated Lung Cancer Segmentation in Tissue Micro Array Analysis Histopathological Images Using a Prototype of Computer-Assisted Diagnosis. J. Pers. Med. 2023, 13, 388. https://doi.org/10.3390/jpm13030388
Althubaity DD, Alotaibi FF, Osman AMA, Al-khadher MA, Abdalla YHA, Alwesabi SA, Abdulrahman EEH, Alhemairy MA. Automated Lung Cancer Segmentation in Tissue Micro Array Analysis Histopathological Images Using a Prototype of Computer-Assisted Diagnosis. Journal of Personalized Medicine. 2023; 13(3):388. https://doi.org/10.3390/jpm13030388
Chicago/Turabian StyleAlthubaity, DaifAllah D., Faisal Fahad Alotaibi, Abdalla Mohamed Ahmed Osman, Mugahed Ali Al-khadher, Yahya Hussein Ahmed Abdalla, Sadeq Abdo Alwesabi, Elsadig Eltaher Hamed Abdulrahman, and Maram Abdulkhalek Alhemairy. 2023. "Automated Lung Cancer Segmentation in Tissue Micro Array Analysis Histopathological Images Using a Prototype of Computer-Assisted Diagnosis" Journal of Personalized Medicine 13, no. 3: 388. https://doi.org/10.3390/jpm13030388
APA StyleAlthubaity, D. D., Alotaibi, F. F., Osman, A. M. A., Al-khadher, M. A., Abdalla, Y. H. A., Alwesabi, S. A., Abdulrahman, E. E. H., & Alhemairy, M. A. (2023). Automated Lung Cancer Segmentation in Tissue Micro Array Analysis Histopathological Images Using a Prototype of Computer-Assisted Diagnosis. Journal of Personalized Medicine, 13(3), 388. https://doi.org/10.3390/jpm13030388







