Plasma Brain-Derived Neurotrophic Factor (BDNF) Concentration and BDNF/TrkB Gene Polymorphisms in Croatian Adults with Asthma
Abstract
1. Introduction
2. Materials and Methods
2.1. Participants
2.2. Data and Blood Collection
2.3. Determination of Plasma BDNF Concentration
2.4. DNA Extraction and Genotyping
2.5. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Barrios, J.; Ai, X. Neurotrophins in asthma. Curr. Allergy Asthma Rep. 2018, 18, 10. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization Asthma. Global Prevalence. Available online: https://www.who.int/news-room/q-a-detail/asthma (accessed on 4 February 2020).
- Braido, F. Failure in asthma control: Reasons and consequences. Scientifica 2013, 2013, 549252. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, P.W.; Ghushchyan, V.H.; Slejko, J.F.; Belozeroff, V.; Globe, D.R.; Lin, S.L. The burden of adult asthma in the United States: Evidence from the Medical Expenditure Panel Survey. J. Allergy Clin. Immunol. 2011, 127, 363–369.e3. [Google Scholar] [CrossRef] [PubMed]
- Godar, M.; Blanchetot, C.; de Haard, H.; Lambrecht, B.N.; Brusselle, G. Personalized medicine with biologics for severe type 2 asthma: Current status and future prospects. MAbs 2018, 10, 34–45. [Google Scholar] [CrossRef]
- Vercelli, D. Discovering susceptibility genes for asthma and allergy. Nat. Rev. Immunol. 2008, 8, 169–182. [Google Scholar] [CrossRef]
- Wang, J.Y.; Wang, A.L.; Han, W.; Mu, Z.L. Association between a functional single nucleotide polymorphism in the brain-derived neurotrophic factor gene and risk of child asthma. Genet. Mol. Res. 2015, 14, 16233–16240. [Google Scholar] [CrossRef]
- Klein, R.; Nanduri, V.; Jing, S.A.; Lamballe, F.; Tapley, P.; Bryant, S.; Cordon-Cardo, C.; Jones, K.R.; Reichardt, L.F.; Barbacid, M. The trkB tyrosine protein kinase is a receptor for brain-derived neurotrophic factor and neurotrophin-3. Cell 1991, 66, 395–403. [Google Scholar] [CrossRef]
- Soppet, D.; Escandon, E.; Maragos, J.; Middlemas, D.S.; Reid, S.W.; Blair, J.; Burton, L.E.; Stanton, B.R.; Kaplan, D.R.; Hunter, T.; et al. The neurotrophic factors brain-derived neurotrophic factor and neurotrophin-3 are ligands for the trkB tyrosine kinase receptor. Cell 1991, 65, 895–903. [Google Scholar] [CrossRef]
- Squinto, S.P.; Stitt, T.N.; Aldrich, T.H.; Davis, S.; Bianco, S.M.; Radziejewski, C.; Glass, D.J.; Masiakowski, P.; Furth, M.E.; Valenzuela, D.M.; et al. TrkB encodes a functional receptor for brain-derived neurotrophic factor and neurotrophin-3 but not nerve growth factor. Cell 1991, 65, 885–893. [Google Scholar] [CrossRef]
- Kaplan, D.R.; Miller, F.D. Neurotrophin signal transduction in the nervous system. Curr. Opin. Neurobiol. 2000, 10, 381–391. [Google Scholar] [CrossRef]
- Minichiello, L. TrkB signalling pathways in LTP and learning. Nat. Rev. Neurosci. 2009, 10, 850–860. [Google Scholar] [CrossRef] [PubMed]
- Numakawa, T.; Suzuki, S.; Kumamaru, E.; Adachi, N.; Richards, M.; Kunugi, H. BDNF function and intracellular signaling in neurons. Histol. Histopathol. 2010, 25, 237–258. [Google Scholar] [CrossRef]
- Braun, A.; Lommatzsch, M.; Renz, H. The role of neurotrophins in allergic bronchial asthma. Clin. Exp. Allergy 2000, 30, 178–186. [Google Scholar] [CrossRef]
- Jesenak, M.; Babusikova, E.; Evinova, A.; Banovcin, P.; Dobrota, D. Val66Met polymorphism in the BDNF gene in children with bronchial asthma. Pediatric Pulmonol. 2015, 50, 631–637. [Google Scholar] [CrossRef] [PubMed]
- Wetmore, C.; Olson, L. Neuronal and nonneuronal expression of neurotrophins and their receptors in sensory and sympathetic ganglia suggest new intercellular trophic interactions. J. Comp. Neurol. 1995, 353, 143–159. [Google Scholar] [CrossRef]
- Prakash, Y.S.; Martin, R.J. Brain-derived neurotrophic factor in the airways. Pharmacol. Ther. 2014, 143, 74–86. [Google Scholar] [CrossRef]
- Ricci, A.; Felici, L.; Mariotta, S.; Mannino, F.; Schmid, G.; Terzano, C.; Cardillo, G.; Amenta, F.; Bronzetti, E. Neurotrophin and neurotrophin receptor protein expression in the human lung. Am. J. Respir. Cell Mol. Biol. 2004, 30, 12–19. [Google Scholar] [CrossRef]
- Brigadski, T.; Leßmann, V. The physiology of regulated BDNF release. Cell Tissue Res. 2020, 382, 15–45. [Google Scholar] [CrossRef] [PubMed]
- Dragunas, G.; Woest, M.E.; Nijboer, S.; Bos, S.T.; van Asselt, J.; de Groot, A.P.; Vohlídalová, E.; Vermeulen, C.J.; Ditz, B.; Vonk, J.M.; et al. Cholinergic neuroplasticity in asthma driven by TrkB signaling. FASEB J. 2020, 34, 7703–7717. [Google Scholar] [CrossRef] [PubMed]
- Lommatzsch, M.; Schloetcke, K.; Klotz, J.; Schuhbaeck, K.; Zingler, D.; Zingler, C.; Schulte-Herbrüggen, O.; Gill, H.; Schuff-Werner, P.; Virchow, J.C. Brain-derived neurotrophic factor in platelets and airflow limitation in asthma. Am. J. Respir. Crit. Care Med. 2005, 171, 115–120. [Google Scholar] [CrossRef]
- Muller, G.C.; Pitrez, P.M.; Teixeira, A.L.; Pires, P.S.; Jones, M.H.; Stein, R.T.; Bauer, M.E. Plasma brain-derived neurotrophic factor levels are associated with clinical severity in school age children with asthma. Clin. Exp. Allergy 2010, 40, 1755–1759. [Google Scholar] [CrossRef] [PubMed]
- Joachim, R.A.; Noga, O.; Sagach, V.; Hanf, G.; Fliege, H.; Kocalevent, R.D.; Peters, E.M.; Klapp, B.F. Correlation between immune and neuronal parameters and stress perception in allergic asthmatics. Clin. Exp. Allergy 2008, 38, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Koskela, H.O.; Purokivi, M.K.; Romppanen, J. Neurotrophins in chronic cough: Association with asthma but not with cough severity. Clin. Respir. J. 2010, 4, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Egan, M.F.; Kojima, M.; Callicott, J.H.; Goldberg, T.E.; Kolachana, B.S.; Bertolino, A.; Zaitsev, E.; Gold, B.; Goldman, D.; Dean, M.; et al. The BDNF val66met polymorphism affects activity-dependent secretion of BDNF and human memory and hippocampal function. Cell 2003, 112, 257–269. [Google Scholar] [CrossRef]
- Baj, G.; Carlino, D.; Gardossi, L.; Tongiorgi, E. Toward a unified biological hypothesis for the BDNF Val66Met-associated memory deficits in humans: A model of impaired dendritic mRNA trafficking. Front. Neurosci. 2013, 7, 188. [Google Scholar] [CrossRef]
- Voegeli, G.; Ramoz, N.; Shekhtman, T.; Courtet, P.; Gorwood, P.; Kelsoe, J.R. Neurotrophin Genes and Antidepressant-Worsening Suicidal Ideation: A Prospective Case-Control Study. Int. J. Neuropsychopharmacol. 2016, 19, pyw059. [Google Scholar] [CrossRef]
- Szczepankiewicz, A.; Breborowicz, A.; Skibińska, M.; Wiłkość, M.; Tomaszewska, M.; Hauser, J. Association analysis of brain-derived neurotrophic factor gene polymorphisms in asthmatic children. Pediatr. Allergy Immunol. 2007, 18, 293–297. [Google Scholar] [CrossRef]
- Szczepankiewicz, A.; Rose-Zerilli, M.J.; Barton, S.J.; Holgate, S.T.; Holloway, J.W. Association analysis of brain-derived neurotrophic factor gene polymorphisms in asthmatic families. Int. Arch. Allergy Immunol. 2009, 149, 343–349. [Google Scholar] [CrossRef]
- Yinli, C.; Jie, H.; Li, Z.; Jun, G.; Peiling, L.; Weihong, Y. Association between brain-derived neurothropic factor variants and asthma in Chinese Han children. Acta Paediatr. 2013, 102, e247–e250. [Google Scholar] [CrossRef]
- Zeilinger, S.; Pinto, L.A.; Nockher, W.A.; Depner, M.; Klopp, N.; Illig, T.; von Mutius, E.; Renz, H.; Kabesch, M. The effect of BDNF gene variants on asthma in German children. Allergy 2009, 64, 1790–1794. [Google Scholar] [CrossRef]
- Szczepankiewicz, A.; Rachel, M.; Sobkowiak, P.; Kycler, Z.; Wojsyk-Banaszak, I.; Schöneich, N.; Szczawińska-Popłonyk, A.; Bręborowicz, A. Neurotrophin serum concentrations and polymorphisms of neurotrophins and their receptors in children with asthma. Respir. Med. 2013, 107, 30–36. [Google Scholar] [CrossRef]
- Global Initiative for Asthma. Global Strategy for Asthma Management and Prevention. Available online: http://ginasthma.org (accessed on 4 February 2020).
- Quanjer, P.H.; Tammeling, G.J.; Cotes, J.E.; Pedersen, O.F.; Peslin, R.; Yernault, J.C. Lung volumes and forced ventilatory flows. Report Working Party Standardization of Lung Function Tests, European Community for Steel and Coal. Official Statement of the European Respiratory Society. Eur. Respir. J. 1993, 16, 5–40. [Google Scholar] [CrossRef] [PubMed]
- Cotes, J.E.; Chinn, D.J.; Quanjer, P.H.; Roca, J.; Yernault, J.C. Standardization of the measurement of transfer factor (diffusing capacity). Report Working Party Standardization of Lung Function Tests, European Community for Steel and Coal. Official Statement of the European Respiratory Society. Eur. Respir. J. 1993, 16, 41–52. [Google Scholar] [CrossRef] [PubMed]
- Eiringhaus, K.; Renz, H.; Matricardi, P.; Skevaki, C. Component-Resolved Diagnosis in Allergic Rhinitis and Asthma. J. Appl. Lab. Med. 2019, 3, 883–898. [Google Scholar] [CrossRef] [PubMed]
- Buhl, R.; Humbert, M.; Bjermer, L.; Chanez, P.; Heaney, L.G.; Pavord, I.; Quirce, S.; Virchow, J.C.; Holgate, S.; Expert Group of the European Consensus Meeting for Severe Eosinophilic Asthma. Severe Eosinophilic Asthma: A roadmap to consensus. Eur. Respir. J. 2017, 49, 1700634. [Google Scholar] [CrossRef]
- Woodruff, P.G.; Modrek, B.; Choy, D.F.; Jia, G.; Abbas, A.R.; Ellwagner, A.; Koth, L.L.; Arron, J.R.; Fahy, J.V. T-helper type 2-driven inflammation defines major subphenotypes of asthma. Am. J. Respir. Crit. Care Med. 2009, 180, 388–395. [Google Scholar] [CrossRef]
- Wenzel, S.E. Asthma phenotypes: The evolution from clinical to molecular approaches. Nat. Med. 2012, 18, 716–725. [Google Scholar] [CrossRef]
- Pavord, I.; Behmer, T.; Braido, F.; Cosio, B.G.; Humbert, M.; Idzko, M.; Adamek, L. Severe T2-high asthma in the biological era: European experts’ opinion. Eur. Respir. Rev. 2019, 28, 190054. [Google Scholar] [CrossRef]
- Fitzpatrick, A.M.; Chipps, B.E.; Holguin, F.; Woodruff, P.G. T2-“low” asthma: Overview and management strategies. J. Allergy Clin. Immunol. Pract. 2020, 8, 452–463. [Google Scholar] [CrossRef]
- Kim, S.D.; Cho, K.S. Samter’s Triad: State of the Art. Clin. Exp. Otorhinolaryngol. 2018, 11, 71–80. [Google Scholar] [CrossRef]
- Miller, S.A.; Dykes, D.D.; Polesky, H.F. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 1988, 16, 1215. [Google Scholar] [CrossRef]
- World Health Organization—Regional Office for Europe. Blood Services in South-Eastern Europe 2007. Available online: http://www.euro.who.int/en/health-topics/Health-systems/blood-safety/publications2/2007/blood-services-in-south-eastern-europe-current-status-and-challenges-2007 (accessed on 4 February 2020).
- Vuk, T.; Ljubičić, J.; Gulan Harcet, J.; Očić, T.; Jukić, I. Post-donation information management—contribution to the safety of transfusion treatment. Transfus. Clin. Biol. 2019, 26, 353–354. [Google Scholar] [CrossRef]
- Fuhlbrigge, A.L.; Jackson, B.; Wright, R. Gender and asthma. Immunol. Allergy Clin. 2002, 22, 10. [Google Scholar] [CrossRef]
- Zein, J.G.; Denson, J.L.; Wechsler, M.E. Asthma over the Adult Life Course: Gender and Hormonal Influences. Clin. Chest. Med. 2019, 40, 149–161. [Google Scholar] [CrossRef] [PubMed]
- Zein, J.G.; Dweik, R.A.; Comhair, S.A.; Bleecker, E.R.; Moore, W.C.; Peters, S.P.; Busse, W.W.; Jarjour, N.N.; Calhoun, W.J.; Castro, M.; et al. Severe Asthma Research Program. Asthma Is More Severe in Older Adults. PLoS ONE 2015, 10, e0133490. [Google Scholar] [CrossRef] [PubMed]
- Chatkin, J.M.; Dullius, C.R. The management of asthmatic smokers. Asthma Res. Pract. 2016, 2, 10. [Google Scholar] [CrossRef] [PubMed]
- Glavak Tkalić, R.; Miletić, G.-M.; Sakoman, S. Prevalence of substance use among the general population: Situation in Croatia and comparison with other European countries. Druš. Istraž. 2013, 22, 557–578. [Google Scholar] [CrossRef]
- Katsaounou, P.; Ioannou, M.; Hyland, M.E.; Odemyr, M.; Spranger, O.; Lindberg, A.; Gasser, M.; Conde, L.G.; Jaumont, X.; Kasujee, I. Smoking asthmatics, a neglected large phenotype of asthmatic patients. Open J. Asthma 2019, 3, 1–8. [Google Scholar] [CrossRef]
- Sustar, A.; Perkovic, M.N.; Erjavec, G.N.; Strac, D.S.; Pivac, N. Association between reduced brain-derived neurotrophic factor concentration & coronary heart disease. Indian J. Med. Res. 2019, 150, 43–49. [Google Scholar] [CrossRef]
- Pillai, A.; Bruno, D.; Sarreal, A.S.; Hernando, R.T.; Saint-Louis, L.A.; Nierenberg, J.; Ginsberg, S.D.; Pomara, N.; Mehta, P.D.; Zetterberg, H.; et al. Plasma BDNF levels vary in relation to body weight in females. PLoS ONE 2012, 7, e39358. [Google Scholar] [CrossRef]
- Lommatzsch, M.; Zingler, D.; Schuhbaeck, K.; Schloetcke, K.; Zingler, C.; Schuff-Werner, P.; Virchow, J.C. The impact of age, weight and gender on BDNF levels in human platelets and plasma. Neurobiol. Aging 2005, 26, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Bhang, S.Y.; Choi, S.W.; Ahn, J.H. Changes in plasma brain-derived neurotrophic factor levels in smokers after smoking cessation. Neurosci. Lett. 2010, 468, 7–11. [Google Scholar] [CrossRef] [PubMed]
- Jamal, M.; Van der Does, W.; Elzinga, B.M.; Molendijk, M.L.; Penninx, B.W. Association between smoking, nicotine dependence, and BDNF Val66Met polymorphism with BDNF concentrations in serum. Nicotine Tob. Res. 2015, 17, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Xia, H.; Du, X.; Yin, G.; Zhang, Y.; Li, X.; Cai, J.; Huang, X.; Ning, Y.; Soares, J.C.; Wu, F.; et al. Effects of smoking on cognition and BDNF levels in a male Chinese population: Relationship with BDNF Val66Met polymorphism. Sci. Rep. 2019, 9, 217. [Google Scholar] [CrossRef]
- Sathish, V.; Vanoosten, S.K.; Miller, B.S.; Aravamudan, B.; Thompson, M.A.; Pabelick, C.M.; Vassallo, R.; Prakash, Y.S. Brain-derived neurotrophic factor in cigarette smoke-induced airway hyperreactivity. Am. J. Respir. Cell. Mol. Biol. 2013, 48, 431–438. [Google Scholar] [CrossRef]
- Noga, O.; Hanf, G.; Schäper, C.; O’Connor, A.; Kunkel, G. The influence of inhalative corticosteroids on circulating Nerve Growth Factor, Brain-Derived Neurotrophic Factor and Neurotrophin-3 in allergic asthmatics. Clin. Exp. Allergy 2001, 31, 1906–1912. [Google Scholar] [CrossRef]
- Freeman, M.R.; Sathish, V.; Manlove, L.; Wang, S.; Britt, R.D., Jr.; Thompson, M.A.; Pabelick, C.M.; Prakash, Y.S. Brain-derived neurotrophic factor and airway fibrosis in asthma. Am. J. Physiol. Lung Cell. Mol. Physiol. 2017, 313, L360–L370. [Google Scholar] [CrossRef]
- Watanabe, T.; Fajt, M.L.; Trudeau, J.B.; Voraphani, N.; Hu, H.; Zhou, X.; Holguin, F.; Wenzel, S.E. Brain-derived neurotrophic factor expression in asthma. Association with severity and type 2 inflammatory processes. Am. J. Respir. Cell. Mol. Biol. 2015, 53, 844–852. [Google Scholar] [CrossRef]
- Szczepankiewicz, A.; Rachel, M.; Sobkowiak, P.; Kycler, Z.; Wojsyk-Banaszak, I.; Schöneich, N.; Skibinska, M.; Bręborowicz, A. Serum neurotrophin-3 and neurotrophin-4 levels are associated with asthma severity in children. Eur. Respir. J. 2012, 39, 1035–1037. [Google Scholar] [CrossRef]
- Salama, A.A.; Mostafa, G.A.; Abd Al-Aziz, M.M.; Ibrahim, M.N. Brain-derived neurotrophic factor in asthmatic children. Egypt. J. Pediatr. Allergy Immunol. 2003, 1, 102–109. [Google Scholar]
- Lommatzsch, M.; Niewerth, A.; Klotz, J.; Schulte-Herbrüggen, O.; Zingler, C.; Schuff-Werner, P.; Virchow, J.C. Platelet and plasma BDNF in lower respiratory tract infections of the adult. Respir. Med. 2007, 101, 1493–1499. [Google Scholar] [CrossRef] [PubMed]
- Azoulay, D.; Shehadeh, M.; Chepa, S.; Shaoul, E.; Baroum, M.; Horowitz, N.A.; Kaykov, E. Recovery from SARS-CoV-2 infection is associated with serum BDNF restoration. J. Infect. 2020, 81, e79–e81. [Google Scholar] [CrossRef]
- Coffey, C.S.; Mulligan, R.M.; Schlosser, R.J. Mucosal expression of nerve growth factor and brain-derived neurotrophic factor in chronic rhinosinusitis. Am. J. Rhinol. Allergy 2009, 23, 571–574. [Google Scholar] [CrossRef] [PubMed]
- Jornot, L.; Grouzmann, E.; Lacroix, J.S.; Rochat, T. BDNF and DPP-IV in polyps and middle turbinates epithelial cells. Rhinology 2007, 45, 129–133. [Google Scholar] [PubMed]
- Rajan, J.P.; Wineinger, N.E.; Stevenson, D.D.; White, A.A. Prevalence of aspirin-exacerbated respiratory disease among asthmatic patients: A meta-analysis of the literature. J. Allergy Clin. Immunol. 2015, 135, 676–681.e1. [Google Scholar] [CrossRef] [PubMed]
- Virchow, J.C.; Julius, P.; Lommatzsch, M.; Luttmann, W.; Renz, H.; Braun, A. Neurotrophins are increased in bronchoalveolar lavage fluid after segmental allergen provocation. Am. J. Respir. Crit. Care. Med. 1998, 158, 2002–2005. [Google Scholar] [CrossRef] [PubMed]
- Noga, O.; Englmann, C.; Hanf, G.; Grützkau, A.; Seybold, J.; Kunkel, G. The production, storage and release of the neurotrophins nerve growth factor, brain-derived neurotrophic factor and neurotrophin-3 by human peripheral eosinophils in allergics and non-allergics. Clin. Exp. Allergy 2003, 33, 649–654. [Google Scholar] [CrossRef]
- Xie, X.; Zhu, Y.; Zhang, J.; Zhai, C.; Feng, W.; Pan, Y.; Liu, L.; Su, X.; Yang, L.; Li, M. Association between Val66Met polymorphisms in brain-derived neurotrophic factor gene and asthma risk: A meta-analysis. Inflamm. Res. 2015, 64, 875–883. [Google Scholar] [CrossRef]
- Szczepankiewicz, A.; Bręborowicz, A.; Sobkowiak, P.; Popiel, A. Association of BDNF gene polymorphism with asthma in polish children. World Allergy Organ. J. 2010, 3, 235–238. [Google Scholar] [CrossRef]
- Chen, Z.Y.; Patel, P.D.; Sant, G.; Meng, C.X.; Teng, K.K.; Hempstead, B.L.; Lee, F.S. Variant brain-derived neurotrophic factor (BDNF) (Met66) alters the intracellular trafficking and activity-dependent secretion of wild-type BDNF in neurosecretory cells and cortical neurons. J. Neurosci. 2004, 24, 4401–4411. [Google Scholar] [CrossRef]
- Bhang, S.; Ahn, J.H.; Choi, S.W. Brain-derived neurotrophic factor and serotonin transporter gene-linked promoter region genes alter serum levels of brain-derived neurotrophic factor in humans. J. Affect. Disord. 2011, 128, 299–304. [Google Scholar] [CrossRef]
- Lang, U.E.; Hellweg, R.; Sander, T.; Gallinat, J. The Met allele of the BDNF Val66Met polymorphism is associated with increased BDNF serum concentrations. Mol. Psychiatry 2009, 14, 120–122. [Google Scholar] [CrossRef] [PubMed]
- Terracciano, A.; Piras, M.G.; Lobina, M.; Mulas, A.; Meirelles, O.; Sutin, A.R.; Chan, W.; Sanna, S.; Uda, M.; Crisponi, L.; et al. Genetics of serum BDNF: Meta-analysis of the Val66Met and genome-wide association study. World J. Biol. Psychiatry 2013, 14, 583–589. [Google Scholar] [CrossRef]
- Andiappan, A.K.; Parate, P.N.; Anantharaman, R.; Suri, B.K.; de Wang, Y.; Chew, F.T. Genetic variation in BDNF is associated with allergic asthma and allergic rhinitis in an ethnic Chinese population in Singapore. Cytokine 2011, 56, 218–223. [Google Scholar] [CrossRef]
- Jin, P.; Andiappan, A.K.; Quek, J.M.; Lee, B.; Au, B.; Sio, Y.Y.; Irwanto, A.; Schurmann, C.; Grabe, H.J.; Suri, B.K.; et al. A functional brain-derived neurotrophic factor (BDNF) gene variant increases the risk of moderate-to-severe allergic rhinitis. J. Allergy Clin. Immunol. 2015, 135, 1486–1493. [Google Scholar] [CrossRef]
- Nassenstein, C.; Braun, A.; Erpenbeck, V.J.; Lommatzsch, M.; Schmidt, S.; Krug, N.; Luttmann, W.; Renz, H.; Virchow, J.C., Jr. The neurotrophins nerve growth factor, brain-derived neurotrophic factor, neurotrophin-3, and neurotrophin-4 are survival and activation factors for eosinophils in patients with allergic bronchial asthma. J. Exp. Med. 2003, 198, 455–467. [Google Scholar] [CrossRef]
- Nassenstein, C.; Möhring, U.H.; Luttmann, W.; Virchow, J.C., Jr.; Braun, A. Differential expression of the neurotrophin receptors p75NTR, TrkA, TrkB and TrkC in human peripheral blood mononuclear cells. Exp. Toxicol. Pathol. 2006, 57, 55–63. [Google Scholar] [CrossRef]
- Nassenstein, C.; Dawbarn, D.; Pollock, K.; Allen, S.J.; Erpenbeck, V.J.; Spies, E.; Krug, N.; Braun, A. Pulmonary distribution, regulation, and functional role of Trk receptors in a murine model of asthma. J. Allergy Clin. Immunol. 2006, 118, 597–605. [Google Scholar] [CrossRef]
- Britt, R.D., Jr.; Thompson, M.A.; Wicher, S.A.; Manlove, L.J.; Roesler, A.; Fang, Y.H.; Roos, C.; Smith, L.; Miller, J.D.; Pabelick, C.M.; et al. Smooth muscle brain-derived neurotrophic factor contributes to airway hyperreactivity in a mouse model of allergic asthma. FASEB J. 2019, 33, 3024–3034. [Google Scholar] [CrossRef] [PubMed]
- Müller, P.; Duderstadt, Y.; Lessmann, V.; Müller, N.G. Lactate and BDNF: Key Mediators of Exercise Induced Neuroplasticity? J. Clin. Med. 2020, 9, 1136. [Google Scholar] [CrossRef]
- Di Marco, F.; Santus, P.; Centanni, S. Anxiety and depression in asthma. Curr. Opin. Pulm. Med. 2011, 17, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhao, M.; Zhang, Y.; Shen, X.; Yuan, Y. Correlation of 5-HTT, BDNF and NPSR1 gene polymorphisms with anxiety and depression in asthmatic patients. Int. J. Mol. Med. 2016, 38, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Szilasi, M.E.; Pak, K.; Kardos, L.; Varga, V.E.; Seres, I.; Mikaczo, A.; Fodor, A.; Szilasi, M.; Tajti, G.; Papp, C.; et al. The Alteration of Irisin-Brain-Derived Neurotrophic Factor Axis Parallels Severity of Distress Disorder in Bronchial Asthma Patients. Front. Neurosci. 2017, 11, 653. [Google Scholar] [CrossRef] [PubMed]
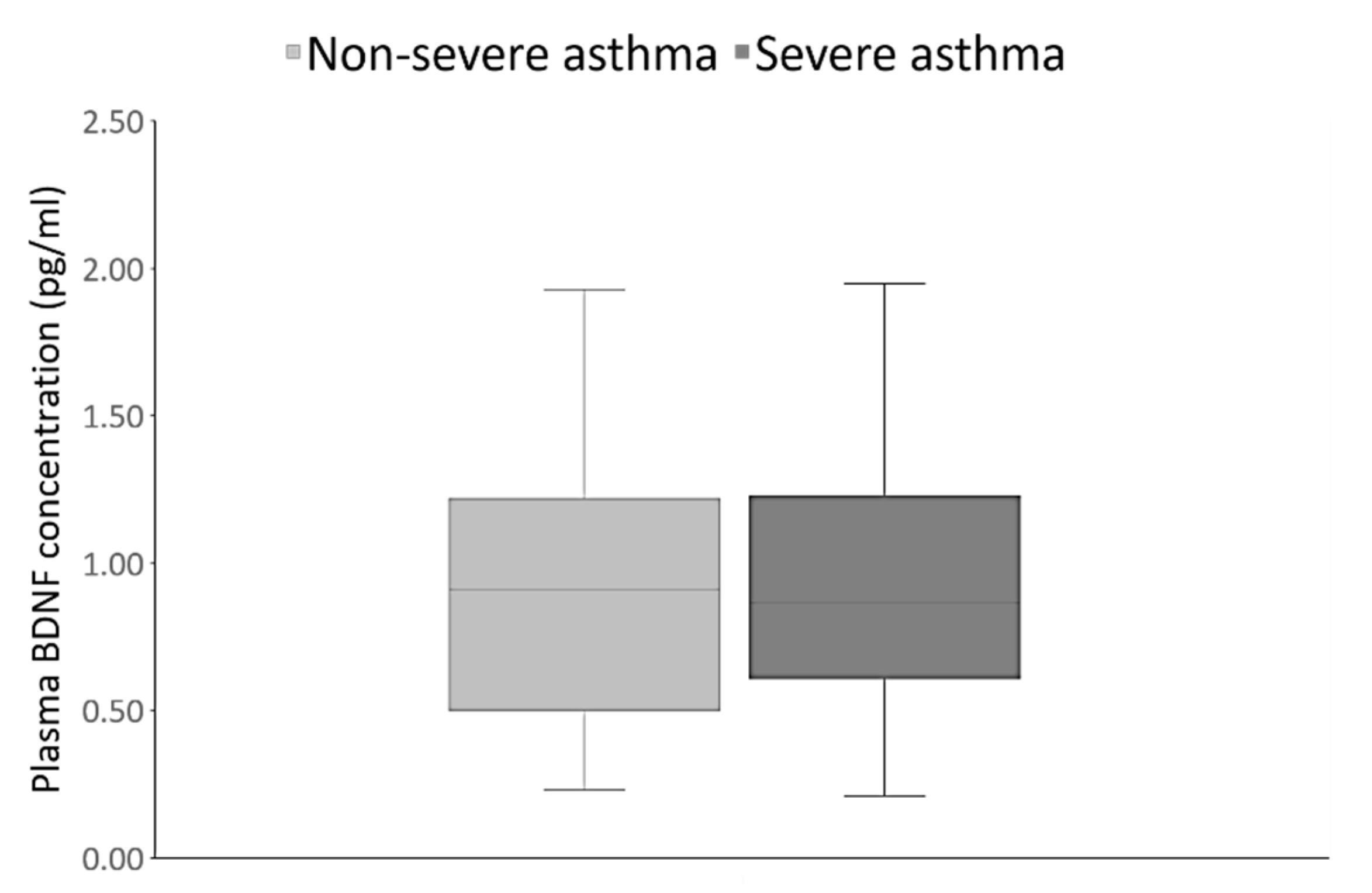

| Parameter | Asthma Patients (n = 120) | Healthy Controls (n = 120) | Statistical Analysis |
|---|---|---|---|
| Age (years) Median (25%; 75%) | 58.00 (40.25; 67.00) | 42.00 (33.25; 51.00) | p < 0.0001; U = 3912.00 Mann-Whitney test |
| Males n (%) | 41 (34.17) | 73 (60.83) | p < 0.0001 Fisher’s exact test |
| Current smokers n (%) | 10 (8.33) | 41 (34.17) | p = 0.0005 Fisher’s exact test |
| BMI (kg/m2) Median (25%; 75%) | 26.65 (23.03; 29.80) | 26.20 (23.33; 28.85) | p = 0.64; U = 6949.00 Mann-Whitney test |
| BMI category: Normal weight (18.5–24.9 kg/m2), n (%) | 44 (36.67) | 43 (35.83) | p = 0.48; χ2 = 4.52 χ2-test |
| BMI category: Obesity (≥30 kg/m2), n (%) | 28 (23.33) | 22 (18.33) | p = 0.43 Fisher’s exact test |
| BDNF Val66Met Polymorphism | Asthma Patients n (%) | Healthy Controls n (%) | Statistical Analysis | |
|---|---|---|---|---|
| Genotypes | AA | 3 (2.50) | 2 (1.67) | p = 0.54; χ2 = 1.23 χ2-test |
| AG | 43 (35.83) | 36 (30.00) | ||
| GG | 74 (61.67) | 82 (68.33) | ||
| Alleles | A | 49 (20.42) | 40 (16.67) | p = 0.35 Fisher’s exact test |
| G | 191 (79.58) | 200 (83.33) | ||
| Carriers | A | 46 (38.33) | 38 (31.67) | p = 0.34 Fisher’s exact test |
| GG | 74 (61.67) | 82 (68.33) | ||
| G | 117 (97.50) | 118 (98.33) | p = 1.00 Fisher’s exact test | |
| AA | 3 (2.50) | 2 (1.67) | ||
| BDNF Val66Met Polymorphism | Asthma Patients | Healthy Subjects | |||
|---|---|---|---|---|---|
| Plasma BDNF Concentration (pg/mL) Median (25%; 75%) | Statistical Analysis | Plasma BDNF Concentration (pg/mL), Median (25%; 75%) | Statistical Analysis | ||
| Genotypes | AA | 0.45 (0.43; 1.24) | p = 0.52 Kruskal-Wallis test | 0.56 (0.47; 0.65) | p = 0.03 Kruskal-Wallis test * |
| AG | 0.81 (0.52; 1.35) | 0.51 (0.32; 0.73) | |||
| GG | 0.94 (0.62; 0.94) | 0.68 (0.49; 1.05) | |||
| Carriers | A | 0.80 (0.46; 1.34) | p = 0.39; U = 1544.00 Mann-Whitney test | 0.51 (0.35; 0.70) | p = 0.008; U = 1089.00 Mann-Whitney test |
| GG | 0.94 (0.62; 1.20) | 0.68 (0.49; 1.05) | |||
| G | 0.91 (0.61; 1.22) | p = 0.37; U = 120.50 Mann-Whitney test | 0.59 (0.45; 0.93) | p = 0.64; U = 94.50 Mann-Whitney test | |
| AA | 0.45 (0.43; 1.24) | 0.56 (0.47; 0.65) | |||
| NTRK2 rs1439050 Polymorphism | Asthma Patients n (%) | Healthy Controls n (%) | Statistical Analysis | |
|---|---|---|---|---|
| Genotypes | GG | 65 (54.17) | 61 (50.83) | p = 0.86; χ2 = 0.29 χ2-test |
| GT | 49 (40.83) | 52 (43.33) | ||
| TT | 6 (5.00) | 7 (5.83) | ||
| Alleles | G | 179 (74.58) | 174 (72.50) | p = 0.68 Fisher’s exact test |
| T | 61 (25.42) | 66 (27.50) | ||
| Carriers | G | 114 (95.00) | 113 (94.17) | p = 1.00 Fisher’s exact test |
| TT | 6 (5.00) | 7 (5.83) | ||
| T | 55 (45.83) | 59 (49.17) | p = 0.70 Fisher’s exact test | |
| GG | 65 (54.17) | 61 (50.83) | ||
| NTRK2 rs1439050 Polymorphism | Asthma Patients | Healthy Subjects | |||
|---|---|---|---|---|---|
| Plasma BDNF Concentration (pg/mL) Median (25%; 75%) | Statistical Analysis | Plasma BDNF Concentration (pg/mL), Median (25%; 75%) | Statistical Analysis | ||
| Genotypes | GG | 0.81 (0.59; 1.15) | p = 0.13 Kruskal-Wallis test | 0.59 (0.49; 0.94) | p = 0.42 Kruskal-Wallis test |
| GT | 0.97 (0.63; 1.37) | 0.59 (0.39; 0.84) | |||
| TT | 0.51 (0.41; 1.37) | 0.65 (0.30; 1.48) | |||
| Carriers | G | 0.91 (0.61; 1.22) | p = 0.23; U = 241.5 Mann-Whitney test | 0.59 (0.45; 0.90) | p = 0.52; U = 336.5 Mann-Whitney test |
| TT | 0.51 (0.41; 1.37) | 0.65 (0.30; 1.48) | |||
| T | 0.97 (0.58; 1.37) | p = 0.21; U = 1550 Mann-Whitney test | 0.60 (0.39; 0.88) | p = 0.36; U = 1623 Mann-Whitney test | |
| GG | 0.81 (0.59; 1.15) | 0.59 (0.49; 0.94) | |||
| BDNF Val66Met Polymorphism | Non-Severe Asthma Patients, n (%) | Severe Asthma Patients, n (%) | Statistical Analysis | |
|---|---|---|---|---|
| Genotypes | AA | 0 (0.00) | 3 (4.92) | p = 0.13; χ2 = 4.04 χ2-test |
| AG | 19 (32.20) | 24 (39.34) | ||
| GG | 40 (67.80) | 34 (55.74) | ||
| Alleles | A | 19 (16.10) | 30 (24.59) | p = 0.11 Fisher’s exact test |
| G | 99 (73.90) | 92 (75.41) | ||
| Carriers | A | 19 (32.20) | 27 (44.26) | p = 0.19 Fisher’s exact test |
| GG | 40 (67.80) | 34 (55.74) | ||
| G | 59 (100.00) | 58 (95.08) | p = 0.24 Fisher’s exact test | |
| AA | 0 (0.00) | 3 (4.92) | ||
| NTRK2 rs1439050 Polymorphism | Non-Severe Asthma Patients, n (%) | Severe Asthma Patients, n (%) | Statistical Analysis | |
|---|---|---|---|---|
| Genotypes | GG | 33 (55.93) | 32 (52.46) | p = 0.56; χ2 = 1.16 χ2-test |
| GT | 22 (37.29) | 27 (44.26) | ||
| TT | 4 (6.78) | 2 (3.28) | ||
| Alleles | G | 88 (74.58) | 91 (74.59) | p = 1.00 Fisher’s exact test |
| T | 30 (25.42) | 31 (25.41) | ||
| Carriers | G | 55 (93.22) | 59 (96.72) | p = 0.43 Fisher’s exact test |
| TT | 4 (6.78) | 2 (3.28) | ||
| T | 26 (44.07) | 29 (47.54) | p = 0.72 Fisher’s exact test | |
| GG | 33 (55.93) | 32 (52.46) | ||
| Clinical Characteristics | Median (25%; 75%) | Plasma BDNF Concentration (pg/mL) |
|---|---|---|
| Total Serum IgE (IU/mL) | 191.00 (43.00; 398.00) | p = 0.50; Spearman correlation r = −0.06 |
| Blood eosinophils (×109/L) | 0.215 (0.10; 0.40) | p = 0.69; Spearman correlation r = 0.04 |
| Blood neutrophils (×109/L) | 4.560 (3.62; 6.30) | p = 0.92; Spearman correlation r = −0.01 |
| FeNO (ppb) | 34.00 (14.28; 67.75) | p = 0.98; Spearman correlation r = −0.002 |
| FEV1 (% of predicted value) | 74.90 (54.28; 90.08) | p = 0.41; Spearman correlation r = 0.07 |
| FVC (% of predicted value) | 88.50 (75.08; 101.80) | p = 0.65; Spearman correlation r = −0.04 |
| PEF (% of predicted value) | 77.05 (57.25; 95.52) | p = 0.58; Spearman correlation r = −0.05 |
| DLCO (%) | 83.30 (73.68; 95.35) | p = 0.055: Spearman correlation r = −0.17 |
| Duration of disease (years) | 13.50 (8.00; 29.00) | p = 0.023; Spearman correlation r = 0.21 |
| Comorbidities (n) | 2.00 (1.25; 4.00) | p = 0.35; Spearman correlation r = 0.08 |
| Clinical Characteristics | n (%) | Plasma BDNF Concentration (pg/mL) |
| Penicillin allergy | 25 (20.83) | p = 0.18; U = 977.50; Mann-Whitney test |
| Nutritive allergy | 12 (10.00) | p = 0.10; U = 458.50; Mann-Whitney test |
| Animal dander/feather allergy | 18 (15.00) | p = 0.31; U = 779.00; Mann-Whitney test |
| Dust allergy | 51 (42.50) | p = 0.06; U = 1408.00; Mann-Whitney test |
| Pollen allergy | 53 (44.17) | p = 0.72; U = 1709.00; Mann-Whitney test |
| Fungal/mold allergy | 9 (7.50) | p = 0.28; U = 390.50; Mann-Whitney test |
| Early onset asthma (age < 12 years) | 21 (17.50) | p = 0.12; U = 817.00; Mann-Whitney test |
| History of pneumonia | 21 (17.50) | p = 0.026; U = 718.00; Mann-Whitney test |
| Emergency intervention (ever) | 89 (74.17) | p = 0.53; U = 1274.00; Mann-Whitney test |
| Hospitalization for asthma (ever) | 58 (48.33) | p = 0.70; U = 1724.00; Mann-Whitney test |
| Nasal polyps | 27 (22.50) | p = 0.025; U = 891.00; Mann-Whitney test |
| Aspirin sensitivity | 11 (9.17) | p = 0.009; U = 314.00; Mann-Whitney test |
| Allergen specific immunotherapy | 15 (12.50) | p = 0.99; U = 787.00; Mann-Whitney test |
| Oral corticosteroid therapy | 30 (25.00) | p = 0.58; U = 1258.00; Mann-Whitney test |
| Biological therapy | 20 (16.67) | p = 0.80; U = 962.50; Mann-Whitney test |
| Asthma Phenotypes | Plasma BDNF Concentration (pg/mL) Median (25%, 75%) | Statistical Analysis |
|---|---|---|
| T2-high (n = 94) | 0.91 (0.60; 1.30) | p = 0.43; U = 1097.00 Mann-Whitney test |
| T2-low (n = 26) | 0.82 (0.42; 1.14) | |
| Non-allergic (n = 42) | 0.98 (0.60; 1.29) | p = 0.41; U = 1487.00 Mann-Whitney test |
| Allergic (n = 78) | 0.82 (0.58; 1.20) | |
| Non-eosinophilic (n = 73) | 0.92 (0.54; 1.20) | p = 0.54; U = 1602.00 Mann-Whitney test |
| Eosinophilic (n = 47) | 0.88 (0.61; 1.34) | |
| Non-AERD (n = 111) | 0.85 (0.58; 1.18) | p = 0.017; U = 262.00 Mann-Whitney test |
| AERD (n = 9) | 1.22 (0.92; 2.05) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sreter, K.B.; Popovic-Grle, S.; Lampalo, M.; Konjevod, M.; Tudor, L.; Nikolac Perkovic, M.; Jukic, I.; Bingulac-Popovic, J.; Safic Stanic, H.; Markeljevic, J.; et al. Plasma Brain-Derived Neurotrophic Factor (BDNF) Concentration and BDNF/TrkB Gene Polymorphisms in Croatian Adults with Asthma. J. Pers. Med. 2020, 10, 189. https://doi.org/10.3390/jpm10040189
Sreter KB, Popovic-Grle S, Lampalo M, Konjevod M, Tudor L, Nikolac Perkovic M, Jukic I, Bingulac-Popovic J, Safic Stanic H, Markeljevic J, et al. Plasma Brain-Derived Neurotrophic Factor (BDNF) Concentration and BDNF/TrkB Gene Polymorphisms in Croatian Adults with Asthma. Journal of Personalized Medicine. 2020; 10(4):189. https://doi.org/10.3390/jpm10040189
Chicago/Turabian StyleSreter, Katherina B., Sanja Popovic-Grle, Marina Lampalo, Marcela Konjevod, Lucija Tudor, Matea Nikolac Perkovic, Irena Jukic, Jasna Bingulac-Popovic, Hana Safic Stanic, Jasenka Markeljevic, and et al. 2020. "Plasma Brain-Derived Neurotrophic Factor (BDNF) Concentration and BDNF/TrkB Gene Polymorphisms in Croatian Adults with Asthma" Journal of Personalized Medicine 10, no. 4: 189. https://doi.org/10.3390/jpm10040189
APA StyleSreter, K. B., Popovic-Grle, S., Lampalo, M., Konjevod, M., Tudor, L., Nikolac Perkovic, M., Jukic, I., Bingulac-Popovic, J., Safic Stanic, H., Markeljevic, J., Pivac, N., & Svob Strac, D. (2020). Plasma Brain-Derived Neurotrophic Factor (BDNF) Concentration and BDNF/TrkB Gene Polymorphisms in Croatian Adults with Asthma. Journal of Personalized Medicine, 10(4), 189. https://doi.org/10.3390/jpm10040189







