Comparative Analysis of Third Molar Segmentation Performance Between Sexes Using Deep Learning Models
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design and Data Source
2.2. Image Review and Annotation
2.3. Inclusion and Exclusion Criteria
2.4. Dataset Split Structure and Class Distribution
2.5. Image Quality Control and Observer Reliability
2.6. Ethics Approval
2.7. Image Preprocessing and Tooth Segmentation
2.8. Feature Extraction
2.9. Model Development and Hyperparameters
2.10. Performance Evaluation Metrics
2.10.1. Precision
2.10.2. Sensitivity (Recall)
2.10.3. F1 Score
2.10.4. Mean Average Precision (mAP)
2.10.5. ROC Curves
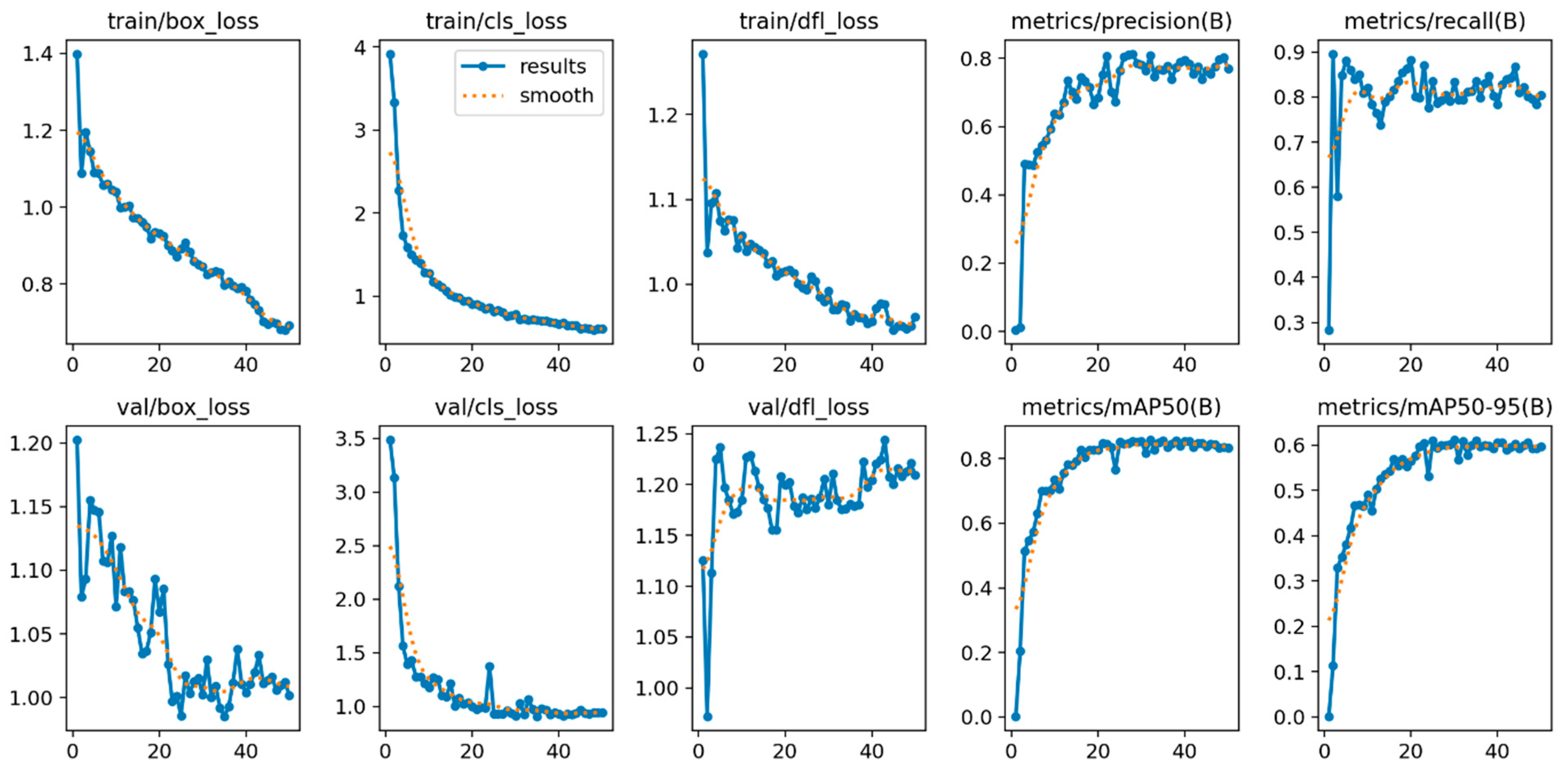
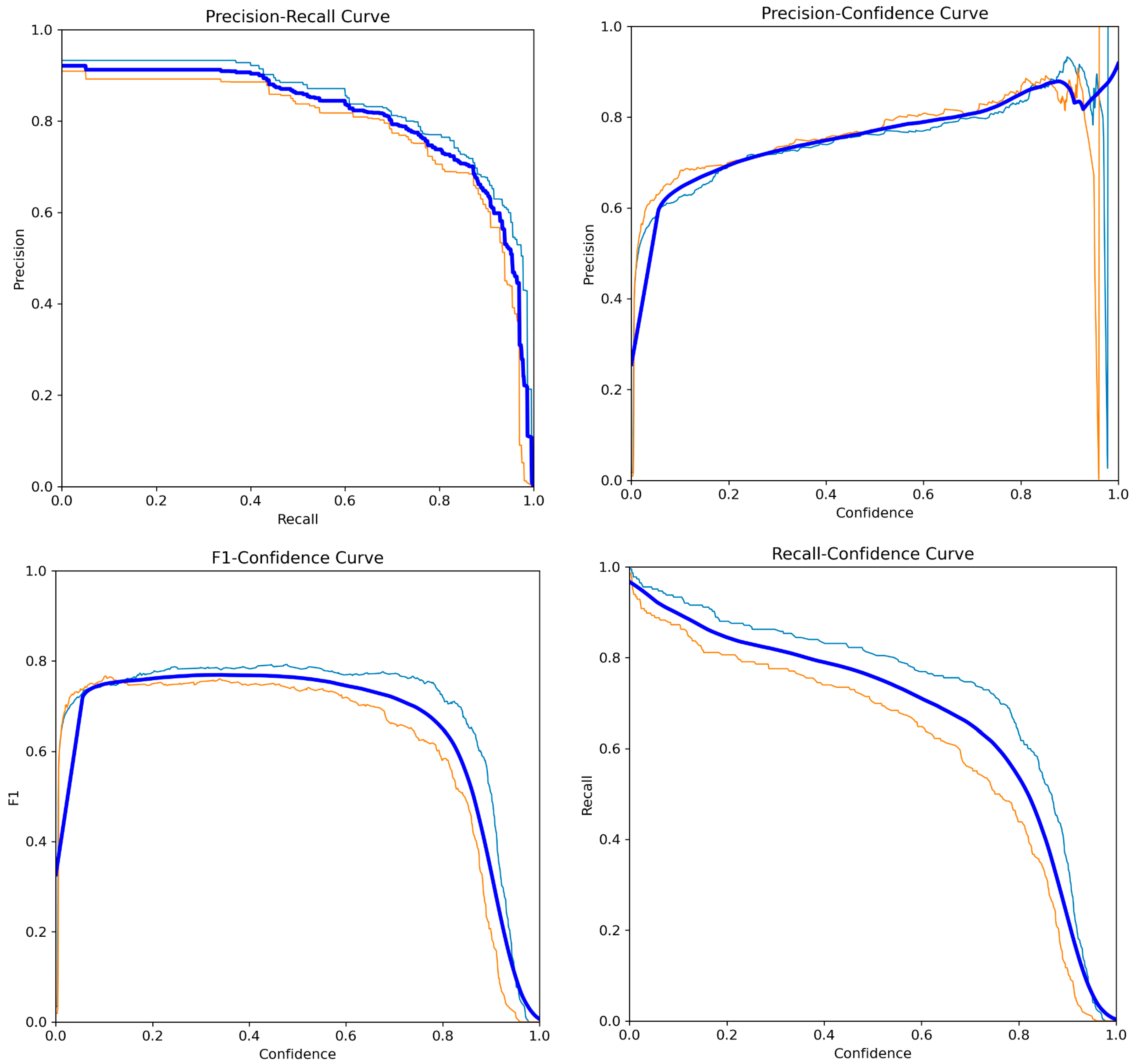
3. Results
Metric Analyses
4. Discussion
4.1. Comparison with Full-Dentition Classification Models
4.2. Influence of Region Restriction
4.3. Role of Segmentation vs. Classification Paradigms
4.4. Differential Performance by Sex
4.5. Population, Training Size, and Generalizability
4.6. Forensic Implications and Model Interpretability
4.7. Synthesis and Future Directions
4.8. Limitations
5. Conclusions
Future Work
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Park, S.J.; Yang, S.; Kim, J.M.; Kang, J.H.; Kim, J.E.; Huh, K.H.; Lee, S.S.; Yi, W.J.; Heo, M.S. Automatic and robust estimation of sex and chronological age from panoramic radiographs using a multi-task deep learning network: A study on a South Korean population. Int. J. Leg. Med. 2024, 138, 1741–1757. [Google Scholar] [CrossRef] [PubMed]
- Bu, W.Q.; Guo, Y.X.; Zhang, D.; Du, S.Y.; Han, M.Q.; Wu, Z.X.; Tang, Y.; Chen, T.; Guo, Y.C.; Meng, H.T. Automatic sex estimation using deep convolutional neural network based on orthopantomogram images. Forensic Sci. Int. 2023, 348, 111704. [Google Scholar] [CrossRef]
- Ciconelle, A.C.M.; da Silva, R.L.B.; Kim, J.H.; Rocha, B.A.; Dos Santos, D.G.; Vianna, L.G.R.; Gomes Ferreira, L.G.; Pereira dos Santos, V.H.; Costa, J.O.; Vicente, R. Deep learning for sex determination: Analyzing over 200,000 panoramic radiographs. J. Forensic Sci. 2023, 68, 2057–2064. [Google Scholar] [CrossRef] [PubMed]
- Patil, V.; Vineetha, R.; Vatsa, S.; Shetty, D.K.; Raju, A.; Naik, N.; Malarout, N. Artificial neural network for gender determination using mandibular morphometric parameters: A comparative retrospective study. Cogent Eng. 2020, 7, 1723783. [Google Scholar] [CrossRef]
- Márquez-Ruiz, A.B.; González-Herrera, L.; Luna, J.D.D.; Valenzuela, A. Radiologic assessment of third molar development and cervical vertebral maturation to validate age of majority in a Mexican population. Int. J. Leg. Med. 2025, 139, 1673–1680. [Google Scholar] [CrossRef]
- Şahin, T.N.; Kölüş, T. Age and sex estimation in children and young adults using panoramic radiographs with convolutional neural networks. Appl. Sci. 2024, 14, 7014. [Google Scholar] [CrossRef]
- Schwartz, G.T.; Dean, M.C. Sexual dimorphism in modern human permanent teeth. Am. J. Phys. Anthropol. Off. Publ. Am. Assoc. Phys. Anthropol. 2005, 128, 312–317. [Google Scholar] [CrossRef]
- Orhan, K.; Ozer, L.; Orhan, A.I.; Dogan, S.; Paksoy, C.S. Radiographic evaluation of third molar development in relation to chronological age among Turkish children and youth. Forensic Sci. Int. 2007, 165, 46–51. [Google Scholar] [CrossRef]
- Levesque, G.Y.; Demirjian, A.; Tanguay, R. Sexual dimorphism in the development, emergence, and agenesis of the mandibular third molar. J. Dent. Res. 1981, 60, 1735–1741. [Google Scholar] [CrossRef] [PubMed]
- Liversidge, H.M.; Marsden, P.H. Estimating age and the likelihood of having attained 18 years of age using mandibular third molars. Br. Dent. J. 2010, 209, E13. [Google Scholar] [CrossRef]
- Liversidge, H.M.; Peariasamy, K.; Folayan, M.O.; Adeniyi, A.O.; Ngom, P.I.; Mikami, Y.; Shimada, Y.; Kuroe, K.; Tvete, I.F.; Kvaal, S.I. A radiographic study of the mandibular third molar root development in different ethnic groups. J. Forensic Odonto-Stomatol. 2017, 35, 97–108. [Google Scholar]
- Gunst, K.; Mesotten, K.; Carbonez, A.; Willems, G. Third molar root development and its relationship to chronological age and sex. Forensic Sci. Int. 2003, 136, 52–57. [Google Scholar] [CrossRef] [PubMed]
- Dashti, M.; Azimi, T.; Khosraviani, F.; Azimian, S.; Bahanan, L.; Zahmatkesh, H.; Ashi, H.; Khurshid, Z. Systematic review and meta-analysis on the accuracy of artificial intelligence algorithms in individuals gender detection using orthopantomograms. Int. Dent. J. 2025, 75, 2157–2168. [Google Scholar] [CrossRef] [PubMed]
- Franco, A.; Porto, L.; Heng, D.; Murray, J.; Lygate, A.; Franco, R.; Bueno, J.; Sobania, M.; Costa, M.M.; Paranhos, L.R.; et al. Diagnostic performance of convolutional neural networks for dental sexual dimorphism. Sci. Rep. 2022, 12, 17279. [Google Scholar] [CrossRef]
- Abdou, M.A. Literature review: Efficient deep neural networks techniques for medical image analysis. Neural Comput. Appl. 2022, 34, 5791–5812. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. In Proceedings of the 6th International Conference on Learning Representations, San Diego, CA, USA, 7–9 May 2015; pp. 1–14. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Huang, G.; Liu, Z.; van der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 2261–2269. [Google Scholar]
- Ren, S.; He, K.; Girshick, R.; Sun, J. Faster R-CNN: Towards real-time object detection with region proposal networks. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 1137–1149. [Google Scholar] [CrossRef]
- Redmon, J.; Divvala, S.; Girshick, R.; Farhadi, A. You only look once: Unified, real-time object detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 779–788. [Google Scholar]
- Hussain, S.; Guo, F.; Li, W.; Shen, Z. DilUnet: A U-net based architecture for blood vessels segmentation. Comput. Methods Programs Biomed. 2022, 218, 106732. [Google Scholar] [CrossRef]
- Ibraheem, W.I.; Jain, S.; Ayoub, M.N.; Namazi, M.A.; Alfaqih, A.I.; Aggarwal, A.; Meshni, A.A.; Almarghlani, A.; Alhumaidan, A.A. Assessment of the Diagnostic Accuracy of Artificial Intelligence Software in Identifying Common Periodontal and Restorative Dental Conditions (Marginal Bone Loss, Periapical Lesion, Crown, Restoration, Dental Caries) in Intraoral Periapical Radiographs. Diagnostics 2025, 15, 1432. [Google Scholar] [CrossRef]
- Guldiken, I.N.; Tekin, A.; Kanbak, T.; Kahraman, E.N.; Özcan, M. Prognostic Evaluation of Lower Third Molar Eruption Status from Panoramic Radiographs Using Artificial Intelligence-Supported Machine and Deep Learning Models. Bioengineering 2025, 12, 1176. [Google Scholar] [CrossRef]
- Ataş, I. Human gender prediction based on deep transfer learning from panoramic radiograph images. arXiv 2022, arXiv:2205.09850. [Google Scholar] [CrossRef]
- Küchler, E.C.; Kirschneck, C.; Marañón-Vásquez, G.A.; Schroder, Â.G.D.; Baratto-Filho, F.; Romano, F.L.; Stuani, M.B.S.; Matsumoto, M.A.N.; de Araujo, C.M. Mandibular and dental measurements for sex determination using machine learning. Sci. Rep. 2024, 14, 9587. [Google Scholar] [CrossRef]
- RoboFlow. “RoboFlow Object Detection Model” RoboFlow. 2025. Available online: https://www.roboflow.com/ (accessed on 20 October 2025).
- Ardelean, A.I.; Ardelean, E.R.; Marginean, A. Can YOLO detect retinal pathologies? A step towards automated OCT analysis. Diagnostics 2025, 15, 1823. [Google Scholar] [CrossRef]
- Khanam, R.; Hussain, M. A Review of YOLOv12: Attention-Based Enhancements vs. Previous Versions. arXiv 2025, arXiv:2504.11995. [Google Scholar] [CrossRef]
- Zhao, Y.; Lv, W.; Xu, S.; Wei, J.; Wang, G.; Dang, Q.; Liu, Y.; Chen, J. Detrs beat yolos on real-time object detection. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Seattle, WA, USA, 17–21 June 2024; pp. 16965–16974. [Google Scholar]
- Lv, W.; Zhao, Y.; Chang, Q.; Huang, K.; Wang, G.; Liu, Y. Rt-detrv2: Improved baseline with bag-of-freebies for real-time detection transformer. arXiv 2024, arXiv:2407.17140. [Google Scholar]
- Esmaeilyfard, R.; Paknahad, M.; Dokohaki, S. Sex classification of first molar teeth in cone beam computed tomography images using data mining. Forensic Sci. Int. 2021, 318, 110633. [Google Scholar] [CrossRef] [PubMed]
- Hemalatha, B.; Bhuvaneswari, P.; Nataraj, M.; Shanmugavadivel, G. Human dental age and gender assessment from dental radiographs using deep convolutional neural network. Inf. Technol. Control 2023, 52, 322–335. [Google Scholar] [CrossRef]
- Hougaz, A.B.; Lima, D.; Peters, B.; Cury, P.; Oliveira, L. Sex estimation on panoramic dental radiographs: A methodological approach. In Simpósio Brasileiro de Computação Aplicada à Saúde (SBCAS); Sociedade Brasileira de Computação: Porto Alegre, Brazil, 2023; pp. 115–125. [Google Scholar]
- Ke, W.; Fan, F.; Liao, P.; Lai, Y.; Wu, Q.; Du, W.; Chen, H.; Deng, Z.; Zhang, Y. Biological gender estimation from panoramic dental x-ray images based on multiple feature fusion model. Sens. Imaging 2020, 21, 54. [Google Scholar] [CrossRef]
- Kim, I.; Yang, S.; Choi, Y.; Kwon, H.; Lee, C.; Park, W. Automated Sex and Age Estimation from Orthopantomograms Using Deep Learning: A Comparison with Human Predictions. Forensic Sci. Int. 2025, 374, 112531. [Google Scholar] [CrossRef]
- Kohinata, K.; Kitano, T.; Nishiyama, W.; Mori, M.; Iida, Y.; Fujita, H.; Katsumata, A. Deep learning for preliminary profiling of panoramic images. Oral Radiol. 2023, 39, 275–281. [Google Scholar] [CrossRef]
- Pertek, H.; Kamaşak, M.; Kotan, S.; Hatipoğlu, F.P.; Hatipoğlu, Ö.; Köse, T.E. Comparison of mandibular morphometric parameters in digital panoramic radiography in gender determination using machine learning. Oral Radiol. 2024, 40, 415–423. [Google Scholar] [CrossRef] [PubMed]
- Vila-Blanco, N.; Varas-Quintana, P.; Aneiros-Ardao, Á.; Tomás, I.; Carreira, M.J. XAS: Automatic yet eXplainable Age and Sex determination by combining imprecise per-tooth predictions. Comput. Biol. Med. 2022, 149, 106072. [Google Scholar] [CrossRef]
- Ortiz, A.G.; Costa, C.; Silva, R.H.A.; Biazevic, M.G.H.; Michel-Crosato, E. Sex estimation: Anatomical references on panoramic radio- graphs using machine learning. Forensic Imaging 2020, 20, 200356. [Google Scholar] [CrossRef]
- Arian, M.S.H.; Rakib, M.T.A.; Ali, S.; Ahmed, S.; Farook, T.H.; Mohammed, N.; Dudley, J. Pseudo labelling workflow, margin losses, hard triplet mining, and PENViT backbone for explainable age and biological gender estimation using dental pan-oramic radiographs. SN Appl. Sci. 2023, 5, 279. [Google Scholar] [CrossRef]
- Ilic, I.; Vodanovic, M.; Subasic, M. Gender estimation from pan-oramic dental X-ray images using deep convolutional networks. In Proceedings of the IEEE EUROCON 2019-18th International Conference on Smart Technologies, Novi Sad, Serbia, 1–4 July 2019. [Google Scholar]
- Scavassini, L.; Silva, R.; Khan, A.; Anees, W.; Murray, J.; Angelakopoulos, N.; Arakelyan, M.; Porto, L.; Abade, A.; Franco, A. YOLO11m-cls applied to sex and age classification based on the radiographic analysis of the nasal aperture. Sci. Rep. 2025, 15, 40784. [Google Scholar] [CrossRef] [PubMed]
- Hirunchavarod, N.; Dangsungnoen, L.; Thongprasant, K.; Phuphatham, P.; Prathansap, N.; Sributsayakarn, N.; Pornprasertsuk-Damrongsri, S.; Jirarattanasopha, V.; Intharah, T. OPG-SHAP: A Dental AI Tool for Explaining Learned Orthopantomogram Image Recognition. In Proceedings of the 2024 International Technical Conference on Circuits/Systems, Computers, and Communications (ITC-CSCC), Okinawa, Japan, 2–5 July 2024; pp. 1–6. [Google Scholar]
- Kp, N. Accuracy Assessment of Annotation Techniques for Gender Determination on Orthopantomogram. Int. Dent. J. 2025, 75, 106103. [Google Scholar] [CrossRef]
- Cameriere, R.; Ferrante, L.; De Angelis, D.; Scarpino, F.; Galli, F. The comparison between measurement of open apices of third molars and Demirjian stages to test chronological age of over 18 year olds in living subjects. Int. J. Leg. Med. 2008, 122, 493–497. [Google Scholar] [CrossRef]
- Olze, A.; Schmeling, A.; Taniguchi, M.; Maeda, H.; van Niekerk, P.; Wernecke, K.D.; Geserick, G. Forensic age estimation in living subjects: The ethnic factor in wisdom tooth mineralization. Int. J. Leg. Med. 2004, 118, 170–173. [Google Scholar] [CrossRef]


| Split | Total Patient | Female Labels | Male Labels | Female % | Male % |
|---|---|---|---|---|---|
| Train | 531 | 951 | 1010 | 48.50 | 51.50 |
| Valid | 115 | 226 | 210 | 51.83 | 48.17 |
| Test | 111 | 225 | 196 | 53.44 | 46.56 |
| Total | 757 | 1402 | 1416 | 49.75 | 50.25 |
| Feature | YOLOv12n | YOLO26n | RT-DETR v2 |
|---|---|---|---|
| Dimension | Nano | Nano | Large |
| Number of Parameters | ~2.5 M | ~2.5 M | ~32.8 M |
| Training Time | ~15 min | ~15 min | ~14 h |
| Architecture Type | CNN | CNN | CNN + Transformer |
| Model Family | YOLO | YOLO | Transformer-based DETR |
| Model Complexity | Low | Low–Medium | High |
| Feature Extraction | Convolutional | Convolutional | Self-attention |
| Computational Cost | Very low | Low | Very high |
| Model | Precision (P) | Recall® | mAP50 | mAP50–95 |
|---|---|---|---|---|
| YOLOv12n | 0.738 | 0.806 | 0.810 | 0.574 |
| RT-DETR v2 | 0.790 | 0.775 | 0.772 | 0.544 |
| YOLO26n | 0.616 | 0.831 | 0.778 | 0.561 |
| Performance (Mean) | 0.714 | 0.804 | 0.786 | 0.559 |
| Model | Predicted/True | Female | Male | Background |
|---|---|---|---|---|
| YOLOv12n | Female | 182 | 48 | 42 |
| Male | 38 | 140 | 40 | |
| Background | 5 | 8 | 0 | |
| RT-DETR v2 | Female | 173 | 55 | 50 |
| Male | 37 | 118 | 44 | |
| Background | 15 | 23 | 0 | |
| YOLO26n | Female | 191 | 49 | 13 |
| Male | 30 | 141 | 14 | |
| Background | 4 | 6 | 0 |
| Class | YOLOv12n (mAP50) | RT-DETR v2 (mAP50) | YOLO26n (mAP50) |
|---|---|---|---|
| Female | 0.837 | 0.766 | 0.806 |
| Male | 0.783 | 0.778 | 0.749 |
| References | Imaging | Tooth/Region | Task | Model/Method | Metric(s) | Key Outcome | Sensitivity | Specificity | Precision | F1-Score |
|---|---|---|---|---|---|---|---|---|---|---|
| Ataş I [24] | OPG | Dentition | Sex estimation | DenseNet121 | Accuracy | 97.25% | - | 96.80 | 96.80 | 97.25 |
| Esmaeilyfard et al. [31] | CBCT | Dentition | Sex estimation | NB | Accuracy | 92.31% | 91.23 | 92.01 | 89.83 | - |
| Hemalatha et al. [32] | OPG | Dentition | Sex estimation | Deep CNN | Accuracy | 91.70% | 100 | 85.70 | - | - |
| Hougaz et al. [33] | OPG | Dentition | Sex estimation | EfficientNet-B0 | Accuracy | - | - | - | - | 91.30% ± 0.47 |
| EfficientNet-B7 | - | - | - | - | 90.00% ± 0.01 | |||||
| EfficientNetV2-Small | - | - | - | - | 89.10% ± 0.67 | |||||
| EfficientNetV2-Large | - | - | - | - | 91.43% ± 0.67 | |||||
| Ke et al. [34] | OPG | Dentition | Sex estimation | VGG16 + MFF | Accuracy | 94.60 ± 0.58% | - | - | - | - |
| Patil et al. [4] | OPG | Dentition | Sex estimation | ANN | Accuracy | 75.00% | - | - | - | - |
| Kim et al. [35] | OPG | Dentition | Sex estimation | EfficientNetV2 | Accuracy | 90.20% | - | - | 93.10 | 90.40 |
| Kohinata et al. [36] | OPG | Dentition | Sex estimation | VGG-Net | Accuracy | 75.50% | - | - | - | - |
| Pertek et al. [37] | OPG | Dentition | Sex estimation | ANN | Accuracy | 83.00 ± 1.90% | - | - | - | - |
| Vila-Blanco et al. [38] | OPG | Dentition | Sex estimation | Two-path CNN(female/male) | Accuracy | 91.82%/89.09% | - | - | 89.65/94.23 | - |
| Franco et al. [14] | OPG | Dentomaxillofacial | Sex estimation | DenseNet121 (TL vs. FS) | Acc/AUC | TL superior 82.18%, AUC 0.91 | - | 92.20 | 80.72 | 80.64 |
| Ortiz et al. [39] | OPG | Dentition | Sex estimation | Neural Network | Accuracy | 89.10% | - | - | - | 79.00 |
| Arian et al. [40] | OPG | Dentition | Sex estimation | PENVIT | Accuracy | 84.49% | - | - | - | - |
| Bu et al. [2] | OPG | Dentition | Sex estimation | ResNeXt/EfficientNet/ViT | Acc/AUC | Acc 86.79%, AUC 90.64 | 86.75 | - | 92.27 | 89.42 |
| Ciconelle et al. [3] | OPG | Dentomaxillofacial | Sex estimation | CNN (ResNet) | Accuracy | Up to 95.22% | - | - | - | - |
| Ilic et al. [41] | OPG | Dentition | Sex estimation | Deep CNN | Accuracy | 94.30% | - | - | - | - |
| Park et al. [1] | OPG | Dentition | Sex estimation | ForensicNet (EffNet-B3 + CBAM) | Accuracy | 99.20% | 99.00 | 99.30 | - | - |
| Scavassini et al. [42] | OPG | Nasal aperture | Sex estimation | YOLO11m-cls | Accuracy | 74.00% | 74.00 | 74.00 | 74.00 | - |
| Hirunchavarod et al. [43] | OPG | Dentition | Oral part detection | YOLOv5/YOLOv8 | mAP@50 | 0.989/ 0.989 | - | - | 98.60/98.60 | 0.987/ 0.985 |
| Kp N [44] | OPG | Specific anatomical landmarks | Sex estimation | YOLO-NAS | mAP | 51.55% | - | - | - | - |
| Roboflow 3.0 | mAP | 57.90% | - | - | 85.40 | - | ||||
| YOLOv8 | mAP | 63.60% | - | - | 85.10 | - | ||||
| Present study | OPG | Only 3rd molars | Sex estimation | YOLOv12 | mAP@50/mAP@50–95 | 0.810/0.574 | - | - | 0.738 | - |
| YOLO26 | 0.778/0.561 | 0.616 | ||||||||
| RT-DETR v2 | 0.772/0.544 | 0.790 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Bulut, A.; Aşkın, M.B.; Çınarer, G. Comparative Analysis of Third Molar Segmentation Performance Between Sexes Using Deep Learning Models. Diagnostics 2026, 16, 977. https://doi.org/10.3390/diagnostics16070977
Bulut A, Aşkın MB, Çınarer G. Comparative Analysis of Third Molar Segmentation Performance Between Sexes Using Deep Learning Models. Diagnostics. 2026; 16(7):977. https://doi.org/10.3390/diagnostics16070977
Chicago/Turabian StyleBulut, Ayşe, Melis Büşra Aşkın, and Gökalp Çınarer. 2026. "Comparative Analysis of Third Molar Segmentation Performance Between Sexes Using Deep Learning Models" Diagnostics 16, no. 7: 977. https://doi.org/10.3390/diagnostics16070977
APA StyleBulut, A., Aşkın, M. B., & Çınarer, G. (2026). Comparative Analysis of Third Molar Segmentation Performance Between Sexes Using Deep Learning Models. Diagnostics, 16(7), 977. https://doi.org/10.3390/diagnostics16070977





