Divergent Evolution of Tuberculosis Lesions During Treatment: A Longitudinal CT-Based Analysis of Progression and Regression Patterns
Abstract
1. Introduction
2. Methods
2.1. Patients and CT Series
2.2. CT Segmentation and Image Annotations
2.3. Longitudinal Tracking of Lesion Volume
- (1)
- Stable: Lesion volume remained consistent across all scans, with no significant enlargement or reduction, and all variations were attributed to measurement error.
- (2)
- Decrease: Lesions showing a significant reduction in volume between two CT scans. For lesions with three or more scans, there was an overall decline in volume that could include stable periods but no instances of volume increase.
- (3)
- Increase: Lesions showing a significant increase in volume between two CT scans. For lesions with three or more scans, an overall enlargement was observed that could include stable periods but no reduction in volume. This category also included new lesions appearing after the initial CT scan that either enlarged or remained stable without shrinking.
- (4)
- Mix-I-D: A fluctuating pattern observed only in lesions with more than two scans, in which the lesion volume first increased and then decreased. Periods of stability between some CT scans were permitted.
- (5)
- Mix-D-I: A fluctuating pattern observed only in lesions with more than two scans, in which the lesion volume first decreased and then increased. Periods of stability between some CT scans were permitted.
2.4. Statistical Analyses
2.5. Patient and Public Involvement
3. Results
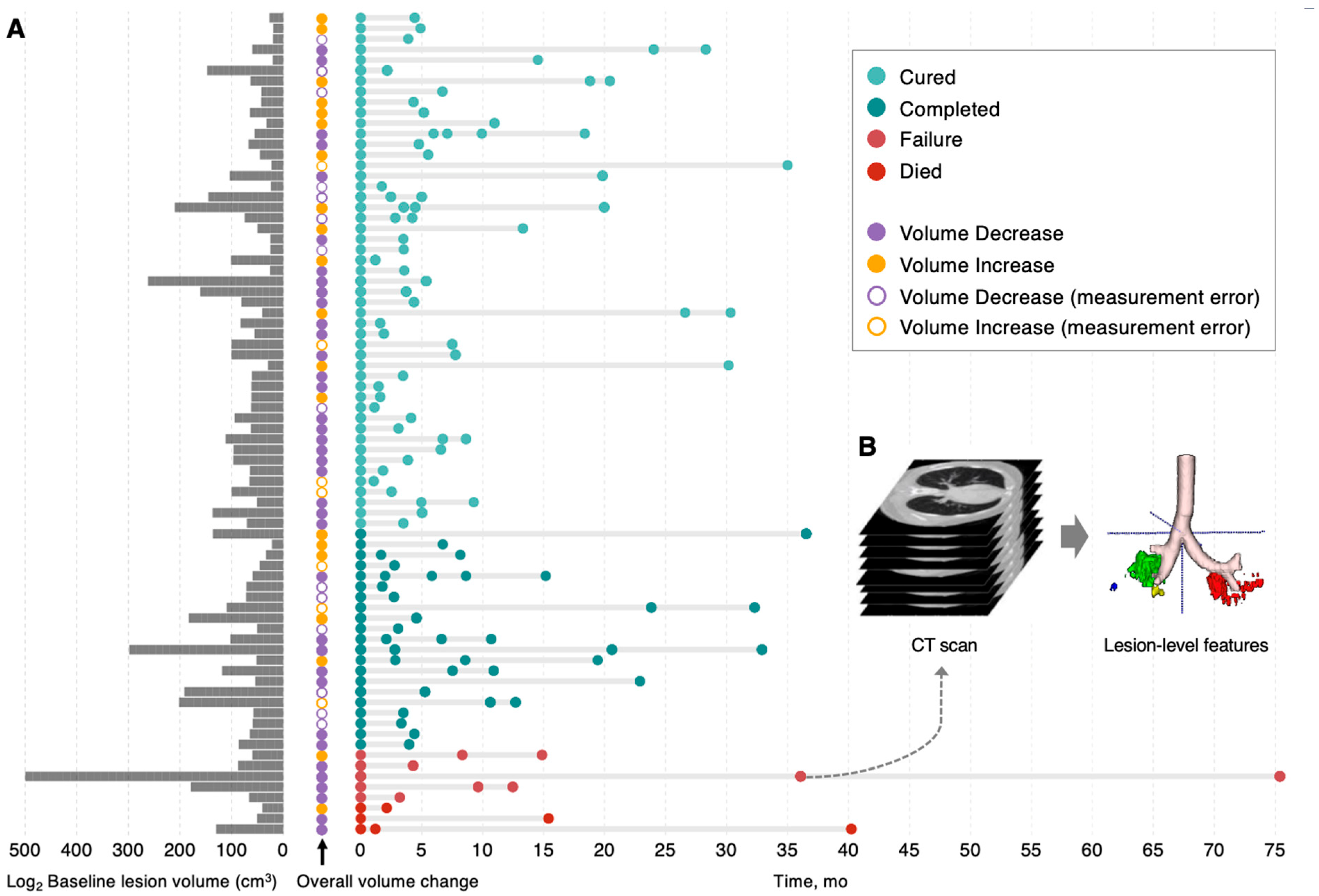
3.1. Patients, CT Scans, and Lesion Capture
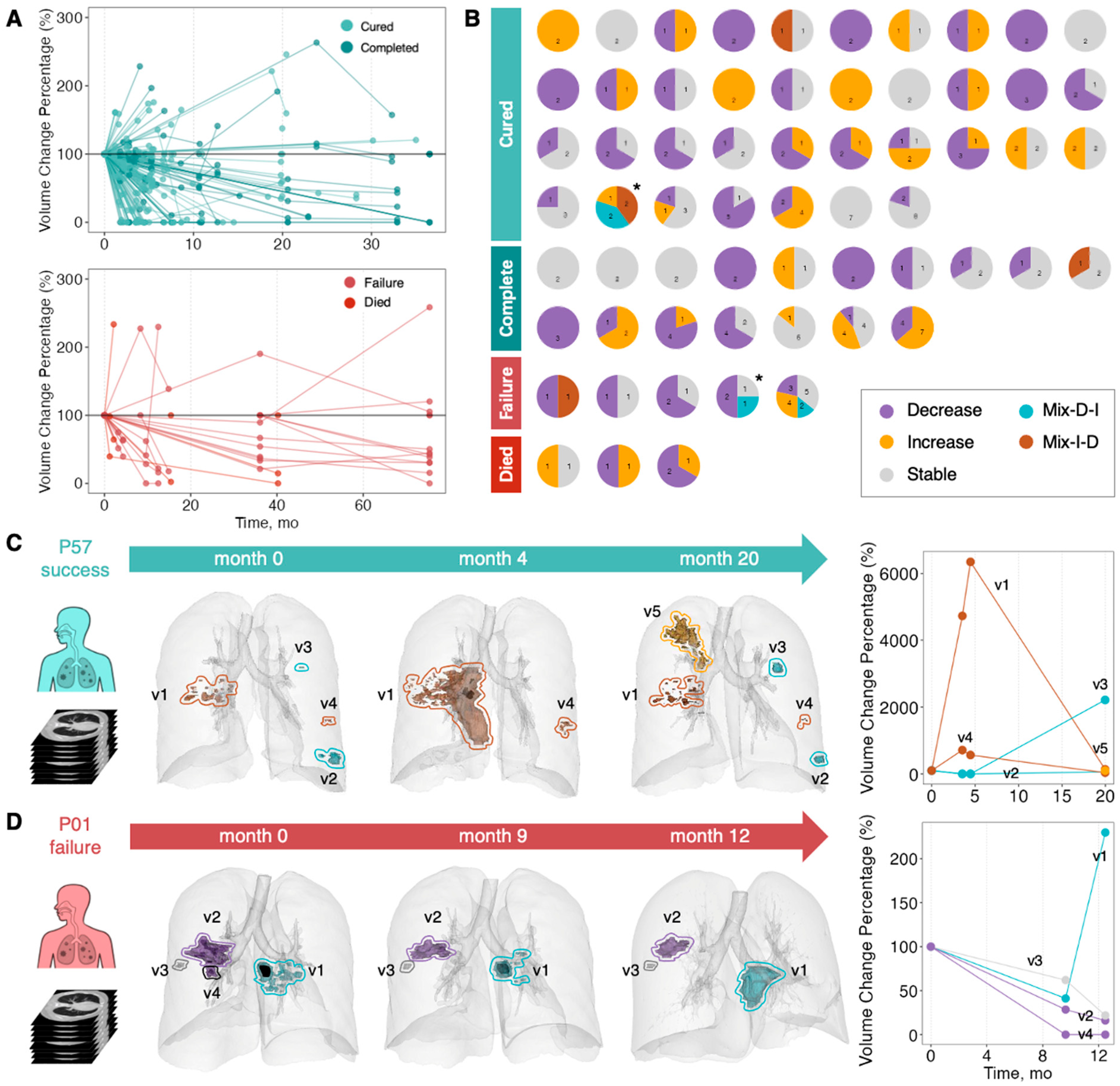
3.2. Disparate Evolutionary Trajectories of Different Lesions Within PTB Patients
3.3. Longitudinal Characteristics of Lesion Changes May Be Associated with Patient Treatment Outcomes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Global Tuberculosis Report 2024; WHO Press: Geneva, Switzerland, 2024. [Google Scholar]
- Wells, G.; Glasgow, J.N.; Nargan, K.; Lumamba, K.; Madansein, R.; Maharaj, K.; Perumal, L.Y.; Matthew, M.; Hunter, R.L.; Pacl, H.; et al. A high-resolution 3D atlas of the spectrum of tuberculous and COVID-19 lung lesions. EMBO Mol. Med. 2022, 14, e16283. [Google Scholar] [CrossRef]
- Lavin, R.C.; Tan, S. Spatial relationships of intra-lesion heterogeneity in Mycobacterium tuberculosis microenvironment, replication status, and drug efficacy. PLoS Pathog. 2022, 18, e1010459. [Google Scholar] [CrossRef]
- Donovan, M.L.; Bielefeldt-Ohmann, H.; Rollo, R.F.; McPherson, S.J.; Schultz, T.E.; Mori, G.; Kling, J.C.; Blumenthal, A. Distinct contributions of the innate immune receptors TLR2 and RP105 to formation and architecture of structured lung granulomas in mice infected with Mycobacterium tuberculosis. Immunology 2023, 169, 13–26. [Google Scholar] [CrossRef] [PubMed]
- Ordonez, A.A.; Wang, H.; Magombedze, G.; Ruiz-Bedoya, C.A.; Srivastava, S.; Chen, A.; Tucker, E.W.; Urbanowski, M.E.; Pieterse, L.; Fabian Cardozo, E.; et al. Dynamic imaging in patients with tuberculosis reveals heterogeneous drug exposures in pulmonary lesions. Nat. Med. 2020, 26, 529–534. [Google Scholar] [CrossRef] [PubMed]
- Lin, P.L.; Ford, C.B.; Coleman, M.T.; Myers, A.J.; Gawande, R.; Ioerger, T.; Sacchettini, J.; Fortune, S.M.; Flynn, J.L. Sterilization of granulomas is common in active and latent tuberculosis despite within-host variability in bacterial killing. Nat. Med. 2014, 20, 75–79. [Google Scholar] [CrossRef]
- Lenaerts, A.; Barry, C.E., 3rd; Dartois, V. Heterogeneity in tuberculosis pathology, microenvironments and therapeutic responses. Immunol. Rev. 2015, 264, 288–307. [Google Scholar] [CrossRef]
- Martin, C.J.; Cadena, A.M.; Leung, V.W.; Lin, P.L.; Maiello, P.; Hicks, N.; Chase, M.R.; Flynn, J.L.; Fortune, S.M. Digitally Barcoding Mycobacterium tuberculosis Reveals In Vivo Infection Dynamics in the Macaque Model of Tuberculosis. mBio 2017, 8, 10-1128. [Google Scholar] [CrossRef] [PubMed]
- Gideon, H.P.; Hughes, T.K.; Tzouanas, C.N.; Wadsworth, M.H.; Tu, A.A.; Gierahn, T.M.; Peters, J.M.; Hopkins, F.F.; Wei, J.R.; Kummerlowe, C.; et al. Multimodal profiling of lung granulomas in macaques reveals cellular correlates of tuberculosis control. Immunity 2022, 55, 827–846.e10. [Google Scholar] [CrossRef]
- Lin, P.L.; Coleman, T.; Carney, J.P.; Lopresti, B.J.; Tomko, J.; Fillmore, D.; Dartois, V.; Scanga, C.; Frye, L.J.; Janssen, C.; et al. Radiologic Responses in Cynomolgus Macaques for Assessing Tuberculosis Chemotherapy Regimens. Antimicrob. Agents Chemother. 2013, 57, 4237–4244. [Google Scholar] [CrossRef]
- Skoura, E.; Zumla, A.; Bomanji, J. Imaging in tuberculosis. Int. J. Infect. Dis. 2015, 32, 87–93. [Google Scholar] [CrossRef]
- Malherbe, S.T.; Chen, R.Y.; Dupont, P.; Kant, I.; Kriel, M.; Loxton, A.G.; Smith, B.; Beltran, C.G.; Van Zyl, S.; McAnda, S.; et al. Quantitative 18F-FDG PET-CT scan characteristics correlate with tuberculosis treatment response. EJNMMI Res. 2020, 10, 8. [Google Scholar] [CrossRef] [PubMed]
- Malherbe, S.T.; Shenai, S.; Ronacher, K.; Loxton, A.G.; Dolganov, G.; Kriel, M.; Van, T.; Chen, R.Y.; Warwick, J.; Via, L.E.; et al. Persisting positron emission tomography lesion activity and Mycobacterium tuberculosis mRNA after tuberculosis cure. Nat. Med. 2016, 22, 1094–1100. [Google Scholar] [CrossRef] [PubMed]
- Mattila, J.T.; Gould, V.A.; Junecko, B.A.; Bellavia, M.C.; Borish, H.J.; White, A.G.; Maiello, P.; Nyiranshuti, L.; Lin, P.L.; Scanga, C.A.; et al. Bacteria load and hypoxia contribute to glucose uptake by macrophages and T cells in cynomolgus macaque granulomas. J. Immunol. 2022, 208, 50.22. [Google Scholar] [CrossRef]
- Chen, R.Y.; Dodd, L.E.; Lee, M.; Paripati, P.; Hammoud, D.A.; Mountz, J.M.; Jeon, D.; Zia, N.; Zahiri, H.; Coleman, M.T.; et al. PET/CT imaging correlates with treatment outcome in patients with multidrug-resistant tuberculosis. Sci. Transl. Med. 2014, 6, 265ra166. [Google Scholar] [CrossRef]
- Jiang, H.; Zhu, C.; Qin, L.; Wu, X.; Yin, J.; Guo, Y.; Ma, H.; Jia, J.; Huo, F.; Xue, Y.; et al. Case report: A 9-year systematic treatment failure of a pulmonary tuberculosis patient. Front. Public Health 2022, 10, 966891. [Google Scholar] [CrossRef]
- Liu, Q.; Via, L.E.; Luo, T.; Liang, L.; Liu, X.; Wu, S.; Shen, Q.; Wei, W.; Ruan, X.; Yuan, X.; et al. Within patient microevolution of Mycobacterium tuberculosis correlates with heterogeneous responses to treatment. Sci. Rep. 2015, 5, 17507. [Google Scholar] [CrossRef]
- Rosenthal, A.; Gabrielian, A.; Engle, E.; Hurt, D.E.; Alexandru, S.; Crudu, V.; Sergueev, E.; Kirichenko, V.; Lapitskii, V.; Snezhko, E.; et al. The TB Portals: An Open-Access, Web-Based Platform for Global Drug-Resistant-Tuberculosis Data Sharing and Analysis. J. Clin. Microbiol. 2017, 55, 3267–3282. [Google Scholar] [CrossRef]
- Sambarey, A.; Smith, K.; Chung, C.; Arora, H.S.; Yang, Z.; Agarwal, P.P.; Chandrasekaran, S. Integrative analysis of multimodal patient data identifies personalized predictors of tuberculosis treatment prognosis. iScience 2024, 27, 109025. [Google Scholar] [CrossRef]
- Goodman, L.R.; Gulsun, M.; Washington, L.; Nagy, P.G.; Piacsek, K.L. Inherent variability of CT lung nodule measurements in vivo using semiautomated volumetric measurements. AJR Am. J. Roentgenol. 2006, 186, 989–994. [Google Scholar] [CrossRef]
- Bartlett, E.C.; Kemp, S.V.; Rawal, B.; Devaraj, A. Defining growth in small pulmonary nodules using volumetry: Results from a “coffee-break” CT study and implications for current nodule management guidelines. Eur. Radiol. 2021, 32, 1912–1920. [Google Scholar] [CrossRef] [PubMed]
- Gietema, H.A.; Schaefer-Prokop, C.M.; Mali, W.P.; Groenewegen, G.; Prokop, M. Pulmonary nodules: Interscan variability of semiautomated volume measurements with multisection CT—influence of inspiration level, nodule size, and segmentation performance. Radiology 2007, 245, 888–894. [Google Scholar] [CrossRef] [PubMed]
- Ingram, S.W.; Yang, J.; Wendt, R., 3rd; Beadle, B.M.; Rao, A.; Wang, X.A.; Court, L.E. The influence of non-rigid anatomy and patient positioning on endoscopy-CT image registration in the head and neck. Med. Phys. 2017, 44, 4159–4168. [Google Scholar] [PubMed]
- Haarburger, C.; Müller-Franzes, G.; Weninger, L.; Kuhl, C.; Truhn, D.; Merhof, D. Radiomics feature reproducibility under inter-rater variability in segmentations of CT images. Sci. Rep. 2020, 10, 12688. [Google Scholar] [CrossRef] [PubMed]
- Heinze, G.; Schemper, M. A solution to the problem of separation in logistic regression. Stat. Med. 2002, 21, 2409–2419. [Google Scholar] [CrossRef]
- Puhr, R.; Heinze, G.; Nold, M.; Lusa, L.; Geroldinger, A. Firth’s logistic regression with rare events: Accurate effect estimates and predictions? Stat. Med. 2017, 36, 2302–2317. [Google Scholar] [CrossRef]
- Cadena, A.M.; Fortune, S.M.; Flynn, J.L. Heterogeneity in tuberculosis. Nat. Rev. Immunol. 2017, 17, 691–702. [Google Scholar] [CrossRef]
- Chen, Y.; Ji, L.; Liu, Q.; Li, J.; Hong, C.; Jiang, Q.; Gan, M.; Takiff, H.E.; Yu, W.; Tan, W.; et al. Lesion Heterogeneity and Long-Term Heteroresistance in Multidrug-Resistant Tuberculosis. J. Infect. Dis. 2021, 224, 889–893. [Google Scholar] [CrossRef]
- Urbanowski, M.E.; Ordonez, A.A.; Ruiz-Bedoya, C.A.; Jain, S.K.; Bishai, W.R. Cavitary tuberculosis: The gateway of disease transmission. Lancet Infect. Dis. 2020, 20, e117–e128. [Google Scholar] [CrossRef]
- Chen, R.Y.; Yu, X.; Smith, B.; Liu, X.; Gao, J.; Diacon, A.H.; Dawson, R.; Tameris, M.; Zhu, H.; Qu, Y.; et al. Radiological and functional evidence of the bronchial spread of tuberculosis: An observational analysis. Lancet Microbe 2021, 2, e518–e526. [Google Scholar] [CrossRef]
- Yoder, M.A.; Lamichhane, G.; Bishai, W.R. Cavitary pulmonary tuberculosis: The Holy Grail of disease transmission. Curr. Sci. 2004, 86, 74–81. [Google Scholar]
- Honeyborne, I.; McHugh, T.D.; Phillips, P.P.; Bannoo, S.; Bateson, A.; Carroll, N.; Perrin, F.M.; Ronacher, K.; Wright, L.; Van Helden, P.D.; et al. Molecular Bacterial Load Assay, a Culture-Free Biomarker for Rapid and Accurate Quantification of Sputum Mycobacterium tuberculosis Bacillary Load during Treatment. J. Clin. Microbiol. 2011, 49, 3905–3911. [Google Scholar] [CrossRef]
- Honeyborne, I.; McHugh, T.D.; Kuittinen, I.; Cichonska, A.; Evangelopoulos, D.; Ronacher, K.; Van Helden, P.D.; Gillespie, S.H.; Fernandez-Reyes, D.; Walzl, G.; et al. Profiling persistent tubercule bacilli from patient sputa during therapy predicts early drug efficacy. BMC Med. 2016, 14, 68. [Google Scholar] [CrossRef]
- Nimmo, C.; Brien, K.; Millard, J.; Grant, A.D.; Padayatchi, N.; Pym, A.S.; O’Donnell, M.; Goldstein, R.; Breuer, J.; Balloux, F. Dynamics of within-host Mycobacterium tuberculosis diversity and heteroresistance during treatment. EBioMedicine 2020, 55, 102747. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Yew, W.W.; Barer, M.R. Targeting Persisters for Tuberculosis Control. Antimicrob. Agents Chemother. 2012, 56, 2223–2230. [Google Scholar] [CrossRef] [PubMed]
- Chang, E.; Cavallo, K.; Behar, S.M. CD4 T cell dysfunction is associated with bacterial recrudescence during chronic tuberculosis. Nat. Commun. 2025, 16, 2636. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Kwak, N.; Yoon, S.H.; Park, N.; Kim, Y.R.; Lee, J.H.; Lee, J.Y.; Park, Y.; Kang, Y.A.; Kim, S.; et al. Artificial intelligence-based radiographic extent analysis to predict tuberculosis treatment outcomes: A multicenter cohort study. Sci. Rep. 2024, 14, 13162. [Google Scholar] [CrossRef]
- Song, Q.; Guo, X.; Zhang, L.; Yang, L.; Lu, X. New Approaches in the Classification and Prognosis of Sign Clusters on Pulmonary CT Images in Patients with Multidrug-Resistant Tuberculosis. Front. Microbiol. 2021, 12, 714617. [Google Scholar] [CrossRef]
- Nam, K.J.; Jeong, Y.J.; Kim, Y.D.; Kim, K.I.; Lee, J.W.; Park, H.K.; Hoseok, I. Chronic destructive pulmonary tuberculosis: Assessment of disease activity by computed tomography. Acta Radiol. 2012, 53, 1014–1019. [Google Scholar] [CrossRef]
- Tiwari, A.; Katiyar, A. Hybrid deep learning assisted chest X-ray image segmentation and classification for tuberculosis disease diagnosis. Intell. Decis. Technol. 2023, 18, 561–569. [Google Scholar] [CrossRef]
- Türk, F. RNGU-NET: A novel efficient approach in Segmenting Tuberculosis using chest X-Ray images. PeerJ Comput. Sci. 2024, 10, e1780. [Google Scholar] [CrossRef]
- Engle, E.; Gabrielian, A.; Long, A.; Hurt, D.E.; Rosenthal, A. Performance of Qure.ai automatic classifiers against a large annotated database of patients with diverse forms of tuberculosis. PLoS ONE 2020, 15, e0224445. [Google Scholar] [CrossRef] [PubMed]
- Liang, S.; Xu, X.; Yang, Z.; Du, Q.; Zhou, L.; Shao, J.; Guo, J.; Ying, B.; Li, W.; Wang, C. Deep learning for precise diagnosis and subtype triage of drug-resistant tuberculosis on chest computed tomography. MedComm 2024, 5, e487. [Google Scholar] [CrossRef] [PubMed]
- Duwairi, R.; Melhem, A.B. A deep learning-based framework for automatic detection of drug resistance in tuberculosis patients. Egypt. Inform. J. 2023, 24, 139–148. [Google Scholar] [CrossRef]
- Lv, X.; Li, Y.; Cai, B.; He, W.; Wang, R.; Chen, M.; Pan, J.; Hou, D. Utility of Machine Learning and Radiomics Based on Cavity for Predicting the Therapeutic Response of MDR-TB. Infect. Drug Resist. 2023, 16, 6893–6904. [Google Scholar] [CrossRef]
- Nijiati, M.; Guo, L.; Abulizi, A.; Fan, S.; Wubuli, A.; Tuersun, A.; Nijiati, P.; Xia, L.; Hong, K.; Zou, X. Deep learning and radiomics of longitudinal CT scans for early prediction of tuberculosis treatment outcomes. Eur. J. Radiol. 2023, 169, 111180. [Google Scholar] [CrossRef]
| Success (n = 70) | Failure (n = 8) | p | Univariable | Multivariable | |||||
|---|---|---|---|---|---|---|---|---|---|
| OR (95%CI) | p | OR (95%CI) | p | ||||||
| Demographic &. Clinical | |||||||||
| Age, years | |||||||||
| ≥35 | 48 | (68.6%) | 5 | (62.5%) | 0.706 | 0.73 (0.17–3.05) | 0.668 | - | |
| Male | 37 | (52.9%) | 6 | (75.0%) | 0.285 | 2.32 (0.50–10.74) | 0.261 | - | |
| Patient-Level Lesion Static Data | |||||||||
| Baseline total lesion | |||||||||
| Baseline total lesion number | 2.0 | (2.0–3.0) | 2.5 | (2.0–3.3) | 0.348 | 1.17 (0.94–1.45) | 0.188 | ||
| Baseline total lesion volume (cm3) * | 7.6 | (2.8–31.2) | 29.0 | (20.3–106.1) | 0.091 | 1.75 (1.06–2.89) | 0.021 | ||
| Baseline total active lesion | |||||||||
| Baseline total active lesion number | 2.0 | (1.0–2.8) | 2.0 | (1.8–3.0) | 0.478 | 1.27 (0.87–1.86) | 0.239 | ||
| Baseline total active lesion volume (cm3) * | 6.6 | (2.4–31.2) | 28.9 | (20.3–106.2) | 0.071 | 1.76 (1.06–2.92) | 0.019 | 1.71 (0.98—3.01) | 0.041 |
| The presence of cavities at baseline | 33 | (47.1%) | 5 | (62.5%) | 0.476 | 1.76 (0.43–7.27) | 0.430 | ||
| The number of cavities at baseline | 0.0 | (0.0–1.0) | 1.0 | (0.0–1.3) | 0.291 | 1.63 (0.81–3.27) | 0.194 | ||
| The presence of a specific primary lesion type | |||||||||
| Consolidation | 31 | (44.3%) | 6 | (75.0%) | 0.141 | 3.26 (0.70–15.09) | 0.111 | ||
| Nodule | 39 | (55.7%) | 4 | (50.0%) | 1.000 | 0.80 (0.20–3.20) | 0.749 | ||
| Cluster of nodules | 30 | (42.9%) | 3 | (37.5%) | 1.000 | 0.85 (0.20–3.50) | 0.815 | ||
| Tree in buds | 0 | (0.0%) | 2 | (25.0%) | 0.009 | 54.23 (2.35–1254.09) | 0.003 | ||
| Strand | 4 | (5.7%) | 0 | (0.0%) | 1.000 | 0.87 (0.04–17.60) | 0.926 | ||
| Atelectasis | 1 | (1.4%) | 0 | (0.0%) | 1.000 | 2.73 (0.10–72.36) | 0.582 | ||
| Presence of satellite lesion | 64 | (91.4%) | 6 | (75.0%) | 0.188 | 0.26 (0.05–1.39) | 0.141 | ||
| The presence of a specific satellite lesion type | |||||||||
| Bronchiectasis | 1 | (1.4%) | 2 | (25.0%) | 0.027 | 17.82 (2.02–157.34) | 0.010 | ||
| Tree in buds | 14 | (20.0%) | 5 | (62.5%) | 0.018 | 6.12 (1.42–26.34) | 0.014 | ||
| Cluster of nodules | 1 | (1.4%) | 0 | (0.0%) | 1.000 | 2.73 (0.10–72.36) | 0.582 | ||
| Reversed halo sign | 1 | (1.4%) | 0 | (0.0%) | 1.000 | 2.73 (0.10–72.36) | 0.582 | ||
| Strand | 1 | (1.4%) | 0 | (0.0%) | 1.000 | 2.73 (0.10–72.36) | 0.582 | ||
| The presence of a specific accompanying characteristics type | |||||||||
| Calcification | 14 | (20.0%) | 1 | (12.5%) | 1.000 | 0.78 (0.12–4.93) | 0.787 | ||
| Cavity | 33 | (47.1%) | 5 | (62.5%) | 0.476 | 1.76 (0.43–7.27) | 0.430 | ||
| Slight to moderate density | 52 | (74.3%) | 5 | (62.5%) | 0.675 | 0.55 (0.13–2.34) | 0.431 | ||
| Fibrosis | 4 | (5.7%) | 0 | (0.0%) | 1.000 | 0.87 (0.04–17.60) | 0.926 | ||
| Patient-Level Lesion Dynamic Data | |||||||||
| New lesion | |||||||||
| The presence of newly emerged lesion | 14.0 | (20.0%) | 4 | (50.0%) | 0.078 | 3.90 (0.93–16.26) | 0.067 | ||
| The number of newly emerged lesion | 0.0 | (0.0–0.0) | 0.5 | (0.0–1.0) | 0.066 | 1.38 (0.96–1.98) | 0.111 | ||
| The presence of a specific volume change pattern | |||||||||
| Decrease | 41 | (58.6%) | 7 | (87.5%) | 0.143 | 3.55 (0.58–21.84) | 0.128 | ||
| Mix-D-I (n = 21) | 1 | (5.9%) | 2 | (50.0%) | 0.080 | 11.00 (0.98–123.98) | 0.046 | 17.49 (1.28—239.87) | 0.024 |
| Stable | 36 | (51.4%) | 5 | (62.5%) | 0.715 | 1.49 (0.36–6.14) | 0.581 | ||
| Increase | 29 | (41.4%) | 4 | (50.0%) | 0.716 | 1.41 (0.35–5.64) | 0.631 | ||
| Mix-I-D (n = 21) | 3 | (17.6%) | 1 | (25.0%) | 1.000 | 1.78 (0.19–16.69) | 0.622 | ||
| CT series | |||||||||
| Number of CT scans per person | 2.0 | (2.0–2.0) | 2.5 | (2.0–3.0) | 0.207 | ||||
| Patients with more than 2 CT scans | 17 | (24.3%) | 4 | (50.0%) | 0.201 | ||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Qin, L.; Jiang, J.; Ma, S.; Liu, X.; Lv, P.; Wang, W.; Takiff, H.E.; Xie, Y.L.; Liu, Q.; Li, W. Divergent Evolution of Tuberculosis Lesions During Treatment: A Longitudinal CT-Based Analysis of Progression and Regression Patterns. Diagnostics 2026, 16, 892. https://doi.org/10.3390/diagnostics16060892
Qin L, Jiang J, Ma S, Liu X, Lv P, Wang W, Takiff HE, Xie YL, Liu Q, Li W. Divergent Evolution of Tuberculosis Lesions During Treatment: A Longitudinal CT-Based Analysis of Progression and Regression Patterns. Diagnostics. 2026; 16(6):892. https://doi.org/10.3390/diagnostics16060892
Chicago/Turabian StyleQin, Liyi, Jiaxin Jiang, Shiran Ma, Xiaoming Liu, Pingxin Lv, Wei Wang, Howard E. Takiff, Yingda L. Xie, Qingyun Liu, and Weimin Li. 2026. "Divergent Evolution of Tuberculosis Lesions During Treatment: A Longitudinal CT-Based Analysis of Progression and Regression Patterns" Diagnostics 16, no. 6: 892. https://doi.org/10.3390/diagnostics16060892
APA StyleQin, L., Jiang, J., Ma, S., Liu, X., Lv, P., Wang, W., Takiff, H. E., Xie, Y. L., Liu, Q., & Li, W. (2026). Divergent Evolution of Tuberculosis Lesions During Treatment: A Longitudinal CT-Based Analysis of Progression and Regression Patterns. Diagnostics, 16(6), 892. https://doi.org/10.3390/diagnostics16060892






