Comparison of Manual, Semi-Automatic, and Automatic CT-Based Methods for Liver Volume Segmentation
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. CT Imaging Protocol
2.3. Segmentation Methods
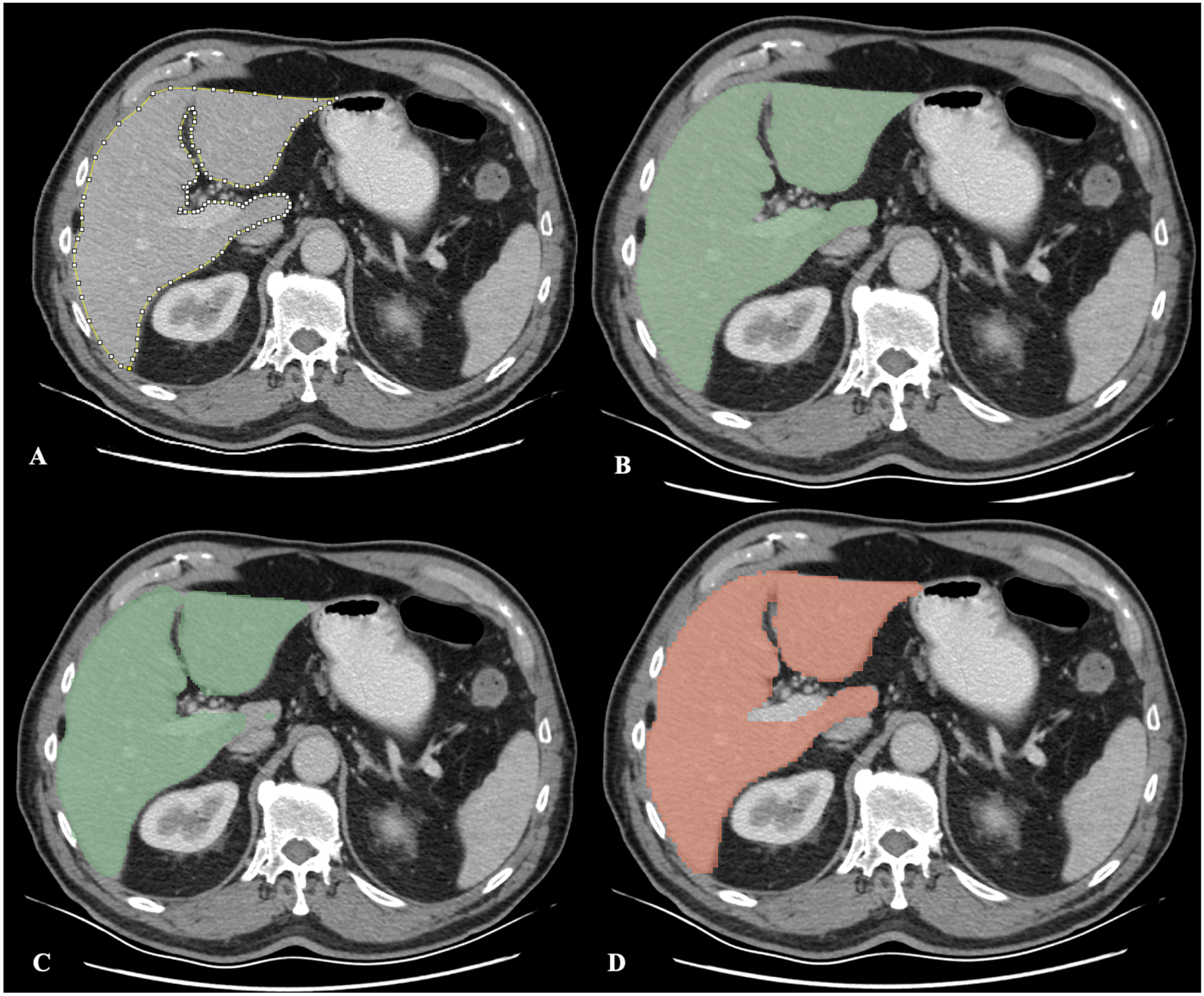
2.3.1. Manual Segmentation
2.3.2. Semi-Automatic Segmentation with the RVX Liver Module
2.3.3. Deep Learning-Based Segmentation with the RVX Liver Module
2.3.4. Automatic Segmentation with the TotalSegmentator Module
2.4. Segmentation Agreement Analysis
2.5. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CT | Computed tomography |
| ICC | Intraclass correlation coefficient |
| ANOVA | Analysis of variance |
| DICE | Dice similarity coefficient |
| HD | Hausdorff distance |
| SD | Standard deviation |
| ROI | Region of interest |
| RVX | Robust vascular network extraction and understanding within hepatic biomedical images |
| DICOM | Digital imaging and communications in medicine |
| PACS | Picture archiving and communication system |
References
- Ferrero, A.; Vigano, L.; Polastri, R.; Muratore, A.; Eminefendic, H.; Regge, D.; Capussotti, L. Postoperative liver dysfunction and future remnant liver: Where is the limit? Results of a prospective study. World J. Surg. 2007, 31, 1643–1651. [Google Scholar] [CrossRef]
- Gotra, A.; Chartrand, G.; Vu, K.N.; Vandenbroucke-Menu, F.; Massicotte-Tisluck, K.; de Guise, J.A.; Tang, A. Comparison of MRI- and CT-based semiautomated liver segmentation: A validation study. Abdom. Radiol. 2017, 42, 478–489. [Google Scholar] [CrossRef] [PubMed]
- Gloger, O.; Kuhn, J.; Stanski, A.; Völzke, H.; Puls, R. A fully automatic three-step liver segmentation method on LDA-based probability maps for multiple contrast MR images. Magn. Reson. Imaging 2010, 28, 882–897. [Google Scholar] [CrossRef] [PubMed]
- Joiner, B.J.; Simpson, A.L.; Leal, J.N.; D’angelica, M.I.; Do, R.K.G. Assessing splenic enlargement on CT by unidimensional measurement changes in patients with colorectal liver metastases. Abdom. Imaging 2015, 40, 2338–2344. [Google Scholar] [CrossRef] [PubMed]
- Pattanayak, P.; Turkbey, E.B.; Summers, R.M. Comparative evaluation of three software packages for liver and spleen segmentation and volumetry. Acad. Radiol. 2017, 24, 831–839. [Google Scholar] [CrossRef]
- Planz, V.B.; Lubner, M.G.; Pickhardt, P.J. Volumetric analysis at abdominal CT: Oncologic and non-oncologic applications. Br. J. Radiol. 2019, 92, 20180631. [Google Scholar] [CrossRef]
- Aljabri, M.; AlAmir, M.; AlGhamdi, M.; Abdel-Mottaleb, M.; Collado-Mesa, F. Towards a better understanding of annotation tools for medical imaging: A survey. Multimed. Tools Appl. 2022, 81, 25877–25911. [Google Scholar] [CrossRef]
- Kothari, G.; Woon, B.; Patrick, C.J.; Korte, J.; Wee, L.; Hanna, G.G.; Kron, T.; Hardcastle, N.; Siva, S. The impact of inter-observer variation in delineation on robustness of radiomics features in non-small cell lung cancer. Sci. Rep. 2022, 12, 12822. [Google Scholar] [CrossRef]
- Lenchik, L.; Heacock, L.; Weaver, A.A.; Boutin, R.D.; Cook, T.S.; Itri, J.; Filippi, C.G.; Gullapalli, R.P.; Lee, J.; Zagurovskaya, M.; et al. Automated segmentation of tissues using CT and MRI: A systematic review. Acad. Radiol. 2019, 26, 1695–1706. [Google Scholar] [CrossRef]
- Farooq, S.; Khan, Z. A survey of computer aided diagnosis (Cad) of liver in medical diagnosis. VAWKUM Trans. Comput. Sci. 2020, 8, 23–29. [Google Scholar] [CrossRef]
- Kalshabay, Y.; Zholdybay, Z.; Di Martino, M.; Medeubekov, U.; Baiguissova, D.; Ainakulova, A.; Doskhanov, M.; Baimakhanov, B. CT volume analysis in living donor liver transplantation: Accuracy of three different approaches. Insights Imaging 2023, 14, 82. [Google Scholar] [CrossRef]
- Khan, R.A.; Luo, Y.; Wu, F.X. Machine learning based liver disease diagnosis: A systematic review. Neurocomputing 2022, 468, 492–509. [Google Scholar] [CrossRef]
- Nayantara, P.V.; Kamath, S.; Manjunath, K.N.; Rajagopal, K. Computer-aided diagnosis of liver lesions using CT images: A systematic review. Comput. Biol. Med. 2020, 127, 104035. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Liu, X.; Zhao, J.; Liu, Y.; Liu, S.; Liu, Y.; Zhao, J. Computer auxiliary diagnosis technique of detecting cholangiocarcinoma based on medical imaging: A review. Comput. Methods Programs Biomed. 2021, 208, 106265. [Google Scholar] [CrossRef] [PubMed]
- Affane, A.; Chetoui, M.A.; Lamy, J.; Lienemann, G.; Peron, R.; Beaurepaire, P.; Dollé, G.; Lebre, M.-A.; Magnin, B.; Merveille, O.; et al. The R-Vessel-X Project. IRBM 2025, 46, 100876. [Google Scholar] [CrossRef]
- Wasserthal, J.; Breit, H.C.; Meyer, M.T.; Pardella, M.; Hinck, D.; Sauter, A.W.; Heye, T.; Boll, D.T.; Cyriac, J.; Yang, S.; et al. TotalSegmentator: Robust segmentation of 104 anatomic structures in CT images. Radiol. Artif. Intell. 2023, 5, e230024. [Google Scholar] [CrossRef]
- Gross, M.; Huber, S.; Arora, S.; Pradella, M.; Hinck, D.; Sauter, A.W.; Heye, T.; Boll, D.T.; Cyriac, J.; Yang, S.; et al. Automated MRI liver segmentation for anatomical segmentation, liver volumetry, and extraction of radiomics. Eur. Radiol. 2024, 34, 5056–5065. [Google Scholar] [CrossRef]
- Dockès, J.; Varoquaux, G.; Poline, J.-B. Preventing dataset shift from breaking machine-learning biomarkers. GigaScience 2021, 10, giab055. [Google Scholar] [CrossRef]
- Lemke, A.J.; Brinkmann, M.J.; Schott, T.; Niehues, S.M.; Settmacher, U.; Neuhaus, P.; Felix, R. Living donor right liver lobes: Preoperative CT volumetric measurement for calculation of intraoperative weight and volume. Radiology 2006, 240, 736–742. [Google Scholar] [CrossRef]
- Heimann, T.; van Ginneken, B.; Styner, M.A.; Arzhaeva, Y.; Aurich, V.; Bauer, C.; Beck, A.; Becker, C.; Beichel, R.; Bekes, G.; et al. Comparison and evaluation of methods for liver segmentation from CT datasets. IEEE Trans. Med. Imaging 2009, 28, 1251–1265. [Google Scholar] [CrossRef]
- Suzuki, K.; Epstein, M.L.; Kohlbrenner, R.; Garg, S.; Hori, M.; Oto, A.; Baron, R.L. Quantitative radiology: Automated CT liver volumetry compared with interactive volumetry and manual volumetry. Am. J. Roentgenol. 2011, 197, W706–W712. [Google Scholar] [CrossRef]
- Gotra, A.; Chartrand, G.; Massicotte-Tisluck, K.; Morin-Roy, F.; Vandenbroucke-Menu, F.; de Guise, J.A.; Tang, A. Validation of a semiautomated liver segmentation method using CT for accurate volumetry. Acad. Radiol. 2015, 22, 1088–1098. [Google Scholar] [CrossRef] [PubMed]
- Gross, M.; Spektor, M.; Jaffe, A.; Kucukkaya, A.S.; Iseke, S.; Haider, S.P.; Strazzabosco, M.; Chapiro, J.; Onofrey, J.A. Improved performance and consistency of deep learning 3D liver segmentation with heterogeneous cancer stages in magnetic resonance imaging. PLoS ONE 2021, 16, e0260630. [Google Scholar] [CrossRef] [PubMed]
- Kavur, A.E.; Gezer, N.S.; Barış, M.; Sahin, Y.; Ozkan, S.; Baydar, B.; Yuksel, U.; Kilikcier, C.; Olut, S.; Akar, G.B.; et al. Comparison of semi-automatic and deep learning-based automatic methods for liver segmentation in living liver transplant donors. Diagn. Interv. Radiol. 2020, 26, 11–21. [Google Scholar] [CrossRef]
- Yu, A.C.; Mohajer, B.; Eng, J. External validation of deep learning algorithms for radiologic diagnosis: A systematic review. Radiol. Artif. Intell. 2022, 4, e210064. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Sun, J.; Ding, H.; Wang, X.; Wang, G. Robust liver vessel extraction using 3D U-Net with variant Dice loss function. Comput. Biol. Med. 2018, 101, 153–162. [Google Scholar] [CrossRef]
- Ibragimov, B.; Toesca, D.; Chang, D.; Koong, A.; Xing, L. Combining deep learning with anatomical analysis for segmentation of the portal vein for liver SBRT planning. Phys. Med. Biol. 2017, 62, 8943–8958. [Google Scholar] [CrossRef]
- Núñez, L.; Ferreira, C.; Mojtahed, A.; Lamb, H.; Cappio, S.; Husainy, M.A.; Dennis, A.; Pansini, M. Assessing the performance of AI-assisted technicians in liver segmentation, Couinaud division, and lesion detection: A pilot study. Abdom. Radiol. 2024, 49, 4264–4272. [Google Scholar] [CrossRef]
- Pettit, R.W.; Marlatt, B.B.; Corr, S.J.; Havelka, J.; Rana, A. nnU-Net deep learning method for segmenting parenchyma and determining liver volume from computed tomography images. Ann. Surg. Open 2022, 3, e0155. [Google Scholar] [CrossRef]
- Nakayama, Y.; Li, Q.; Katsuragawa, S.; Ikeda, R.; Hiai, Y.; Awai, K.; Kusunoki, S.; Yamashita, Y.; Okajima, H.; Inomata, Y.; et al. Automated hepatic volumetry for living related liver transplantation at multisection CT. Radiology 2006, 240, 743–748. [Google Scholar] [CrossRef]
- Luciani, A.; Rusko, L.; Baranes, L.; Pichon, E.; Loze, B.; Deux, J.-F.; Laurent, A.; Tran-Van-Nhieu, J.; Rahmouni, A. Automated liver volumetry in orthotopic liver transplantation using multiphase acquisitions on MDCT. Am. J. Roentgenol. 2012, 198, W568–W574. [Google Scholar] [CrossRef]
- Karlo, C.; Reiner, C.S.; Stolzmann, P.; Breitenstein, S.; Marincek, B.; Weishaupt, D.; Frauenfelder, T. CT- and MRI-based volumetry of resected liver specimens: Comparison with intraoperative volume and weight measurements and calculation of conversion factors. Eur. J. Radiol. 2010, 75, e107–e111. [Google Scholar] [CrossRef]



| Estimates | ||||
|---|---|---|---|---|
| Measure: | Mean (cm3) | Std. Error | 95% Confidence Interval | |
| Lower Bound | Upper Bound | |||
| Manual | 1503.9 | 38.382 | 1427.6 | 1580.2 |
| RVX Semi-Automatic | 1512.6 | 40.235 | 1432.5 | 1592.5 |
| RVX Deep Learning | 1549.8 | 39.629 | 1470.9 | 1628.5 |
| TotalSegmentator | 1518.3 | 39.390 | 1439.9 | 1596.6 |
| Pairwise Comparisons | ||||||
|---|---|---|---|---|---|---|
| Measure: | Mean Difference (I–J) | Std. Error | Sig. b | 95% Confidence Interval for Difference b | ||
| Lower Bound | Upper Bound | |||||
| Manual | ||||||
| RVX Semi-Automatic | 8.63 | 8.247 | 1.000 | −30.916 | 13.644 | |
| RVX Deep Learning | 45.83 * | 6.505 | <0.001 | −63.402 | −28.258 | |
| TotalSegmentator | 14.37 | 5.454 | 0.060 | −29.106 | 0.363 | |
| RVX Semi-Automatic | ||||||
| RVX Deep Learning | 37.19 * | 7.654 | <0.001 | −57.872 | −16.516 | |
| TotalSegmentator | 5.73 | 6.210 | 1.000 | −22.511 | 11.041 | |
| RVX Deep Learning | ||||||
| TotalSegmentator | −31.45 * | 4.639 | <0.001 | 18.927 | 43.991 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Dogan, B.; Simsek, S.B.; Sonmez, S.; Ozgen Sonmez, M.N.; Dasci, O.; Ozmen, Z. Comparison of Manual, Semi-Automatic, and Automatic CT-Based Methods for Liver Volume Segmentation. Diagnostics 2026, 16, 817. https://doi.org/10.3390/diagnostics16050817
Dogan B, Simsek SB, Sonmez S, Ozgen Sonmez MN, Dasci O, Ozmen Z. Comparison of Manual, Semi-Automatic, and Automatic CT-Based Methods for Liver Volume Segmentation. Diagnostics. 2026; 16(5):817. https://doi.org/10.3390/diagnostics16050817
Chicago/Turabian StyleDogan, Berna, Sadik Bugrahan Simsek, Sefa Sonmez, Merve Nur Ozgen Sonmez, Omur Dasci, and Zafer Ozmen. 2026. "Comparison of Manual, Semi-Automatic, and Automatic CT-Based Methods for Liver Volume Segmentation" Diagnostics 16, no. 5: 817. https://doi.org/10.3390/diagnostics16050817
APA StyleDogan, B., Simsek, S. B., Sonmez, S., Ozgen Sonmez, M. N., Dasci, O., & Ozmen, Z. (2026). Comparison of Manual, Semi-Automatic, and Automatic CT-Based Methods for Liver Volume Segmentation. Diagnostics, 16(5), 817. https://doi.org/10.3390/diagnostics16050817






