Opportunities and Challenges of Visual Large Language Models in Imaging Diagnostics: Lessons from Brain Metastasis Detection in Clinical MRI
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Cohort
- Patients ≥ 18 years who were diagnosed with either non-small cell lung cancer (NSCLC), malignant melanoma, breast cancer or renal cell carcinoma and who did not suffer from a secondary malignancy.
- Received a clinically indicated MRI of the brain between 1 January 2017 and 1 August 2024 during routine cancer follow-up.
- (a) For the metastatic group, the presence of brain metastases had to be mentioned in the radiological report. Metastases had to be confirmed either by histopathology or by the prior or follow-up examinations with an increase in size or a shrinkage during treatment.
- (b) For the non-metastatic group, lack of brain metastases had to be mentioned in the radiological report, which was confirmed in a ground truth annotation.
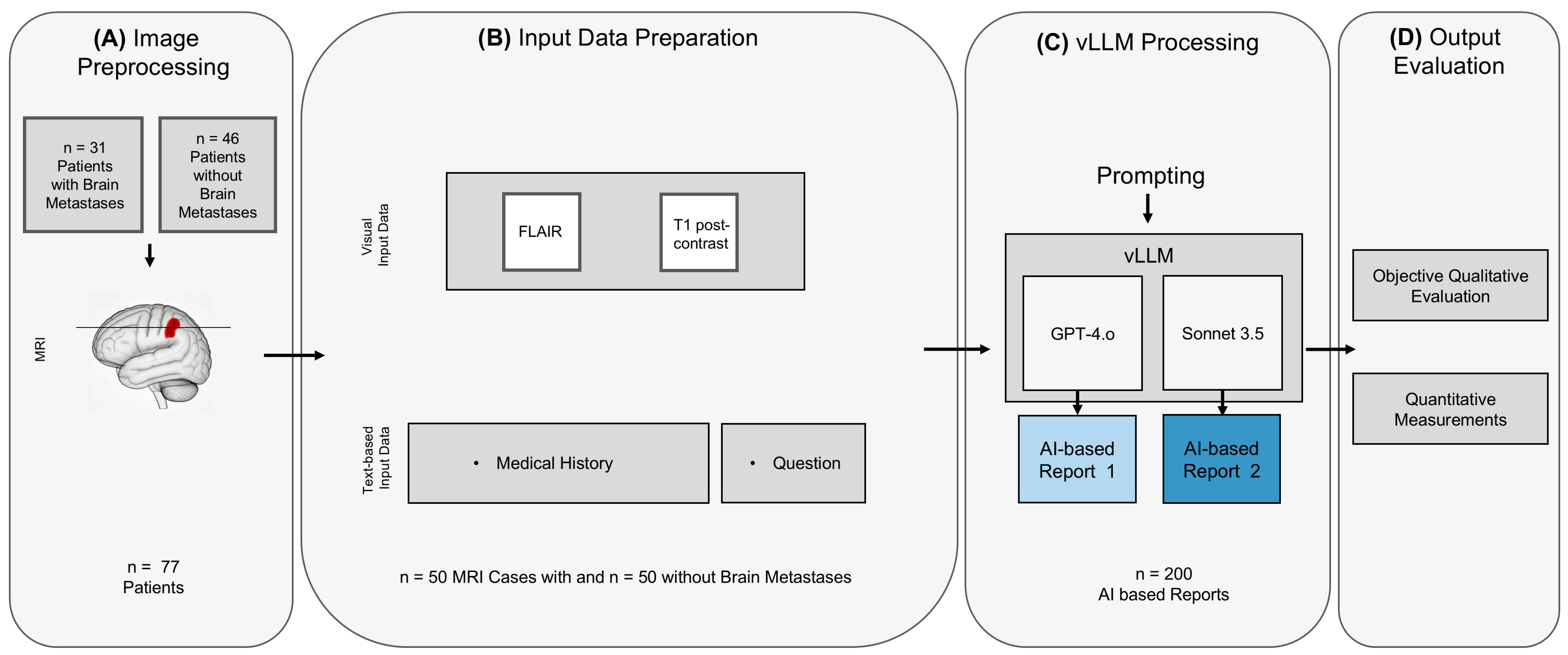
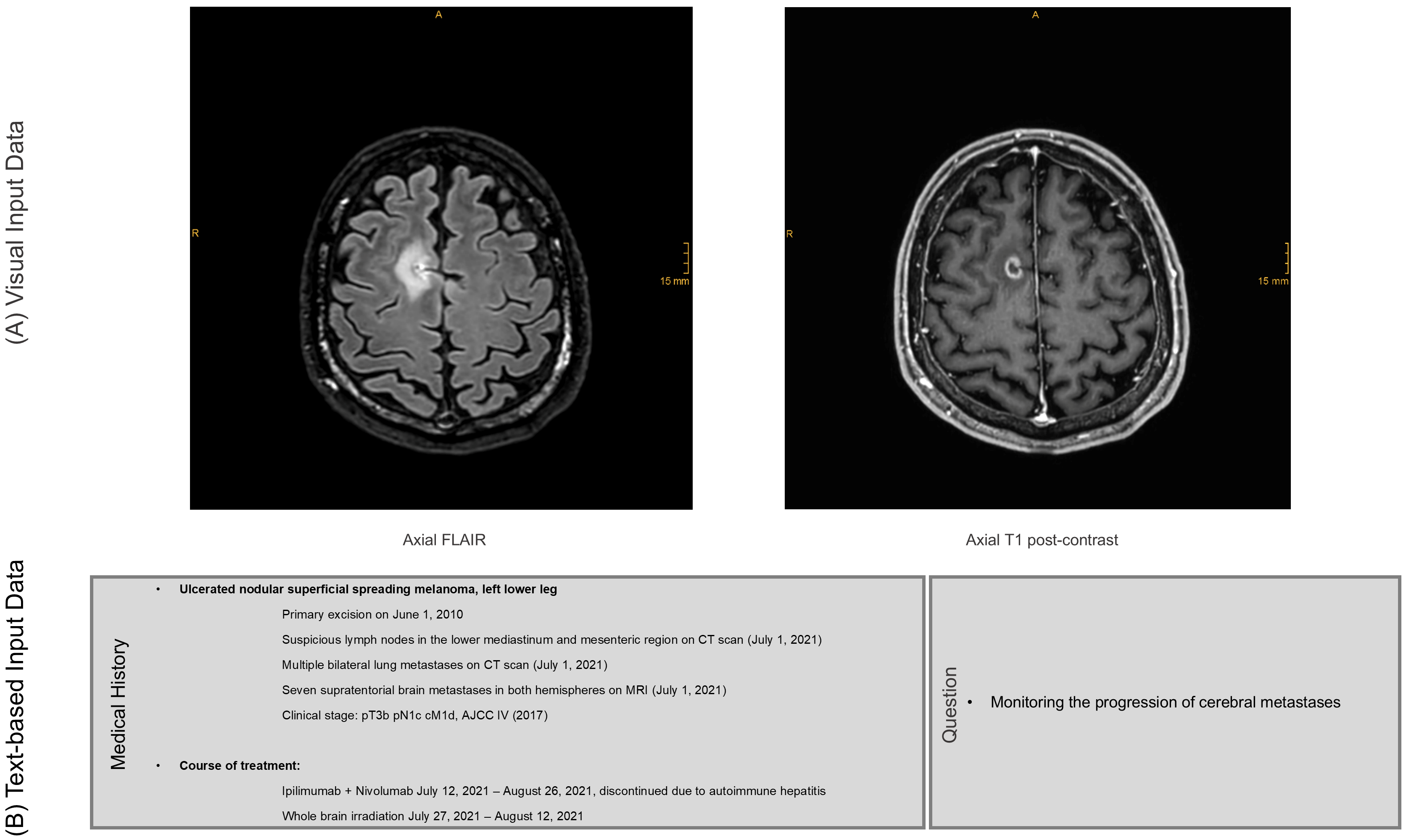
2.2. Image Acquisition, Preprocessing and Input Data
2.3. vLLM Processing
2.3.1. Visual Large Language Models
2.3.2. Prompt Engineering and AI-Generated Reports
- •
- Imagine you are a radiologist. The two images are sequences from the same examination.
- •
- Please identify the sequences. Then, create a report and an assessment in which you address the overall question.
- •
- Please also take the medical history into account. Indicate whether any pathological findings are visible. If so, indicate the size of the lesion (the contrast-enhancing portion in two axes) and indicate where the lesion is localized (anatomical brain region and side).
2.4. Output Evaluation
- •
- Both MRI sequences were correctly identified;
- •
- Presence or absence of a pathological lesion in the presented images was correctly identified;
- •
- Anatomical region of the lesion, if present, was correctly identified (e.g., frontal lobe);
- •
- Side of the lesion, if present, was correctly identified;
- •
- In case of an identified lesion, whether additional lesions were wrongfully detected or not.
2.5. Statistical Methods
3. Results
Study Cohort
- a.
- Evaluation of the AI-generated reports
- b.
- Lesion size estimation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AI | Artificial Intelligence |
| aiRR | AI-generated Radiology Report |
| C | Claude Sonnet 3.5 (Anthropic) |
| CNN | Convolutional Neural Network |
| FLAIR | Fluid Attenuated Inversion Recovery |
| G | GPT-4o (OpenAI) |
| JPEG | Joint Photographic Experts Group |
| LLM | Large Language Model |
| NSCLC | Non-Small Cell Lung Cancer |
| ViTs | Vision Transformers |
| vLLM | Visual Large Language Model |
References
- Wen, P.Y.; Loeffler, J.S. Management of Brain Metastases. Oncology 1999, 13, 941–954, 957–961; discussion 961–962, 969. [Google Scholar]
- Nayak, L.; Lee, E.Q.; Wen, P.Y. Epidemiology of Brain Metastases. Curr. Oncol. Rep. 2012, 14, 48–54. [Google Scholar] [CrossRef]
- Sperduto, P.W.; Kased, N.; Roberge, D.; Xu, Z.; Shanley, R.; Luo, X.; Sneed, P.K.; Chao, S.T.; Weil, R.J.; Suh, J.; et al. Summary Report on the Graded Prognostic Assessment: An Accurate and Facile Diagnosis-Specific Tool to Estimate Survival for Patients with Brain Metastases. J. Clin. Oncol. 2012, 30, 419–425. [Google Scholar] [CrossRef] [PubMed]
- Brown, P.D.; Jaeckle, K.; Ballman, K.V.; Farace, E.; Cerhan, J.H.; Anderson, S.K.; Carrero, X.W.; Barker, F.G.; Deming, R.; Burri, S.H.; et al. Effect of Radiosurgery Alone vs Radiosurgery with Whole Brain Radiation Therapy on Cognitive Function in Patients with 1 to 3 Brain Metastases: A Randomized Clinical Trial. JAMA 2016, 316, 401–409. [Google Scholar] [CrossRef]
- Davis, P.C.; Hudgins, P.A.; Peterman, S.B.; Hoffman, J.C. Diagnosis of Cerebral Metastases: Double-Dose Delayed CT vs Contrast-Enhanced MR Imaging. AJNR Am. J. Neuroradiol. 1991, 12, 293–300. [Google Scholar]
- Schaefer, P.W.; Budzik, R.F.; Gonzalez, R.G. Imaging of Cerebral Metastases. Neurosurg. Clin. N. Am. 1996, 7, 393–423. [Google Scholar] [CrossRef]
- Crivellari, D.; Pagani, O.; Veronesi, A.; Lombardi, D.; Nolè, F.; Thürlimann, B.; Hess, D.; Borner, M.; Bauer, J.; Martinelli, G.; et al. High Incidence of Central Nervous System Involvement in Patients with Metastatic or Locally Advanced Breast Cancer Treated with Epirubicin and Docetaxel. Ann. Oncol. 2001, 12, 353–356. [Google Scholar] [CrossRef] [PubMed]
- Sundermeyer, M.L.; Meropol, N.J.; Rogatko, A.; Wang, H.; Cohen, S.J. Changing Patterns of Bone and Brain Metastases in Patients with Colorectal Cancer. Clin. Color. Cancer 2005, 5, 108–113. [Google Scholar] [CrossRef]
- Mamon, H.J.; Yeap, B.Y.; Jänne, P.A.; Reblando, J.; Shrager, S.; Jaklitsch, M.T.; Mentzer, S.; Lukanich, J.M.; Sugarbaker, D.J.; Baldini, E.H.; et al. High Risk of Brain Metastases in Surgically Staged IIIA Non-Small-Cell Lung Cancer Patients Treated with Surgery, Chemotherapy, and Radiation. J. Clin. Oncol. 2005, 23, 1530–1537. [Google Scholar] [CrossRef]
- Rahman, T.; Chowdhury, M.E.H.; Khandakar, A.; Islam, K.R.; Islam, K.F.; Mahbub, Z.B.; Kadir, M.A.; Kashem, S. Transfer Learning with Deep Convolutional Neural Network (CNN) for Pneumonia Detection Using Chest X-Ray. Appl. Sci. 2020, 10, 3233. [Google Scholar] [CrossRef]
- Katal, S.; York, B.; Gholamrezanezhad, A. AI in Radiology: From Promise to Practice—A Guide to Effective Integration. Eur. J. Radiol. 2024, 181, 111798. [Google Scholar] [CrossRef]
- Abadia, A.F.; Yacoub, B.; Stringer, N.; Snoddy, M.; Kocher, M.; Schoepf, U.J.; Aquino, G.J.; Kabakus, I.; Dargis, D.; Hoelzer, P.; et al. Diagnostic Accuracy and Performance of Artificial Intelligence in Detecting Lung Nodules in Patients with Complex Lung Disease: A Noninferiority Study. J. Thorac. Imaging 2022, 37, 154. [Google Scholar] [CrossRef]
- Lakhani, P.; Sundaram, B. Deep Learning at Chest Radiography: Automated Classification of Pulmonary Tuberculosis by Using Convolutional Neural Networks. Radiology 2017, 284, 574–582. [Google Scholar] [CrossRef]
- Bhayana, R. Chatbots and Large Language Models in Radiology: A Practical Primer for Clinical and Research Applications. Radiology 2024, 310, e232756. [Google Scholar] [CrossRef]
- Sun, Z.; Ong, H.; Kennedy, P.; Tang, L.; Chen, S.; Elias, J.; Lucas, E.; Shih, G.; Peng, Y. Evaluating GPT4 on Impressions Generation in Radiology Reports. Radiology 2023, 307, e231259. [Google Scholar] [CrossRef] [PubMed]
- Kottlors, J.; Bratke, G.; Rauen, P.; Kabbasch, C.; Persigehl, T.; Schlamann, M.; Lennartz, S. Feasibility of Differential Diagnosis Based on Imaging Patterns Using a Large Language Model. Radiology 2023, 308, e231167. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, S.; Sakaguchi, Y.; Kouno, N.; Takasawa, K.; Ishizu, K.; Akagi, Y.; Aoyama, R.; Teraya, N.; Bolatkan, A.; Shinkai, N.; et al. Comparison of Vision Transformers and Convolutional Neural Networks in Medical Image Analysis: A Systematic Review. J. Med. Syst. 2024, 48, 84. [Google Scholar] [CrossRef]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An Image Is Worth 16 × 16 Words: Transformers for Image Recognition at Scale 2021. arXiv 2020, arXiv:2010.11929. [Google Scholar]
- Günay, S.; Öztürk, A.; Yiğit, Y. The Accuracy of Gemini, GPT-4, and GPT-4o in ECG Analysis: A Comparison with Cardiologists and Emergency Medicine Specialists. Am. J. Emerg. Med. 2024, 84, 68–73. [Google Scholar] [CrossRef]
- Suh, P.S.; Shim, W.H.; Suh, C.H.; Heo, H.; Park, C.R.; Eom, H.J.; Park, K.J.; Choe, J.; Kim, P.H.; Park, H.J.; et al. Comparing Diagnostic Accuracy of Radiologists versus GPT-4V and Gemini Pro Vision Using Image Inputs from Diagnosis Please Cases. Radiology 2024, 312, e240273. [Google Scholar] [CrossRef] [PubMed]
- Hayden, N.; Gilbert, S.; Poisson, L.M.; Griffith, B.; Klochko, C. Performance of GPT-4 with Vision on Text- and Image-Based ACR Diagnostic Radiology In-Training Examination Questions. Radiology 2024, 312, e240153. [Google Scholar] [CrossRef]
- Brin, D.; Sorin, V.; Barash, Y.; Konen, E.; Glicksberg, B.S.; Nadkarni, G.N.; Klang, E. Assessing GPT-4 Multimodal Performance in Radiological Image Analysis. Eur. Radiol. 2025, 35, 1959–1965. [Google Scholar] [CrossRef]
- Han, T.; Adams, L.C.; Bressem, K.K.; Busch, F.; Nebelung, S.; Truhn, D. Comparative Analysis of Multimodal Large Language Model Performance on Clinical Vignette Questions. JAMA 2024, 331, 1320–1321. [Google Scholar] [CrossRef] [PubMed]
- Huppertz, M.S.; Siepmann, R.; Topp, D.; Nikoubashman, O.; Yüksel, C.; Kuhl, C.K.; Truhn, D.; Nebelung, S. Revolution or Risk?—Assessing the Potential and Challenges of GPT-4V in Radiologic Image Interpretation. Eur. Radiol. 2025, 35, 1111–1121. [Google Scholar] [CrossRef]
- Zhu, L.; Mou, W.; Lai, Y.; Chen, J.; Lin, S.; Xu, L.; Lin, J.; Guo, Z.; Yang, T.; Lin, A.; et al. Step into the Era of Large Multimodal Models: A Pilot Study on ChatGPT-4V(Ision)’s Ability to Interpret Radiological Images. Int. J. Surg. 2024, 110, 4096–4102. [Google Scholar] [CrossRef]
- Deng, J.; Heybati, K.; Shammas-Toma, M. When Vision Meets Reality: Exploring the Clinical Applicability of GPT-4 with Vision. Clin. Imaging 2024, 108, 110101. [Google Scholar] [CrossRef]
- Sarangi, P.K.; Narayan, R.K.; Mohakud, S.; Vats, A.; Sahani, D.; Mondal, H. Assessing the Capability of ChatGPT, Google Bard, and Microsoft Bing in Solving Radiology Case Vignettes. Indian J. Radiol. Imaging 2024, 34, 276–282. [Google Scholar] [CrossRef] [PubMed]
- Agbareia, R.; Omar, M.; Soffer, S.; Glicksberg, B.S.; Nadkarni, G.N.; Klang, E. Visual-Textual Integration in LLMs for Medical Diagnosis: A Preliminary Quantitative Analysis. Comput. Struct. Biotechnol. J. 2025, 27, 184–189. [Google Scholar] [CrossRef] [PubMed]
- Ozkara, B.B.; Chen, M.M.; Federau, C.; Karabacak, M.; Briere, T.M.; Li, J.; Wintermark, M. Deep Learning for Detecting Brain Metastases on MRI: A Systematic Review and Meta-Analysis. Cancers 2023, 15, 334. [Google Scholar] [CrossRef]
- Pennig, L.; Shahzad, R.; Caldeira, L.; Lennartz, S.; Thiele, F.; Goertz, L.; Zopfs, D.; Meißner, A.-K.; Fürtjes, G.; Perkuhn, M.; et al. Automated Detection and Segmentation of Brain Metastases in Malignant Melanoma: Evaluation of a Dedicated Deep Learning Model. AJNR Am. J. Neuroradiol. 2021, 42, 655–662. [Google Scholar] [CrossRef]
- Yin, S.; Luo, X.; Yang, Y.; Shao, Y.; Ma, L.; Lin, C.; Yang, Q.; Wang, D.; Luo, Y.; Mai, Z.; et al. Development and Validation of a Deep-Learning Model for Detecting Brain Metastases on 3D Post-Contrast MRI: A Multi-Center Multi-Reader Evaluation Study. Neuro-Oncology 2022, 24, 1559–1570. [Google Scholar] [CrossRef] [PubMed]
- Rudie, J.D.; Weiss, D.A.; Colby, J.B.; Rauschecker, A.M.; Laguna, B.; Braunstein, S.; Sugrue, L.P.; Hess, C.P.; Villanueva-Meyer, J.E. Three-Dimensional U-Net Convolutional Neural Network for Detection and Segmentation of Intracranial Metastases. Radiol. Artif. Intell. 2021, 3, e200204. [Google Scholar] [CrossRef] [PubMed]
- Ong, J.C.L.; Chang, S.Y.-H.; William, W.; Butte, A.J.; Shah, N.H.; Chew, L.S.T.; Liu, N.; Doshi-Velez, F.; Lu, W.; Savulescu, J.; et al. Ethical and Regulatory Challenges of Large Language Models in Medicine. Lancet Digit. Health 2024, 6, e428–e432. [Google Scholar] [CrossRef] [PubMed]
- Elbattah, M.; Arnaud, E.; Ghazali, D.A.; Dequen, G. Exploring the Ethical Challenges of Large Language Models in Emergency Medicine: A Comparative International Review. In Proceedings of the 2024 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Lisbon, Portugal, 3–6 December 2024; pp. 5750–5755. [Google Scholar]


| Variable | ||
| Mean age; SD | 58.8 (y); SD ± 12.7 | |
| Sex | female 45/77 (58.4%); male 32/77 (41.6%) | |
| Primary tumor | Patients without brain metastases (46/77; 59.7%) | Patients with brain metastases (31/77; 40.3%) |
| Melanoma | 15/46 (32.6%) | 9/31 (29.0%) |
| NSCLC | 15/46 (32.6%) | 8/31 (25.8%) |
| Breast Cancer | 10/46 (21.7%) | 7/31 (22.6%) |
| RCC | 6/46 (13.0%) | 7/31 (22.6%) |
| Category | Accuracy GPT-4o (%) | Accuracy Sonnet 3.5 (%) | p-Value |
|---|---|---|---|
| Sensitivity | 100% (92.9%, 100%) [50/50] | 100% (92.9%, 100%) [50/50] | 1.00 |
| Specificity | 8% (3.2%, 18.8%) [4/50] | 4% (1.1%, 13.5%) [2/50] | 0.625 |
| Diagnostic Accuracy | 54% [54/100] | 52% [52/100] | 0.625 |
| Sequence Correctly Detected | 100% (96.3%, 100%) [100/100] | 93% (86.3%, 96.6%) [93/100] | 0.0156 |
| Too Many Lesions Detected * | 12% (5.6%, 23.8%) [6/50] | 12% (5.6%, 23.8%) [6/50] | 1.00 |
| Localization Correctly Detected * | 70% (56.2%, 80.9%) [35/50] | 72% (58.3%, 82.5%) [36/50] | 1.00 |
| Side Correctly Detected * | 62% (48.2%, 74.1%) [31/50] | 76% (62.6%, 85.7%) [38/50] | 0.189 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Nelles, C.; Abou Zeid, N.; Terzis, R.; Iuga, A.-I.; Görtz, L.; Spurek, M.A.; Maintz, D.; Lennartz, S.; Kottlors, J. Opportunities and Challenges of Visual Large Language Models in Imaging Diagnostics: Lessons from Brain Metastasis Detection in Clinical MRI. Diagnostics 2026, 16, 749. https://doi.org/10.3390/diagnostics16050749
Nelles C, Abou Zeid N, Terzis R, Iuga A-I, Görtz L, Spurek MA, Maintz D, Lennartz S, Kottlors J. Opportunities and Challenges of Visual Large Language Models in Imaging Diagnostics: Lessons from Brain Metastasis Detection in Clinical MRI. Diagnostics. 2026; 16(5):749. https://doi.org/10.3390/diagnostics16050749
Chicago/Turabian StyleNelles, Christian, Nour Abou Zeid, Robert Terzis, Andra-Iza Iuga, Lukas Görtz, Marvin A. Spurek, David Maintz, Simon Lennartz, and Jonathan Kottlors. 2026. "Opportunities and Challenges of Visual Large Language Models in Imaging Diagnostics: Lessons from Brain Metastasis Detection in Clinical MRI" Diagnostics 16, no. 5: 749. https://doi.org/10.3390/diagnostics16050749
APA StyleNelles, C., Abou Zeid, N., Terzis, R., Iuga, A.-I., Görtz, L., Spurek, M. A., Maintz, D., Lennartz, S., & Kottlors, J. (2026). Opportunities and Challenges of Visual Large Language Models in Imaging Diagnostics: Lessons from Brain Metastasis Detection in Clinical MRI. Diagnostics, 16(5), 749. https://doi.org/10.3390/diagnostics16050749





