An Interpretable Ensemble Transformer Framework for Breast Cancer Detection in Ultrasound Images
Abstract
1. Introduction
2. Related Works
2.1. Traditional Deep CNN Architectures for BUS Classification
2.2. Ensemble and Transfer Learning Approaches
2.3. Hybrid and Multi-Task Architectures
2.4. Real-Time and Clinical Workflow-Oriented Applications
3. Materials and Methods
3.1. Data Acquisition
3.2. Data Preparation and Preprocessing
3.3. Data Splitting
3.4. Data Augmentation
- Resizing: Images were resized to match the input resolution required by the pre-trained feature extractor models.
- Random flipping: Horizontal and vertical flips were applied with a probability of 0.5 to simulate variability in lesion orientation.
- Random rotation: A rotation factor of 0.2 was used, allowing for image rotations of up to ±36 degrees, simulating the potential rotation of ultrasound images.
- Random contrast adjustment: A contrast factor of 0.2 was applied to simulate variations in image intensity and lighting conditions.
- Random cropping: A target height and width of 20% of the original dimensions were used to introduce local occlusions and simulate positional variability in lesions.
- Random zooming: Zoom transformations with both height and width factors set to 0.2 were used to reflect differences in imaging distance and magnification.
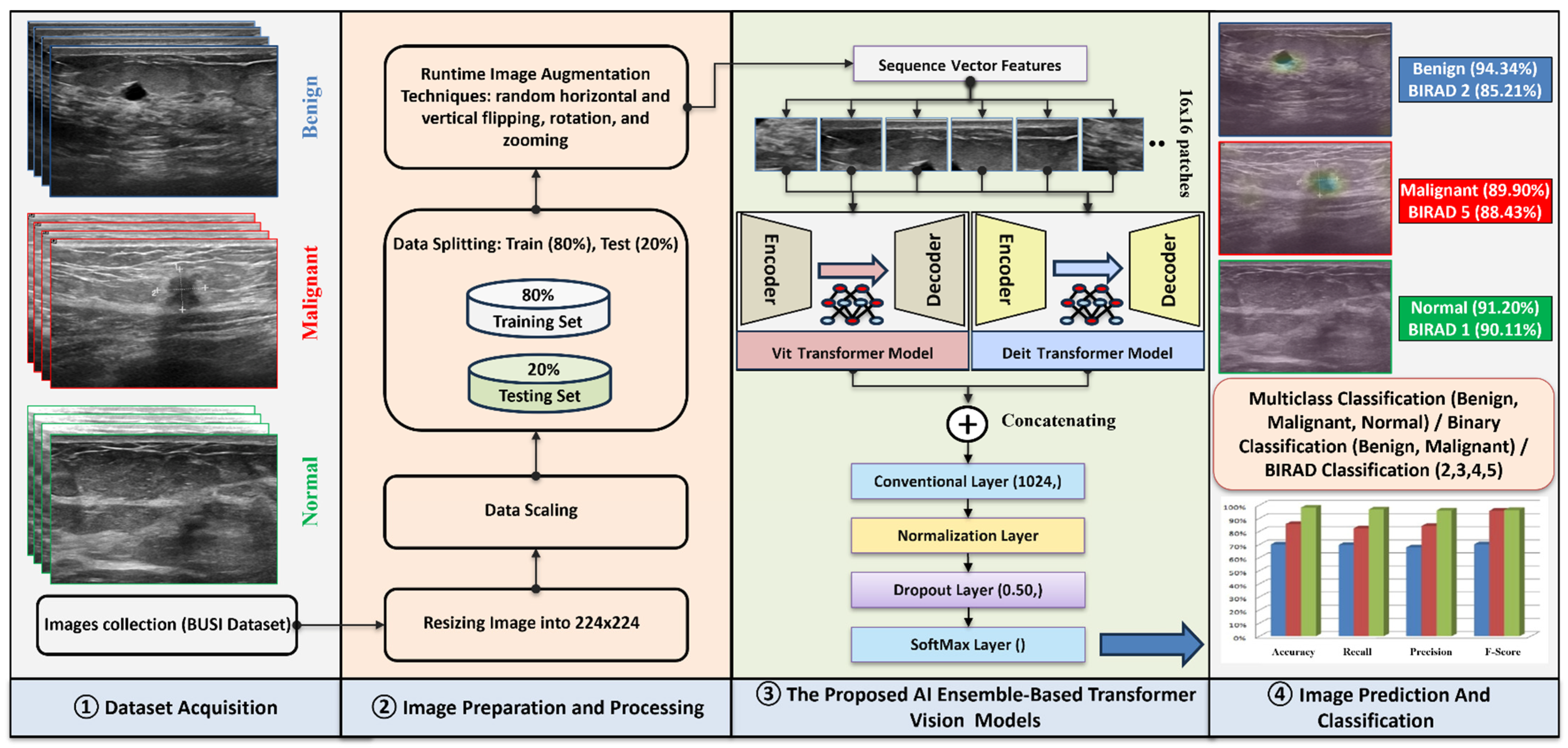
3.5. The Proposed Deep Learning Models
3.5.1. AI-Based Individual Models
3.5.2. AI-Based Ensemble of Individual Models
3.5.3. AI-Based Ensemble of Deit and ViT Model
- ViT excels at capturing global contextual relationships through pure self-attention mechanisms, enabling robust high-level feature abstraction [63].
- Deit enhances model efficiency through knowledge distillation and data optimization techniques, demonstrating superior performance in limited-data scenarios [68].
3.6. Fine-Tuning Models
- A fully connected (dense) layer with 1024 neurons.
- A batch normalization layer to stabilize learning.
- A dropout layer with a rate of 0.5 to reduce overfitting.
- A final dense output layer whose configuration is task-dependent:
- ➢
- 3 neurons for multiclass classification (normal, benign, malignant),
- ➢
- 2 neurons for binary classification (benign vs. malignant),
- ➢
- 4 or 6 neurons for BI-RADS scoring, depending on the dataset used.
3.7. Environment Setup
4. Results
- Scenario A investigates traditional CNN-based models. Eight popular pre-trained architectures—VGG19, VGG16, MobileNetV2, ResNet50, Xception, InceptionResNetV2, DenseNet201, and InceptionV3—are fine-tuned for a 3-class classification task (benign, malignant, normal).
- Scenario B examines six cutting-edge transformer-based models: ViT-Hybrid, ViT, Deit, DiT, Swin, and Beit. These models are unified under a consistent classification framework and evaluated on the same task.
- Scenario C focuses on ensemble learning. It introduces a novel ViT + Deit ensemble that exploits complementary transformer features for improved classification performance. This ensemble is compared against seven CNN-based ensembles, including combinations like DenseNet201 + VGG19 + Xception, VGG16 + ResNet50, and others.
4.1. Feature Space Analysis
4.2. Scenario A: Breast Cancer Classification Using Individual AI Models
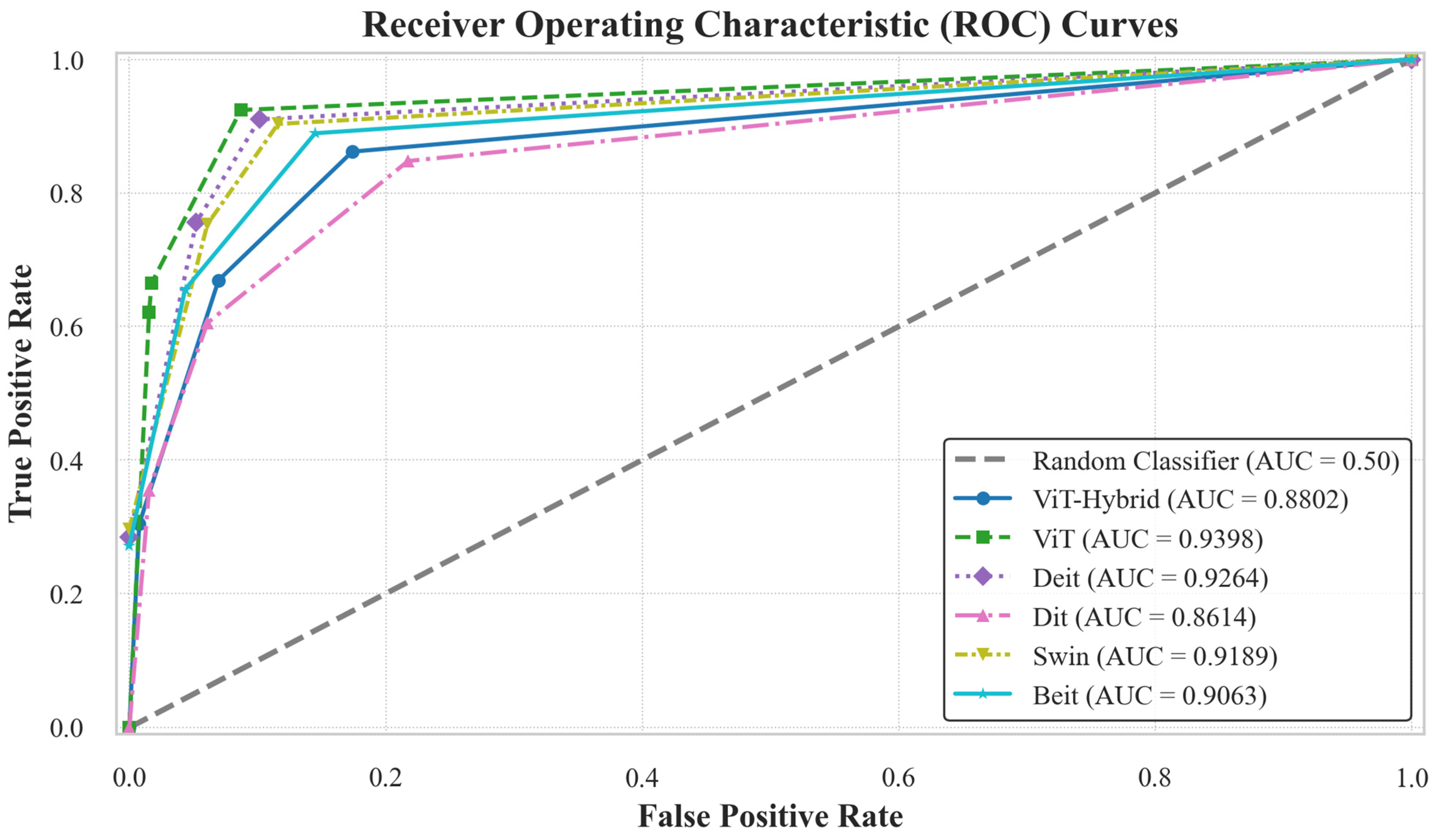
4.3. Scenario B: Breast Cancer Classification Using Vision Transformer Models
4.4. Scenario C: Breast Cancer Classification-Based AI Ensemble Classifier
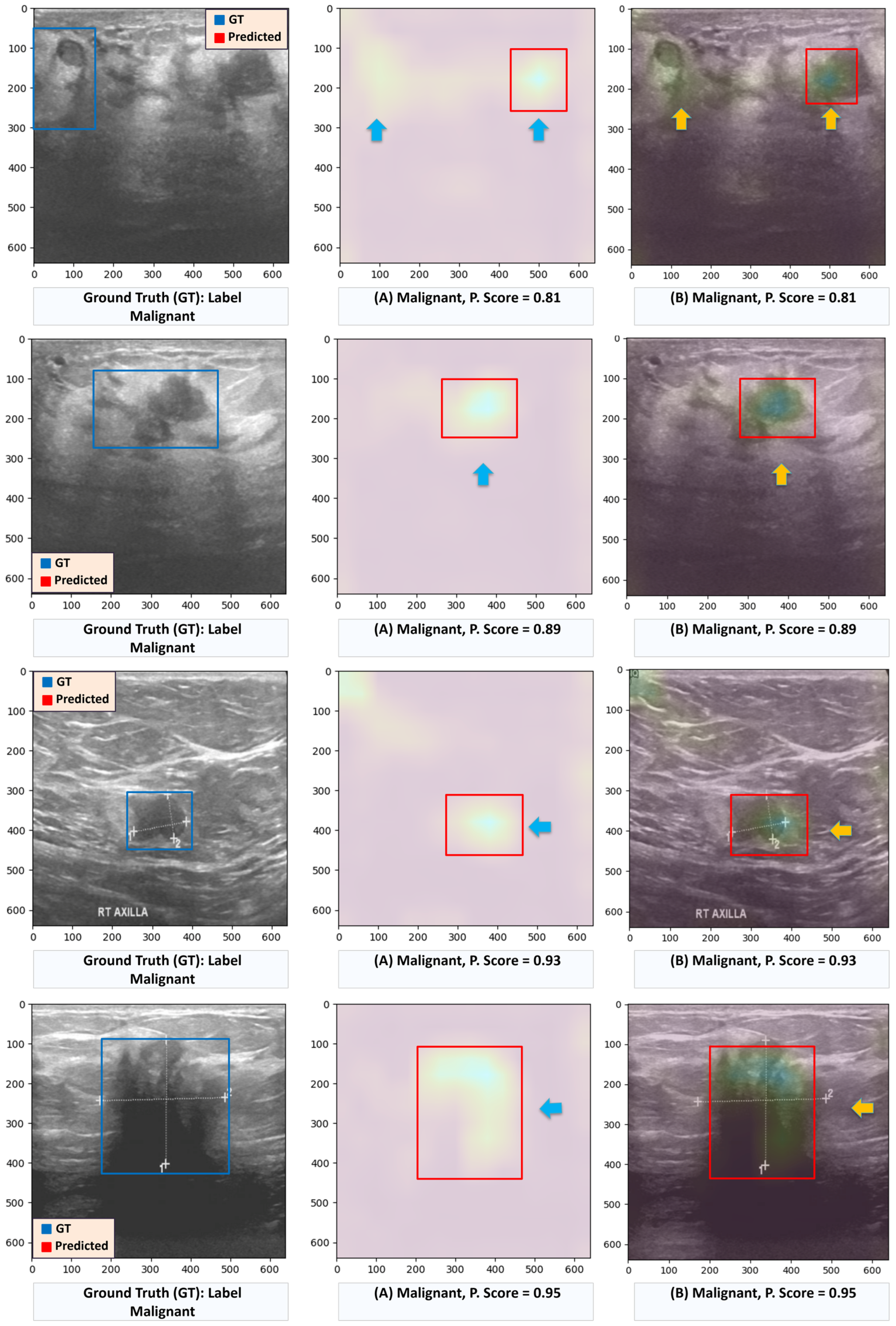
4.4.1. Detailed Analysis of Misclassified Samples for the Deit + ViT Ensemble Model
- (i)
- Model-based similarity, computed using cosine similarity between deep feature embeddings extracted from the trained network; and
- (ii)
- Pixel-based similarity, computed directly from normalized raw image intensities.
Model-Based Similarity Reveals Latent Feature Overlap
Pixel-Based Similarity Confirms Visual Distinctiveness
Recommendations for Future Improvements
4.5. Ablation Study
5. Discussion
5.1. Performance Evaluation of the Proposed AI Models
5.2. Clinical Applicability and Deployment Considerations
5.3. The Complexity Time of the Proposed CAD Framework
5.4. Comparison with Related Work on Breast Cancer Classification
5.5. Limitation and Future Work
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Iacob, R.; Iacob, E.R.; Stoicescu, E.R.; Ghenciu, D.M.; Cocolea, D.M.; Constantinescu, A.; Ghenciu, L.A.; Manolescu, D.L. Evaluating the role of breast ultrasound in early detection of breast cancer in low-and middle-income countries: A comprehensive narrative review. Bioengineering 2024, 11, 262. [Google Scholar] [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- World Health Organization. Breast Cancer. www.who.int. 2025. Available online: https://www.who.int/news-room/fact-sheets/detail/breast-cancer (accessed on 31 August 2025).
- World Health Organization. The Global Breast Cancer Initiative. www.who.int. 2021. Available online: https://www.who.int/initiatives/global-breast-cancer-initiative (accessed on 31 August 2025).
- Xu, H.; Xu, B. Breast cancer: Epidemiology, risk factors and screening. Chin. J. Cancer Res. 2023, 35, 565–583. [Google Scholar] [CrossRef] [PubMed]
- Al-Tam, R.M. Diversifying Medical Imaging of Breast Lesions. Master’s Thesis, University of Algarve, Faro, Portugal, 2015. [Google Scholar]
- Roheel, A.; Khan, A.; Anwar, F.; Akbar, Z.; Akhtar, M.F.; Imran Khan, M.; Sohail, M.F.; Ahmad, R. Global epidemiology of breast cancer based on risk factors: A systematic review. Front. Oncol. 2023, 13, 1240098. [Google Scholar] [CrossRef]
- Rakha, E.A.; Tse, G.M.; Quinn, C.M. An update on the pathological classification of breast cancer. Histopathology 2023, 82, 5–16. [Google Scholar] [CrossRef]
- Burciu, O.M.; Sas, I.; Popoiu, T.-A.; Merce, A.-G.; Moleriu, L.; Cobec, I.M. Correlations of imaging and therapy in breast cancer based on molecular patterns: An important issue in the diagnosis of breast cancer. Int. J. Mol. Sci. 2024, 25, 8506. [Google Scholar] [CrossRef] [PubMed]
- Al-Tam, R.M.; Al-Hejri, A.M.; Narangale, S.M.; Samee, N.A.; Mahmoud, N.F.; Al-masni, M.A.; Al-antari, M.A. A Hybrid Workflow of Residual Convolutional Transformer Encoder for Breast Cancer Classification Using Digital X-ray Mammograms. Biomedicines 2022, 10, 2971. [Google Scholar] [CrossRef]
- Abdel Samee, N.; Houssein, E.H.; Mohamed, O.; Mahmoud, N.F.; Talaat, R.; Al-Hejri, A.M.; Al-Tam, R.M. Using Deep DenseNet with Cyclical Learning Rate to Classify Leukocytes for Leukemia Identification. Front. Oncol. 2023, 13, 1230434. [Google Scholar] [CrossRef]
- Açar, Ç.R.; Orguc, S. Comparison of Performance in Diagnosis and Characterization of Breast Lesions: Contrast-Enhanced Mammography Versus Breast Magnetic Resonance Imaging. Clin. Breast Cancer 2024, 24, 481–493. [Google Scholar] [PubMed]
- Mann, R.M.; Cho, N.; Moy, L. Breast MRI: State of the art. Radiology 2019, 292, 520–536. [Google Scholar] [CrossRef]
- Malherbe, K.; Tafti, D. Breast ultrasound. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2024. [Google Scholar]
- Candelaria, R.P.; Hwang, L.; Bouchard, R.R.; Whitman, G.J. Breast ultrasound: Current concepts. Semin Ultrasound CT MRI 2013, 34, 213–225. [Google Scholar] [CrossRef]
- Zanotel, M.; Bednarova, I.; Londero, V.; Linda, A.; Lorenzon, M.; Girometti, R.; Zuiani, C. Automated breast ultrasound: Basic principles and emerging clinical applications. Radiol. Med. 2018, 123, 1–12. [Google Scholar] [CrossRef]
- Liu, X.; Dai, Y.; Wu, Y.; Li, F.; Liang, M.; Wu, Q. Diagnostic accuracy of automated breast volume scanning, hand-held ultrasound and molybdenum-target mammography for breast lesions: A systematic review and meta-analysis. Gland Surg. 2025, 14, 294. [Google Scholar] [CrossRef]
- Dan, Q.; Xu, Z.; Burrows, H.; Bissram, J.; Stringer, J.S.A.; Li, Y. Diagnostic performance of deep learning in ultrasound diagnosis of breast cancer: A systematic review. npj Precis. Oncol. 2024, 8, 21. [Google Scholar] [CrossRef]
- Al-Tam, R.M.; Narangale, S.M. Breast Cancer Detection and Diagnosis Using Machine Learning: A Survey. J. Sci. Res. 2021, 65, 265–285. [Google Scholar] [CrossRef]
- Carriero, A.; Groenhoff, L.; Vologina, E.; Basile, P.; Albera, M. Deep Learning in Breast Cancer Imaging: State of the Art and Recent Advancements in Early 2024. Diagnostics 2024, 14, 848. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.; Misra, S.; Managuli, R.; Barr, R.G.; Lee, J.; Kim, C. Attention-based fusion network for breast cancer segmentation and classification using multi-modal ultrasound images. Ultrasound Med. Biol. 2025, 51, 568–577. [Google Scholar] [CrossRef] [PubMed]
- Munteanu, B.Ş.; Murariu, A.; Nichitean, M.; Nichitean, M.; Pitac, L.-G.; Dioşan, L. Value of Original and Generated Ultrasound Data Towards Training Robust Classifiers for Breast Cancer Identification. Inf. Syst. Front. 2025, 27, 75–96. [Google Scholar] [CrossRef]
- Al-Tam, R.M.; Al-Hejri, A.M.; Alshamrani, S.S.; Al-antari, M.A.; Narangale, S.M. Multimodal breast cancer hybrid explainable computer-aided diagnosis using medical mammograms and ultrasound Images. Biocybern. Biomed. Eng. 2024, 44, 731–758. [Google Scholar] [CrossRef]
- Alotaibi, M.; Aljouie, A.; Alluhaidan, N.; Qureshi, W.; Almatar, H.; Alduhayan, R.; Alsomaie, B.; Almazroa, A. Breast cancer classification based on convolutional neural network and image fusion approaches using ultrasound images. Heliyon 2023, 9, e22406. [Google Scholar] [CrossRef]
- AlZoubi, A.; Lu, F.; Zhu, Y.; Ying, T.; Ahmed, M.; Du, H. Classification of breast lesions in ultrasound images using deep convolutional neural networks: Transfer learning versus automatic architecture design. Med. Biol. Eng. Comput. 2024, 62, 135–149. [Google Scholar] [CrossRef]
- Alom, M.R.; Farid, F.A.; Rahaman, M.A.; Rahman, A.; Debnath, T.; Miah, A.S.M.; Mansor, S. An explainable AI-driven deep neural network for accurate breast cancer detection from histopathological and ultrasound images. Sci. Rep. 2025, 15, 17531. [Google Scholar] [CrossRef]
- Gu, Y.; Xu, W.; Lin, B.; An, X.; Tian, J.; Ran, H.; Ren, W.; Chang, C.; Yuan, J.; Kang, C.; et al. Deep learning based on ultrasound images assists breast lesion diagnosis in China: A multicenter diagnostic study. Insights Imaging 2022, 13, 124. [Google Scholar] [CrossRef] [PubMed]
- Podda, A.S.; Balia, R.; Barra, S.; Carta, S.; Fenu, G.; Piano, L. Fully-automated deep learning pipeline for segmentation and classification of breast ultrasound images. J. Comput. Sci. 2022, 63, 101816. [Google Scholar] [CrossRef]
- Zhang, H.; Han, L.; Chen, K.; Peng, Y.; Lin, J. Diagnostic efficiency of the breast ultrasound computer-aided prediction model based on convolutional neural network in breast cancer. J. Digit. Imaging 2020, 33, 1218–1223. [Google Scholar] [CrossRef]
- Liao, W.-X.; He, P.; Hao, J.; Wang, X.-Y.; Yang, R.-L.; An, D.; Cui, L.-G. Automatic identification of breast ultrasound image based on supervised block-based region segmentation algorithm and features combination migration deep learning model. IEEE J. Biomed. Health Inform. 2019, 24, 984–993. [Google Scholar] [CrossRef]
- Zhou, G.; Mosadegh, B. Distilling knowledge from an ensemble of vision transformers for improved classification of breast ultrasound. Acad. Radiol. 2024, 31, 104–120. [Google Scholar] [CrossRef]
- Islam, M.R.; Rahman, M.M.; Ali, M.S.; Nafi, A.A.N.; Alam, M.S.; Godder, T.K.; Miah, M.S.; Islam, M.K. Enhancing breast cancer segmentation and classification: An Ensemble Deep Convolutional Neural Network and U-net approach on ultrasound images. Mach. Learn. Appl. 2024, 16, 100555. [Google Scholar] [CrossRef]
- Xiao, T.; Liu, L.; Li, K.; Qin, W.; Yu, S.; Li, Z. Comparison of transferred deep neural networks in ultrasonic breast masses discrimination. BioMed Res. Int. 2018, 2018, 4605191. [Google Scholar] [CrossRef]
- Becker, A.S.; Mueller, M.; Stoffel, E.; Marcon, M.; Ghafoor, S.; Boss, A. Classification of breast cancer in ultrasound imaging using a generic deep learning analysis software: A pilot study. Br. J. Radiol. 2018, 91, 20170576. [Google Scholar] [CrossRef] [PubMed]
- Wan, K.W.; Wong, C.H.; Ip, H.F.; Fan, D.; Yuen, P.L.; Fong, H.Y.; Ying, M. Evaluation of the performance of traditional machine learning algorithms, convolutional neural network and AutoML Vision in ultrasound breast lesions classification: A comparative study. Quant. Imaging Med. Surg. 2021, 11, 1381. [Google Scholar] [CrossRef]
- Ejiyi, C.J.; Qin, Z.; Ukwuoma, C.; Agbesi, V.K.; Oluwasanmi, A.; Al-antari, M.A.; Bamisile, O. A unified 2D medical image segmentation network (SegmentNet) through distance-awareness and local feature extraction. Biocybern. Biomed. Eng. 2024, 44, 431–449. [Google Scholar] [CrossRef]
- Sahu, A.; Das, P.K.; Meher, S. An efficient deep learning scheme to detect breast cancer using mammogram and ultrasound breast images. Biomed. Signal Process. Control 2024, 87, 105377. [Google Scholar]
- Chen, J.; Pan, T.; Zhu, Z.; Liu, L.; Zhao, N.; Feng, X.; Zhang, W.; Wu, Y.; Cai, C.; Luo, X.; et al. A deep learning-based multimodal medical imaging model for breast cancer screening. Sci. Rep. 2025, 15, 14696. [Google Scholar] [CrossRef]
- Aumente-Maestro, C.; Díez, J.; Remeseiro, B. A multi-task framework for breast cancer segmentation and classification in ultrasound imaging. Comput. Methods Programs Biomed. 2025, 260, 108540. [Google Scholar]
- Lee, S.E.; Han, K.; Youk, J.H.; Lee, J.E.; Hwang, J.-Y.; Rho, M.; Yoon, J.; Kim, E.-K.; Yoon, J.H. Differing benefits of artificial intelligence-based computer-aided diagnosis for breast US according to workflow and experience level. Ultrasonography 2022, 41, 718–727. [Google Scholar] [CrossRef]
- Wang, N.; Bian, C.; Wang, Y.; Xu, M.; Qin, C.; Yang, X.; Wang, T.; Li, A.; Shen, D.; Ni, D. Densely deep supervised networks with threshold loss for cancer detection in automated breast ultrasound. In Proceedings of the Medical Image Computing and Computer Assisted Intervention—MICCAI 2018: 21st International Conference, Granada, Spain, 16–20 September 2018; Proceedings, Part IV 11. pp. 641–648. [Google Scholar]
- Zhang, S.; Liao, M.; Wang, J.; Zhu, Y.; Zhang, Y.; Zhang, J.; Zheng, R.; Lv, L.; Zhu, D.; Chen, H.; et al. Fully automatic tumor segmentation of breast ultrasound images with deep learning. J. Appl. Clin. Med. Phys. 2023, 24, e13863. [Google Scholar] [PubMed]
- Zhao, Y.; Li, X.; Zhou, C.; Peng, H.; Zheng, Z.; Chen, J.; Ding, W. A review of cancer data fusion methods based on deep learning. Inf. Fusion 2024, 108, 102361. [Google Scholar] [CrossRef]
- Al-Hejri, A.M.; Al-Tam, R.M.; Fazea, M.; Sable, A.H.; Lee, S.; Al-antari, M.A. ETECADx: Ensemble Self-Attention Transformer Encoder for Breast Cancer Diagnosis Using Full-Field Digital X-ray Breast Images. Diagnostics 2022, 13, 89. [Google Scholar] [CrossRef]
- Maruf, N.A.; Basuhail, A.; Ramzan, M.U. Enhanced Breast Cancer Diagnosis Using Multimodal Feature Fusion with Radiomics and Transfer Learning. Diagnostics 2025, 15, 2170. [Google Scholar] [CrossRef]
- Al-Tam, R.M.; Al-Hejri, A.M.; Naji, E.; Hashim, F.A.; Alshamrani, S.S.; Alshehri, A.; Narangale, S.M. A Hybrid Framework of Transformer Encoder and Residential Conventional for Cardiovascular Disease Recognition Using Heart Sounds. IEEE Access 2024, 12, 123099–123113. [Google Scholar] [CrossRef]
- Gu, Z.; Huang, J.; Zhou, C.; Wang, Q.; Kong, J.; You, X.; Zhang, Z.; Zhao, H. Assessing breast cancer volume alterations post-neoadjuvant chemotherapy through DenseNet-201 deep learning analysis on DCE-MRI. J. Radiat. Res. Appl. Sci. 2024, 17, 100971. [Google Scholar] [CrossRef]
- Sharma, S.; Kumar, S. The Xception model: A potential feature extractor in breast cancer histology images classification. ICT Express 2022, 8, 101–108. [Google Scholar] [CrossRef]
- Soulami, K.B.; Kaabouch, N.; Saidi, M.N. Breast cancer: Classification of suspicious regions in digital mammograms based on capsule network. Biomed. Signal Process. Control 2022, 76, 103696. [Google Scholar]
- Yu, X.; Tian, J.; Chen, Z.; Meng, Y.; Zhang, J. Predictive breast cancer diagnosis using ensemble fuzzy model. Image Vis. Comput. 2024, 148, 105146. [Google Scholar] [CrossRef]
- Fan, Z.; Wu, X.; Li, C.; Chen, H.; Liu, W.; Zheng, Y.; Chen, J.; Li, X.; Sun, H.; Jiang, T.; et al. CAM-VT: A weakly supervised cervical cancer nest image identification approach using conjugated attention mechanism and visual transformer. Comput. Biol. Med. 2023, 162, 107070. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Xu, Y.; Lv, T.; Cui, L.; Zhang, C.; Wei, F. Dit: Self-supervised pre-training for document image transformer. In Proceedings of the 30th ACM International Conference on Multimedia, Lisbon, Portugal, 10–14 October 2022; pp. 3530–3539. [Google Scholar]
- Bao, H.; Dong, L.; Piao, S.; Wei, F. Beit: Bert pre-training of image transformers. arXiv 2021, arXiv:2106.08254. [Google Scholar]
- Iqbal, A.; Sharif, M. BTS-ST: Swin transformer network for segmentation and classification of multimodality breast cancer images. Knowl.-Based Syst. 2023, 267, 110393. [Google Scholar] [CrossRef]
- Li, L.; Mei, Z.; Li, Y.; Yu, Y.; Liu, M. A dual data stream hybrid neural network for classifying pathological images of lung adenocarcinoma. Comput. Biol. Med. 2024, 175, 108519. [Google Scholar] [CrossRef]
- Al-Dhabyani, W.; Gomaa, M.; Khaled, H.; Fahmy, A. Dataset of breast ultrasound images. Data Br. 2020, 28, 104863. [Google Scholar] [CrossRef]
- Schwarzhans, F.; George, G.; Sanchez, L.E.; Zaric, O.; Abraham, J.E.; Woitek, R.; Hatamikia, S. Image normalization techniques and their effect on the robustness and predictive power of breast MRI radiomics. Eur. J. Radiol. 2025, 187, 112086. [Google Scholar] [CrossRef]
- Wang, J.; Perez, L. The effectiveness of data augmentation in image classification using deep learning. Convolutional Neural Netw. Vis. Recognit. 2017, 11, 1–8. [Google Scholar]
- Mumuni, A.; Mumuni, F. Data augmentation: A comprehensive survey of modern approaches. Array 2022, 16, 100258. [Google Scholar] [CrossRef]
- Islam, T.; Hafiz, M.S.; Jim, J.R.; Kabir, M.M.; Mridha, M.F. A systematic review of deep learning data augmentation in medical imaging: Recent advances and future research directions. Healthc. Anal. 2024, 5, 100340. [Google Scholar] [CrossRef]
- Tupper, A.; Gagné, C. Analyzing Data Augmentation for Medical Images: A Case Study in Ultrasound Images. arXiv 2024, arXiv:2403.09828. [Google Scholar] [CrossRef]
- Xu, M.; Yoon, S.; Fuentes, A.; Park, D.S. A comprehensive survey of image augmentation techniques for deep learning. Pattern Recognit. 2023, 137, 109347. [Google Scholar] [CrossRef]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv 2020, arXiv:2010.11929. [Google Scholar]
- Han, K.; Wang, Y.; Chen, H.; Chen, X.; Guo, J.; Liu, Z.; Tang, Y.; Xiao, A.; Xu, C.; Xu, Y.; et al. A survey on vision transformer. IEEE Trans. Pattern Anal. Mach. Intell. 2022, 45, 87–110. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. Adv. Neural Inf. Process. Syst. 2017, 30, 1–8. [Google Scholar]
- Pati, A.; Parhi, M.; Pattanayak, B.K.; Singh, D.; Singh, V.; Kadry, S.; Nam, Y.; Kang, B.-G. Breast cancer diagnosis based on IoT and deep transfer learning enabled by fog computing. Diagnostics 2023, 13, 2191. [Google Scholar] [CrossRef]
- Al-antari, M.A.; Al-Tam, R.M.; Al-Hejri, A.M.; Al-Huda, Z.; Lee, S.; Yıldırım, Ö.; Gu, Y.H. A hybrid segmentation and classification CAD framework for automated myocardial infarction prediction from MRI images. Sci. Rep. 2025, 15, 14196. [Google Scholar] [CrossRef]
- Alotaibi, A.; Alafif, T.; Alkhilaiwi, F.; Alatawi, Y.; Althobaiti, H.; Alrefaei, A.; Hawsawi, Y.; Nguyen, T. ViT-DeiT: An Ensemble Model for Breast Cancer Histopathological Images Classification. In Proceedings of the 2023 1st International Conference on Advanced Innovations in Smart Cities (ICAISC), Jeddah, Saudi Arabia, 23–25 January 2023; pp. 1–6. [Google Scholar]
- Seeland, M.; Mäder, P. Multi-view classification with convolutional neural networks. PLoS ONE 2021, 16, e0245230. [Google Scholar] [CrossRef]
- Aldakhil, L.A.; Alhasson, H.F.; Alharbi, S.S. Attention-based deep learning approach for breast cancer histopathological image multi-classification. Diagnostics 2024, 14, 1402. [Google Scholar] [CrossRef]
- Gómez-Flores, W.; Gregorio-Calas, M.J.; de Albuquerque Pereira, W. BUS-BRA: A breast ultrasound dataset for assessing computer-aided diagnosis systems. Med. Phys. 2024, 51, 3110–3123. [Google Scholar] [CrossRef] [PubMed]
- Pawłowska, A.; Ćwierz-Pieńkowska, A.; Domalik, A.; Jaguś, D.; Kasprzak, P.; Matkowski Rafałand Fura, Ł.; Nowicki, A.; Żołek, N. Curated benchmark dataset for ultrasound based breast lesion analysis. Sci. Data 2024, 11, 148. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhang, J.; Zhang, Y.; Li, X.; Ma, X.; Deng, J.; Shen, H.; Wang, D.; Mei, L.; Lei, C. BUSI_WHU: Breast Cancer Ultrasound Image Dataset. Mendeley Data V3 2025, 15, 1751. [Google Scholar] [CrossRef]
- Güler, M.; Sart, G.; Algorabi, Ö.; Adıguzel Tuylu, A.N.; Türkan, Y.S. Breast Cancer Classification with Various Optimized Deep Learning Methods. Diagnostics 2025, 15, 1751. [Google Scholar] [CrossRef]
- Piddubnyi, A.; Kolomiiets, O.; Danilchenko, S.; Stepanenko, A.; Moskalenko, Y.; Moskalenko, R. The prospects of using structural phase analysis of microcalcifications in breast cancer diagnostics. Diagnostics 2023, 13, 737. [Google Scholar] [CrossRef]
- Al-Tam, R.M.; Hashim, F.A.; Maqsood, S.; Abualigah, L.; Alwhaibi, R.M. Enhancing Parkinson’s Disease Diagnosis Through Stacking Ensemble-Based Machine Learning Approach. IEEE Access 2024, 12, 79549–79567. [Google Scholar] [CrossRef]
- Al-Hejri, A.M.; Al-Tam, R.M.; Sable, A.H.; Almuhaya, B.; Alshamrani, S.S.; Alshmrany, K.M. A hybrid vision transformer with ensemble CNN framework for cervical cancer diagnosis. BMC Med. Inform. Decis. Mak. 2025, 25, 411. [Google Scholar] [CrossRef]
- Radosavovic, I.; Kosaraju, R.P.; Girshick, R.; He, K.; Dollár, P. Designing network design spaces. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Seattle, WA, USA, 13–19 June 2020; pp. 10428–10436. [Google Scholar]
- Graham, B.; El-Nouby, A.; Touvron, H.; Stock, P.; Joulin, A.; Jégou, H.; Douze, M. Levit: A vision transformer in convnet’s clothing for faster inference. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, QC, Canada, 10–17 October 2021; pp. 12259–12269. [Google Scholar]
- Al-Hejri, A.M.; Sable, A.H.; Al-Tam, R.M.; Al-Antari, M.A.; Alshamrani, S.S.; Alshmrany, K.M.; Alatebi, W. A hybrid explainable federated-based vision transformer framework for breast cancer prediction via risk factors. Sci. Rep. 2025, 15, 18453. [Google Scholar] [CrossRef] [PubMed]
- Miao, H.; Jia, J.; Cao, Y.; Zhou, Y.; Jiang, Y.; Liu, Z.; Zhai, G. Ultrasound-qbench: Can llms aid in quality assessment of ultrasound imaging? arXiv 2025, arXiv:2501.02751. [Google Scholar] [CrossRef]









| Model | Pre-Trained Variant | Trainable Layers (From Index) | Custom Classification Layers |
|---|---|---|---|
| VGG16 | ImageNet | From layer 17 | 1024 FC → BatchNorm → Dropout (0.5) → Dense (3) |
| VGG19 | ImageNet | From layer 17 | Same as above |
| MobileNetV2 | ImageNet | From layer 131 | Same as above |
| ResNet50 | ImageNet | From layer 123 | Same as above |
| Xception | ImageNet | From layer 96 | Same as above |
| InceptionResNetV2 | ImageNet | From layer 720 | Same as above |
| InceptionV3 | ImageNet | From layer 252 | Same as above |
| DenseNet201 | ImageNet | From layer 481 | Same as above |
| ViT-Hybrid | vit-hybrid-base-bit-384 | Frozen | Same as above |
| DIT | dit-base-finetuned-rvlcdip | Frozen | Same as above |
| Swin | swin-tiny-patch4-window7-224 | Frozen | Same as above |
| Beit | beit-base-patch16-224-pt22k-ft22k | Frozen | Same as above |
| Deit | deit-base-patch16-224 | Frozen | Same as above |
| ViT | vit-base-patch16-224-in21k | Frozen | Same as above |
| Ensemble 1 | DenseNet201 + VGG19 + Xception | Frozen and used as Feature extractor only | Shared: 1024 FC → BatchNorm → Dropout (0.5) → Dense (3) |
| Ensemble 2 | DenseNet201+ VGG16+ Xception | Frozen and used as Feature extractor only | Same as above |
| Ensemble 3 | DenseNet201 + VGG19 + InceptionResNetV2 | Frozen and used as Feature extractor only | Same as above |
| Ensemble 4 | DenseNet201 + VGG16 + InceptionResNetV2 | Frozen and used as Feature extractor only | Same as above |
| Ensemble 5 | DenseNet201 + ResNet50 | Frozen and used as Feature extractor only | Same as above |
| Ensemble 6 | VGG19 + ResNet50 | Frozen and used as Feature extractor only | Same as above |
| Ensemble 7 | VGG16 + ResNet50 | Frozen and used as Feature extractor only | Same as above |
| The proposed Ensemble | Deit + ViT | Frozen and used as Feature extractor only | 1024 FC → BatchNorm → Dropout (0.5) → Dense (2/3/4/6) |
| Model | Silhouette Score | Inter-Class Distance |
|---|---|---|
| ViT | 0.50 | 1.5 |
| Deit | 0.48 | 1.4 |
| Deit + ViT | 0.72 | 2.8 |
| AI Model | Class | FP | Acc. | AUC | Evaluation Matrices (%) | ||
|---|---|---|---|---|---|---|---|
| PRE. | SE. | F1. | |||||
| VGG19 | Benign | 8 | 86.62 | 88.24 | 91.00 | 91.00 | 91.00 |
| Malignant | 7 | 78.00 | 83.00 | 80.00 | |||
| Normal | 6 | 88.00 | 78.00 | 82.00 | |||
| VGG16 | Benign | 8 | 86.62 | 88.14 | 90.00 | 91.00 | 90.00 |
| Malignant | 7 | 80.00 | 83.00 | 81.00 | |||
| Normal | 6 | 88.00 | 78.00 | 82.00 | |||
| MobileNetV2 | Benign | 7 | 78.98 | 78.68 | 77.00 | 92.00 | 84.00 |
| Malignant | 14 | 85.00 | 67.00 | 75.00 | |||
| Normal | 12 | 79.00 | 56.00 | 65.00 | |||
| ResNet50 | Benign | 11 | 88.54 | 91.65 | 96.00 | 88.00 | 92.00 |
| Malignant | 4 | 78.00 | 90.00 | 84.00 | |||
| Normal | 3 | 86.00 | 89.00 | 87.00 | |||
| DenseNet201 | Benign | 8 | 86.62 | 87.98 | 88.00 | 91.00 | 89.00 |
| Malignant | 7 | 88.00 | 83.00 | 85.00 | |||
| Normal | 6 | 81.00 | 78.00 | 79.00 | |||
| Xception | Benign | 11 | 80.89 | 82.30 | 82.00 | 88.00 | 85.00 |
| Malignant | 9 | 85.00 | 79.00 | 81.00 | |||
| Normal | 10 | 71.00 | 63.00 | 67.00 | |||
| InceptionResNetV2 | Benign | 20 | 70.70 | 74.72 | 83.00 | 77.00 | 80.00 |
| Malignant | 11 | 57.00 | 74.00 | 65.00 | |||
| Normal | 15 | 57.00 | 44.00 | 50.00 | |||
| InceptionV3 | Benign | 10 | 80.89 | 82.09 | 80.00 | 89.00 | 84.00 |
| Malignant | 11 | 84.00 | 74.00 | 78.00 | |||
| Normal | 9 | 78.00 | 67.00 | 62.00 | |||
| AI Model | Class | FP | Acc. | AUC | Evaluation Matrices (%) | ||
|---|---|---|---|---|---|---|---|
| PRE. | SE. | F1. | |||||
| ViT-Hybrid | Benign | 9 | 86.62 | 88.02 | 87.00 | 90.00 | 88.00 |
| Malignant | 6 | 82.00 | 86.00 | 84.00 | |||
| Normal | 6 | 95.00 | 78.00 | 86.00 | |||
| ViT | Benign | 4 | 93.63 | 93.98 | 93.00 | 95.00 | 94.00 |
| Malignant | 2 | 95.00 | 95.00 | 95.00 | |||
| Normal | 4 | 92.00 | 85.00 | 88.00 | |||
| Deit | Benign | 6 | 91.72 | 92.64 | 92.00 | 93.00 | 93.00 |
| Malignant | 3 | 87.00 | 93.00 | 90.00 | |||
| Normal | 4 | 100.0 | 85.00 | 92.00 | |||
| Dit | Benign | 8 | 84.71 | 86.14 | 84.00 | 91.00 | 87.00 |
| Malignant | 11 | 82.00 | 74.00 | 78.00 | |||
| Normal | 5 | 92.00 | 81.00 | 86.00 | |||
| Swin | Benign | 7 | 90.45 | 91.89 | 91.00 | 92.00 | 92.00 |
| Malignant | 5 | 84.00 | 88.00 | 86.00 | |||
| Normal | 3 | 100.0 | 89.00 | 94.00 | |||
| Beit | Benign | 4 | 90.45 | 90.63 | 89.00 | 95.00 | 92.00 |
| Malignant | 6 | 88.00 | 86.00 | 87.00 | |||
| Normal | 5 | 100.0 | 81.00 | 90.00 | |||
| AI Model | Class | FP | Acc. | AUC | Evaluation Matrices (%) | ||
|---|---|---|---|---|---|---|---|
| PRE. | SE. | F1. | |||||
| DenseNet201 + VGG19 + Xception | Benign | 5 | 89.81 | 89.75 | 90.00 | 94.00 | 92.00 |
| Malignant | 3 | 87.00 | 93.00 | 90.00 | |||
| Normal | 8 | 95.00 | 70.00 | 81.00 | |||
| DenseNet201+ VGG16+ Xception | Benign | 3 | 88.54 | 87.63 | 87.00 | 97.00 | 91.00 |
| Malignant | 6 | 88.00 | 68.00 | 87.00 | |||
| Normal | 9 | 100.0 | 67.00 | 81.00 | |||
| DenseNet201 + VGG19 + InceptionResNetV2 | Benign | 5 | 90.45 | 91.09 | 91.00 | 94.00 | 93.00 |
| Malignant | 5 | 88.00 | 88.00 | 88.00 | |||
| Normal | 5 | 92.00 | 81.00 | 86.00 | |||
| DenseNet201 + VGG16 + InceptionResNetV2 | Benign | 5 | 89.81 | 89.96 | 90.00 | 94.00 | 92.00 |
| Malignant | 4 | 84.00 | 90.00 | 87.00 | |||
| Normal | 7 | 100.0 | 74.00 | 85.00 | |||
| DenseNet201 + ResNet50 | Benign | 6 | 89.31 | 91.00 | 89.00 | 93.00 | 91.00 |
| Malignant | 7 | 92.00 | 83.00 | 88.00 | |||
| Normal | 3 | 89.00 | 89.00 | 89.00 | |||
| VGG19 + ResNet50 | Benign | 6 | 89.17 | 89.97 | 89.00 | 93.00 | 91.00 |
| Malignant | 6 | 84.00 | 86.00 | 85.00 | |||
| Normal | 5 | 100.0 | 81.00 | 90.00 | |||
| VGG16 + ResNet50 | Benign | 4 | 91.08 | 91.49 | 90.00 | 95.00 | 93.00 |
| Malignant | 6 | 88.00 | 86.00 | 87.00 | |||
| Normal | 4 | 100.0 | 85.00 | 92.00 | |||
| Deit + ViT | Benign | 4 | 94.27 | 94.81 | 94.00 | 95.00 | 95.00 |
| Malignant | 2 | 91.00 | 95.00 | 93.00 | |||
| Normal | 3 | 100.0 | 89.00 | 94.00 | |||
| AI Model | Dataset | Class | FP | Acc. | AUC | Evaluation Matrices (%) | ||
|---|---|---|---|---|---|---|---|---|
| PRE. | SE. | F1. | ||||||
| Deit + ViT | BUSI | Benign | 3 | 96.92 | 97.10 | 99.00 | 97.00 | 98.00 |
| Malignant | 1 | 93.00 | 98.00 | 95.00 | ||||
| BUS-BRA | Benign | 30 | 86.77 | 85.70 | 92.00 | 88.00 | 90.00 | |
| Malignant | 25 | 77.00 | 83.00 | 80.00 | ||||
| BrEaST | Benign | 4 | 87.76 | 88.07 | 93.00 | 87.00 | 90.00 | |
| Malignant | 2 | 81.00 | 89.00 | 85.00 | ||||
| BUSI_WHU | Benign | 8 | 86.99 | 86.99 | 86.00 | 89.00 | 87.00 | |
| Malignant | 11 | 90.00 | 85.00 | 87.00 | ||||
| AI Model | Dataset | Class | FP | Acc. | AUC | Evaluation Matrices (%) | ||
|---|---|---|---|---|---|---|---|---|
| PRE. | SE. | F1. | ||||||
| Deit + ViT | BrEaST | 2 | 1 | 68.75 | 81.32 | 71.00 | 83.00 | 77.00 |
| 3 | 2 | 71.00 | 71.00 | 71.00 | ||||
| 4a | 3 | 83.00 | 62.00 | 71.00 | ||||
| 4b | 3 | 60.00 | 67.00 | 63.00 | ||||
| 4c | 3 | 55.00 | 67.00 | 60.00 | ||||
| 5 | 3 | 86.00 | 67.00 | 75.00 | ||||
| BUS-BRA | 2 | 26 | 76.68 | 84.76 | 80.00 | 77.00 | 78.00 | |
| 3 | 20 | 70.00 | 76.00 | 73.00 | ||||
| 4 | 28 | 91.00 | 80.00 | 85.00 | ||||
| 5 | 11 | 45.00 | 65.00 | 53.00 | ||||
| Model | No. of Parameters (Million) | Training Time/ Epoch (Msec) | Testing Time/ Image (s) | Frame Per Second (FPS) |
|---|---|---|---|---|
| ResNet50 | 50.39 | 151 | 0.0110 | 90.90 |
| ViT | 87.18 | 218 | 0.0180 | 55.55 |
| Deit | 87.18 | 218 | 0.0180 | 55.55 |
| VGG16 + ResNet50 | 70.17 | 380 | 0.025 | 40.00 |
| The proposed CAD (ensemble of Deit + ViT) | 174.36 | 490 | 0.032 | 31.25 |
| Reference | Dataset | Labels | Methodology | Accuracy (%) |
|---|---|---|---|---|
| Becker A.S. et al. (2018) [34] | Private dataset/BUSI | Benign/Malignant | Generic DL | 96 (AUC) |
| Xiao T. et al. (2018) [33] | Private dataset/BUSI | Benign/Malignant | CNN | 74.44 (Acc.) 78 (AUC) |
| Wang Y. et al. (2018) [41] | Private dataset/BUSI | Benign/Malignant | DCNN | 95 (Se.) |
| Liao W.X. et al. (2020) [30] | Data were collected in the Peking University Third Hospital. /BUSI | Benign/Malignant | VGG19 | 90.38 (Acc.) 97 (AUC) |
| Zhang H. et al. (2020) [29] | Data were collected from different hospitals./BUSI | Benign/Malignant | InceptionV3 | 82.8 (Acc.) 90.5 (AUC) |
| Wan K.W. et al. (2021) [35] | Mendeley dataset and dataset from Baheya hospital./BUSI | Benign/Malignant/Normal | CNN | 91 (Acc.) |
| Gu Y. et al. (2022) [27] | Data were collected from 32 hospitals./BUSI | Benign/Malignant | VGG-DCNN | 86.40 (Acc.) 91.3 (AUC) |
| Lee S.E. et al. (2022) [40] | Private dataset/BUSI | Benign/Malignant | AI-CAD | 85.4 (Acc.) 85.5 (AUC) |
| Alotaibi, Mohammed et al. (2023) [24] | BUSI | Benign/Malignant | VGG19 | 87.8 (Acc.), 83.8 (F1-score), 94.63 (AUC) |
| Zhang S. et al. (2023) [42] | Private dataset/BUSI | Normal/abnormal | U-NET + DenseNet | 96 (Acc.) 99 (AUC) |
| Ejiyi, Chukwuebuka Joseph et al.(2024) [36] | BUSI | Benign/Malignant/Normal | SegmentNet | 93.88 (Acc.) |
| Islam Rakibul et al. (2024) [32] | BUSI | Benign/Malignant/Normal | EDCNN | 87.82 (Acc.) |
| Sahu, Adyasha et al. (2024) [37] | BUSI | Benign/Malignant | Ensemble of (AlexNet + ResNet + MobileNetV2) | 94.62 (Acc.) in identifying malignancies. |
| Altameemi et al. (2025) [26] | BUSI | Benign/Malignant/Normal | DenseNet121 with custom CNN | 89.87 (Acc.) 90.00 (F1-score) 89.87(Se.) |
| Carlos A. et al.(2025) [39] | BUSI | Benign/Malignant/Normal | UNet++ | 80.20 (Acc.) |
| The proposed model | BUSI | Benign/Malignant/Normal | Ensemble-based Deit + ViT | 94.27 (Acc.), 93.19 (F1-s., Pre., Se.), 94.81 (AUC) |
| Benign/Malignant | 96.92 (Acc.), 93.18 (Pre.), 97.62 (Se.), 95.35 (F1-s.), 97.10 (AUC) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Al-Tam, R.M.; Al-Hejri, A.M.; Hashim, F.A.; Narangale, S.M.; Al-Antari, M.A.; Alzakari, S.A. An Interpretable Ensemble Transformer Framework for Breast Cancer Detection in Ultrasound Images. Diagnostics 2026, 16, 622. https://doi.org/10.3390/diagnostics16040622
Al-Tam RM, Al-Hejri AM, Hashim FA, Narangale SM, Al-Antari MA, Alzakari SA. An Interpretable Ensemble Transformer Framework for Breast Cancer Detection in Ultrasound Images. Diagnostics. 2026; 16(4):622. https://doi.org/10.3390/diagnostics16040622
Chicago/Turabian StyleAl-Tam, Riyadh M., Aymen M. Al-Hejri, Fatma A. Hashim, Sachin M. Narangale, Mugahed A. Al-Antari, and Sarah A. Alzakari. 2026. "An Interpretable Ensemble Transformer Framework for Breast Cancer Detection in Ultrasound Images" Diagnostics 16, no. 4: 622. https://doi.org/10.3390/diagnostics16040622
APA StyleAl-Tam, R. M., Al-Hejri, A. M., Hashim, F. A., Narangale, S. M., Al-Antari, M. A., & Alzakari, S. A. (2026). An Interpretable Ensemble Transformer Framework for Breast Cancer Detection in Ultrasound Images. Diagnostics, 16(4), 622. https://doi.org/10.3390/diagnostics16040622








