Advances and Challenges in SARS-CoV-2 Detection: A Review of Molecular and Serological Technologies
Abstract
1. Introduction
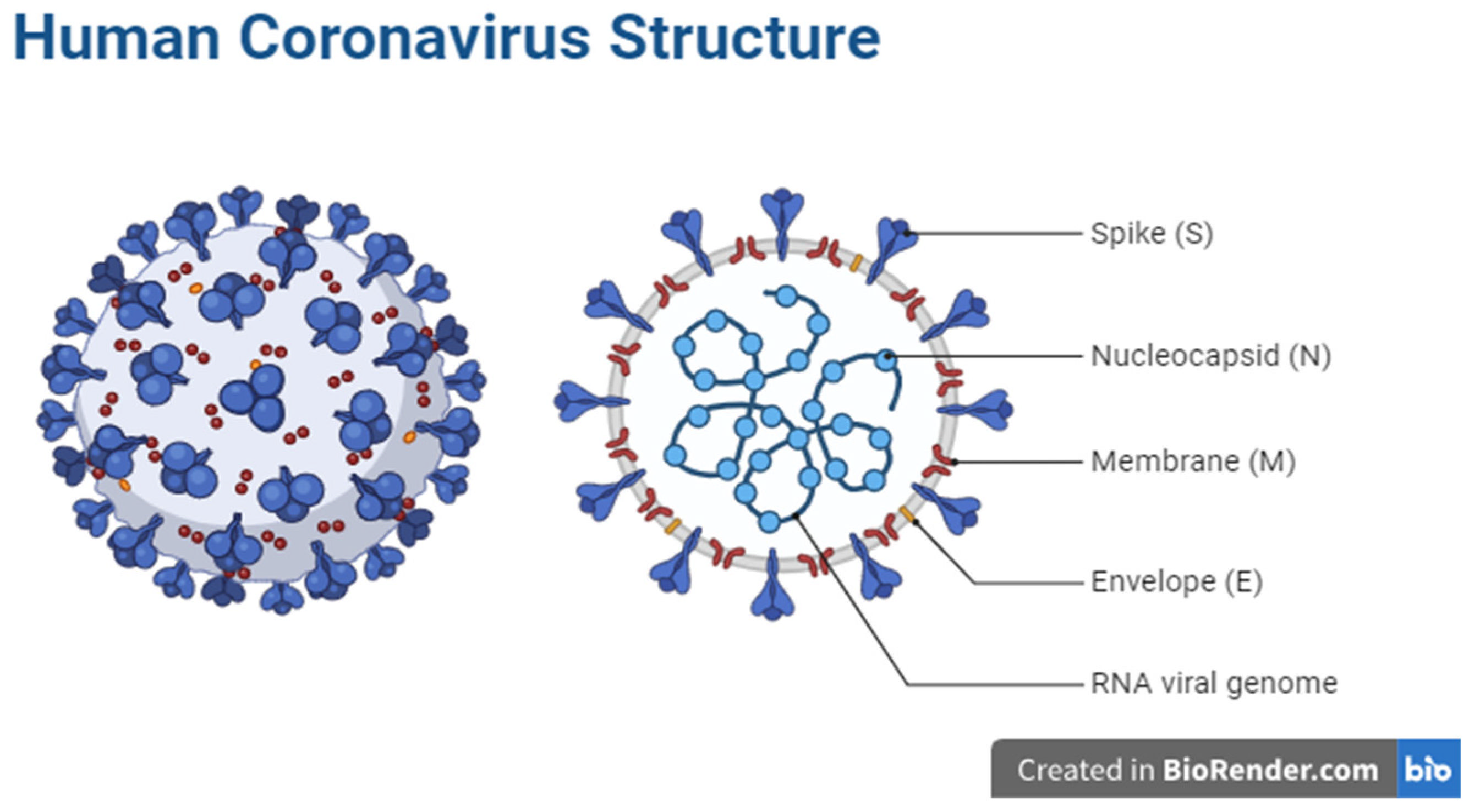
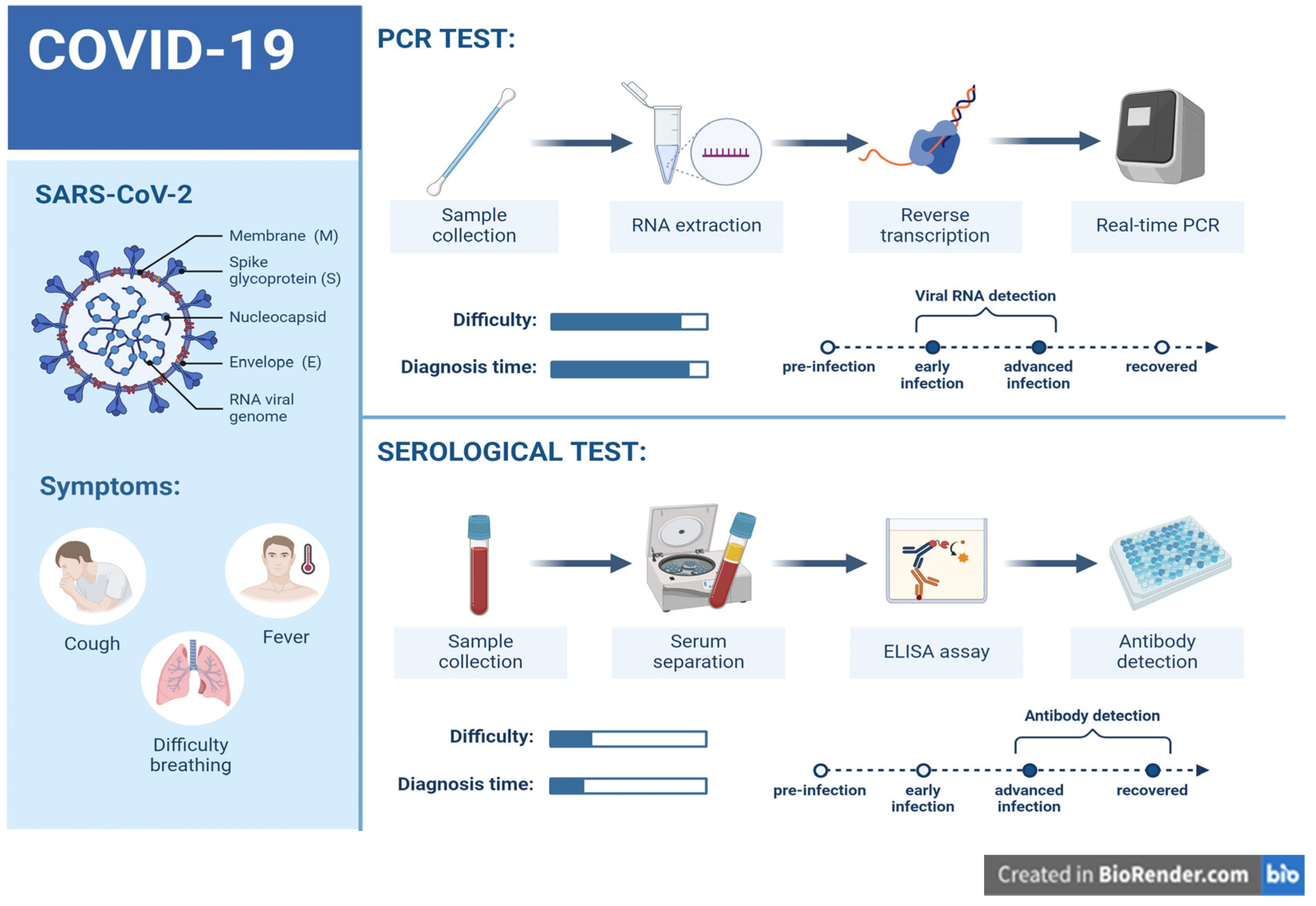
- SARS-CoV-2 Genome and Structure
- Distribution of SARS-CoV-2 in Body Fluids and Tissues
- Collection of COVID-19 Samples
- SARS-CoV-2 Detection Methods
- A. Viral Gene Detection by Molecular Methods
2. Nucleic Acid Amplification Techniques (NAATs)
2.1. Real-Time Reverse Transcriptase Polymerase Chain Reaction (RT-qPCR)
2.2. Sequencing for Diagnostic Purposes
2.3. Digital PCR
3. Isothermal Amplification Techniques
3.1. Reverse Transcriptase Loop-Mediated Isothermal Amplification (RT-LAMP)
3.2. Nucleic Acid Sequence-Based Amplification (NASBA)
3.3. Recombinase Polymerase Amplification (RPA)
3.4. CRISPR-Based Detection Technique
- B. Serological assays
- Detection of SARS-CoV-2 antibodies
- C. SARS-CoV-2 Antigens
4. Conclusions
Funding
Conflicts of Interest
References
- Rai, P.; Kumar, B.K.; Deekshit, V.K.; Karunasagar, I.; Karunasagar, I. Detection technologies and recent developments in the diagnosis of COVID-19 infection. Appl. Microbiol. Biotechnol. 2021, 105, 441–455. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhang, L.; Xie, Y.-H.; Wu, J. Advancements in detection of SARS-CoV-2 infection for confronting COVID-19 pandemics. Lab. Investig. 2022, 102, 4–13. [Google Scholar] [CrossRef]
- Su, S.; Wong, G.; Shi, W.; Liu, J.; Lai, A.C.K.; Zhou, J.; Liu, W.; Bi, Y.; Gao, G.F. Epidemiology, Genetic Recombination, and Pathogenesis of Coronaviruses. Trends Microbiol. 2016, 24, 490–502. [Google Scholar] [CrossRef]
- Perlman, S.; Netland, J. Coronaviruses post-SARS: Update on replication and pathogenesis. Nat. Rev. Microbiol. 2009, 7, 439–450. [Google Scholar] [CrossRef]
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.-M.; Wang, W.; Song, Z.-G.; Hu, Y.; Tao, Z.-W.; Tian, J.-H.; Pei, Y.-Y.; et al. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef]
- Abduljalil, J.; Abduljalil, B. Epidemiology, genome, and clinical features of the pandemic SARS-CoV-2: A recent view. New Microbes New Infect. 2020, 35, 100672. [Google Scholar] [CrossRef]
- Parsamanesh, N.; Pezeshgi, A.; Hemmati, M.; Jameshorani, M.; Saboory, E. Neurological manifestations of coronavirus infections: Role of angiotensin-converting enzyme 2 in COVID-19. Int. J. Neurosci. 2022, 132, 917–924. [Google Scholar] [CrossRef]
- Naqvi, A.A.T.; Fatima, K.; Mohammad, T.; Fatima, U.; Singh, I.K.; Singh, A.; Atif, S.M.; Hariprasad, G.; Hasan, G.M.; Hassan, I. Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: Structural genomics approach. Biochim. Biophys. Acta (BBA)-Mol. Basis Dis. 2020, 1866, 165878. [Google Scholar] [CrossRef]
- Krichel, B.; Falke, S.; Hilgenfeld, R.; Redecke, L.; Uetrecht, C. Processing of the SARS-CoV pp1a/ab nsp7–10 region. Biochem. J. 2020, 477, 1009–1019. [Google Scholar] [CrossRef]
- Khailany, R.A.; Safdar, M.; Ozaslan, M. Genomic characterization of a novel SARS-CoV-2. Gene Rep. 2020, 19, 100682. [Google Scholar] [CrossRef]
- Mu, J.; Xu, J.; Zhang, L.; Shu, T.; Wu, D.; Huang, M.; Ren, Y.; Li, X.; Geng, Q.; Xu, Y.; et al. SARS-CoV-2-encoded nucleocapsid protein acts as a viral suppressor of RNA interference in cells. Sci. China Life Sci. 2020, 63, 1413–1416. [Google Scholar] [CrossRef]
- Voß, D.; Pfefferle, S.; Drosten, C.; Stevermann, L.; Traggiai, E.; Lanzavecchia, A.; Becker, S. Studies on membrane topology, N-glycosylation and functionality of SARS-CoV membrane protein. Virol. J. 2009, 6, 79. [Google Scholar] [CrossRef]
- Korber, B.; Fischer, W.M.; Gnanakaran, S.; Yoon, H.; Theiler, J.; Abfalterer, W.; Hengartner, N.; Giorgi, E.E.; Bhattacharya, T.; Foley, B.; et al. Tracking Changes in SARS-CoV-2 Spike: Evidence that D614G Increases Infectivity of the COVID-19 Virus. Cell 2020, 182, 812–827.e19. [Google Scholar] [CrossRef]
- Kevadiya, B.D.; Machhi, J.; Herskovitz, J.; Oleynikov, M.D.; Blomberg, W.R.; Bajwa, N.; Soni, D.; Das, S.; Hasan, M.; Patel, M.; et al. Diagnostics for SARS-CoV-2 infections. Nat. Mater. 2021, 20, 593–605. [Google Scholar] [CrossRef]
- Zhang, W.; Du, R.-H.; Li, B.; Zheng, X.-S.; Yang, X.-L.; Hu, B.; Wang, Y.-Y.; Xiao, G.-F.; Yan, B.; Shi, Z.-L.; et al. Molecular and serological investigation of 2019-nCoV infected patients: Implication of multiple shedding routes. Emerg. Microbes Infect. 2020, 9, 386–389. [Google Scholar] [CrossRef]
- Yongchen, Z.; Shen, H.; Wang, X.; Shi, X.; Li, Y.; Yan, J.; Chen, Y.; Gu, B. Different longitudinal patterns of nucleic acid and serology testing results based on disease severity of COVID-19 patients. Emerg. Microbes Infect. 2020, 9, 833–836. [Google Scholar] [CrossRef]
- Azzi, L.; Carcano, G.; Gianfagna, F.; Grossi, P.; Gasperina, D.D.; Genoni, A.; Fasano, M.; Sessa, F.; Tettamanti, L.; Carinci, F.; et al. Saliva is a reliable tool to detect SARS-CoV-2. J. Infect. 2020, 81, e45–e50. [Google Scholar] [CrossRef]
- Pfaffe, T.; Cooper-White, J.; Beyerlein, P.; Kostner, K.; Punyadeera, C. Diagnostic potential of saliva: Current state and future applications. Clin. Chem. 2011, 57, 675–687. [Google Scholar] [CrossRef]
- Punyadeera, C.; Dimeski, G.; Kostner, K.; Beyerlein, P.; Cooper-White, J. One-step homogeneous C-reactive protein assay for saliva. J. Immunol. Methods 2011, 373, 19–25. [Google Scholar] [CrossRef]
- Cao, R.; Bao, L.; Pan, M.; Zhang, C.; Liao, H.; Liu, L.; Li, Y.; Li, M. Detection of SARS-CoV-2 in fecal samples with different pretreatment methods and PCR kits. BMC Microbiol. 2021, 21, 56. [Google Scholar] [CrossRef]
- Wyllie, A.L.; Fournier, J.; Casanovas-Massana, A.; Campbell, M.; Tokuyama, M.; Vijayakumar, P.; Warren, J.L.; Geng, B.; Muenker, M.C.; Moore, A.J.; et al. Saliva or Nasopharyngeal Swab Specimens for Detection of SARS-CoV-2. N. Engl. J. Med. 2020, 383, 1283–1286. [Google Scholar] [CrossRef]
- Petruzzi, G.; De Virgilio, A.; Pichi, B.; Mazzola, F.; Zocchi, J.; Mercante, G.; Spriano, G.; Pellini, R. COVID-19: Nasal and oropharyngeal swab. Head Neck 2020, 42, 1303–1304. [Google Scholar] [CrossRef]
- Kinloch, N.N.; Ritchie, G.; Brumme, C.J.; Dong, W.; Dong, W.; Lawson, T.; Jones, R.B.; Montaner, J.S.G.; Leung, V.; Romney, M.G.; et al. Suboptimal Biological Sampling as a Probable Cause of False-Negative COVID-19 Diagnostic Test Results. J. Infect. Dis. 2020, 222, 899–902. [Google Scholar] [CrossRef]
- Piras, A.; Rizzo, D.; Uzzau, S.; De Riu, G.; Rubino, S.; Bussu, F. Inappropriate Nasopharyngeal Sampling for SARS-CoV-2 Detection Is a Relevant Cause of False-Negative Reports. Otolaryngol. Neck Surg. 2020, 163, 459–461. [Google Scholar] [CrossRef]
- Irifune, S.; Ashizawa, N.; Takazono, T.; Mutantu, P.; Nabeshima, T.; Tun, M.M.N.; Ota, K.; Hirayama, T.; Fujita, A.; Tashiro, M.; et al. Discrepancy of SARS-CoV-2 PCR results due to the sample collection sites and possible improper sampling. J. Infect. Chemother. 2021, 27, 1525–1528. [Google Scholar] [CrossRef]
- Pondaven-Letourmy, S.; Alvin, F.; Boumghit, Y.; Simon, F. How to perform a nasopharyngeal swab in adults and children in the COVID-19 era. Eur. Ann. Otorhinolaryngol. Head Neck Dis. 2020, 137, 325–327. [Google Scholar] [CrossRef]
- Basso, D.; Aita, A.; Navaglia, F.; Franchin, E.; Fioretto, P.; Moz, S.; Bozzato, D.; Zambon, C.-F.; Martin, B.; Prà, C.D.; et al. SARS-CoV-2 RNA identification in nasopharyngeal swabs: Issues in pre-analytics. Clin. Chem. Lab. Med. 2020, 58, 1579–1586. [Google Scholar] [CrossRef]
- Filchakova, O.; Dossym, D.; Ilyas, A.; Kuanysheva, T.; Abdizhamil, A.; Bukasov, R. Review of COVID-19 testing and diagnostic methods. Talanta 2022, 244, 123409. [Google Scholar] [CrossRef]
- Pascarella, G.; Strumia, A.; Piliego, C.; Bruno, F.; Del Buono, R.; Costa, F.; Scarlata, S.; Agrò, F.E. COVID-19 diagnosis and management: A comprehensive review. J. Intern. Med. 2020, 288, 192–206. [Google Scholar] [CrossRef]
- Kang, T.; Lu, J.; Yu, T.; Long, Y.; Liu, G. Advances in nucleic acid amplification techniques (NAATs): COVID-19 point-of-care diagnostics as an example. Biosens. Bioelectron. 2022, 206, 114109. [Google Scholar] [CrossRef]
- Xiao, A.T.; Tong, Y.X.; Zhang, S. Profile of RT-PCR for SARS-CoV-2: A Preliminary Study From 56 COVID-19 Patients. Clin. Infect. Dis. 2020, 71, 2249–2251. [Google Scholar] [CrossRef]
- Feng, W.; Newbigging, A.M.; Le, C.; Pang, B.; Peng, H.; Cao, Y.; Wu, J.; Abbas, G.; Song, J.; Wang, D.-B.; et al. Molecular Diagnosis of COVID-19: Challenges and Research Needs. Anal. Chem. 2020, 92, 10196–10209. [Google Scholar] [CrossRef]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.W.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 2020, 25, 2000045. [Google Scholar] [CrossRef]
- Perveen, S.; Negi, A.; Gopalakrishnan, V.; Panda, S.; Sharma, V.; Sharma, R. COVID-19 diagnostics: Molecular biology to nanomaterials. Clin. Chim. Acta 2023, 538, 139–156. [Google Scholar] [CrossRef]
- Tang, Y.-W.; Schmitz, J.E.; Persing, D.H.; Stratton, C.W. The Laboratory Diagnosis of COVID-19 Infection: Current Issues and Challenges. J. Clin. Microbiol. 2020, 58, e00512-20. [Google Scholar] [CrossRef]
- Chan, J.F.-W.; Yip, C.C.-Y.; To, K.K.-W.; Tang, T.H.-C.; Wong, S.C.-Y.; Leung, K.-H.; Fung, A.Y.-F.; Ng, A.C.-K.; Zou, Z.; Tsoi, H.-W.; et al. Improved Molecular Diagnosis of COVID-19 by the Novel, Highly Sensitive and Specific COVID-19-RdRp/Hel Real-Time Reverse Transcription-PCR Assay Validated In Vitro and with Clinical Specimens. J. Clin. Microbiol. 2020, 58. [Google Scholar] [CrossRef]
- Bru, D.; Martin-Laurent, F.; Philippot, L. Quantification of the detrimental effect of a single primer-template mismatch by real-time PCR using the 16S rRNA gene as an example. Appl. Environ. Microbiol. 2008, 74, 1660–1663. [Google Scholar] [CrossRef]
- Farkas, C.; Fuentes-Villalobos, F.; Garrido, J.L.; Haigh, J.; Barría, M.I. Insights on early mutational events in SARS-CoV-2 virus reveal founder effects across geographical regions. PeerJ 2020, 8, e9255. [Google Scholar] [CrossRef] [PubMed]
- Koyama, T.; Platt, D.; Parida, L. Variant analysis of SARS-CoV-2 genomes. Bull. World Health Organ. 2020, 98, 495–504. [Google Scholar] [CrossRef] [PubMed]
- Kaden, R. Early Phylogenetic Diversification of SARS-CoV-2: Determination of Variants and the Effect on Epidemiology, Immunology, and Diagnostics. J. Clin. Med. 2020, 9, 1615. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef] [PubMed]
- Udugama, B.; Kadhiresan, P.; Kozlowski, H.N.; Malekjahani, A.; Osborne, M.; Li, V.Y.C.; Chen, H.; Mubareka, S.; Gubbay, J.B.; Chan, W.C.W. Diagnosing COVID-19: The Disease and Tools for Detection. ACS Nano 2020, 14, 3822–3835. [Google Scholar] [CrossRef] [PubMed]
- Hadfield, J.; Megill, C.; Bell, S.M.; Huddleston, J.; Potter, B.; Callender, C.; Sagulenko, P.; Bedford, T.; Neher, R.A. Nextstrain: Real-time tracking of pathogen evolution. Bioinformatics 2018, 34, 4121–4123. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.W.; Tambyah, P.A.; Hui, D.S. Emergence of a new SARS-CoV-2 variant in the UK. J. Infect. 2021, 82, e27–e28. [Google Scholar] [CrossRef] [PubMed]
- Makoni, M. South Africa responds to new SARS-CoV-2 variant. Lancet 2021, 397, 267. [Google Scholar] [CrossRef] [PubMed]
- Lei, S.; Chen, S.; Zhong, Q. Digital PCR for accurate quantification of pathogens: Principles, applications, challenges and future prospects. Int. J. Biol. Macromol. 2021, 184, 750–759. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Balda, S.; Apreja, M.; Kataria, K.; Capalash, N.; Sharma, P. COVID-19 Diagnosis: Current and Future Techniques. Int. J. Biol. Macromol. 2021, 193, 1835–1844. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Ding, C.; Chen, Q.; Xie, J.; Yu, J.; Shi, Y.; Jiang, C.; Zhang, Z.; He, H.; Ge, Y.; et al. Digital PCR assay for the effective detection of COVID-19 patients with SARS-CoV-2 low viral load. J. Virol. Methods 2021, 295, 114185. [Google Scholar] [CrossRef]
- Dong, L.; Zhou, J.; Niu, C.; Wang, Q.; Pan, Y.; Sheng, S.; Wang, X.; Zhang, Y.; Yang, J.; Liu, M.; et al. Highly accurate and sensitive diagnostic detection of SARS-CoV-2 by digital PCR. Talanta 2021, 224, 121726. [Google Scholar] [CrossRef]
- Ternovoi, V.A.; Lutkovsky, R.Y.; Ponomareva, E.P.; Gladysheva, A.V.; Chub, E.V.; Tupota, N.L.; Smirnova, A.M.; Nazarenko, A.A.; Loktev, V.B.; Gavrilova, E.V.; et al. Detection of SARS-CoV-2 RNA in nasopharyngeal swabs from COVID-19 patients and asymptomatic cases of infection by real-time and digital PCR. Russ. Clin. Lab. Diagn. 2020, 65, 785–792. [Google Scholar] [CrossRef]
- Kim, K.B.; Choi, H.; Lee, G.D.; Lee, J.; Lee, S.; Kim, Y.; Cho, S.-Y.; Lee, D.-G.; Kim, M. Analytical and Clinical Performance of Droplet Digital PCR in the Detection and Quantification of SARS-CoV-2. Mol. Diagn. Ther. 2021, 25, 617–628. [Google Scholar] [CrossRef]
- Suo, T.; Liu, X.; Feng, J.; Guo, M.; Hu, W.; Guo, D.; Ullah, H.; Yang, Y.; Zhang, Q.; Wang, X.; et al. ddPCR: A more accurate tool for SARS-CoV-2 detection in low viral load specimens. Emerg. Microbes Infect. 2020, 9, 1259–1268. [Google Scholar] [CrossRef]
- Poggio, P.; Songia, P.; Vavassori, C.; Ricci, V.; Banfi, C.; Barbieri, S.S.; Garoffolo, G.; Myasoedova, V.A.; Piacentini, L.; Raucci, A.; et al. Digital PCR for high sensitivity viral detection in false-negative SARS-CoV-2 patients. Sci. Rep. 2021, 11, 4310. [Google Scholar] [CrossRef]
- Huang, W.E.; Lim, B.; Hsu, C.; Xiong, D.; Wu, W.; Yu, Y.; Jia, H.; Wang, Y.; Zeng, Y.; Ji, M.; et al. RT-LAMP for rapid diagnosis of coronavirus SARS-CoV-2. Microb. Biotechnol. 2020, 13, 950–961. [Google Scholar] [CrossRef] [PubMed]
- El-Kafrawy, S.A.; El-Daly, M.M.; Hassan, A.M.; Kaki, R.M.; Abuzenadah, A.M.; Kamal, M.A.; Azhar, E.I. A Direct Method for RT-PCR Detection of SARS-CoV-2 in Clinical Samples. Healthcare 2021, 9, 37. [Google Scholar] [CrossRef] [PubMed]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, E63. [Google Scholar] [CrossRef] [PubMed]
- Mathuria, J.P.; Yadav, R. Rajkumar Laboratory diagnosis of SARS-CoV-2—A review of current methods. J. Infect. Public Health 2020, 13, 901–905. [Google Scholar] [CrossRef] [PubMed]
- Augustine, R.; Hasan, A.; Das, S.; Ahmed, R.; Mori, Y.; Notomi, T.; Kevadiya, B.D.; Thakor, A.S. Loop-Mediated Isothermal Amplification (LAMP): A Rapid, Sensitive, Specific, and Cost-Effective Point-of-Care Test for Coronaviruses in the Context of COVID-19 Pandemic. Biology 2020, 9, 182. [Google Scholar] [CrossRef] [PubMed]
- Amaral, C.; Antunes, W.; Moe, E.; Duarte, A.G.; Lima, L.M.P.; Santos, C.; Gomes, I.L.; Afonso, G.S.; Vieira, R.; Teles, H.S.S.; et al. A molecular test based on RT-LAMP for rapid, sensitive and inexpensive colorimetric detection of SARS-CoV-2 in clinical samples. Sci. Rep. 2021, 11, 16430. [Google Scholar] [CrossRef] [PubMed]
- Thi, V.L.D.; Herbst, K.; Boerner, K.; Meurer, M.; Kremer, L.P.; Kirrmaier, D.; Freistaedter, A.; Papagiannidis, D.; Galmozzi, C.; Stanifer, M.L.; et al. A colorimetric RT-LAMP assay and LAMP-sequencing for detecting SARS-CoV-2 RNA in clinical samples. Sci. Transl. Med. 2020, 12, eabc7075. [Google Scholar] [CrossRef]
- Nagamine, K.; Hase, T.; Notomi, T. Accelerated reaction by loop-mediated isothermal amplification using loop primers. Mol. Cell. Probes 2002, 16, 223–229. [Google Scholar] [CrossRef] [PubMed]
- Thai, H.T.C.; Le, M.Q.; Vuong, C.D.; Parida, M.; Minekawa, H.; Notomi, T.; Hasebe, F.; Morita, K. Development and evaluation of a novel loop-mediated isothermal amplification method for rapid detection of severe acute respiratory syndrome coronavirus. J. Clin. Microbiol. 2004, 42, 1956–1961. [Google Scholar] [CrossRef] [PubMed]
- Mori, Y.; Notomi, T. Loop-mediated isothermal amplification (LAMP): A rapid, accurate, and cost-effective diagnostic method for infectious diseases. J. Infect. Chemother. 2009, 15, 62–69. [Google Scholar] [CrossRef] [PubMed]
- Huang, P.; Wang, H.; Cao, Z.; Jin, H.; Chi, H.; Zhao, J.; Yu, B.; Yan, F.; Hu, X.; Wu, F.; et al. A Rapid and Specific Assay for the Detection of MERS-CoV. Front. Microbiol. 2018, 9, 1101. [Google Scholar] [CrossRef] [PubMed]
- Cao, Z.; Wang, H.; Wang, L.; Li, L.; Jin, H.; Xu, C.; Feng, N.; Wang, J.; Li, Q.; Zhao, Y.; et al. Visual Detection of West Nile Virus Using Reverse Transcription Loop-Mediated Isothermal Amplification Combined with a Vertical Flow Visualization Strip. Front. Microbiol. 2016, 7, 554. [Google Scholar] [CrossRef] [PubMed]
- Chotiwan, N.; Brewster, C.D.; Magalhaes, T.; Weger-Lucarelli, J.; Duggal, N.K.; Rückert, C.; Nguyen, C.; Luna, S.M.G.; Fauver, J.R.; Andre, B.; et al. Rapid and specific detection of Asian- and African-lineage Zika viruses. Sci. Transl. Med. 2017, 9, eaag0538. [Google Scholar] [CrossRef] [PubMed]
- Kwallah, A.O.; Inoue, S.; Muigai, A.W.; Kubo, T.; Sang, R.; Morita, K.; Mwau, M. A real-time reverse transcription loop-mediated isothermal amplification assay for the rapid detection of yellow fever virus. J. Virol. Methods 2013, 193, 23–27. [Google Scholar] [CrossRef]
- Shirato, K.; Semba, S.; El-Kafrawy, S.A.; Hassan, A.M.; Tolah, A.M.; Takayama, I.; Kageyama, T.; Notomi, T.; Kamitani, W.; Matsuyama, S.; et al. Development of fluorescent reverse transcription loop-mediated isothermal amplification (RT-LAMP) using quenching probes for the detection of the Middle East respiratory syndrome coronavirus. J. Virol. Methods 2018, 258, 41–48. [Google Scholar] [CrossRef]
- El-Kafrawy, S.A.; El-Daly, M.M.; Hassan, A.M.; Harakeh, S.M.; Alandijany, T.A.; Azhar, E.I. Rapid and Reliable Detection of SARS-CoV-2 Using Direct RT-LAMP. Diagnostics 2022, 12, 828. [Google Scholar] [CrossRef]
- Floriano, I.; Silvinato, A.; Bernardo, W.M.; Reis, J.C.; Soledade, G. Accuracy of the Polymerase Chain Reaction (PCR) test in the diagnosis of acute respiratory syndrome due to coronavirus: A systematic review and meta-analysis. Rev. Assoc. Med. Bras. 2020, 66, 880–888. [Google Scholar] [CrossRef]
- Xing, W.; Liu, Y.; Wang, H.; Li, S.; Lin, Y.; Chen, L.; Zhao, Y.; Chao, S.; Huang, X.; Ge, S.; et al. A High-Throughput, Multi-Index Isothermal Amplification Platform for Rapid Detection of 19 Types of Common Respiratory Viruses Including SARS-CoV-2. Engineering 2020, 6, 1130–1140. [Google Scholar] [CrossRef]
- Kilic, T.; Weissleder, R.; Lee, H. Molecular and Immunological Diagnostic Tests of COVID-19: Current Status and Challenges. iScience 2020, 23, 101406. [Google Scholar] [CrossRef] [PubMed]
- Lau, Y.L.; Ismail, I.B.; Mustapa, N.I.B.; Lai, M.Y.; Soh, T.S.T.; Hassan, A.H.; Peariasamy, K.M.; Lee, Y.L.; Kahar, M.K.B.A.; Chong, J.; et al. Development of a reverse transcription recombinase polymerase amplification assay for rapid and direct visual detection of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). PLoS ONE 2021, 16, e0245164. [Google Scholar] [CrossRef]
- Wang, P.; Ma, C.; Zhang, X.; Chen, L.; Yi, L.; Liu, X.; Lu, Q.; Cao, Y.; Gao, S. A Ligation/Recombinase Polymerase Amplification Assay for Rapid Detection of SARS-CoV-2. Front. Cell. Infect. Microbiol. 2021, 11, 680728. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Shen, H.; Zhang, Y.; Shen, D.; Zhu, M.; Song, Y.; Zhu, Z.; Yang, C.J. A microfluidic-integrated lateral flow recombinase polymerase amplification (MI-IF-RPA) assay for rapid COVID-19 detection. Lab Chip 2021, 21, 2019–2026. [Google Scholar] [CrossRef] [PubMed]
- Carter, L.J.; Garner, L.V.; Smoot, J.W.; Li, Y.; Zhou, Q.; Saveson, C.J.; Sasso, J.M.; Gregg, A.C.; Soares, D.J.; Beskid, T.R.; et al. Assay Techniques and Test Development for COVID-19 Diagnosis. ACS Cent. Sci. 2020, 6, 591–605. [Google Scholar] [CrossRef] [PubMed]
- Palaz, F.; Kalkan, A.K.; Tozluyurt, A.; Ozsoz, M. CRISPR-based tools: Alternative methods for the diagnosis of COVID-19. Clin. Biochem. 2021, 89, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Broughton, J.P.; Deng, X.; Yu, G.; Fasching, C.L.; Servellita, V.; Singh, J.; Miao, X.; Streithorst, J.A.; Granados, A.; Sotomayor-Gonzalez, A.; et al. CRISPR–Cas12-based detection of SARS-CoV-2. Nat. Biotechnol. 2020, 38, 870–874. [Google Scholar] [CrossRef]
- Hou, T.; Zeng, W.; Yang, M.; Chen, W.; Ren, L.; Ai, J.; Wu, J.; Liao, Y.; Gou, X.; Li, Y.; et al. Development and evaluation of a rapid CRISPR-based diagnostic for COVID-19. PLOS Pathog. 2020, 16, e1008705. [Google Scholar] [CrossRef]
- Moitra, P.; Alafeef, M.; Dighe, K.; Frieman, M.B.; Pan, D. Selective Naked-Eye Detection of SARS-CoV-2 Mediated by N Gene Targeted Antisense Oligonucleotide Capped Plasmonic Nanoparticles. ACS Nano 2020, 14, 7617–7627. [Google Scholar] [CrossRef]
- Pisanic, N.; Randad, P.R.; Kruczynski, K.; Manabe, Y.C.; Thomas, D.L.; Pekosz, A.; Klein, S.L.; Betenbaugh, M.J.; Clarke, W.A.; Laeyendecker, O.; et al. COVID-19 Serology at Population Scale: SARS-CoV-2-Specific Antibody Responses in Saliva. J. Clin. Microbiol. 2020, 59. [Google Scholar] [CrossRef]
- Hou, H.; Wang, T.; Zhang, B.; Luo, Y.; Mao, L.; Wang, F.; Wu, S.; Sun, Z. Detection of IgM and IgG antibodies in patients with coronavirus disease 2019. Clin. Transl. Immunol. 2020, 9, e01136. [Google Scholar] [CrossRef]
- Padoan, A.; Sciacovelli, L.; Basso, D.; Negrini, D.; Zuin, S.; Cosma, C.; Faggian, D.; Matricardi, P.; Plebani, M. IgA-Ab response to spike glycoprotein of SARS-CoV-2 in patients with COVID-19: A longitudinal study. Clin. Chim. Acta 2020, 507, 164–166. [Google Scholar] [CrossRef]
- Long, Q.-X.; Liu, B.-Z.; Deng, H.-J.; Wu, G.-C.; Deng, K.; Chen, Y.-K.; Liao, P.; Qiu, J.-F.; Lin, Y.; Cai, X.-F.; et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat. Med. 2020, 26, 845–848. [Google Scholar] [CrossRef]
- Varadhachary, A.; Chatterjee, D.; Garza, J.; Garr, R.P.; Foley, C.; Letkeman, A.; Dean, J.; Haug, D.; Breeze, J.; Traylor, R.; et al. Salivary anti-SARS-CoV-2 IgA as an accessible biomarker of mucosal immunity against COVID-19. MedRxiv 2020. [Google Scholar] [CrossRef]
- Petherick, A. Developing antibody tests for SARS-CoV-2. Lancet 2020, 395, 1101–1102. [Google Scholar] [CrossRef] [PubMed]
- Wolters, F.; van de Bovenkamp, J.; Bosch, B.v.D.; Brink, S.v.D.; Broeders, M.; Chung, N.H.; Favié, B.; Goderski, G.; Kuijpers, J.; Overdevest, I.; et al. Multi-center evaluation of cepheid xpert® xpress SARS-CoV-2 point-of-care test during the SARS-CoV-2 pandemic. J. Clin. Virol. 2020, 128, 104426. [Google Scholar] [CrossRef] [PubMed]
- Kohmer, N.; Westhaus, S.; Rühl, C.; Ciesek, S.; Rabenau, H.F. Brief clinical evaluation of six high-throughput SARS-CoV-2 IgG antibody assays. J. Clin. Virol. 2020, 129, 104480. [Google Scholar] [CrossRef] [PubMed]
- Qiu, M.; Shi, Y.; Guo, Z.; Chen, Z.; He, R.; Chen, R.; Zhou, D.; Dai, E.; Wang, X.; Si, B.; et al. Antibody responses to individual proteins of SARS coronavirus and their neutralization activities. Microbes Infect. 2005, 7, 882–889. [Google Scholar] [CrossRef] [PubMed]
- Porte, L.; Legarraga, P.; Vollrath, V.; Aguilera, X.; Munita, J.M.; Araos, R.; Pizarro, G.; Vial, P.; Iruretagoyena, M.; Dittrich, S.; et al. Evaluation of a novel antigen-based rapid detection test for the diagnosis of SARS-CoV-2 in respiratory samples. Int. J. Infect. Dis. 2020, 99, 328–333. [Google Scholar] [CrossRef] [PubMed]
- Diao, B.; Wen, K.; Zhang, J.; Chen, J.; Han, C.; Chen, Y.; Wang, S.; Deng, G.; Zhou, H.; Wu, Y. Accuracy of a nucleocapsid protein antigen rapid test in the diagnosis of SARS-CoV-2 infection. Clin. Microbiol. Infect. 2021, 27, 289.e1–289.e4. [Google Scholar] [CrossRef]
- Ihling, C.; Tänzler, D.; Hagemann, S.; Kehlen, A.; Hüttelmaier, S.; Arlt, C.; Sinz, A. Mass Spectrometric Identification of SARS-CoV-2 Proteins from Gargle Solution Samples of COVID-19 Patients. J. Proteome Res. 2020, 19, 4389–4392. [Google Scholar] [CrossRef]
- Arike, L.; Peil, L. Spectral counting label-free proteomics. Methods Mol. Biol. 2014, 1156, 213–222. [Google Scholar] [CrossRef]
- Petrillo, M.; Brogna, C.; Cristoni, S.; Querci, M.; Piazza, O.; Eede, G.V.D. Increase of SARS-CoV-2 RNA load in faecal samples prompts for rethinking of SARS-CoV-2 biology and COVID-19 epidemiology. F1000Research 2021, 10, 370. [Google Scholar] [CrossRef] [PubMed]
- Brogna, C.; Brogna, B.; Bisaccia, D.R.; Lauritano, F.; Marino, G.; Montano, L.; Cristoni, S.; Prisco, M.; Piscopo, M. Could SARS-CoV-2 Have Bacteriophage Behavior or Induce the Activity of Other Bacteriophages? Vaccines 2022, 10, 708. [Google Scholar] [CrossRef]
- Brogna, C.; Costanzo, V.; Brogna, B.; Bisaccia, D.R.; Brogna, G.; Giuliano, M.; Montano, L.; Viduto, V.; Cristoni, S.; Fabrowski, M.; et al. Analysis of Bacteriophage Behavior of a Human RNA Virus, SARS-CoV-2, through the Integrated Approach of Immunofluorescence Microscopy, Proteomics and D-Amino Acid Quantification. Int. J. Mol. Sci. 2023, 24, 3929. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.-J.; Goh, P.-Y.; Fielding, B.C.; Shen, S.; Chou, C.-F.; Fu, J.-L.; Leong, H.N.; Leo, Y.S.; Ooi, E.E.; Ling, A.E.; et al. Profiles of antibody responses against severe acute respiratory syndrome coronavirus recombinant proteins and their potential use as diagnostic markers. Clin. Diagn. Lab Immunol. 2004, 11, 362–371. [Google Scholar] [CrossRef] [PubMed]
- Kyosei, Y.; Namba, M.; Yamura, S.; Takeuchi, R.; Aoki, N.; Nakaishi, K.; Watabe, S.; Ito, E. Proposal of De Novo Antigen Test for COVID-19: Ultrasensitive Detection of Spike Proteins of SARS-CoV-2. Diagnostics 2020, 10, 594. [Google Scholar] [CrossRef] [PubMed]
- Grant, B.D.; Anderson, C.E.; Williford, J.R.; Alonzo, L.F.; Glukhova, V.A.; Boyle, D.S.; Weigl, B.H.; Nichols, K.P. SARS-CoV-2 Coronavirus Nucleocapsid Antigen-Detecting Half-Strip Lateral Flow Assay Toward the Development of Point of Care Tests Using Commercially Available Reagents. Anal. Chem. 2020, 92, 11305–11309. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Ju, C.; Han, C.; Shi, R.; Chen, X.; Duan, D.; Yan, J.; Yan, X. Nanozyme chemiluminescence paper test for rapid and sensitive detection of SARS-CoV-2 antigen. Biosens. Bioelectron. 2020, 173, 112817. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
El-Daly, M.M. Advances and Challenges in SARS-CoV-2 Detection: A Review of Molecular and Serological Technologies. Diagnostics 2024, 14, 519. https://doi.org/10.3390/diagnostics14050519
El-Daly MM. Advances and Challenges in SARS-CoV-2 Detection: A Review of Molecular and Serological Technologies. Diagnostics. 2024; 14(5):519. https://doi.org/10.3390/diagnostics14050519
Chicago/Turabian StyleEl-Daly, Mai M. 2024. "Advances and Challenges in SARS-CoV-2 Detection: A Review of Molecular and Serological Technologies" Diagnostics 14, no. 5: 519. https://doi.org/10.3390/diagnostics14050519
APA StyleEl-Daly, M. M. (2024). Advances and Challenges in SARS-CoV-2 Detection: A Review of Molecular and Serological Technologies. Diagnostics, 14(5), 519. https://doi.org/10.3390/diagnostics14050519




