Accuracy Analysis of Deep Learning Methods in Breast Cancer Classification: A Structured Review
Abstract
:1. Introduction
2. Review Method
2.1. Preliminary Identification
2.2. Screening
2.3. Eligibility
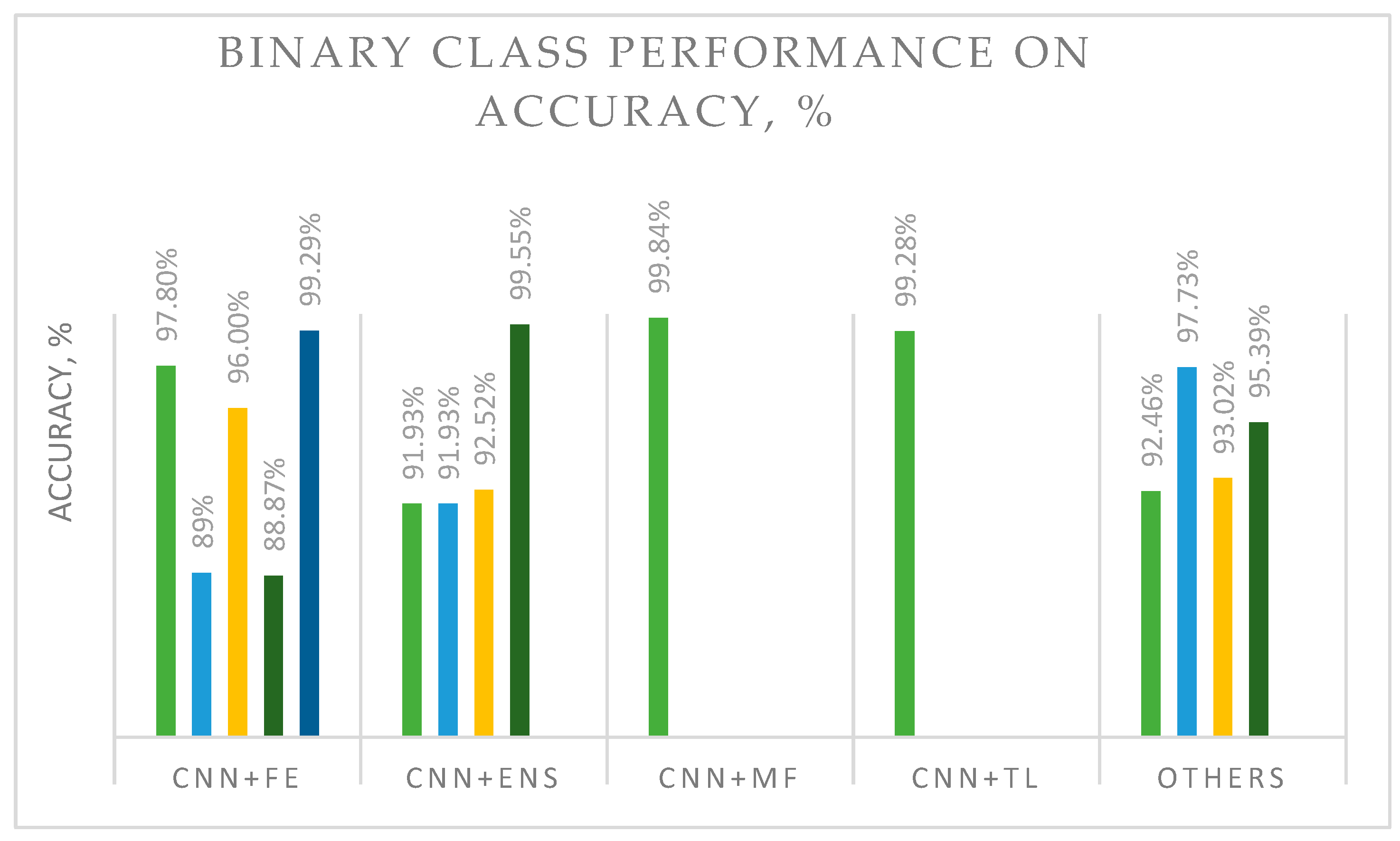
3. Results and Findings
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Li, G.; Li, C.; Wu, G.; Xu, G.; Zhou, Y.; Zhang, H. MF-OMKT: Model Fusion Based on Online Mutual Knowledge Transfer for Breast Cancer Histopathological Image Classification. Artif. Intell. Med. 2022, 134, 102433. [Google Scholar] [CrossRef] [PubMed]
- Dar, R.A.; Rasool, M.; Assad, A. Breast Cancer Detection Using Deep Learning: Datasets, Methods, and Challenges Ahead. Comput. Biol. Med. 2022, 149, 106073. [Google Scholar] [CrossRef]
- Ukwuoma, C.C.; Hossain, M.A.; Jackson, J.K.; Nneji, G.U.; Monday, H.N.; Qin, Z. Multi-Classification of Breast Cancer Lesions in Histopathological Images Using DEEP_Pachi: Multiple Self-Attention Head. Diagnostics 2022, 12, 1152. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Mehra, R. Conventional Machine Learning and Deep Learning Approach for Multi-Classification of Breast Cancer Histopathology Images—A Comparative Insight. J. Digit. Imaging 2020, 33, 632–654. [Google Scholar] [CrossRef] [PubMed]
- Ameh Joseph, A.; Abdullahi, M.; Junaidu, S.B.; Hassan Ibrahim, H.; Chiroma, H. Improved Multi-Classification of Breast Cancer Histopathological Images Using Handcrafted Features and Deep Neural Network (Dense Layer). Intell. Syst. Appl. 2022, 14, 200066. [Google Scholar] [CrossRef]
- Sali, R.; Adewole, S.; Ehsan, L.; Denson, L.A.; Kelly, P.; Amadi, B.C.; Holtz, L.; Ali, S.A.; Moore, S.R.; Syed, S.; et al. Hierarchical Deep Convolutional Neural Networks for Multi-Category Diagnosis of Gastrointestinal Disorders on Histopathological Images. In Proceedings of the 2020 IEEE International Conference on Healthcare Informatics, ICHI, Oldenburg, Germany, 30 November–3 December 2020; IEEE: Piscatvie, NJ, USA, 2020. [Google Scholar]
- Saturi, R.; Prem Chand, P. A Novel Variant-Optimized Search Algorithm for Nuclei Detection in Histopathology Breast Cancer Images. In Lecture Notes in Networks and Systems; Springer: Singapore, 2022; Volume 286. [Google Scholar]
- Liu, M.; He, Y.; Wu, M.; Zeng, C. Breast Histopathological Image Classification Method Based on Autoencoder and Siamese Framework. Information 2022, 13, 107. [Google Scholar] [CrossRef]
- Yang, J.; Ju, J.; Guo, L.; Ji, B.; Shi, S.; Yang, Z.; Gao, S.; Yuan, X.; Tian, G.; Liang, Y.; et al. Prediction of HER2-Positive Breast Cancer Recurrence and Metastasis Risk from Histopathological Images and Clinical Information via Multimodal Deep Learning. Comput. Struct. Biotechnol. J. 2022, 20, 333–342. [Google Scholar] [CrossRef]
- Yang, Y.; Guan, C. Classification of Histopathological Images of Breast Cancer Using an Improved Convolutional Neural Network Model. J. Xray Sci. Technol. 2022, 30, 33–44. [Google Scholar] [CrossRef]
- Mohamed, A.; Amer, E.; Eldin, N.; Hossam, M.; Elmasry, N.; Adnan, G.T. The Impact of Data Processing and Ensemble on Breast Cancer Detection Using Deep Learning. J. Comput. Commun. 2022, 1, 27–37. [Google Scholar] [CrossRef]
- Munappy, A.R.; Bosch, J.; Olsson, H.H.; Arpteg, A.; Brinne, B. Data Management for Production Quality Deep Learning Models: Challenges and Solutions. J. Syst. Softw. 2022, 191, 111359. [Google Scholar] [CrossRef]
- Franceschini, S.; Grelet, C.; Leblois, J.; Gengler, N.; Soyeurt, H. Can Unsupervised Learning Methods Applied to Milk Recording Big Data Provide New Insights into Dairy Cow Health? J. Dairy Sci. 2022, 105, 6760–6772. [Google Scholar] [CrossRef]
- Yang, C.-T.; Kristiani, E.; Leong, Y.K.; Chang, J.-S. Big Data and Machine Learning Driven Bioprocessing—Recent Trends and Critical Analysis. Bioresour. Technol. 2023, 372, 128625. [Google Scholar] [CrossRef]
- Nwonye, M.J.; Narasimhan, V.L.; Mbero, Z.A. Sensitivity Analysis of Coronary Heart Disease Using Two Deep Learning Algorithms CNN RNN. In 2021 IST-Africa Conference (IST-Africa); IEEE: Piscatvie, NJ, USA, 2021. [Google Scholar]
- Chauhan, N.K.; Singh, K. A Review on Conventional Machine Learning vs. deep learning. In 2018 International Conference on Computing, Power and Communication Technologies (GUCON); IEEE: Piscatvie, NJ, USA, 2019; pp. 347–352. [Google Scholar]
- Vankdothu, R.; Hameed, M.A. Brain Tumor MRI Images Identification and Classification Based on the Recurrent Convolutional Neural Network. Meas. Sens. 2022, 24, 100412. [Google Scholar] [CrossRef]
- Bagherzadeh, S.; Maghooli, K.; Shalbaf, A.; Maghsoudi, A. Recognition of Emotional States Using Frequency Effective Connectivity Maps through Transfer Learning Approach from Electroencephalogram Signals. Biomed. Signal Process Control 2022, 75, 103544. [Google Scholar] [CrossRef]
- Elangovan, P.; Nath, M.K. En-ConvNet: A Novel Approach for Glaucoma Detection from Color Fundus Images Using Ensemble of Deep Convolutional Neural Networks. Int. J. Imaging Syst. Technol. 2022, 32, 2034–2048. [Google Scholar] [CrossRef]
- Cui, R.; Liu, M.; Alzheimer’s Disease Neuroimaging Initiative. RNN-Based Longitudinal Analysis for Diagnosis of Alzheimer’s Disease. Comput. Med. Imaging Graph. 2019, 73, 1–10. [Google Scholar] [CrossRef]
- Freeborough, W.; van Zyl, T. Investigating Explainability Methods in Recurrent Neural Network Architectures for Financial Time Series Data. Appl. Sci. 2022, 12, 1427. [Google Scholar] [CrossRef]
- Amin, J.; Sharif, M.; Fernandes, S.L.; Wang, S.-H.; Saba, T.; Khan, A.R. Breast Microscopic Cancer Segmentation and Classification Using Unique 4-Qubit-Quantum Model. Microsc. Res. Tech. 2022, 85, 1926–1936. [Google Scholar] [CrossRef]
- Alqahtani, Y.; Mandawkar, U.; Sharma, A.; Hasan, M.N.S.; Kulkarni, M.H.; Sugumar, R. Breast Cancer Pathological Image Classification Based on the Multiscale CNN Squeeze Model. Comput. Intell. Neurosci. 2022, 2022, 7075408. [Google Scholar] [CrossRef]
- Gour, M.; Jain, S.; Sunil Kumar, T. Residual Learning Based CNN for Breast Cancer Histopathological Image Classification. Int. J. Imaging Syst. Technol. 2020, 30, 621–635. [Google Scholar] [CrossRef]
- Munn, Z.; Peters, M.D.J.; Stern, C.; Tufanaru, C.; McArthur, A.; Aromataris, E. Systematic Review or Scoping Review? Guidance for Authors When Choosing between a Systematic or Scoping Review Approach. BMC Med. Res. Methodol. 2018, 18, 143. [Google Scholar] [CrossRef] [PubMed]
- Gough, D.; Oliver, S.; Thomas, J. (Eds.) An Introduction to Systematic Reviews(2017); SAGE: New York, NY, USA, 2017. [Google Scholar]
- Cho, S.H.; Shin, I.S. A Reporting Quality Assessment of Systematic Reviews and Meta-Analyses in Sports Physical Therapy: A Review of Reviews. Healthcare 2021, 9, 1368. [Google Scholar] [CrossRef] [PubMed]
- Alias, N.A.; Mustafa, W.A.; Jamlos, M.A.; Alquran, H.; Hanafi, H.F.; Ismail, S.; Ab Rahman, K.S. Pap Smear Images Classification Using Machine Learning: A Literature Matrix. Diagnostics 2022, 12, 2900. [Google Scholar] [CrossRef] [PubMed]
- Chattopadhyay, S.; Dey, A.; Singh, P.K.; Oliva, D.; Cuevas, E.; Sarkar, R. MTRRE-Net: A Deep Learning Model for Detection of Breast Cancer from Histopathological Images. Comput. Biol. Med. 2022, 150, 106155. [Google Scholar] [CrossRef] [PubMed]
- Chattopadhyay, S.; Dey, A.; Singh, P.K.; Sarkar, R. DRDA-Net: Dense Residual Dual-Shuffle Attention Network for Breast Cancer Classification Using Histopathological Images. Comput. Biol. Med. 2022, 145, 105437. [Google Scholar] [CrossRef]
- Sharma, M.; Mandloi, A.; Bhattacharya, M. A Novel DeepML Framework for Multi-Classification of Breast Cancer Based on Transfer Learning. Int. J. Imaging Syst. Technol. 2022, 32, 1963–1977. [Google Scholar] [CrossRef]
- Nakach, F.-Z.; Zerouaoui, H.; Idri, A. Hybrid Deep Boosting Ensembles for Histopathological Breast Cancer Classification. Health Technol. 2022, 12, 1043–1060. [Google Scholar] [CrossRef]
- Kim, S.; Yoon, K. Deep and Lightweight Neural Network for Histopathological Image Classification. J. Mob. Multimed. 2022, 18, 1913–1930. [Google Scholar] [CrossRef]
- Alkhathlan, L.; Saudagar, A.K.J. Predicting and Classifying Breast Cancer Using Machine Learning. J. Comput. Biol. 2022, 29, 497–514. [Google Scholar] [CrossRef]
- Xu, Y.; dos Santos, M.A.; Souza, L.F.F.; Marques, A.G.; Zhang, L.; da Costa Nascimento, J.J.; de Albuquerque, V.H.C.; Rebouças Filho, P.P. New Fully Automatic Approach for Tissue Identification in Histopathological Examinations Using Transfer Learning. IET Image Process 2022, 16, 2875–2889. [Google Scholar] [CrossRef]
- Rashmi, R.; Prasad, K.; Udupa, C.B.K. Region-Based Feature Enhancement Using Channel-Wise Attention for Classification of Breast Histopathological Images. Neural Comput. Appl. 2022, 1–16. [Google Scholar] [CrossRef]
- Wakili, M.A.; Shehu, H.A.; Sharif, M.H.; Sharif, M.H.U.; Umar, A.; Kusetogullari, H.; Ince, I.F.; Uyaver, S. Classification of Breast Cancer Histopathological Images Using DenseNet and Transfer Learning. Comput. Intell. Neurosci. 2022, 2022, 8904768. [Google Scholar] [CrossRef]
- Iqbal, S.; Qureshi, A.N. Deep-Hist: Breast Cancer Diagnosis through Histopathological Images Using Convolution Neural Network. J. Intell. Fuzzy Syst. 2022, 43, 1347–1364. [Google Scholar] [CrossRef]
- Kumar, S.; Sharma, S. Sub-Classification of Invasive and Non-Invasive Cancer from Magnification Independent Histopathological Images Using Hybrid Neural Networks. Evol. Intell. 2022, 15, 1531–1543. [Google Scholar] [CrossRef]
- Abbasniya, M.R.; Sheikholeslamzadeh, S.A.; Nasiri, H.; Emami, S. Classification of Breast Tumors Based on Histopathology Images Using Deep Features and Ensemble of Gradient Boosting Methods. Comput. Electr. Eng. 2022, 103, 108382. [Google Scholar] [CrossRef]
- Karthik, R.; Menaka, R.; Siddharth, M. v Classification of Breast Cancer from Histopathology Images Using an Ensemble of Deep Multiscale Networks. Biocybern. Biomed. Eng. 2022, 42, 963–976. [Google Scholar] [CrossRef]
- el Agouri, H.; Azizi, M.; el Attar, H.; el Khannoussi, M.; Ibrahimi, A.; Kabbaj, R.; Kadiri, H.; BekarSabein, S.; EchCharif, S.; Mounjid, C.; et al. Assessment of Deep Learning Algorithms to Predict Histopathological Diagnosis of Breast Cancer: First Moroccan Prospective Study on a Private Dataset. BMC Res. Notes 2022, 15, 66. [Google Scholar] [CrossRef]
- He, Z.; Lin, M.; Xu, Z.; Yao, Z.; Chen, H.; Alhudhaif, A.; Alenezi, F. Deconv-Transformer (DecT): A Histopathological Image Classification Model for Breast Cancer Based on Color Deconvolution and Transformer Architecture. Inf. Sci. 2022, 608, 1093–1112. [Google Scholar] [CrossRef]
- Zou, Y.; Zhang, J.; Huang, S.; Liu, B. Breast Cancer Histopathological Image Classification Using Attention High-Order Deep Network. Int. J. Imaging Syst. Technol. 2022, 32, 266–279. [Google Scholar] [CrossRef]
- Zerouaoui, H.; Idri, A. Deep Hybrid Architectures for Binary Classification of Medical Breast Cancer Images. Biomed. Signal Process Control 2022, 71, 103226. [Google Scholar] [CrossRef]
- Luz, D.S.; Lima, T.J.B.; Silva, R.R.V.; Magalhães, D.M.V.; Araujo, F.H.D. Automatic Detection Metastasis in Breast Histopathological Images Based on Ensemble Learning and Color Adjustment. Biomed. Signal Process Control 2022, 75, 103564. [Google Scholar] [CrossRef]
- Wang, X.; Ahmad, I.; Javeed, D.; Zaidi, S.A.; Alotaibi, F.M.; Ghoneim, M.E.; Daradkeh, Y.I.; Asghar, J.; Eldin, E.T. Intelligent Hybrid Deep Learning Model for Breast Cancer Detection. Electronics 2022, 11, 2767. [Google Scholar] [CrossRef]
- Baghdadi, N.A.; Malki, A.; Balaha, H.M.; AbdulAzeem, Y.; Badawy, M.; Elhosseini, M. Classification of Breast Cancer Using a Manta-Ray Foraging Optimized Transfer Learning Framework. PeerJ Comput. Sci. 2022, 8, e1054. [Google Scholar] [CrossRef] [PubMed]
- Deepa, B.G.; Senthil, S. Predicting Invasive Ductal Carcinoma Tissues in Whole Slide Images of Breast Cancer by Using Convolutional Neural Network Model and Multiple Classifiers. Multimed. Tools Appl. 2022, 81, 8575–8596. [Google Scholar] [CrossRef]
- Abdul Jawad, M.; Khursheed, F. Deep and Dense Convolutional Neural Network for Multi Category Classification of Magnification Specific and Magnification Independent Breast Cancer Histopathological Images. Biomed. Signal Process Control 2022, 78, 103935. [Google Scholar] [CrossRef]
- Cengiz, E.; Kelek, M.M.; Oǧuz, Y.; Yllmaz, C. Classification of Breast Cancer with Deep Learning from Noisy Images Using Wavelet Transform. Biomed. Tech. 2022, 67, 143–150. [Google Scholar] [CrossRef]
- Burçak, K.C.; Uğuz, H. A New Hybrid Breast Cancer Diagnosis Model Using Deep Learning Model and ReliefF. Traitement Du Signal 2022, 39, 521–529. [Google Scholar] [CrossRef]
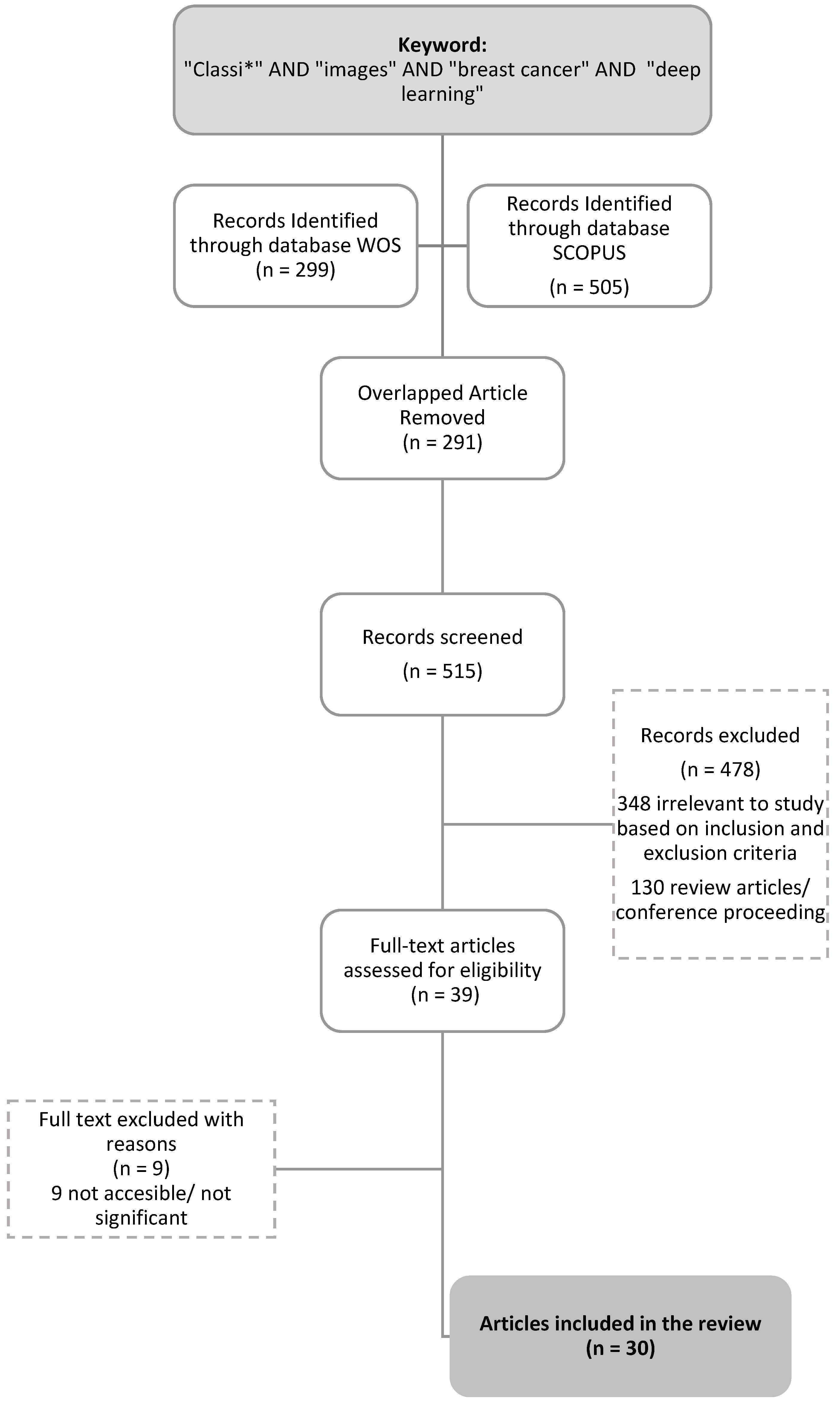

| Search strings from the Scopus database | TITLE-ABS-KEY “classi*” AND “images” AND “breast cancer” AND “deep learning” (LIMIT-TO PUBYEAR, 2022) | Results = 503 Articles |
| Search strings from the WOS (Web of Science) database | TS = “classi*” AND “images” AND “breast cancer” AND “deep learning” AB = “classi*” AND “images” AND “breast cancer” AND “deep learning” Refined by: DOCUMENT TYPES: (ARTICLE) AND PUBLICATION YEAR: 2022 | Results = 299 Articles |
| Criterion | Inclusion | Exclusion |
| Language | English | Non-English |
| Published Year | 2022 | <2022 |
| Source type | Journal (only research articles) | Conference proceeding |
| Document Type | Article | Letter, review, conference, and note |
| Research Area | Computer Science and Engineering | Besides Computer Science and Engineering |
| Reference | Dataset | Class | Methodology | Results | Advantages/Shortcomings |
|---|---|---|---|---|---|
| [5] | BreakHis | Multi-class | Data augmentation and handcrafted feature extraction (FE) techniques (Hu moment, Haralick textures, color histograms, and deep neural networks (DNNs)). Hybrid DL method: CNN + feature extraction. | Accuracy: 40×: 97.87%; 100×: 97.60%; 200×: 96.10%; 400×: 96.84%. Average accuracy: 97.10% | The proposed handcrafted feature extraction and DNN showed the best performance. |
| [29] | Dual residual block with a multi-scale dual residual recurrent network (MTRRE-Net74). DL Method: RNN. | Accuracy: 40×: 97.12%; 100×: 95.2%; 200×: 96.8%; 400×: 97.81%. | Better accuracy at four magnification levels than previously described models. | ||
| [30] | NetDense residual dual-shuffle attention network (DRDA-Net) inspired by the bottleneck unit of the ShuffleNet architecture. DL Method: CNN. | Accuracy: 40×: 95.72%; 100×: 94.41%; 200×: 97.43%; 400×: 98.1%/ | The proposed model showed acceptable accuracy. Densely connected blocks addressed the overfitting and vanishing gradient problems. | ||
| [3] | DenseNet201 and VGG16 architecture models as an ensemble model used to extract global features. DEEP Pachi extracts spatial information on the region of interest. Hybrid DL method: CNN + pre-processing + augmentation + ensemble. | BreakHis Malignant Accuracy: 99%. Benign Accuracy: 100%. | A significant result. | ||
| [31] | Wide-scale data | DeepML framework to achieve multi-class classification. DL Method: CNN. | Average accuracy: 98% (90–10% train–test split) and 89% (80–20% train–test). | Acceptable accuracy. | |
| [32] | BreakHis | Binary | DL are DenseNet_201, MobileNet_V2, and Inception_V3. Ensemble or boosting methods are AdaBoost (ADB), Gradient Boosting Machine (GBM), LightGBM (LGBM), and XGBoost (XGB) with Decision Tree (DT) as a base learner. Hybrid DL method: CNN + ensemble. | XGB + Inception_V3 showed the best accuracy of 92.52%. | Feature extraction and boosting ensembles were shown to be a good combination for image classification. |
| [10] | Pre-processed by multiple scaling decompositions to prevent overfitting due to the large dimension. It improved the DenseNet-201-MSD network model. DL Method: CNN + preprocessing. | Accuracy: 40×: 99.4%; 100×: 98.8%; 200×: 98.2; 400×: 99.4%. | The mode demonstrated very good results and can be used with other image data. | ||
| [33] | LightXception is based on cutting off layers at the bottom of the Xception network, which reduces the number of convolution filter channels. LightXception only has about 35% of the parameters of the original. Hybrid DL method: CNN + filter. | At 100× magnification: Xception Accuracy: 97.31%; Xception Recall: 98.67; Xception Precision: 99.26%; LightXception Accuracy: 97.42%; LightXception Recall: 97.42%; LightXception Precision: 97.42%. | Acceptable classification solution; the model needs improvement. | ||
| [34] | Support vector machine (SVM), CNN, and CNN with transfer learning (TL). Hybrid DL method: CNN + TL. | Accuracy: SVM: 92%; CNN: 94%; CNN + TL: 97%. | Improved results for CNN + transfer learning compared with a single method. | ||
| [35] | DenseNet201 with the SVM RBF classifier. Hybrid DL method: CNN + classifier. | At 200× magnification: Accuracy of 95.39% and precision of 95.43%. | Acceptable accuracy could have been due to the use of traditional machine learning as a classifier. | ||
| [23] | BreakHis | Binary | The attention model features multi-scale channel recalibration and msSE-ResNet convolutional neural network (msSE-ResNet34). Hybrid DL method: CNN + FE. | Accuracy: 88.87%. | Accuracy was low, although the approach was embedded with FE. |
| [36] | CNN and color channel with attention module (CWA-Net). Hybrid DL method: CNN + FE. | At 400× magnification: Private dataset accuracy: 95%; BreakHis dataset accuracy: 96% | Acceptable accuracy. | ||
| [37] | DenseNet as a backbone model and transfer learning (DenTnet). Hybrid DL method: CNN + TL. | Accuracy: 99.28%. | Good generalization ability and computational speed. | ||
| [38] | “Deep-Hist” with pre-trained and Stochastic Gradient Descent (SGD). Type: CNN + optimization. | Accuracy: 92.46%. | The proposed model required accuracy improvements. | ||
| [39] | Pre-trained Xception model with VGG16 and enhanced with logistic regression. Uses real-time data augmentation (AUG). Hybrid DL method: CNN + TL + AUG. | Xception + VGG16: Precision of 78.67% and recall of 0.76. F-score of 0.75, and AUC of 0.86. Xception + VGG16 + logistic regression: precision of 82.45%, recall of 0.82. F-score of 0.82, and AUC of 0.90. | The proposed model outperformed a conventional CNN. Augmentation could reduce the problem of overfitting. | ||
| [22] | Breaches’ | Binary | Xception and deeplabv3+. DL method: CNN. | Binary accuracy: 95%. Malignant accuracy: 99%. | The proposed framework demonstrated remarkable performance on a malignant dataset. |
| [40] | Inception-ResNet-v2 and Categorical Boosting (CatBoost), Extreme Gradient Boosting (XGBoost), and Light Gradient Boosting Machine (LightGBM). DL method: CNN. | Accuracy: 40×: 96.82%; 100×: 95.84%; 200×: 97.01%; 400×: 96.15%; Average: 96.46%. | The proposed Inception-ResNet-v2 and Categorical Boosting (CatBoost) models outperformed other methods. | ||
| [41] | Two custom deep architectures, CSAResnet and DAMCNN, integrated with channel and spatial attention. Type: CNN + ensemble. | Accuracy: 99.55%. | The proposed model demonstrated outstanding accuracy, showing that the employed ensemble method could be successfully used on the studied type of images. | ||
| [42] | Invasive breast carcinoma | CNN with Resnet50 and Xception. Type: CNN. | Xception was better than Restnet50, with an accuracy of 88% and a sensitivity of 95%. | Model performance was still low for Xception. The model could be improved by embedding ensemble and classifier programs. | |
| [43] | BACH, UC, and BreakHis | Deconv-Transformer (DecT) with color jitter data augmentation. Hybrid DL method: CNN + AUG. | BreakHis dataset accuracy: 93.02%. BACH dataset accuracy: 79.06%. UC dataset accuracy: 81.36%. | Performance was low even though augmentation was added. A hyperparameter tuning and pre-processing method would improve performance. | |
| [44] | Embeds attention mechanism and high-order statistical representation into a residual convolutional network (attention high-order deep network (AHoNet). Adds non-dimensionality reduction and local cross-channel interaction to achieve local salient deep features with normalization (NORM). Hybrid DL method: CNN + FE + NORM. | BreakHis dataset accuracy: 99.29%. BACH dataset accuracy: 85%. | Performance was more competitive than previously described models. A very good model to be tested with other types of image data. | ||
| [45] | BreakHis and FNAC | Binary | Twenty-eight hybrid architectures combining seven recent deep learning techniques for feature extraction (DenseNet 201, Inception V3, Inception ResNet V2, MobileNet V2, ResNet 50, VGG16, and VGG19) and four classifiers (MLP, SVM, DT, and KNN) for binary classification. Hybrid DL method: CNN + classifier. | DenseNet 201 (MDEN) with MLP showed the best performance. FNAC accuracy: 99.29%. BreakHis accuracy: 40×: 92.61%; 100×: 92%; 200×: 93.93%; 400×: 91.73%. | The proposed model requires accuracy improvements. |
| [46] | Public image database | Ensemble (ENS) for color adjustment methods with VGG-19 architectures. Hybrid DL method: CNN + ENS. | Accuracy: 91.93%. AUC: 97.72%. | The proposed model with color adjustment was shown to be an acceptable classification solution. | |
| [47] | PCam Kaggle | Hybrid deep learning (CNN-GRU). Hybrid DL method: CNN and GRU. | Accuracy: 86.21%. Precision: 85.50%. Sensitivity: 85.60%. Specificity: 84.71%. F1-score: 88%. AUC: 0.89. | The proposed model showed low accuracy. GRU seemingly could not improve the method’s performance. | |
| [48] | Histopathological data and ultrasound data | Automatic framework for reliable breast cancer classification CNN and TL. Embeds Manta Ray Foraging Optimization (MRFO) as metaheuristic optimization (OPT) to improve the framework’s adaptability. Hybrid DL method: CNN + TL + OPT. | Histopathological data accuracy: 97.73%. Ultrasound data accuracy: 99.01%. | The proposed framework was superior to previously tested methods. MRFO has the potential to be applied to other types of images. | |
| [49] | BreakHis invasive ductal carcinoma (IDC) dataset | CNN with logistic regression (LR), random forest (RF), k-nearest neighbor (K-NN), support vector machine (SVM), linear SVM, Gaussian Naïve Bayesian (GNB), and decision tree (DT) processes. Hybrid DL method: CNN + classifier. | Invasive ductal carcinoma: Range accuracy: 80–86%; range precision: 92–94%; range recall: 91–96%; range F1-score: 94–96%. BreakHis: range accuracy: 91–94%; range precision: 91–95%; range recall: 93–96%; F1-score: 95–98%. | Improvements in accuracy, precision, recall, and F1 score were shown. Hyperparameter tuning, pre-processing, and image augmentation may be used to achieve better classification performance. | |
| [1] | BreakHis | Binary and multi-class | The novel model fusion framework utilizes online mutual knowledge transfer (MF-OMKT) to classify histopathological breast cancer images. Imitates mutual communication and learning. Hybrid DL method: CNN + MF. | Accuracy range: Binary [99.27%, 99.84%]; Multi-class [96.14%, 97.53%]. | The proposed framework demonstrated good accuracy. The authors suggested evaluating a fusion strategy with other cancerous image data. |
| [50] | BreakHis histopathological dataset—ICIAR 2018 | DenseNet architecture and DenseNet architecture with image level mean. Hybrid DL method: CNN + NORM. | BreakHis: Binary DenseNet accuracy: 96.55%. Multi-class DenseNet accuracy: 91.82%. Binary DenseNet with image level mean accuracy: 91.72%. Multi-class with image level mean DenseNet accuracy: 96.72%. Competitive accuracies of 93.25% and 92.3% at the patient and image levels, respectively. | The proposed framework outperformed previously assessed methods. The normalization method seemed to influence improvements in results. | |
| [51] | Cancer images | Not available | Wavelet transform (WT) process is applied to noisy images; VggNet-16 model Hybrid DL method: CNN + preprocessing. | Best accuracy for the dataset with Gaussian noise of a 0.3 intensity: 86.9%. | Accuracy was low. WT was not found to be suitable for image data, though it may be applicable to other types of data. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yusoff, M.; Haryanto, T.; Suhartanto, H.; Mustafa, W.A.; Zain, J.M.; Kusmardi, K. Accuracy Analysis of Deep Learning Methods in Breast Cancer Classification: A Structured Review. Diagnostics 2023, 13, 683. https://doi.org/10.3390/diagnostics13040683
Yusoff M, Haryanto T, Suhartanto H, Mustafa WA, Zain JM, Kusmardi K. Accuracy Analysis of Deep Learning Methods in Breast Cancer Classification: A Structured Review. Diagnostics. 2023; 13(4):683. https://doi.org/10.3390/diagnostics13040683
Chicago/Turabian StyleYusoff, Marina, Toto Haryanto, Heru Suhartanto, Wan Azani Mustafa, Jasni Mohamad Zain, and Kusmardi Kusmardi. 2023. "Accuracy Analysis of Deep Learning Methods in Breast Cancer Classification: A Structured Review" Diagnostics 13, no. 4: 683. https://doi.org/10.3390/diagnostics13040683
APA StyleYusoff, M., Haryanto, T., Suhartanto, H., Mustafa, W. A., Zain, J. M., & Kusmardi, K. (2023). Accuracy Analysis of Deep Learning Methods in Breast Cancer Classification: A Structured Review. Diagnostics, 13(4), 683. https://doi.org/10.3390/diagnostics13040683








