Performance Analysis of Segmentation and Classification of CT-Scanned Ovarian Tumours Using U-Net and Deep Convolutional Neural Networks
Abstract
1. Introduction
2. Literature Review
2.1. Medical Imaging Classification Using CNN
2.2. Medical Imaging Classification Using Ensemble Deep Learning
2.3. Deep Learning in Medical Imaging Segmentation
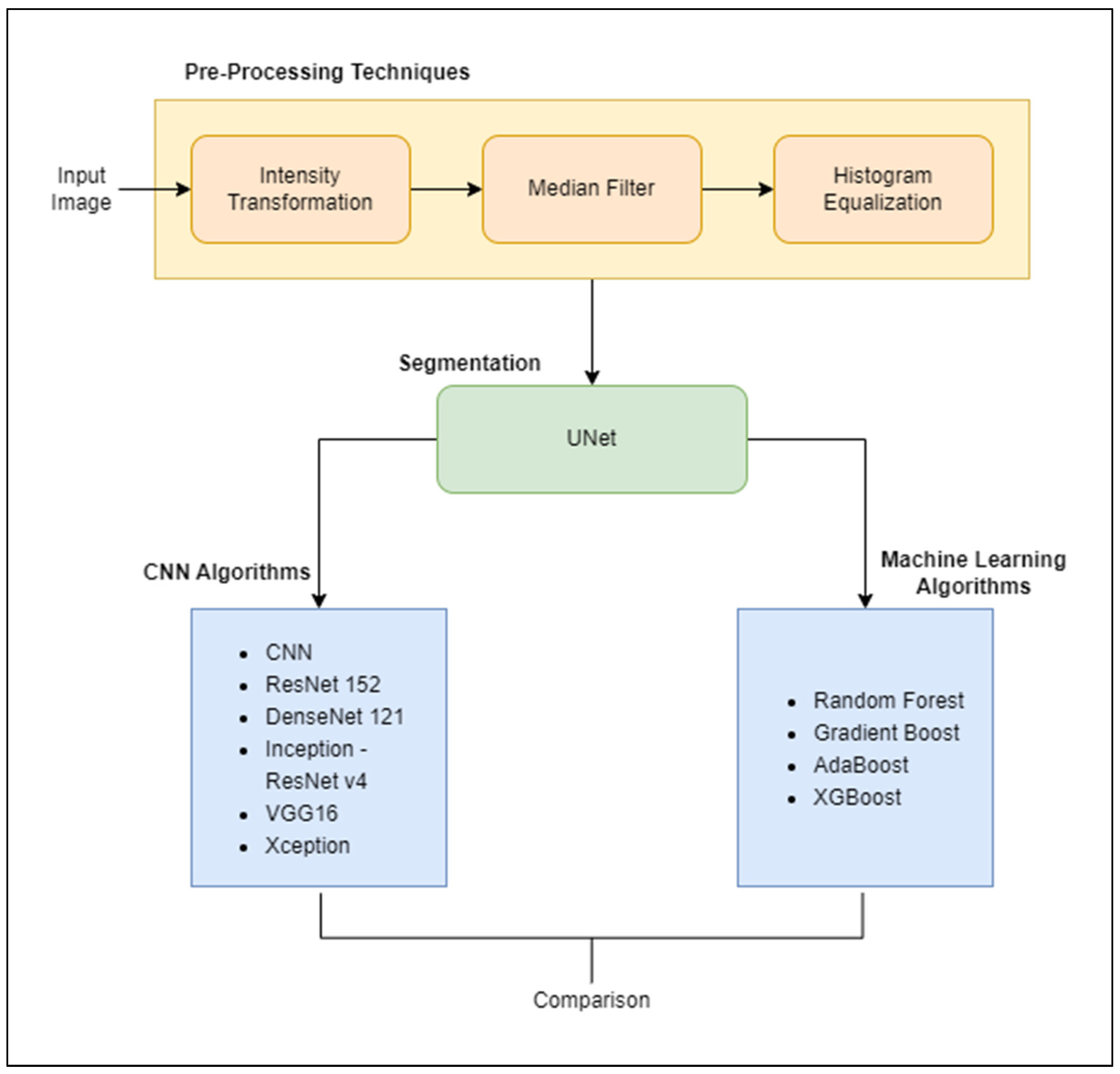
3. Methodology
3.1. Dataset Description
3.2. Segmentation Using U-Net Model
3.3. Detailed Methodology
4. Results and Discussions
4.1. Experimental Settings
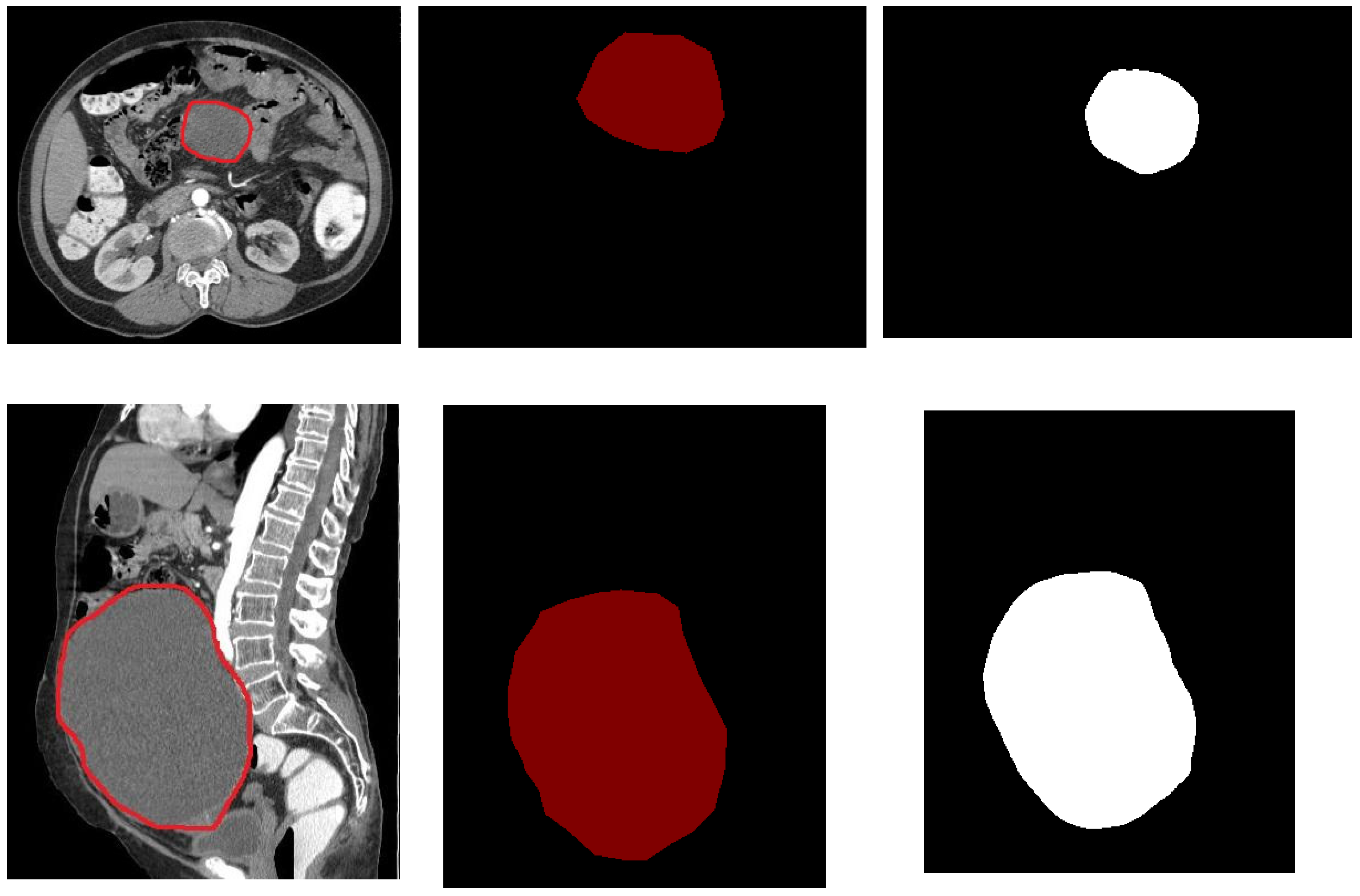
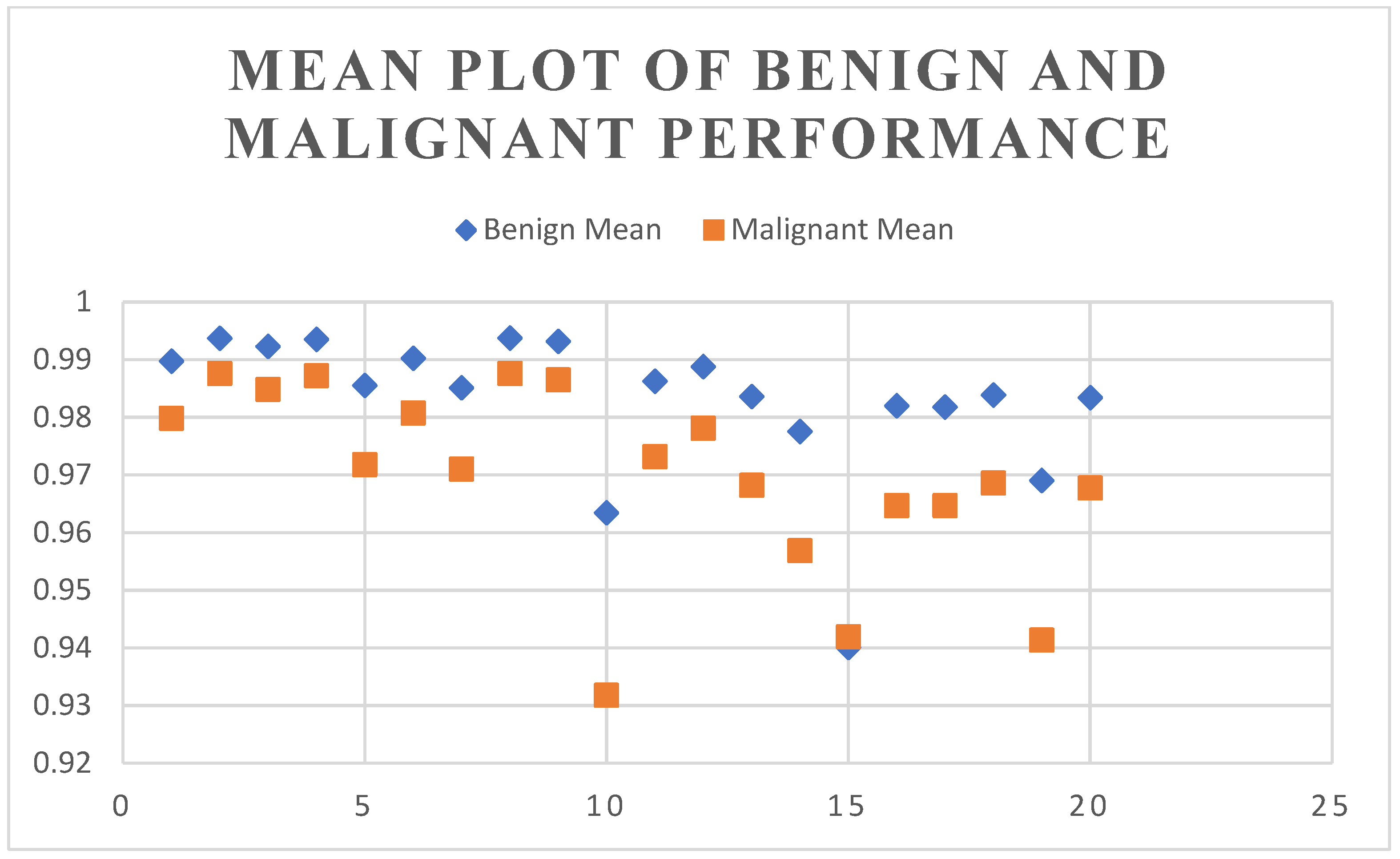
4.2. Segmentation Results Using the UNet Model
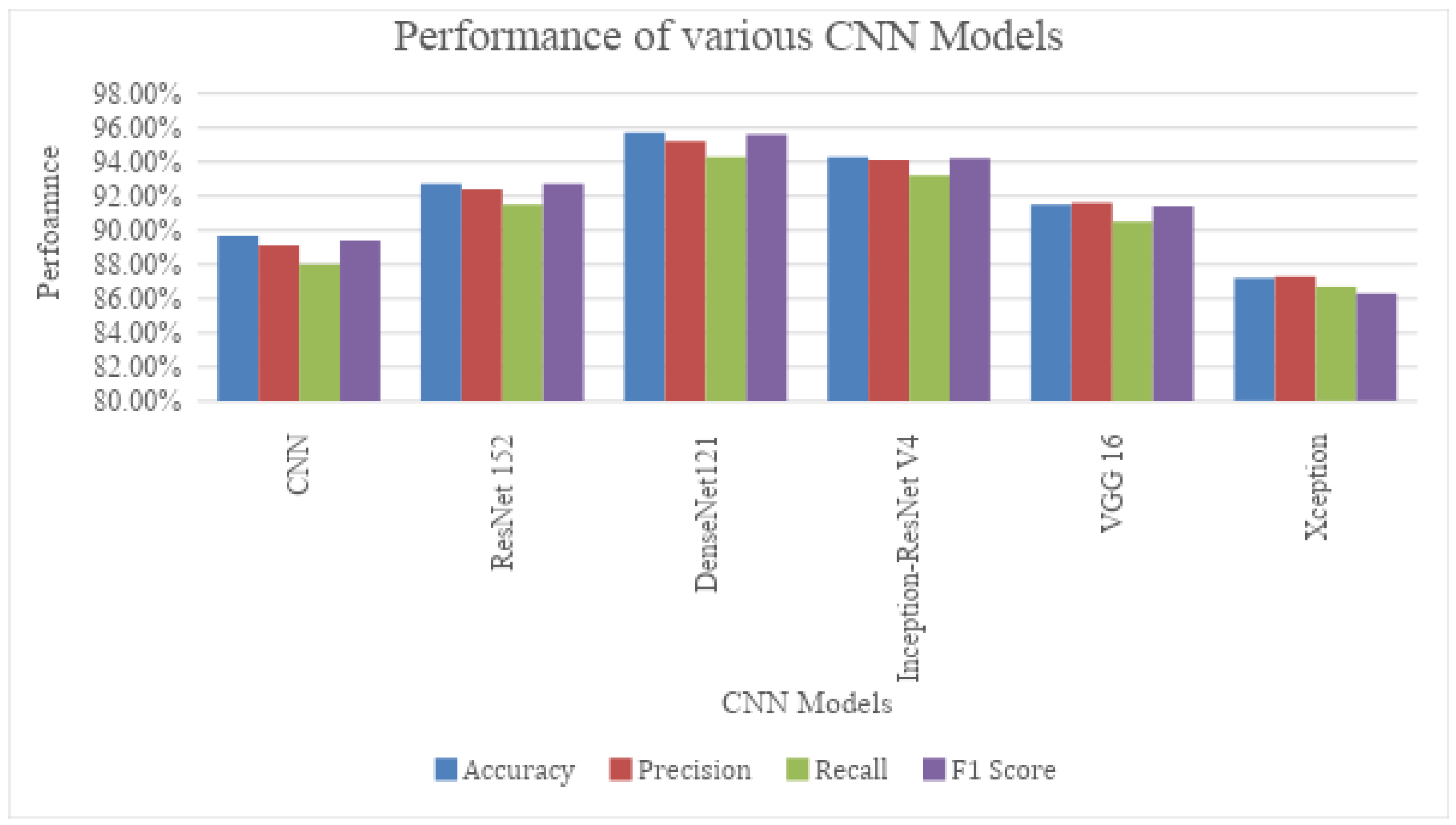
4.3. Classification Results Using Variants of the CNN Model
5. Conclusions and Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Labidi-Galy, S.I.; Treilleux, I.; Goddard-Leon, S.; Combes, J.-D.; Blay, J.-Y.; Ray-Coquard, I.; Caux, C.; Bendriss-Vermare, N. Plasmacytoid dendritic cells infiltrating ovarian cancer are associated with poor prognosis. Oncoimmunology 2012, 1, 380–382. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Wagle, N.S.; Jemal, A. Cancer statistics. CA Cancer J. Clin. 2023, 73, 17–48. [Google Scholar] [CrossRef]
- Jung, Y.; Kim, T.; Han, M.R.; Kim, S.; Kim, G.; Lee, S.; Choi, Y.J. Ovarian tumour diagnosis using deep convolutional neural networks and a denoising convolutional autoencoder. Sci. Rep. 2022, 12, 17024. [Google Scholar] [CrossRef]
- Wang, X.; Li, H.; Zheng, P. Automatic Detection and Segmentation of Ovarian Cancer Using a Multitask Model in Pelvic CT Images. Oxidative Med. Cell. Longev. 2022, 2022, 6009107. [Google Scholar] [CrossRef]
- Mahmood, F.; Borders, D.; Chen, R.J.; Mckay, G.N.; Salimian, K.J.; Baras, A.; Durr, N.J. Deep Adversarial Training for Multi-Organ Nuclei Segmentation in Histopathology Images. IEEE Trans. Med. Imaging 2019, 39, 3257–3267. [Google Scholar] [CrossRef]
- Guan, S.; Loew, M. Breast cancer detection using synthetic mammograms from generative adversarial networks in convolutional neural networks. J. Med. Imaging 2019, 6, 031411. [Google Scholar] [CrossRef] [PubMed]
- Karimi, D.; Dou, H.; Gholipour, A. Medical Image Segmentation Using Transformer Networks. IEEE Access 2022, 10, 29322–29332. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Farahani, H.; Bashashati, A. Multi-Resolution Vision Transformer for Subtype Classification in Ovarian Cancer Whole-Slide Histopathology Images. 2022. Available online: http://hdl.handle.net/2429/81390 (accessed on 2 February 2023).
- Li, H.; Fang, J.; Liu, S.; Liang, X.; Yang, X.; Mai, Z.; Van, M.T.; Wang, T.; Chen, Z.; Ni, D. CR-Unet: A Composite Network for Ovary and Follicle Segmentation in Ultrasound Images. IEEE J. Biomed. Health Inform. 2019, 24, 974–983. [Google Scholar] [CrossRef] [PubMed]
- Goodfellow, I.J.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative Adversarial Networks. arXiv 2014, arXiv:1406.2661v1. [Google Scholar] [CrossRef]
- Nagarajan, P.H.; Tajunisha, N. Automatic Classification of Ovarian Cancer Types from CT Images Using Deep Semi-Supervised Generative Learning and Convolutional Neural Network. Rev. D’intelligence Artif. 2021, 35, 273–280. [Google Scholar] [CrossRef]
- Zhao, Q.; Lyu, S.; Bai, W.; Cai, L.; Liu, B.; Wu, M.; Sang, X.; Yang, M.; Chen, L. A Multi-Modality Ovarian Tumour Ultrasound Image Dataset for Unsupervised Cross-Domain Semantic Segmentation. arXiv 2022, arXiv:2207.06799. [Google Scholar]
- Saha, D.; Mandal, A.; Ghosh, R. MU Net: Ovarian Follicle Segmentation Using Modified U-Net Architecture. Int. J. Eng. Adv. Technol. 2022, 11, 30–35. [Google Scholar] [CrossRef]
- Jin, J.; Zhu, H.; Zhang, J.; Ai, Y.; Zhang, J.; Teng, Y.; Xie, C.; Jin, X. Multiple U-Net-Based Automatic Segmentations and Radiomics Feature Stability on Ultrasound Images for Patients With Ovarian Cancer. Front. Oncol. 2021, 10, 614201. [Google Scholar] [CrossRef] [PubMed]
- Thangamma, N.G.; Prasanna, D.S. Analyzing ovarian tumour and cancer cells using image processing algorithms K means & fuzzy C-means. Int. J. Eng. Technol. 2018, 7, 510–512. [Google Scholar]
- Hema, L.K.; Manikandan, R.; Alhomrani, M.; Pradeep, N.; Alamri, A.S.; Sharma, S.; Alhassan, M. Region-Based Segmentation and Classification for Ovarian Cancer Detection Using Convolution Neural Network. Contrast Media Mol. Imaging 2022, 2022, 5968939. [Google Scholar] [CrossRef]
- Ahamad, M.; Aktar, S.; Uddin, J.; Rahman, T.; Alyami, S.A.; Al-Ashhab, S.; Akhdar, H.F.; Azad, A.; Moni, M.A. Early-Stage Detection of Ovarian Cancer Based on Clinical Data Using Machine Learning Approaches. J. Pers. Med. 2022, 12, 1211. [Google Scholar] [CrossRef]
- Kodipalli, A.; Guha, S.; Dasar, S.; Ismail, T. An inception-ResNet deep learning approach to classify tumours in the ovary as benign and malignant. Expert Syst. 2022, e13215. [Google Scholar] [CrossRef]
- Kodipalli, A.; Devi, S.; Dasar, S.; Ismail, T. Segmentation and classification of ovarian cancer based on conditional adversarial image to image translation approach. Expert Syst. 2022, e13193. [Google Scholar] [CrossRef]
- Ruchitha, P.J.; Sai, R.Y.; Kodipalli, A.; Martis, R.J.; Dasar, S.; Ismail, T. Comparative analysis of active contour random walker and watershed algorithms in segmentation of ovarian cancer. In Proceedings of the 2022 International Conference on Distributed Computing, VLSI, Electrical Circuits and Robotics (DISCOVER), Shivamogga, India, 14–15 October 2022; IEEE: Piscataway, NJ, USA, 2022; pp. 234–238. [Google Scholar]
- Liaqat, A.; Khan, M.A.; Shah, J.H.; Sharif, M.; Yasmin, M.; Fernandes, S.L. Automated ulcer and bleeding classification from WCE images using multiple features fusion and selection. J. Mech. Med. Biol. 2018, 18, 1850038. [Google Scholar] [CrossRef]
- Fernandes, S.L.; Tanik, U.J.; Rajinikanth, V.; Karthik, K.A. A reliable framework for accurate brain image examination and treatment planning based on early diagnosis support for clinicians. Neural Comput. Appl. 2020, 32, 15897–15908. [Google Scholar] [CrossRef]
- Koonce, B. EfficientNet. In Convolutional Neural Networks with Swift for Tensorflow; Apress: Berkeley, CA, USA, 2021. [Google Scholar] [CrossRef]
- Rehman, M.U.; Cho, S.; Kim, J.H.; Chong, K.T. Bu-net: Brain tumour segmentation using modified u-net architecture. Electronics 2020, 9, 2203. [Google Scholar] [CrossRef]
- Rehman, M.U.; Cho, S.; Kim, J.; Chong, K.T. Brainseg-net: Brain tumour mr image segmentation via enhanced encoder–decoder network. Diagnostics 2021, 11, 169. [Google Scholar] [CrossRef] [PubMed]
- Jalali, Y.; Fateh, M.; Rezvani, M.; Abolghasemi, V.; Anisi, M.H. ResBCDU-Net: A deep learning framework for lung CT image segmentation. Sensors 2021, 21, 268. [Google Scholar] [CrossRef] [PubMed]
- van Eijnatten, M.; van Dijk, R.; Dobbe, J.; Streekstra, G.; Koivisto, J.; Wolff, J. CT image segmentation methods for bone used in medical additive manufacturing. Med. Eng. Phys. 2018, 51, 6–16. [Google Scholar] [CrossRef]
- Neelima, G.; Chigurukota, D.R.; Maram, B.; Girirajan, B. Optimal DeepMRSeg based tumour segmentation with GAN for brain tumour classification. Biomed. Signal Process. Control 2022, 74, 103537. [Google Scholar] [CrossRef]
- Minnema, J.; van Eijnatten, M.; Kouw, W.; Diblen, F.; Mendrik, A.; Wolff, J. CT image segmentation of bone for medical additive manufacturing using a convolutional neural network. Comput. Biol. Med. 2018, 103, 130–139. [Google Scholar] [CrossRef]
- Rehman, M.U.; Ryu, J.; Nizami, I.F.; Chong, K.T. RAAGR2-Net: A brain tumour segmentation network using parallel processing of multiple spatial frames. Comput. Biol. Med. 2023, 152, 106426. [Google Scholar] [CrossRef]
- Islam, K.T.; Wijewickrema, S.; O’leary, S. A deep learning framework for segmenting brain tumours using MRI and synthetically generated CT images. Sensors 2022, 22, 523. [Google Scholar] [CrossRef]
- Rukundo, O. Effects of Image Size on Deep Learning. Electronics 2023, 12, 985. [Google Scholar] [CrossRef]






| Image | Benign | |||
|---|---|---|---|---|
| Class0_Dice | Class1_Dice | Class0_Jaccard | Class1_Jaccard | |
| CT_1 | 0.99756 | 0.98193 | 0.99514 | 0.96451 |
| CT_2 | 0.99775 | 0.98975 | 0.99552 | 0.97972 |
| CT_3 | 0.99758 | 0.98710 | 0.99518 | 0.97454 |
| CT_4 | 0.99738 | 0.98974 | 0.99479 | 0.97969 |
| CT_5 | 0.99777 | 0.97334 | 0.99556 | 0.94807 |
| CT_6 | 0.99701 | 0.98348 | 0.99404 | 0.96751 |
| CT_7 | 0.99761 | 0.97269 | 0.99523 | 0.94684 |
| CT_8 | 0.99729 | 0.99024 | 0.99459 | 0.98068 |
| CT_9 | 0.99895 | 0.98740 | 0.99790 | 0.97512 |
| CT_10 | 0.99882 | 0.92817 | 0.99765 | 0.86597 |
| CT_11 | 0.99786 | 0.97470 | 0.99577 | 0.95066 |
| CT_12 | 0.99814 | 0.97956 | 0.99637 | 0.95994 |
| CT_13 | 0.99862 | 0.96864 | 0.99724 | 0.93920 |
| CT_14 | 0.99718 | 0.95798 | 0.99439 | 0.91935 |
| CT_15 | 0.99460 | 0.88555 | 0.98926 | 0.89461 |
| CT_16 | 0.99696 | 0.96709 | 0.99395 | 0.93623 |
| CT_17 | 0.99686 | 0.96678 | 0.99375 | 0.93569 |
| CT_18 | 0.99777 | 0.96998 | 0.99553 | 0.94171 |
| CT_19 | 0.99643 | 0.94171 | 0.99290 | 0.88984 |
| CT_20 | 0.99724 | 0.96965 | 0.99450 | 0.94101 |
| Image | Malignant | |||
|---|---|---|---|---|
| Class0_Dice | Class1_Dice | Class0_Jaccard | Class1_Jaccard | |
| CT_1 | 0.99485 | 0.917209 | 0.989753 | 0.847079 |
| CT_2 | 0.996533 | 0.950731 | 0.993091 | 0.906089 |
| CT_3 | 0.994843 | 0.912769 | 0.989738 | 0.839535 |
| CT_4 | 0.996412 | 0.881644 | 0.992849 | 0.788339 |
| CT_5 | 0.998503 | 0.9179 | 0.99701 | 0.848258 |
| CT_6 | 0.99661 | 0.951691 | 0.993242 | 0.907835 |
| CT_7 | 0.997797 | 0.927598 | 0.995603 | 0.864972 |
| CT_8 | 0.997502 | 0.969651 | 0.995017 | 0.941091 |
| CT_9 | 0.996653 | 0.961937 | 0.993328 | 0.926665 |
| CT_10 | 0.995213 | 0.926677 | 0.990471 | 0.863373 |
| CT_11 | 0.983532 | 0.836238 | 0.967598 | 0.718565 |
| CT_12 | 0.995118 | 0.866793 | 0.990284 | 0.764902 |
| CT_13 | 0.999053 | 0.966729 | 0.998107 | 0.9356 |
| CT_14 | 0.995317 | 0.861702 | 0.990677 | 0.757008 |
| CT_15 | 0.997093 | 0.952554 | 0.994203 | 0.909406 |
| CT_16 | 0.997117 | 0.965229 | 0.99425 | 0.932794 |
| CT_17 | 0.995457 | 0.727209 | 0.990954 | 0.57135 |
| CT_18 | 0.986163 | 0.701614 | 0.972704 | 0.540374 |
| CT_19 | 0.998494 | 0.914519 | 0.996992 | 0.842501 |
| CT_20 | 0.998775 | 0.918228 | 0.997552 | 0.848818 |
| Sl. No | CNN Architectures | Accuracy | Precision | Recall | F1 Score |
|---|---|---|---|---|---|
| 1. | CNN | 89.7% | 89.1% | 88.0% | 89.4% |
| 2. | ResNet 152 | 92.7% | 92.4% | 91.5% | 92.7% |
| 3. | DenseNet121 | 95.7% | 95.2% | 94.3% | 95.6% |
| 4. | Inception-ResNet V4 | 94.3% | 94.1% | 93.2% | 94.2% |
| 5. | VGG 16 | 91.5% | 91.6% | 90.5% | 91.4% |
| 6. | Xception | 87.2% | 87.3% | 86.7% | 86.3% |
| Sl. No | Ensemble ML Models | Accuracy | Precision | Recall | F1 Score |
|---|---|---|---|---|---|
| 1. | Random Forest | 82.4% | 82.7% | 83.6% | 81.5% |
| 2. | Gradient boosting | 80.9% | 80.3% | 80.88% | 79.5% |
| 3. | AdaBoosting | 89.4% | 88.5% | 89.1% | 87.6% |
| 4. | XGBoosting | 79.5% | 80.2% | 79.8% | 80.3% |
| Sl. No | Ensemble ML Models | Accuracy | Precision | Recall | F1 Score |
|---|---|---|---|---|---|
| 1. | Random Forest | 86.3% | 86.3% | 87.9% | 86.2% |
| 2. | Gradient boosting | 82.7% | 82.7% | 83.7% | 81.3% |
| 3. | AdaBoosting | 91.37% | 91.87% | 90.3% | 91.78% |
| 4. | XGBoosting | 80.34% | 81.67% | 80.48% | 81.78% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kodipalli, A.; Fernandes, S.L.; Gururaj, V.; Varada Rameshbabu, S.; Dasar, S. Performance Analysis of Segmentation and Classification of CT-Scanned Ovarian Tumours Using U-Net and Deep Convolutional Neural Networks. Diagnostics 2023, 13, 2282. https://doi.org/10.3390/diagnostics13132282
Kodipalli A, Fernandes SL, Gururaj V, Varada Rameshbabu S, Dasar S. Performance Analysis of Segmentation and Classification of CT-Scanned Ovarian Tumours Using U-Net and Deep Convolutional Neural Networks. Diagnostics. 2023; 13(13):2282. https://doi.org/10.3390/diagnostics13132282
Chicago/Turabian StyleKodipalli, Ashwini, Steven L. Fernandes, Vaishnavi Gururaj, Shriya Varada Rameshbabu, and Santosh Dasar. 2023. "Performance Analysis of Segmentation and Classification of CT-Scanned Ovarian Tumours Using U-Net and Deep Convolutional Neural Networks" Diagnostics 13, no. 13: 2282. https://doi.org/10.3390/diagnostics13132282
APA StyleKodipalli, A., Fernandes, S. L., Gururaj, V., Varada Rameshbabu, S., & Dasar, S. (2023). Performance Analysis of Segmentation and Classification of CT-Scanned Ovarian Tumours Using U-Net and Deep Convolutional Neural Networks. Diagnostics, 13(13), 2282. https://doi.org/10.3390/diagnostics13132282






