GAR-Net: Guided Attention Residual Network for Polyp Segmentation from Colonoscopy Video Frames
Abstract
1. Introduction
- A novel end-to-end deep learning framework for segmenting polyps from colonoscopy video frames.
- A modified and enhanced Residual Block is proposed that suppresses the noise and preserves the low-level feature maps for a more accurate semantic segmentation.
- A special learning technique is introduced with a novel attention mechanism for obtaining an accurate segmentation map.
- A novel attention mechanism to capture the refined attention maps regardless of the size and shape of the polyp, also under improper illumination conditions.
- Design of a competitive and robust model with consistent performance over the benchmark CVC-ClinicDB dataset and the Kvasir-SEG Dataset.
2. Related Works
3. Proposed Methodology
3.1. Building Blocks of GAR-Net
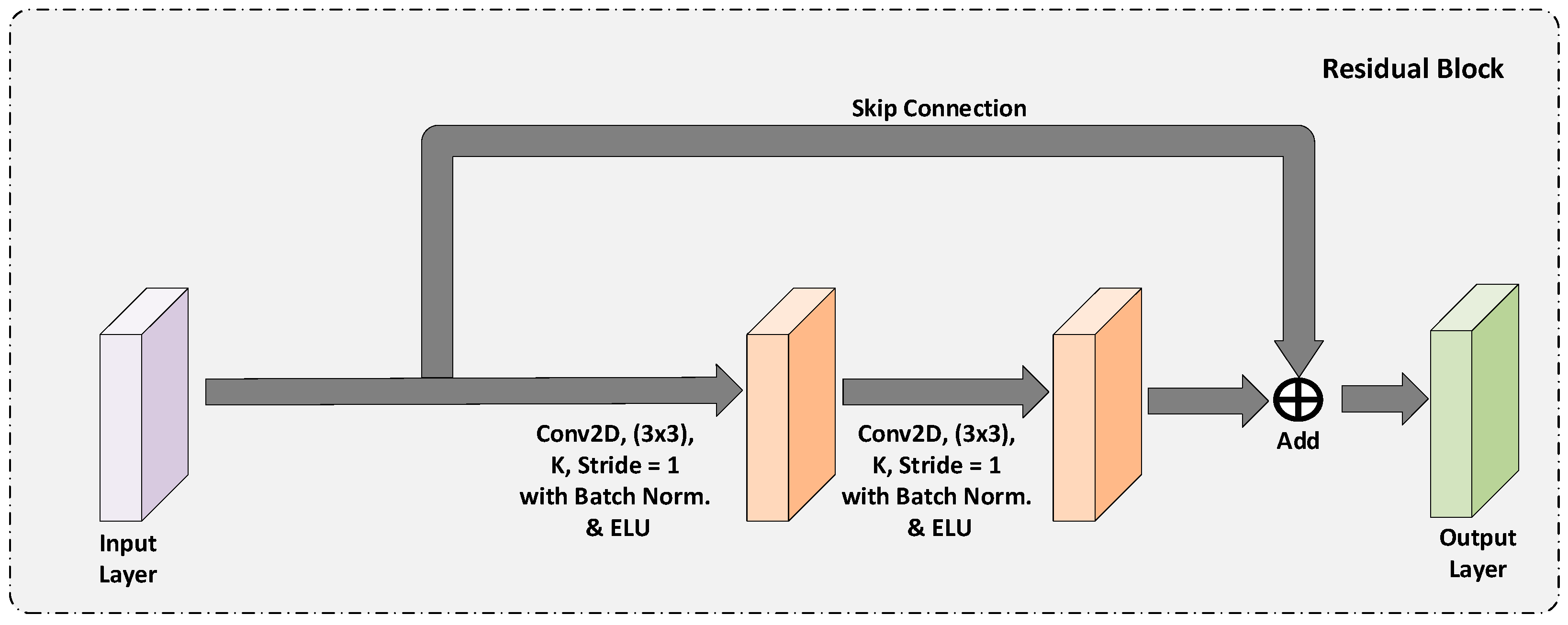
3.1.1. Residual Block
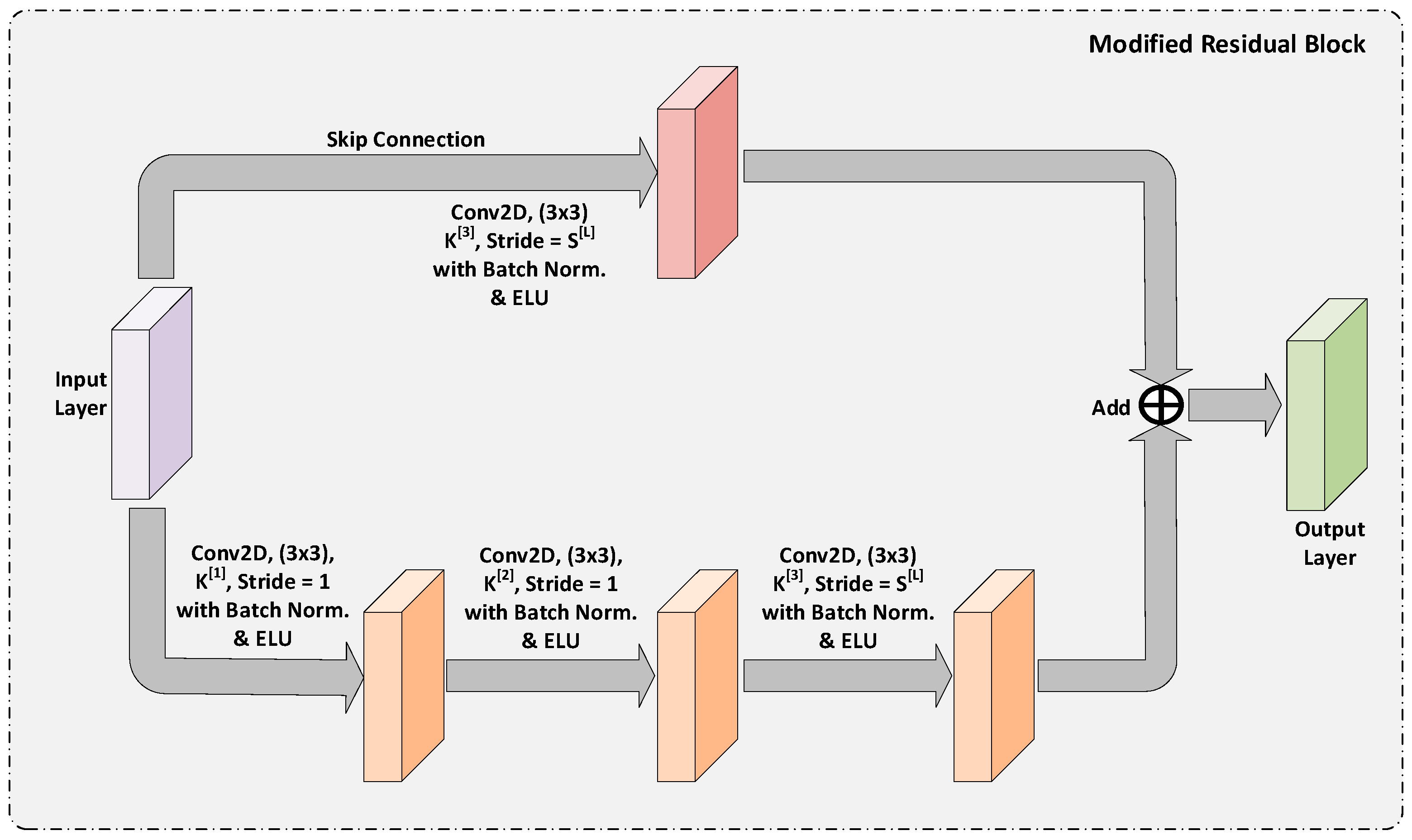
3.1.2. Modified Residual Block
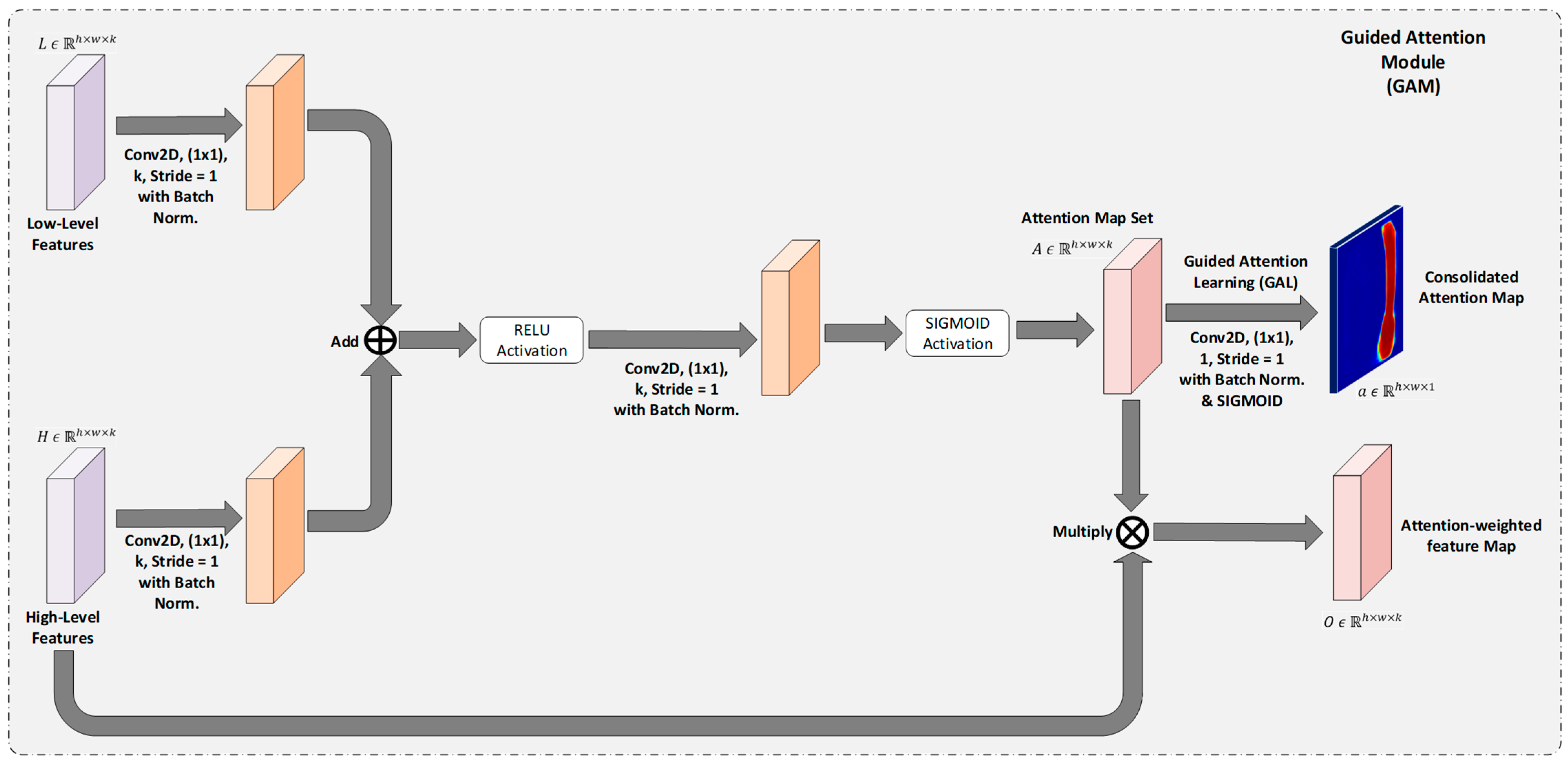
3.1.3. Guided Attention Module
3.2. GAR-Net for Polyp Segmentation
3.2.1. GAR-Net Architecture
3.2.2. Guided Attention Learning
3.2.3. Loss Function for GAR-Net
4. Results and Discussion:
4.1. Dataset
4.2. Experimental Setup
4.3. Training Setup
4.4. Results
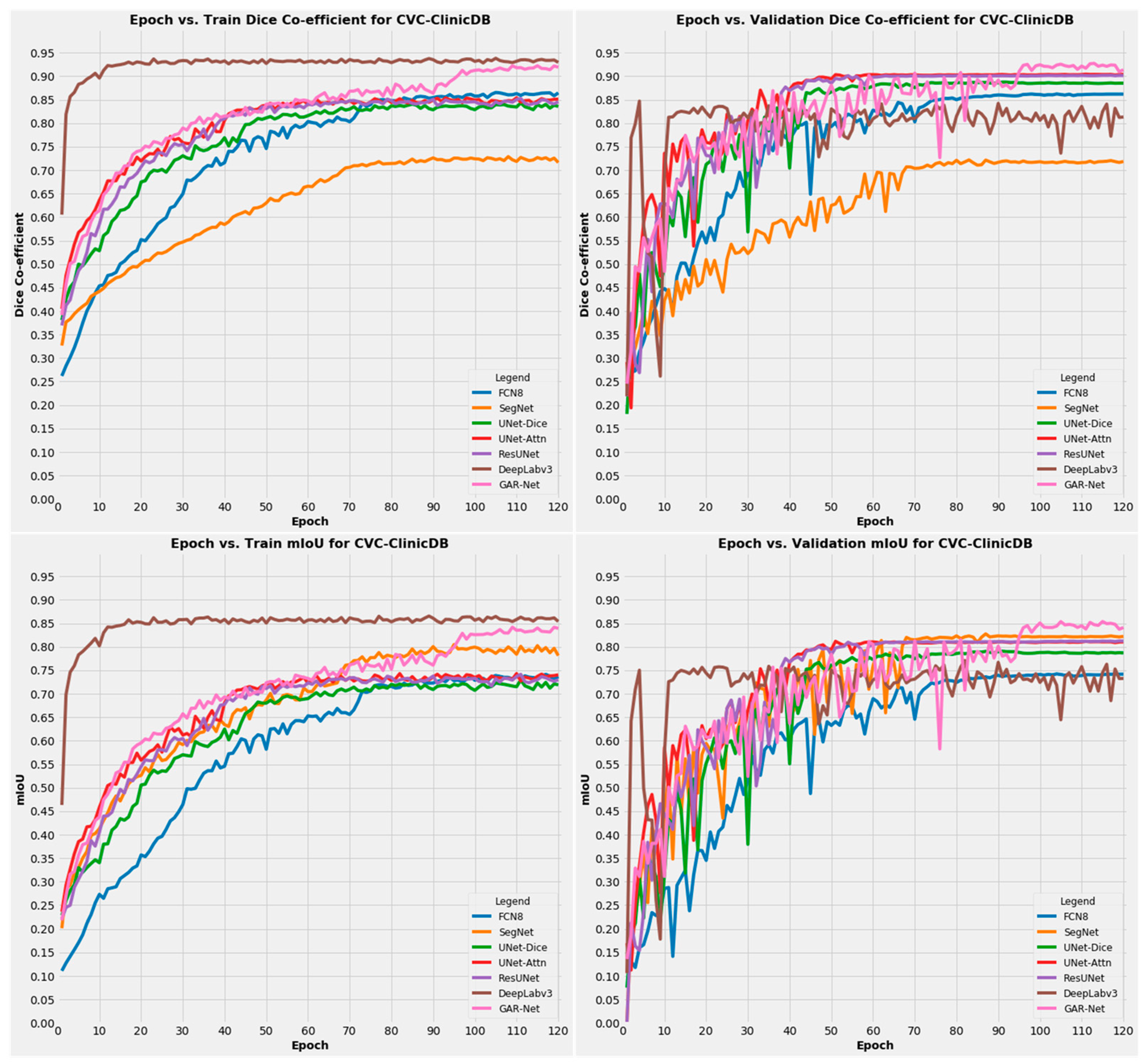
4.4.1. Results on CVC-ClinicDB Dataset
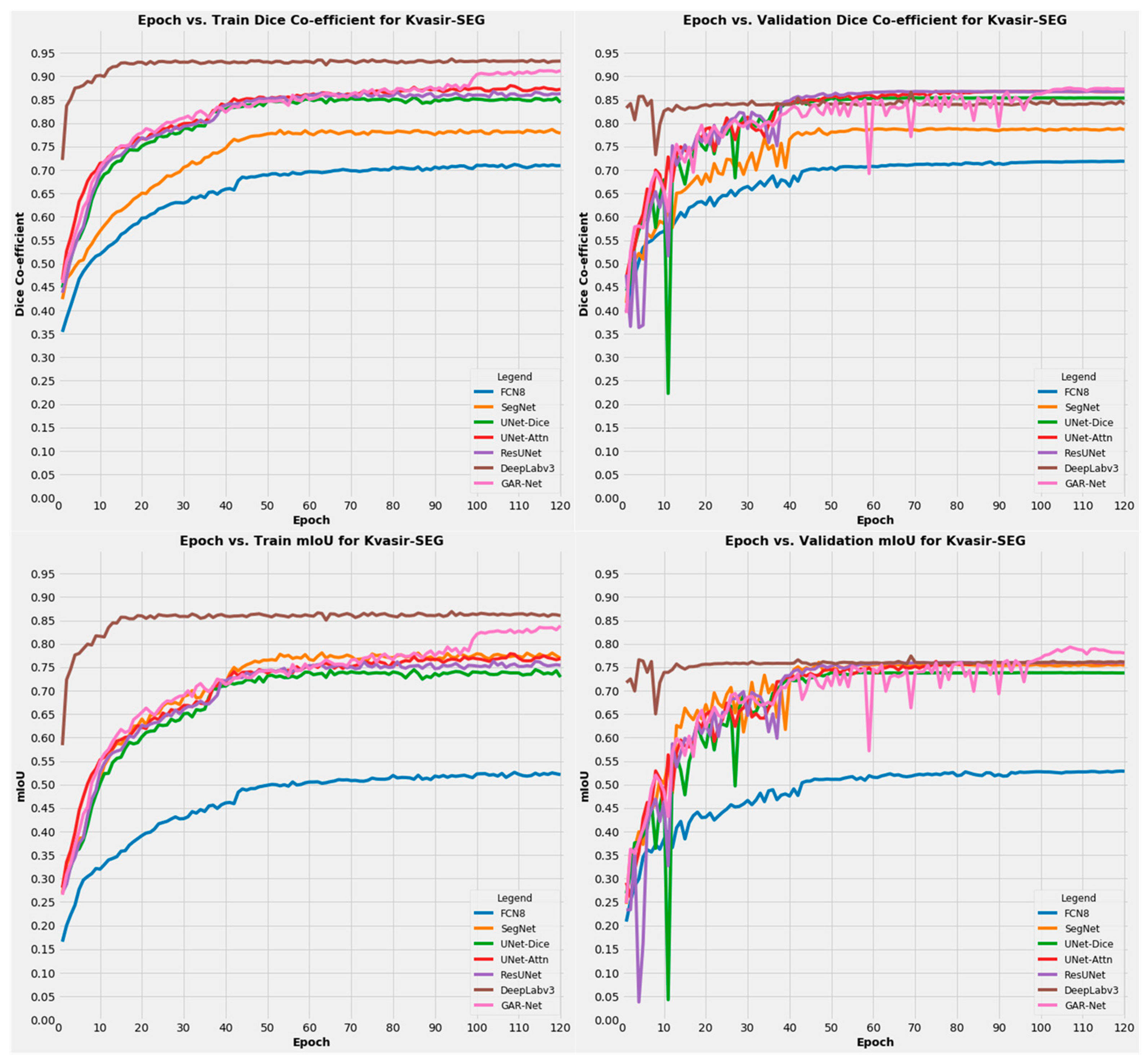
4.4.2. Results on Kvasir-SEG Dataset
4.5. Further Discussions
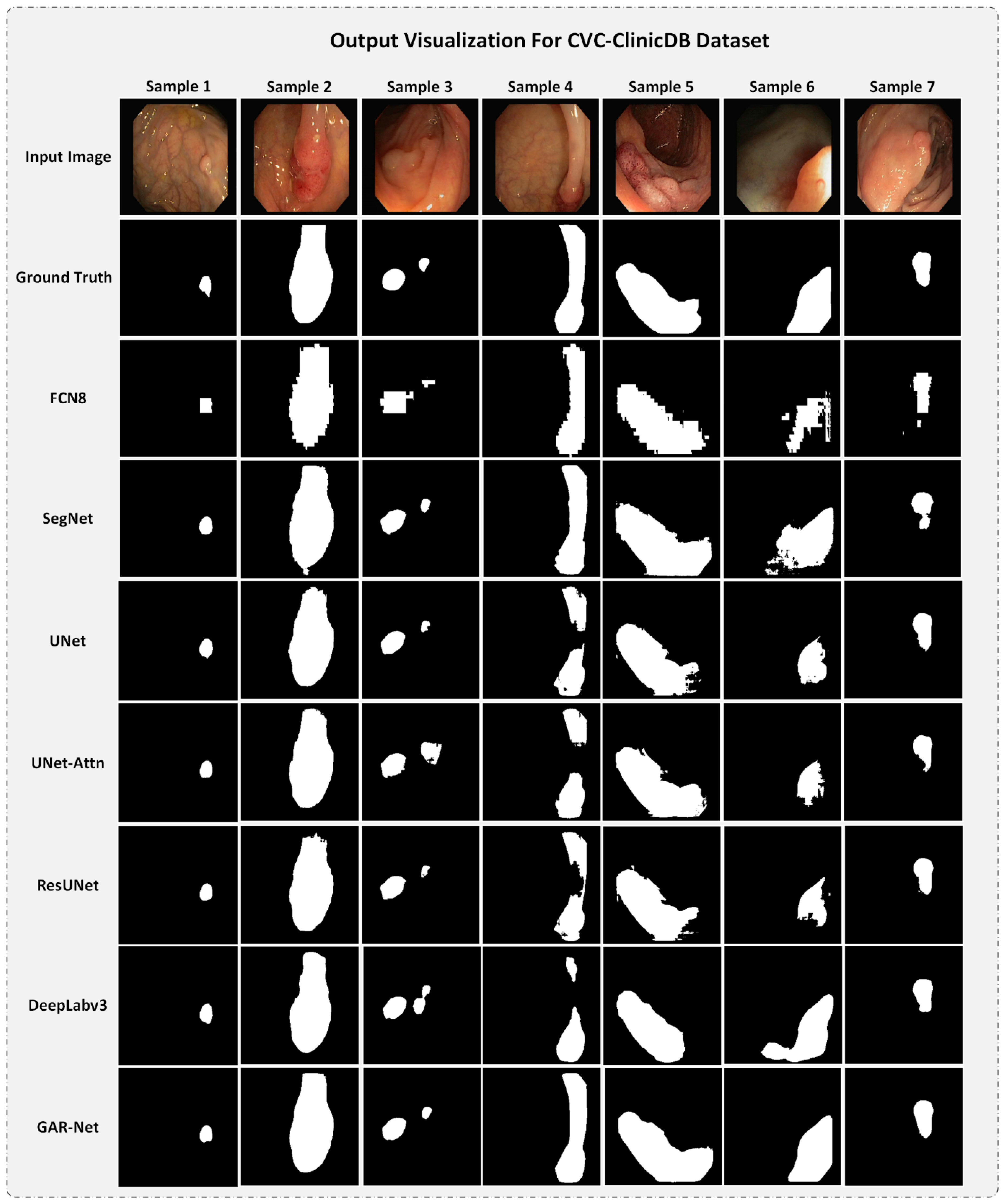
4.5.1. Output Visualization for CVC-ClinicDB Dataset
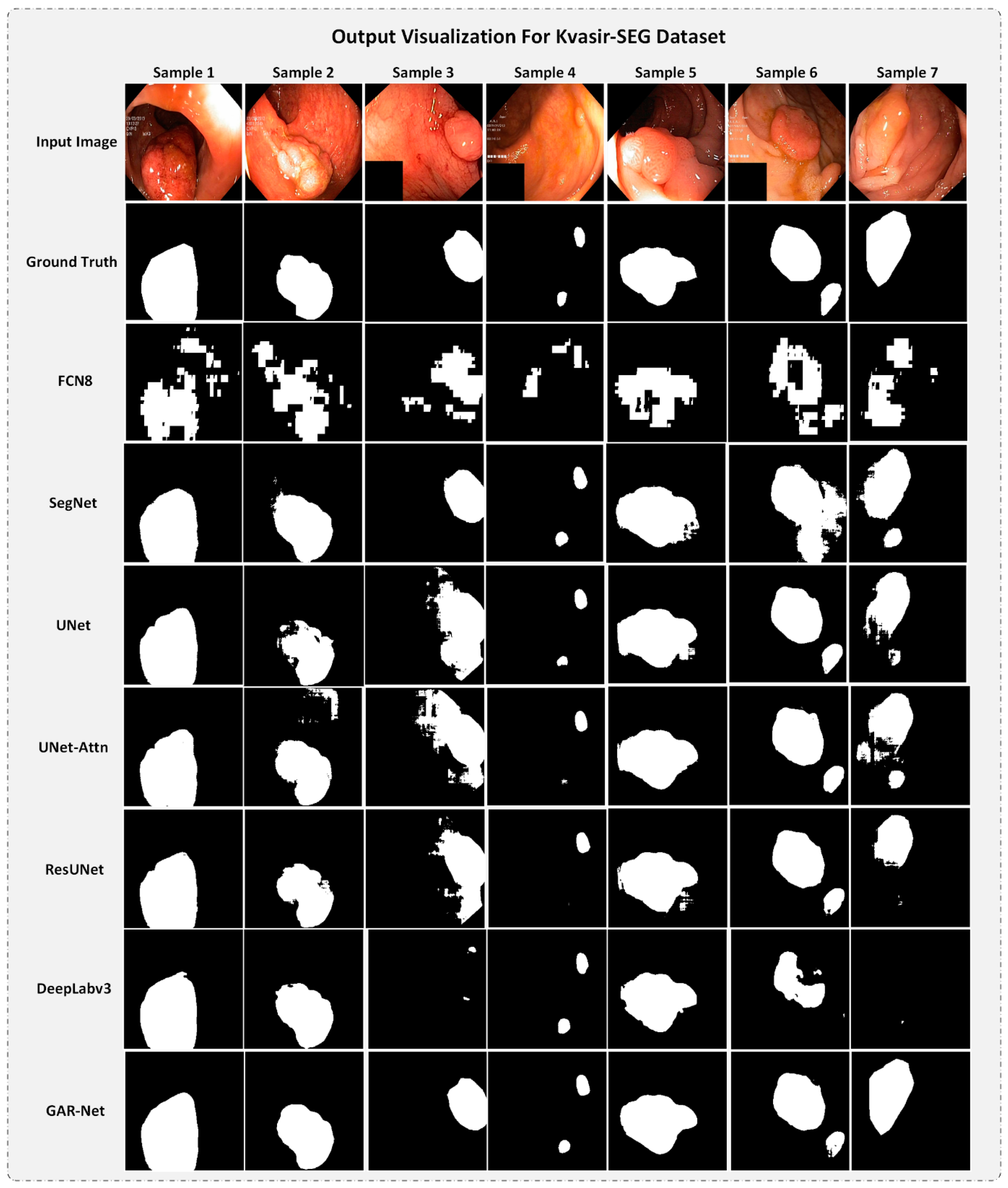
4.5.2. Output Visualization for Kvasir-SEG dataset
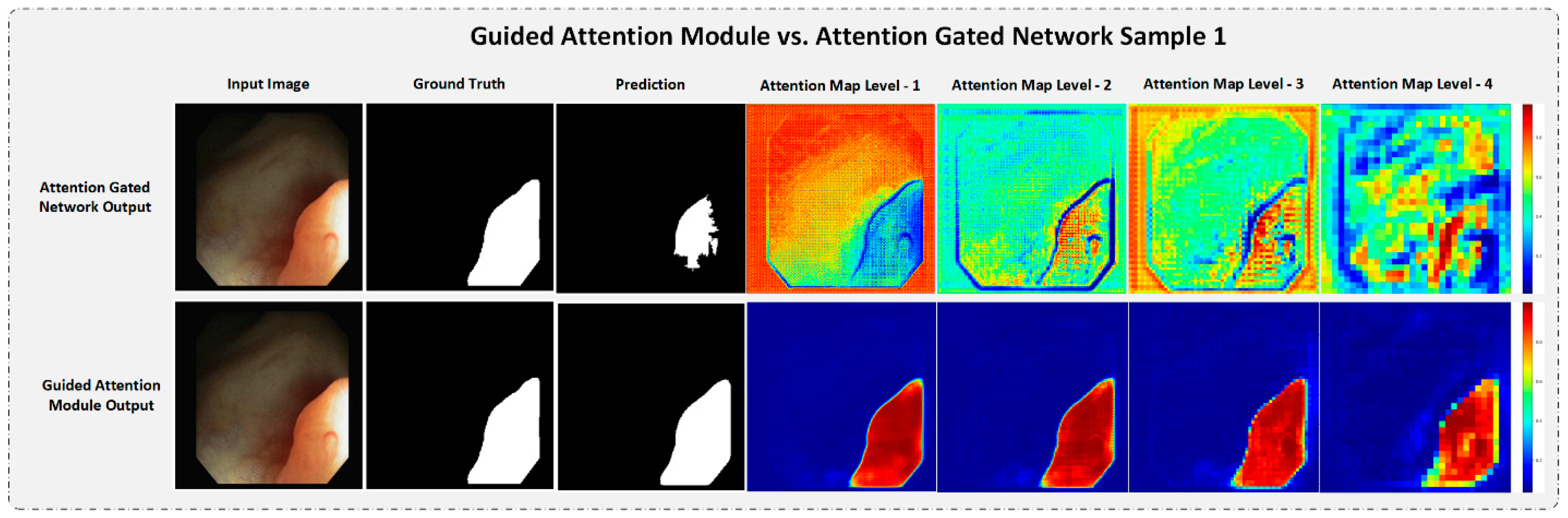
4.5.3. Strength of the Proposed Guided Attention Learning
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lawton, C. Colonoscopic Polypectomy and Long-Term Prevention of Colorectal-Cancer Deaths. Yearb. Oncol. 2013, 2013, 128–129. [Google Scholar] [CrossRef]
- van Rijn, J.C.; Reitsma, J.B.; Stoker, J.; Bossuyt, P.M.; van Deventer, S.J.; Dekker, E. Polyp Miss Rate Determined by Tandem Colonoscopy: A Systematic Review. Am. J. Gastroenterol. 2006, 101, 343–350. [Google Scholar] [CrossRef]
- Breier, M.; Gross, S.; Behrens, A.; Stehle, T.; Aach, T. Active contours for localizing polyps in colonoscopic NBI image data. Med. Imaging 2011 Comput.-Aided Diagn. 2011, 7963, 79632M. [Google Scholar] [CrossRef]
- Bernal, J.; Sánchez, J.; Vilariño, F. Towards automatic polyp detection with a polyp appearance model. Pattern Recognit. 2012, 45, 3166–3182. [Google Scholar] [CrossRef]
- Manjunath, K.N.; Prabhu, G.K.; Siddalingaswamy, P.C. A quantitative validation of segmented colon in virtual colonoscopy using image moments. Biomed. J. 2020, 43, 74–82. [Google Scholar] [CrossRef]
- Diakogiannis, F.I.; Waldner, F.; Caccetta, P.; Wu, C. ResUNet-a: A deep learning framework for semantic segmentation of remotely sensed data. ISPRS J. Photogramm. Remote Sens. 2020, 162, 94–114. [Google Scholar] [CrossRef]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. SegNet: A Deep Convolutional Encoder-Decoder Architecture for Image Segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention; Lecture Notes in Computer Science (Including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Cham, Switzerland, 2015; Volume 9351, pp. 234–241. [Google Scholar] [CrossRef]
- Vázquez, D.; Bernal, J.; Sánchez, F.J.; Fernández-Esparrach, G.; López, A.M.; Romero, A.; Drozdzal, M.; Courville, A. A Benchmark for Endoluminal Scene Segmentation of Colonoscopy Images. J. Healthc. Eng. 2017, 2017, 4037190. [Google Scholar] [CrossRef]
- Haj-Manouchehri, A.; Mohammadi, H.M. Polyp detection using CNNs in colonoscopy video. IET Comput. Vis. 2020, 14, 241–247. [Google Scholar] [CrossRef]
- Hao, D.; Ding, S.; Qiu, L.; Lv, Y.; Fei, B.; Zhu, Y.; Qin, B. Sequential vessel segmentation via deep channel attention network. Neural Netw. 2020, 128, 172–187. [Google Scholar] [CrossRef]
- Chung, M.; Lee, J.; Park, S.; Lee, C.E.; Lee, J.; Shin, Y.-G. Liver segmentation in abdominal CT images via auto-context neural network and self-supervised contour. Artif. Intellingence Med. 2021, 113, 102023. [Google Scholar] [CrossRef]
- Jha, D.; Smedsrud, P.H.; Riegler, M.A.; Halvorsen, P.; de Lange, T.; Johansen, D.; Johansen, H.D. Kvasir-SEG: A Segmented Polyp Dataset. In MultiMedia Modeling; Springer: Cham, Switzerland, 2020; pp. 451–462. [Google Scholar] [CrossRef]
- Aina, O.E.; Adeshina, S.A.; Aibinu, A.M. Deep learning for image-based cervical cancer detection and diagnosis—A survey. In Proceedings of the 2019 15th International Conference on Electronics, Computer and Computation, ICECCO 2019, Abuja, Nigeria, 10–12 December 2019. [Google Scholar] [CrossRef]
- Zaiane, O.; Liu, X.; Zhao, D.; Li, W. A multi-kernel based framework for heterogeneous feature selection and over-sampling for computer-aided detection of pulmonary nodules. Pattern Recognit. 2017, 64, 327–346. [Google Scholar] [CrossRef]
- Tajbakhsh, N.; Suzuki, K. Comparing two classes of end-to-end machine-learning models in lung nodule detection and classification: MTANNs vs. CNNs. Pattern Recognit. 2017, 63, 476–486. [Google Scholar] [CrossRef]
- Gross, S.; Kennel, M.; Stehle, T.; Wulff, J.; Tischendorf, J.; Trautwein, C.; Aach, T. Polyp segmentation in NBI colonoscopy. In Informatik Aktuell; Springer: Berlin/Heidelberg, Germany, 2009; pp. 252–256. [Google Scholar] [CrossRef]
- Karkanis, S.A.; Iakovidis, D.K.; Maroulis, D.E.; Karras, D.A.; Tzivras, M. Computer-Aided Tumor Detection in Endoscopic Video Using Color Wavelet Features. IEEE Trans. Inf. Technol. Biomed. 2003, 7, 141–152. [Google Scholar] [CrossRef]
- Cong, Y.; Wang, S.; Liu, J.; Cao, J.; Yang, Y.; Luo, J. Deep sparse feature selection for computer aided endoscopy diagnosis. Pattern Recognit. 2015, 48, 907–917. [Google Scholar] [CrossRef]
- Sánchez-González, A.; García-Zapirain, B.; Sierra-Sosa, D.; Elmaghraby, A. Automatized colon polyp segmentation via contour region analysis. Comput. Biol. Med. 2018, 100, 152–164. [Google Scholar] [CrossRef]
- Yao, J.; Li, J.; Summers, R.M. Employing topographical height map in colonic polyp measurement and false positive reduction. Pattern Recognit. 2009, 42, 1029–1040. [Google Scholar] [CrossRef][Green Version]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. Adv. Neural Inf. Process. Syst. 2017, 2017, 5999–6009. [Google Scholar]
- Schlemper, J.; Oktay, O.; Schaap, M.; Heinrich, M.; Kainz, B.; Glocker, B.; Rueckert, D. Attention gated networks: Learning to leverage salient regions in medical images. Med. Image Anal. 2019, 53, 197–207. [Google Scholar] [CrossRef]
- Thambawita, V.; Jha, D.; Riegler, M.; Halvorsen, P.; Hammer, H.L.; Johansen, H.D.; Johansen, D. The Medico-Task 2018: Disease detection in the gastrointestinal tract using global features and deep learning. CEUR Workshop Proc. 2018, 2283. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar] [CrossRef]
- Ioffe, S.; Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. In Proceedings of the 32nd International Conference on Machine Learning, ICML 2015, Lille, France, 6–11 July 2015; Volume 1, pp. 448–456. [Google Scholar]
- Bernal, J.; Sánchez, F.J.; Fernández-Esparrach, G.; Gil, D.; Rodríguez, C.; Vilariño, F. WM-DOVA maps for accurate polyp highlighting in colonoscopy: Validation vs. saliency maps from physicians. Comput. Med. Imaging Graph. 2015, 43, 99–111. [Google Scholar] [CrossRef]
- Dong, H.; Supratak, A.; Mai, L.; Liu, F.; Oehmichen, A.; Yu, S.; Guo, Y. TensorLayer: A versatile library for efficient deep learning development. In Proceedings of the 2017 ACM Multimedia Conference, Mountain View, CA, USA, 23–27 October 2017; pp. 1201–1204. [Google Scholar] [CrossRef]
- Keys, R. Cubic convolution interpolation for digital image processing. IEEE Trans. Acoust. 1981, 29, 1153–1160. [Google Scholar] [CrossRef]
- Chen, L.C.; Papandreou, G.; Kokkinos, I.; Murphy, K.; Yuille, A.L. DeepLab: Semantic Image Segmentation with Deep Convolutional Nets, Atrous Convolution, and Fully Connected CRFs. IEEE Trans. Pattern Anal. Mach. Intell. 2018, 40, 834–848. [Google Scholar] [CrossRef]


| S No | Model | Dice Score | mIoU | Pixel Accuracy |
|---|---|---|---|---|
| 1 | FCN8 | 0.8724981 | 0.74308 | 0.9757281 |
| 2 | SegNet | 0.73168665 | 0.811956 | 0.97676927 |
| 3 | UNet | 0.8803434 | 0.767874 | 0.97531813 |
| 4 | UNet-Attn | 0.89131886 | 0.783651 | 0.97713906 |
| 5 | ResUNet | 0.89080334 | 0.781462 | 0.97680587 |
| 6 | DeepLabv3 | 0.90001955 | 0.819477 | 0.9817561 |
| 7 | GAR-Net | 0.9100929 | 0.831234 | 0.9831491 |
| S No | Model | Dice Score | mIoU | Pixel Accuracy |
|---|---|---|---|---|
| 1 | FCN8 | 0.73635364 | 0.548514 | 0.9176181 |
| 2 | SegNet | 0.78634053 | 0.784731 | 0.9607266 |
| 3 | UNet | 0.85862905 | 0.750284 | 0.959373 |
| 4 | UNet-Attn | 0.8637741 | 0.754757 | 0.9587085 |
| 5 | ResUNet | 0.86858636 | 0.761282 | 0.9630264 |
| 6 | DeepLabv3 | 0.87866044 | 0.805872 | 0.9693878 |
| 7 | GAR-Net | 0.8915458 | 0.815802 | 0.9717203 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Raymann, J.; Rajalakshmi, R. GAR-Net: Guided Attention Residual Network for Polyp Segmentation from Colonoscopy Video Frames. Diagnostics 2023, 13, 123. https://doi.org/10.3390/diagnostics13010123
Raymann J, Rajalakshmi R. GAR-Net: Guided Attention Residual Network for Polyp Segmentation from Colonoscopy Video Frames. Diagnostics. 2023; 13(1):123. https://doi.org/10.3390/diagnostics13010123
Chicago/Turabian StyleRaymann, Joel, and Ratnavel Rajalakshmi. 2023. "GAR-Net: Guided Attention Residual Network for Polyp Segmentation from Colonoscopy Video Frames" Diagnostics 13, no. 1: 123. https://doi.org/10.3390/diagnostics13010123
APA StyleRaymann, J., & Rajalakshmi, R. (2023). GAR-Net: Guided Attention Residual Network for Polyp Segmentation from Colonoscopy Video Frames. Diagnostics, 13(1), 123. https://doi.org/10.3390/diagnostics13010123






