Hamlet-Pattern-Based Automated COVID-19 and Influenza Detection Model Using Protein Sequences
Abstract
:1. Introduction
- A novel feature extraction method based on a novel pattern that was inspired by a literary work. The presented feature extraction method is the first text-based feature extraction function creation methodology.
- Using protein sequences, a classification model incorporating the novel pattern was applied for the binary classification of SARS-CoV-2 versus Influenza-A diagnosis. The model attained excellent classification performance, supporting its potential use as an adjunctive screening tool for suspected viral respiratory infections in the current pandemic.
2. Materials and Methods
2.1. Materials
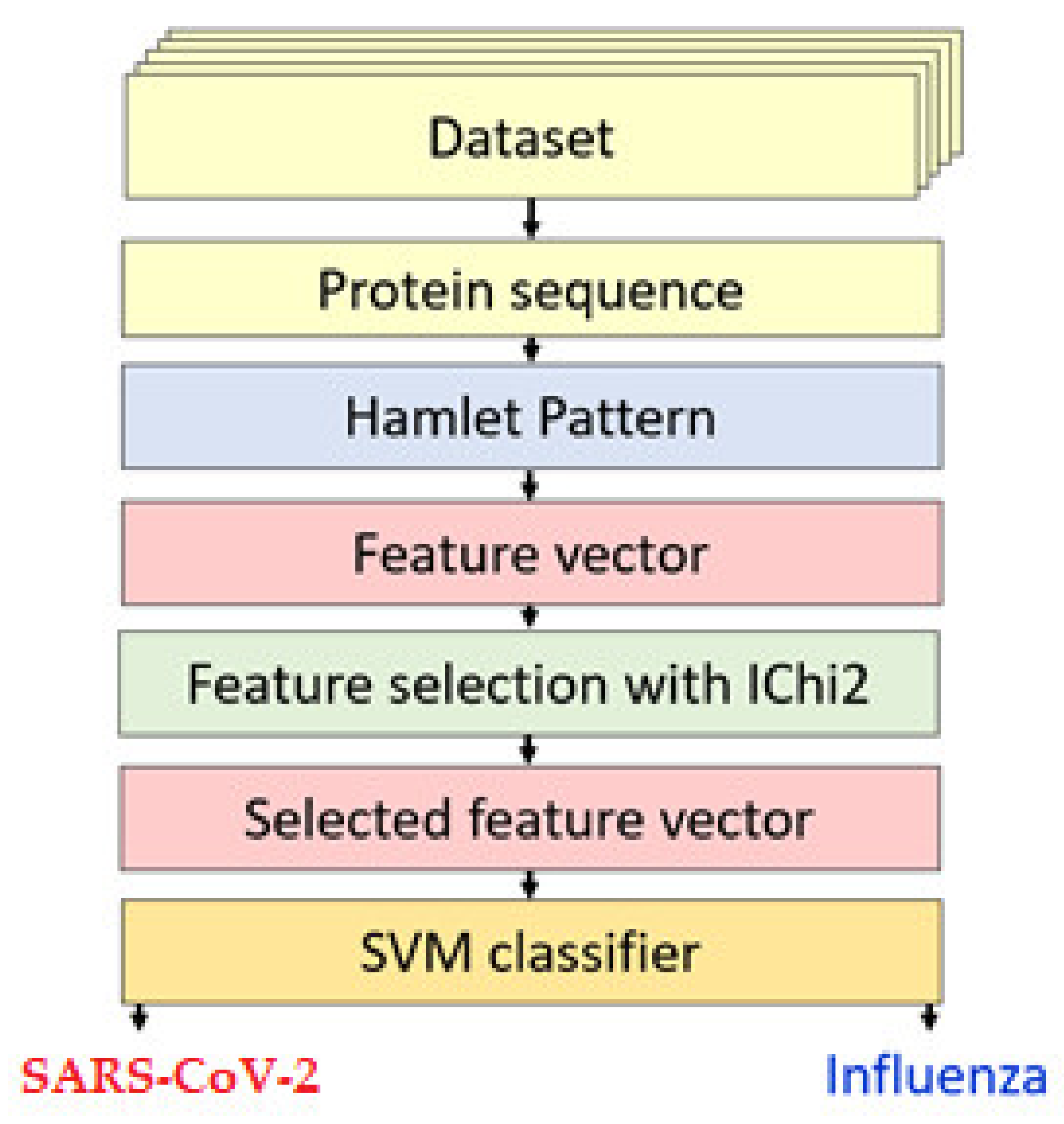
2.2. Our Proposed Protein Sequence Classification Model
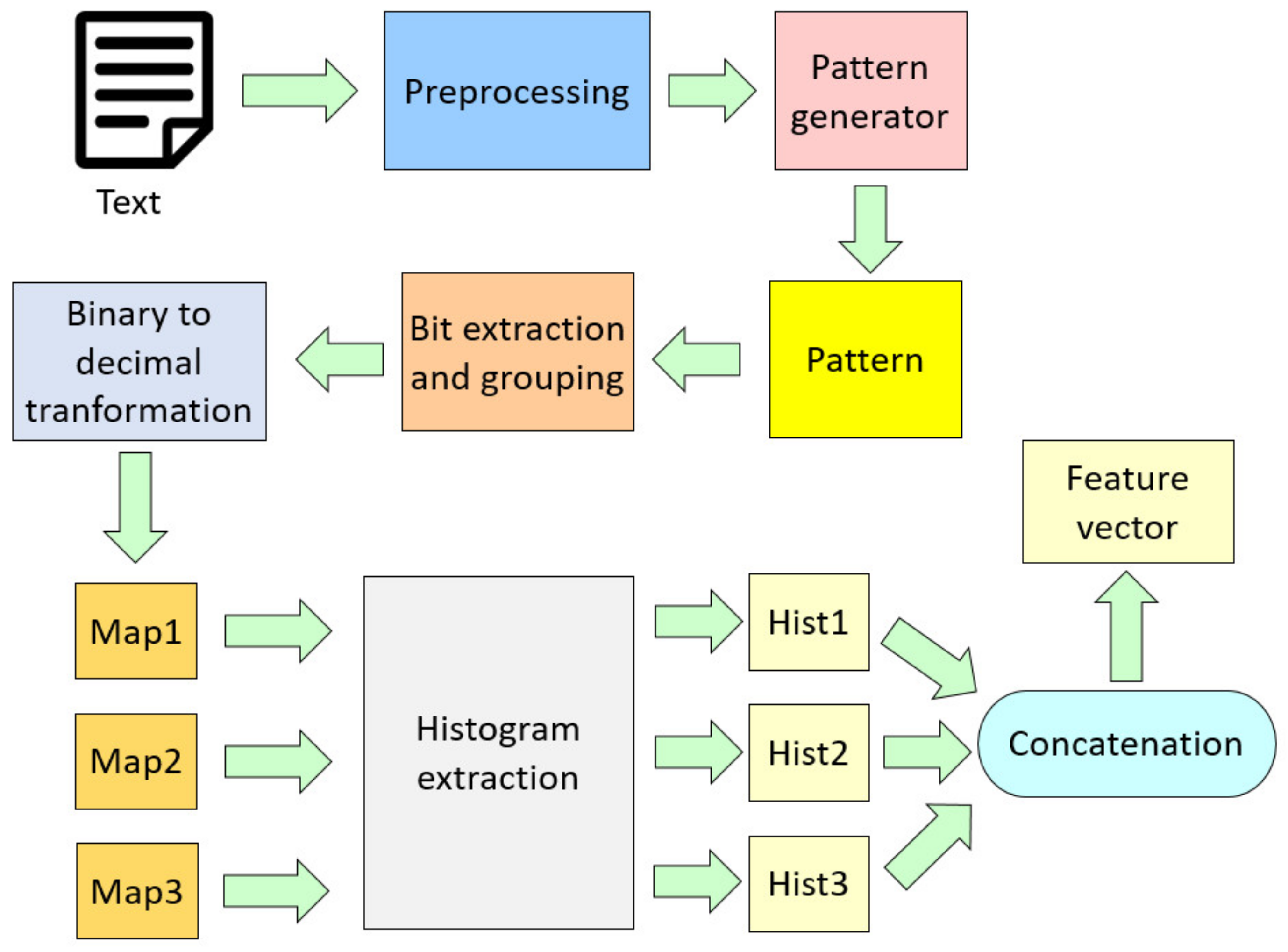
2.2.1. Feature Extraction Using HamletPat
| Algorithm 1. Pattern generator using enumerated letters. |
| Input: The calculated values of the letters Output: Pattern |
| 01: for i = 1 to 26 do // Assign counter |
| 02: ; |
| 03: end for i |
| 04: i = 1; j = 1; // Define variables |
| 05: |
| 06: while < 26 do |
| 07: |
| 08: if then |
| 09: ; |
| 10: ; |
| 11: |
| 12: end if |
| 13: |
| 14: |
| 15: end while |
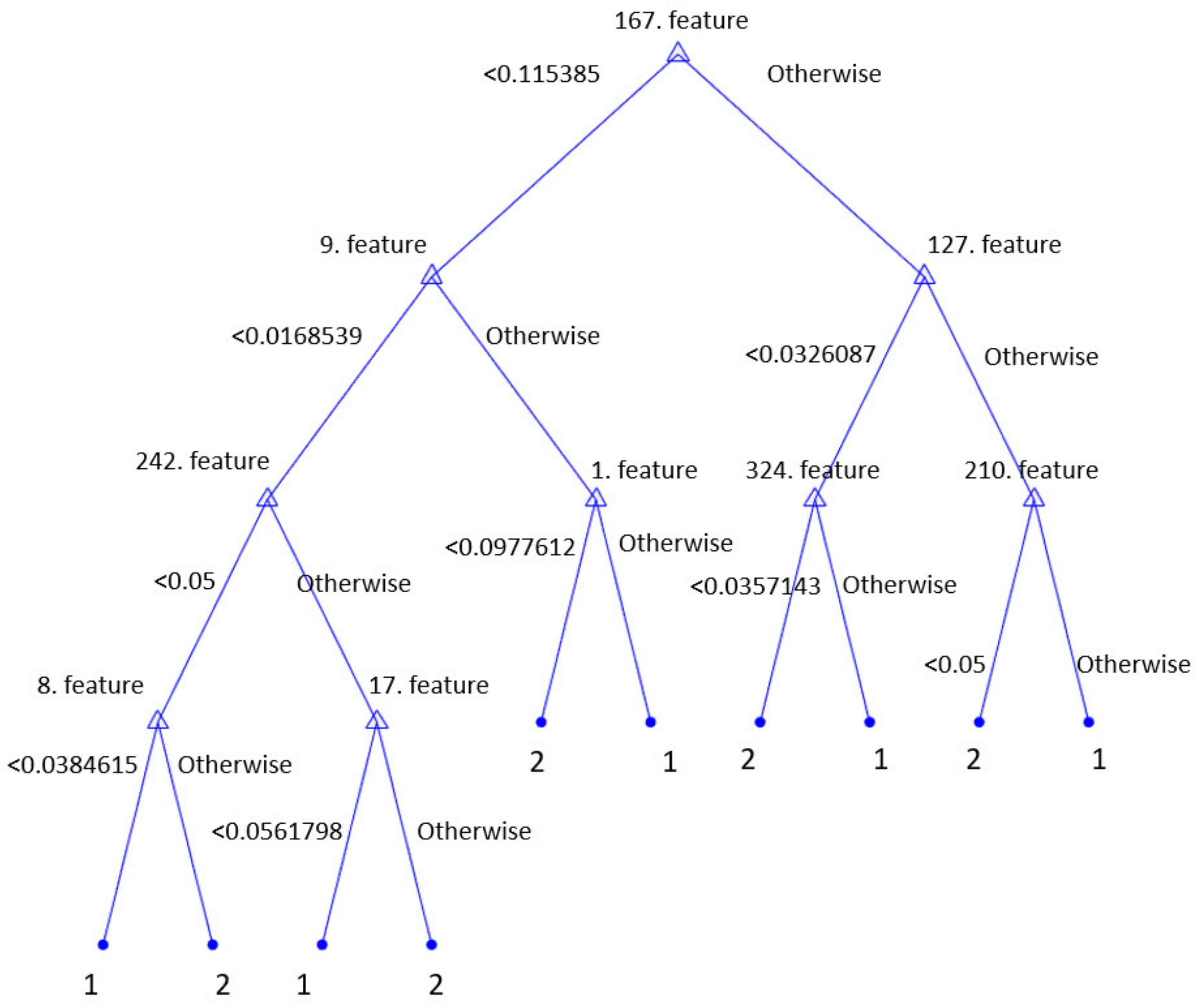
2.2.2. Iterative Chi-Square Feature Selection
2.2.3. Classification
3. Results
3.1. Experimental Setup
3.2. Evaluation Metrics
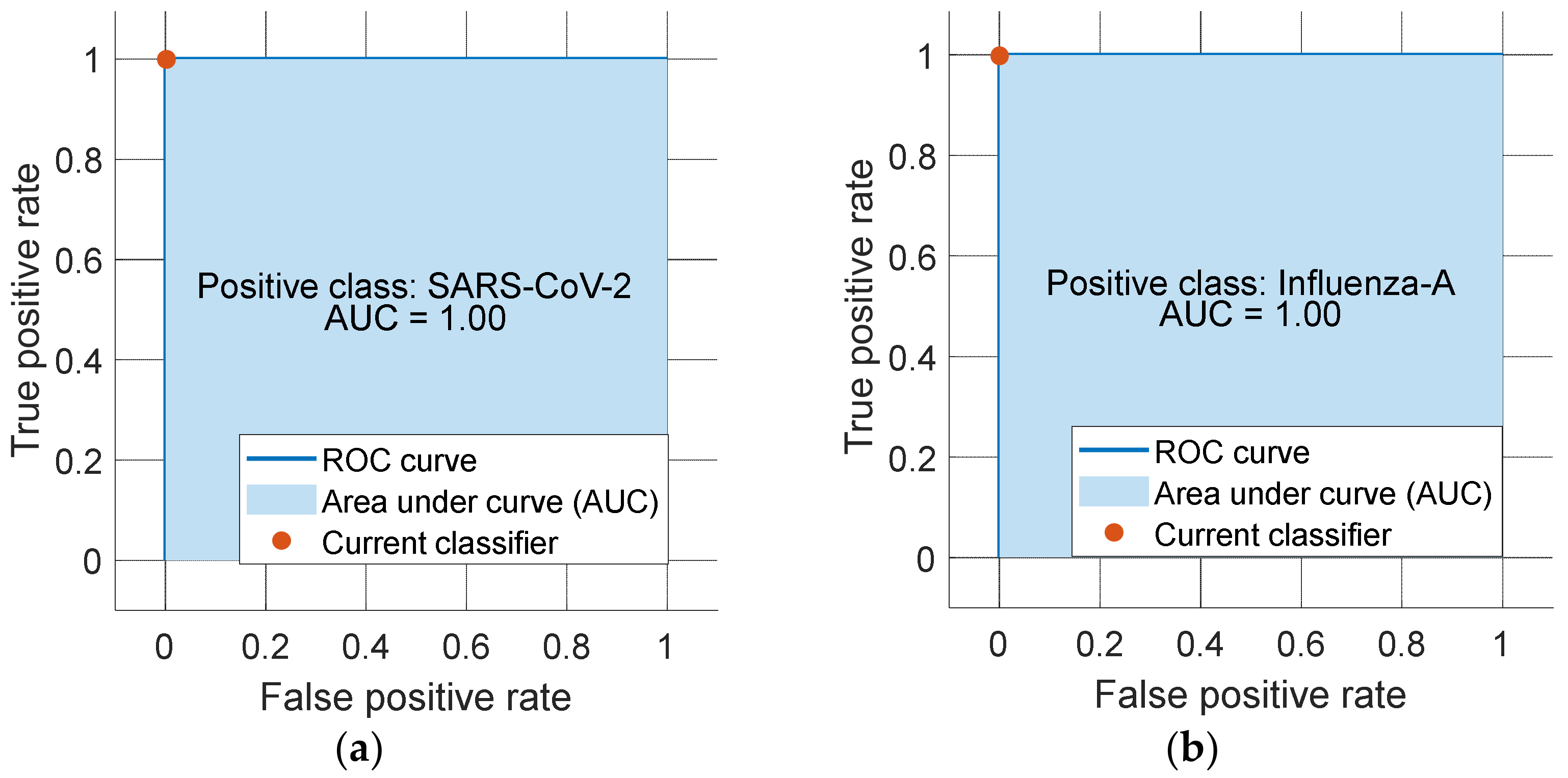
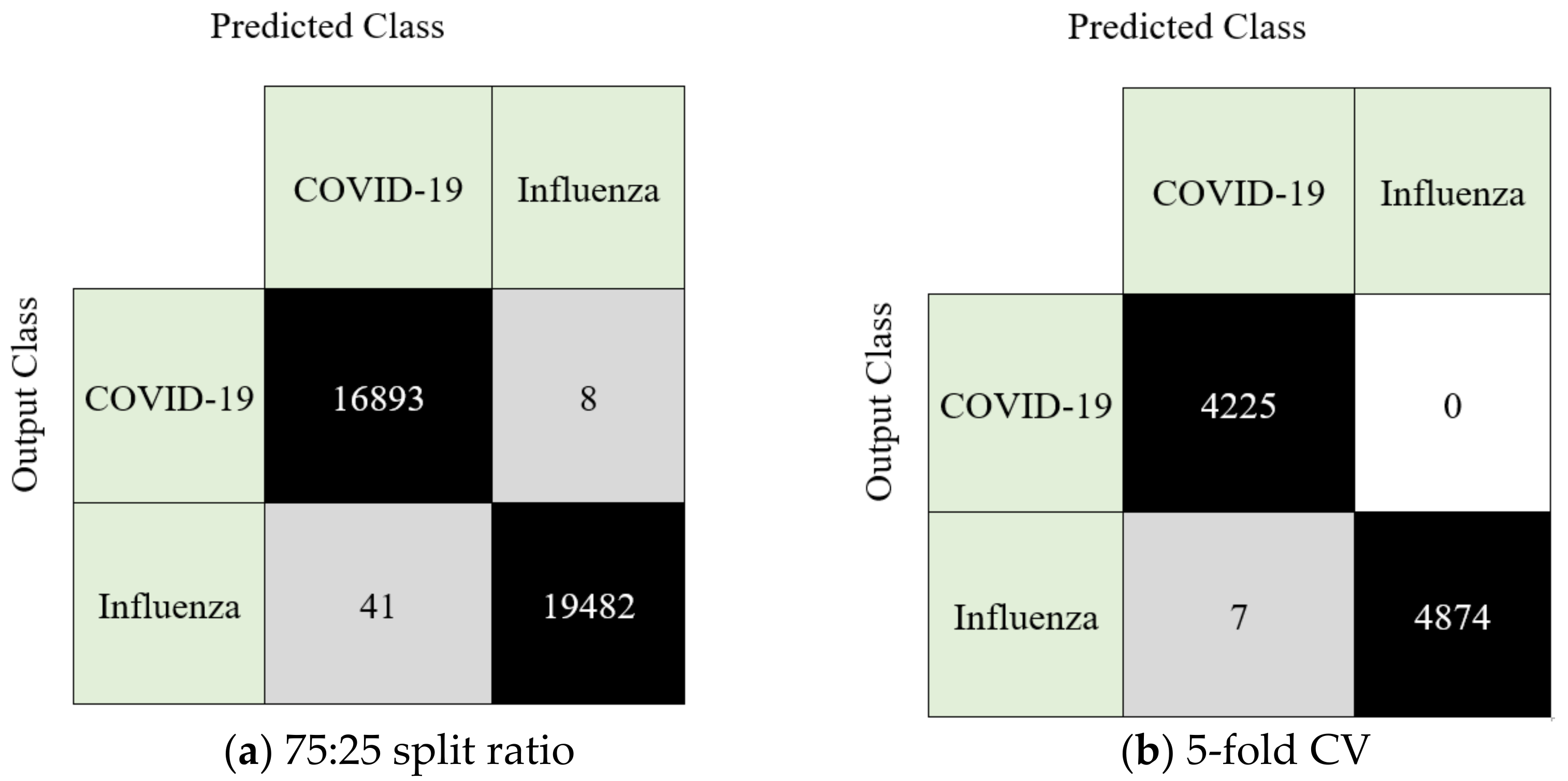
3.3. Performance of the Proposed Model
3.4. Time Complexity Analysis
4. Discussion
- Influenza and COVID-19 share similar symptoms, and clinical discrimination is difficult. Therefore, an automated protein-sequence-based model was developed to differentiate the disorders automatically.
- To our knowledge, HamletPat is the first text-based pattern utilized to create a new feature extraction function.
- The novel HamletPat-based classification model was trained on a two-class dataset and attained 99.87% and 99.92% accuracy rates by deploying a five-fold CV and hold-out (split ratio 75:25) CV, respectively.
- The model is simple, it has a low time complexity of and is easy to implement.
- The model used overlapping blocks with a fixed length of 27. Therefore, the minimum length of the studied protein sequence should be 27 (we used a protein sequence with a length of 100 or greater in the study).
- We used the SVM classifier with default hyperparameters in the study. The hyperparameters can be further optimized using a metaheuristic optimization model.
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| clc,clear all,close all text = ‘bernardowhostherefrancisconayanswermestandandunfoldyourselfbernardolonglivethekingfranciscobernardobernardohefranciscoyoucomemostcarefullyuponyourhourbernardotisnowstrucktwelvegettheetobedfranciscofranciscoforthisreliefmuchthankstisbittercoldandiamsickatheartbernardohaveyouhadquietguardfrancisconotamousestirringbernardowellgoodnightifyoudomeethoratioandmarcellustherivalsofmywatchbidthemmakehastefranciscoithinkihearthemstandhowhosthereenterhoratioandmarcellushoratiofriendstothisgroundmarcellusandliegementothedanefranciscogiveyougoodnightmarcellusofarewellhonestsoldierwhohathrelievedyoufranciscobernardohasmyplacegiveyougoodnightexitmarcellushollabernardobernardosaywhatishoratiotherehoratioapieceofhimbernardowelcomehoratiowelomegoodmarcellusmarcelluswhathasthisthingappeardagaintonightbernardoihaveseennothingmarcellushoratiosaystisbutourfantasyandwillnotletbelieftakeholdofhimtouchingthisdreadedsighttwiceseenofusthereforeihaveentreatedhimalongwithustowatchtheminutesofthisnightthatifagainthisapparitioncomehemayapproveoureyesandspeaktoithoratiotushtushtwillnotappearbernardositdownawhileandletusonceagainassailyourearsthataresofortifiedagainstourstorywhatwehavetwonightsseenhoratiowellsitwedownandletushearbernardospeakofthisbernardolastnightofallwhenyondsamestarthatswestwardfromthepolehadmadehiscoursetoillumethatpartofheavenwherenowitburnsmarcellusandmyselfthebellthenbeatingoneenterghostmarcelluspeacebreaktheeofflookwhereitcomesagainbernardointhesamefigurelikethekingthatsdeadmarcellusthouartascholarspeaktoithoratiobernardolooksitnotlikethekingmarkithoratiohoratiomostlikeitharrowsmewithfearandwonderbernardoitwouldbespoketomarcellusquestionithoratiohoratiowhatartthouthatusurpstthistimeofnighttogetherwiththatfairandwarlikeforminwhichthemajestyofburieddenmarkdidsometimesmarchbyheavenichargetheespeakmarcellusitisoffendedbernardoseeitstalksawayhoratiostayspeakspeakichargetheespeakexitghostmarcellustisgoneandwillntanswerbernardohownowhoratioyoutrembleandlookpaleisnotthissomethingmorethanfantasywhatthinkyouonthoratiobeforemygodimightnotthisbelievewithoutthesensibleandtrueavouchofmineowneyesmarcellusisitnotlikethekinghoratioasthouarttothyselfsuchwastheveryarmourhehadonwhenhetheambitiousnorwaycombatedsofrowndheoncewheninanangryparlehesmotethesleddedpolacksontheicetisstrangemarcellusthustwicebeforeandjumpatthisdeadhourwithmartialstalkhathhegonebyourwatchhoratioinwhatparticularthoughttoworkiknownotbutinthegrossandscopeofmyopinionthisbodessomestrangeeruptiontoourstatemarcellusgoodnowsitdownandtellmehethatknowswhythissamestrictandmostobservantwatchsonightlytoilsthesubjectofthelandandwhysuchdailycastofbrazencannonandforeignmartforimplementsofwarwhysuchimpressofshipwrightswhosesoretaskdoesnotdividethesundayfromtheweekwhatmightbetowardthatthissweatyhastedothmakethenightjointlabourerwiththedaywhoistthatcaninformmehoratiothatcaniatleastthewhispergoessoourlastkingwhoseimageevenbutnowappeardtouswasasyouknowbyfortinbrasofnorwaytheretoprickdonbyamostemulatepridedaredtothecombatinwhichourvalianthamletforsothissideofourknownworldesteemdhimdidslaythisfortinbraswhobyasealdcompactwellratifiedbylawandheraldrydidforfeitwithhislifeallthosehislandswhichhestoodseizedoftotheconqueroragainstthewhichamoietycompetentwasgagedbyourkingwhichhadreturndtotheinheritanceoffortinbrashadhebeenvanquisherasbythesamecovenantandcarriageofthearticledesigndhisfelltohamletnowsiryoungfortinbrasofunimprovedmettlehotandfullhathintheskirtsofnorwayhereandtheresharkdupalistoflawlessresolutesforfoodanddiettosomeenterprisethathathastomachintwhichisnootherasitdothwellappearuntoourstatebuttorecoverofusbystronghandandtermscompulsatorythoseforesaidlandssobyhisfatherlostandthisitakeitisthemainmotiveofourpreparationsthesourceofthisourwatchandthechiefheadofthisposthasteandromageinthelandbernardoithinkitbenootherbuteensowellmayitsortthatthisportentousfigurecomesarmedthroughourwatchsolikethekingthatwasandisthequestionofthesewarshoratioamoteitistotroublethemindseyeinthemosthighandpalmystateofromealittleerethemightiestjuliusfellthegravesstoodtenantlessandthesheeteddeaddidsqueakandgibberintheromanstreetsasstarswithtrainsoffireanddewsofblooddisastersinthesunandthemoiststaruponwhoseinfluenceneptunesempirestandswassickalmosttodoomsdaywitheclipseandeventhelikeprecurseoffierceeventsasharbingersprecedingstillthefatesandprologuetotheomencomingonhaveheavenandearthtogetherdemonstrateduntoourclimaturesandcountrymenbutsoftbeholdlowhereitcomesagainreenterghostillcrossitthoughitblastmestayillusionifthouhastanysoundoruseofvoicespeaktomeiftherebeanygoodthingtobedonethatmaytotheedoeasandgracetomespeaktomecockcrowsifthouartprivytothycountrysfatewhichhappilyforeknowingmayavoidospeakorifthouhastuphoardedinthylifeextortedtreasureinthewombofearthforwhichtheysayyouspiritsoftwalkindeathspeakofitstayandspeakstopitmarcellusmarcellusshallistrikeatitwithmypartisanhoratiodoifitwillnotstandbernardotisherehoratiotisheremarcellustisgoneexitghostwedoitwrongbeingsomajesticaltoofferittheshowofviolenceforitisastheairinvulnerableandourvainblowsmaliciousmockerybernardoitwasabouttospeakwhenthecockcrewhoratioandthenitstartedlikeaguiltythinguponafearfulsummonsihaveheardthecockthatisthetrumpettothemorndothwithhisloftyandshrillsoundingthroatawakethegodofdayandathiswarningwhetherinseaorfireinearthorairtheextravagantanderringspirithiestohisconfineandofthetruthhereinthispresentobjectmadeprobationmarcellusitfadedonthecrowingofthecocksomesaythatevergainstthatseasoncomeswhereinoursavioursbirthiscelebratedthebirdofdawningsingethallnightlongandthentheysaynospiritdaresstirabroadthenightsarewholesomethennoplanetsstrikenofairytakesnorwitchhathpowertocharmsohallowdandsograciousisthetimehoratiosohaveiheardanddoinpartbelieveitbutlookthemorninrussetmantlecladwalksoerthedewofyonhigheastwardhillbreakweourwatchupandbymyadviceletusimpartwhatwehaveseentonightuntoyounghamletforuponmylifethisspiritdumbtouswillspeaktohimdoyouconsentweshallacquainthimwithitasneedfulinourlovesfittingourdutymarcellusletsdotiprayandithismorningknowwhereweshallfindhimmostconveniently’; number = double(text)-96; histo = zeros(1,26); for j = 1:length(number) histo(number (j)) = histo(number (j)) + 1; end plot(histo) % Pattern Generation counter = zeros(1,26); summ = sum(counter); i = 1; j = 1; while(summ < 26) sy = number(i); if (counter(sy) == 0) counter(sy) = 1; pattern(j) = sy; j = j + 1; end summ = sum(counter); i = i + 1; end |
| function histo = hamlet_pat(sinyal) h1 = zeros(1512); h2 = zeros(1256); h3 = h1; for i = 1:length(sinyal)-26 blok = sinyal(i:i + 26); m = blok(14); deger(1:13) = blok(1:13); deger(14:26) = blok(15:27); for j = 1:26 bit(j) = deger(j) >= m; end b1(1:9) = bit(1:9); b2(1:8) = bit(10:17); b3(1:9) = bit(18:26); m1(i) = 0; m2(i) = 0; m3(i) = 0; for j = 1:9 m1(i) = m1(i) + b1(j)*2^(j-1); m3(i) = m3(i) + b3(j)*2^(j-1); end for j = 1:8 m2(i) = m2(i) + b2(j)*2^(j-1); end h1(m1(i) + 1) = h1(m1(i) + 1) + 1; h2(m2(i) + 1) = h2(m2(i) + 1) + 1; h3(m3(i) + 1) = h3(m3(i) + 1) + 1; end histo = [h1 h2 h3]; |
References
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [Green Version]
- Su, S.; Wong, G.; Shi, W.; Liu, J.; Lai, A.C.; Zhou, J.; Liu, W.; Bi, Y.; Gao, G.F. Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol. 2016, 24, 490–502. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khorramdelazad, H.; Kazemi, M.H.; Najafi, A.; Keykhaee, M.; Emameh, R.Z.; Falak, R. Immunopathological similarities between COVID-19 and influenza: Investigating the consequences of Co-infection. Microb. Pathog. 2021, 152, 104554. [Google Scholar]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; Meng, J.; Zhu, Z.; Zhang, Z.; Wang, J. Genome composition and divergence of the novel coronavirus (2019-nCoV) originating in China. Cell Host Microbe 2020, 27, 325–328. [Google Scholar] [CrossRef] [Green Version]
- Ge, X.-Y.; Li, J.-L.; Yang, X.-L.; Chmura, A.A.; Zhu, G.; Epstein, J.H.; Mazet, J.K.; Hu, B.; Zhang, W.; Peng, C. Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor. Nature 2013, 503, 535–538. [Google Scholar] [CrossRef] [Green Version]
- He, X.; Yang, X.; Zhang, S.; Zhao, J.; Zhang, Y.; Xing, E.; Xie, P. Sample-efficient deep learning for COVID-19 diagnosis based on CT scans. medRxiv 2006. [Google Scholar] [CrossRef]
- Li, F. Structure, function, and evolution of coronavirus spike proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef] [Green Version]
- Afify, H.M.; Zanaty, M.S. A Comparative Study of Protein Sequences Classification-Based Machine Learning Methods for COVID-19 Virus against HIV-1. Appl. Artif. Intell. 2021, 35, 1733–1745. [Google Scholar] [CrossRef]
- Long, J.S.; Mistry, B.; Haslam, S.M.; Barclay, W.S. Host and viral determinants of influenza A virus species specificity. Nat. Rev. Microbiol. 2019, 17, 67–81. [Google Scholar] [CrossRef]
- Vasin, A.; Temkina, O.; Egorov, V.; Klotchenko, S.; Plotnikova, M.; Kiselev, O. Molecular mechanisms enhancing the proteome of influenza A viruses: An overview of recently discovered proteins. Virus Res. 2014, 185, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Kumlin, U.; Olofsson, S.; Dimock, K.; Arnberg, N. Sialic acid tissue distribution and influenza virus tropism. Influenza Other Respir. Viruses 2008, 2, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Robson, B. Bioinformatics studies on a function of the SARS-CoV-2 spike glycoprotein as the binding of host sialic acid glycans. Comput. Biol. Med. 2020, 122, 103849. [Google Scholar] [CrossRef] [PubMed]
- Jones, T.C.; Mühlemann, B.; Veith, T.; Biele, G.; Zuchowski, M.; Hofmann, J.; Stein, A.; Edelmann, A.; Corman, V.M.; Drosten, C. An analysis of SARS-CoV-2 viral load by patient age. medRxiv 2012. [Google Scholar] [CrossRef]
- Li, D.; Wang, D.; Dong, J.; Wang, N.; Huang, H.; Xu, H.; Xia, C. False-negative results of real-time reverse-transcriptase polymerase chain reaction for severe acute respiratory syndrome coronavirus 2: Role of deep-learning-based CT diagnosis and insights from two cases. Korean J. Radiol. 2020, 21, 505–508. [Google Scholar] [CrossRef]
- Baygin, M.; Yaman, O.; Barua, P.D.; Dogan, S.; Tuncer, T.; Acharya, U.R. Exemplar Darknet19 feature generation technique for automated kidney stone detection with coronal CT images. Artif. Intell. Med. 2022, 127, 102274. [Google Scholar]
- Barua, P.D.; Dogan, S.; Tuncer, T.; Baygin, M.; Acharya, U.R. Novel automated PD detection system using aspirin pattern with EEG signals. Comput. Biol. Med. 2021, 137, 104841. [Google Scholar]
- Kobat, M.A.; Kivrak, T.; Barua, P.D.; Tuncer, T.; Dogan, S.; Tan, R.-S.; Ciaccio, E.J.; Acharya, U.R. Automated COVID-19 and Heart Failure Detection Using DNA Pattern Technique with Cough Sounds. Diagnostics 2021, 11, 1962. [Google Scholar] [CrossRef]
- Dong, G.; Liu, H. Feature Engineering for Machine Learning and Data Analytics; CRC Press: New York, NY, USA, 2018. [Google Scholar]
- Zheng, A.; Casari, A. Feature Engineering for Machine Learning: Principles and Techniques for Data Scientists; O’Reilly Media, Inc.: Sebastopol, CA, USA, 2018. [Google Scholar]
- Ojala, T.; Pietikainen, M.; Maenpaa, T. Multiresolution gray-scale and rotation invariant texture classification with local binary patterns. IEEE Trans. Pattern Anal. Mach. Intell. 2002, 24, 971–987. [Google Scholar] [CrossRef]
- NCBI Virus. 2022. Available online: https://www.ncbi.nlm.nih.gov (accessed on 3 January 2022).
- Shakespeare, W.; Scene, I. Elsinore. A Platform before the Castle. Available online: https://shakespeare.mit.edu/hamlet/hamlet.1.1.html (accessed on 3 January 2022).
- Baygin, M.; Yaman, O.; Tuncer, T.; Dogan, S.; Barua, P.D.; Acharya, U.R. Automated accurate schizophrenia detection system using Collatz pattern technique with EEG signals. Biomed. Signal Process. Control 2021, 70, 102936. [Google Scholar] [CrossRef]
- Vapnik, V. The support vector method of function estimation. In Nonlinear Modeling; Springer: New York, NY, USA, 1998; pp. 55–85. [Google Scholar]
- Vapnik, V. The Nature of Statistical Learning Theory; Springer Science & Business Media: New York, NY, USA, 2013. [Google Scholar]
- Liu, H.; Setiono, R. Chi2: Feature selection and discretization of numeric attributes. In Proceedings of the 7th IEEE International Conference on Tools with Artificial Intelligence, Herndon, VA, USA, 5–8 November 1995; pp. 388–391. [Google Scholar]
- Powers, D.M. Evaluation: From precision, recall and F-measure to ROC, informedness, markedness and correlation. arXiv 2020, arXiv:2010.16061. [Google Scholar]
- Warrens, M.J. On the equivalence of Cohen’s kappa and the Hubert-Arabie adjusted Rand index. J. Classif. 2008, 25, 177–183. [Google Scholar] [CrossRef] [Green Version]
- Taubenberger, J.K.; Kash, J.C.; Morens, D.M. The 1918 influenza pandemic: 100 years of questions answered and unanswered. Sci. Transl. Med. 2019, 11, eaau5485. [Google Scholar] [CrossRef] [PubMed]
- Jester, B.; Uyeki, T.; Jernigan, D. Readiness for responding to a severe pandemic 100 years after 1918. Am. J. Epidemiol. 2018, 187, 2596. [Google Scholar] [CrossRef] [PubMed]
- Solomon, D.A.; Sherman, A.C.; Kanjilal, S. Influenza in the COVID-19 Era. JAMA 2020, 324, 1342–1343. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.M.; Iqbal, T. Hamlet: A hierarchical multimodal attention-based human activity recognition algorithm. In Proceedings of the 2020 IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS), Las Vegas, NV, USA, 25–29 October 2020; pp. 10285–10292. [Google Scholar]
- Ren, F.; Zhang, Z.; Yan, Y.; Wang, Z.; Su, S.; Philip, S.Y. HAMLET: Hierarchical Attention-based Model with muLti-task sElf-Training for user profiling. In Proceedings of the 2021 IEEE International Conference on Big Data (Big Data), Orlando, FL, USA, 15–18 December 2021; pp. 500–509. [Google Scholar]
- Loh, H.W.; Ooi, C.P.; Seoni, S.; Barua, P.D.; Molinari, F.; Acharya, U.R. Application of Explainable Artificial Intelligence for Healthcare: A Systematic Review of the Last Decade (2011–2022). Comput. Methods Programs Biomed. 2022, 226, 107161. [Google Scholar] [CrossRef] [PubMed]

| id | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ind. | 2 | 5 | 18 | 14 | 1 | 4 | 15 | 23 | 8 | 19 | 20 | 6 | 3 |
| id | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | 23 | 24 | 25 | 26 |
| Ind. | 9 | 25 | 13 | 21 | 12 | 7 | 22 | 11 | 16 | 17 | 24 | 10 | 26 |
| Metric | Cross Validation | SARS-CoV-2 | Influenza-A |
|---|---|---|---|
| Sensitivity (%) | 5-fold CV | 99.95 | 99.79 |
| 75:25 | 100 | 99.86 | |
| Specificity (%) | 5-fold CV | 99.79 | 99.95 |
| 75:25 | 99.86 | 100 | |
| Precision (%) | 5-fold CV | 99.76 | 99.96 |
| 75:25 | 99.83 | 100 | |
| F1-score (%) | 5-fold CV | 99.86 | 99.87 |
| 75:25 | 99.92 | 99.93 | |
| Overall accuracy (%) | 5-fold CV | 99.87 | |
| 75:25 | 99.92 | ||
| Overall geometric mean (%) | 5-fold CV | 99.87 | |
| 75:25 | 99.93 | ||
| Model | Dataset | Number of Observations | Method | Result |
|---|---|---|---|---|
| Afify and Zanaty [9] | NCBI | 18,476 protein sequences: 9238 COVID-19 9238 HIV | Conjoint triad feature extraction and Random Forest classification with hold-out validation (80:20) | Accuracy: 99.80% |
| Our model | NCBI | 36,424 protein sequences: 16,901 COVID-19 19,523 Influenza-A | HamletPat feature extraction, IChi2 feature selection, and SVM classification with hold-out validation (75:25) and 5-fold CV | Accuracy: hold-out: 99.92% 5-fold CV: 99.87% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Erten, M.; Acharya, M.R.; Kamath, A.P.; Sampathila, N.; Bairy, G.M.; Aydemir, E.; Barua, P.D.; Baygin, M.; Tuncer, I.; Dogan, S.; et al. Hamlet-Pattern-Based Automated COVID-19 and Influenza Detection Model Using Protein Sequences. Diagnostics 2022, 12, 3181. https://doi.org/10.3390/diagnostics12123181
Erten M, Acharya MR, Kamath AP, Sampathila N, Bairy GM, Aydemir E, Barua PD, Baygin M, Tuncer I, Dogan S, et al. Hamlet-Pattern-Based Automated COVID-19 and Influenza Detection Model Using Protein Sequences. Diagnostics. 2022; 12(12):3181. https://doi.org/10.3390/diagnostics12123181
Chicago/Turabian StyleErten, Mehmet, Madhav R. Acharya, Aditya P. Kamath, Niranjana Sampathila, G. Muralidhar Bairy, Emrah Aydemir, Prabal Datta Barua, Mehmet Baygin, Ilknur Tuncer, Sengul Dogan, and et al. 2022. "Hamlet-Pattern-Based Automated COVID-19 and Influenza Detection Model Using Protein Sequences" Diagnostics 12, no. 12: 3181. https://doi.org/10.3390/diagnostics12123181









