SD-UNet: Stripping down U-Net for Segmentation of Biomedical Images on Platforms with Low Computational Budgets
Abstract
1. Introduction
1.1. Motivation
1.2. Contributions
- We propose the use of depthwise separable convolution layers to replace all standard CNN layers except the first CNN layer in the original U-Net model
- Depthwise separable convolution layers are known to achieve lower performance compared to standard convolution layers. We demonstrate that performance drop due to the process can be recovered with a method of weight standardization and group normalization.
- SD-UNet model has 8x fewer parameters and requires 23x less storage space. The computational complexity or number of floating point operations (FLOPs) required by SD-UNet is 8x less than is required by the original U-Net model and shows great performance on the segmentation of biomedical images.
2. Related Work
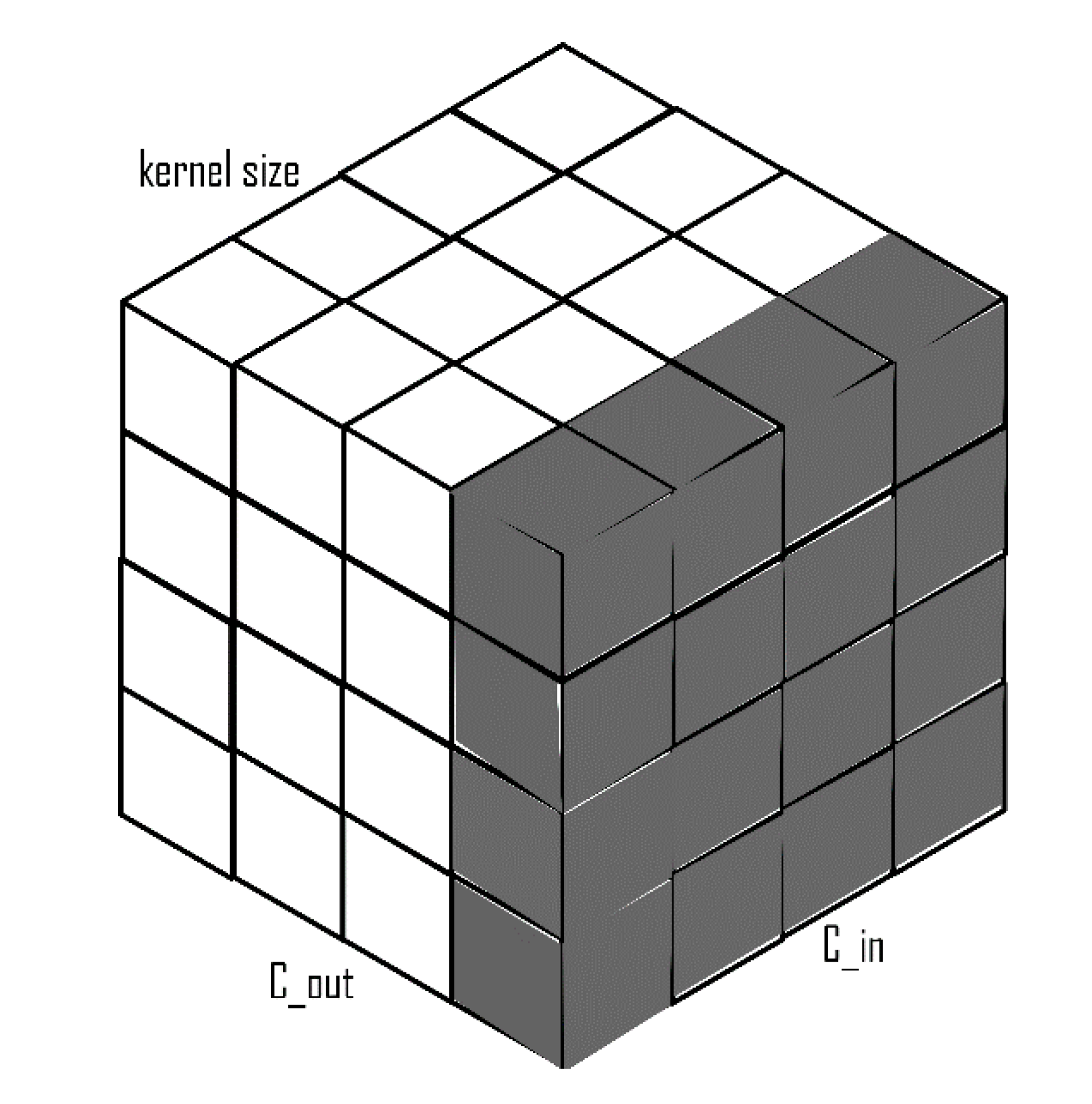
2.1. Depthwise Separable Convolutions
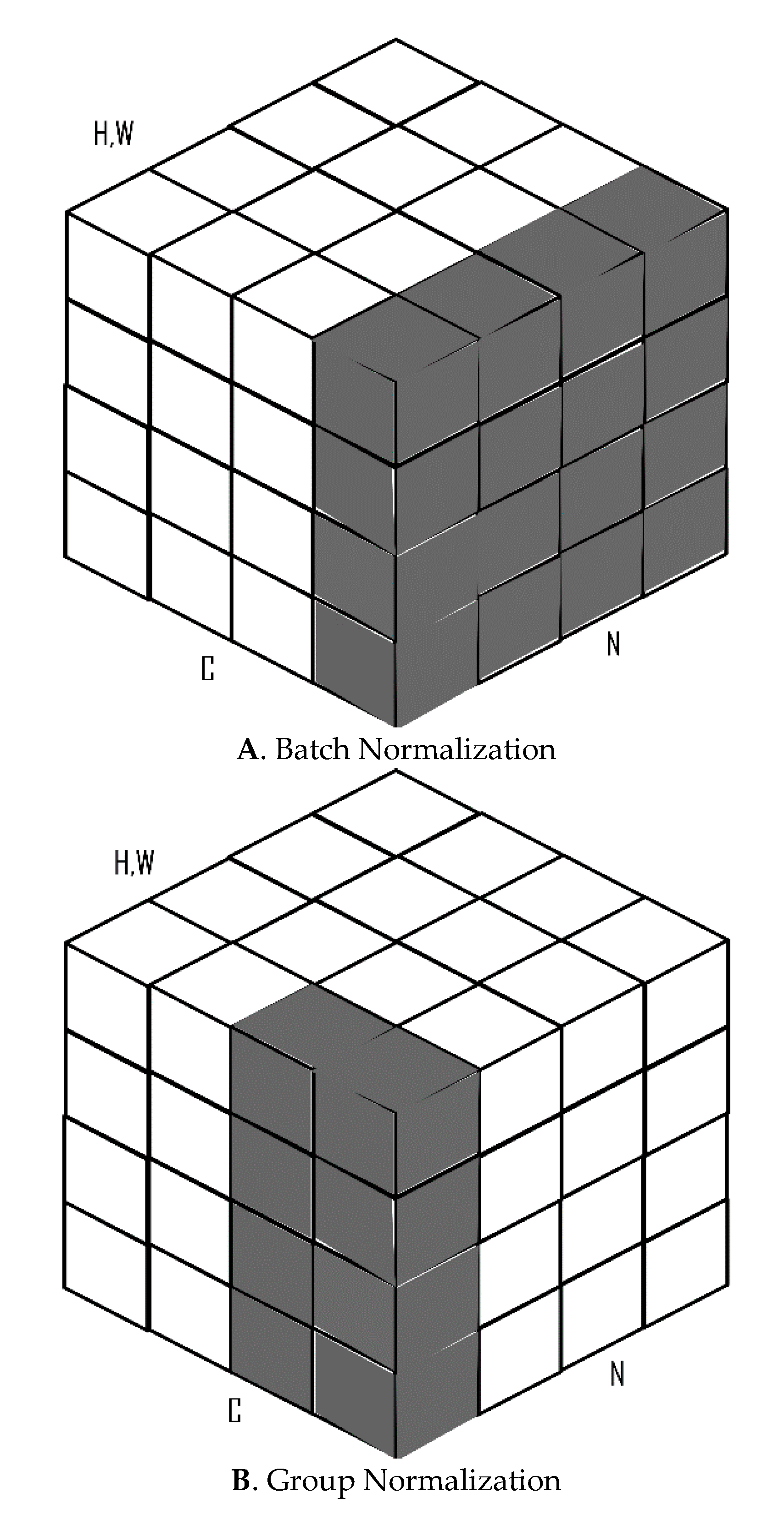
2.2. Batch and Group Normalization
2.3. Weight Standardization
2.4. Fully Convolutional Networks (FCNs)
- Downsampling/Contraction/Encoding Path: On this path, the model extracts and interprets the contextual information on the input image.
- Upsampling/Expanding/Decoding Path: The specific localization or construction of segmentation maps from the extracted context in the encoding path.
- Skip Connections/Bottlenecks: Combines information from encoding and decoding paths by summing feature maps
2.5. U-Net
3. Materials and Methods
3.1. WS with Depthwise Separable Convolutions
3.2. SD-UNet (Proposed Architecture)
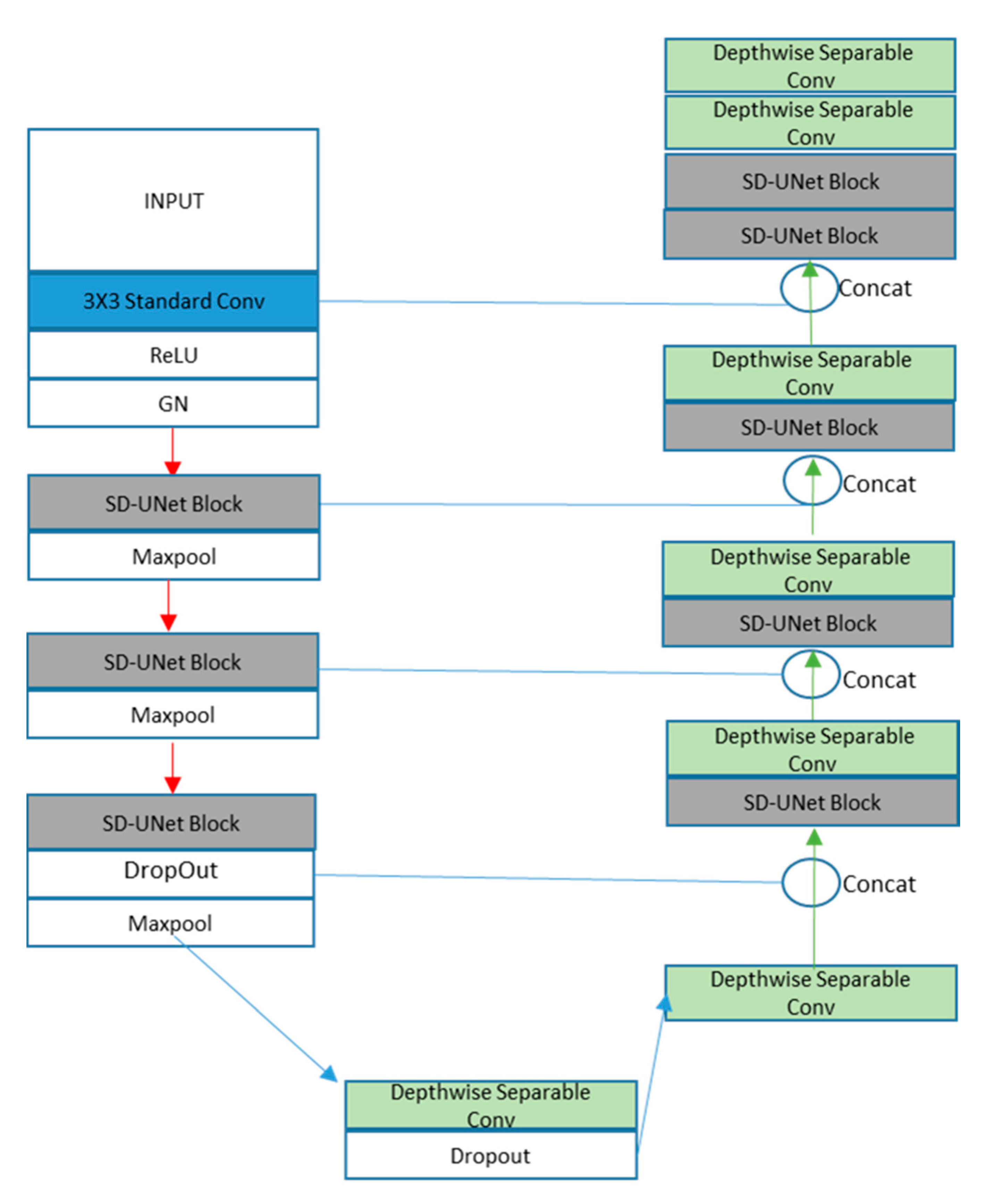
- Block1: A standard convolution layer, a ReLu activation function, and a GN layer
- Block2 and Block3: One SD-UNet block and a max-pooling layer. An SD-UNet block is made up of two depthwise separable convolution layers, two activation layers, and one GN layer (Figure 5).
- Block4: One SD-UNet block, a dropout layer to introduce regularization [39], and a max-pooling layer. All depthwise (3 × 3) convolution layers are weight standardized.
- Block5: A final depthwise separable layer with a dropout layer.
- Block1: A depthwise separable convolution layer with its features concatenated with the dropout layer from Block4 of the encoding path.
- Blocks 2, 3, 4: An SD-UNet block and a depthwise separable layer concatenated with corresponding blocks from the encoding path
- Block 5: Two SD-UNet blocks and two depthwise separable layers with the last one as the final prediction layer (Figure 6).
3.3. Setup
3.4. Datasets
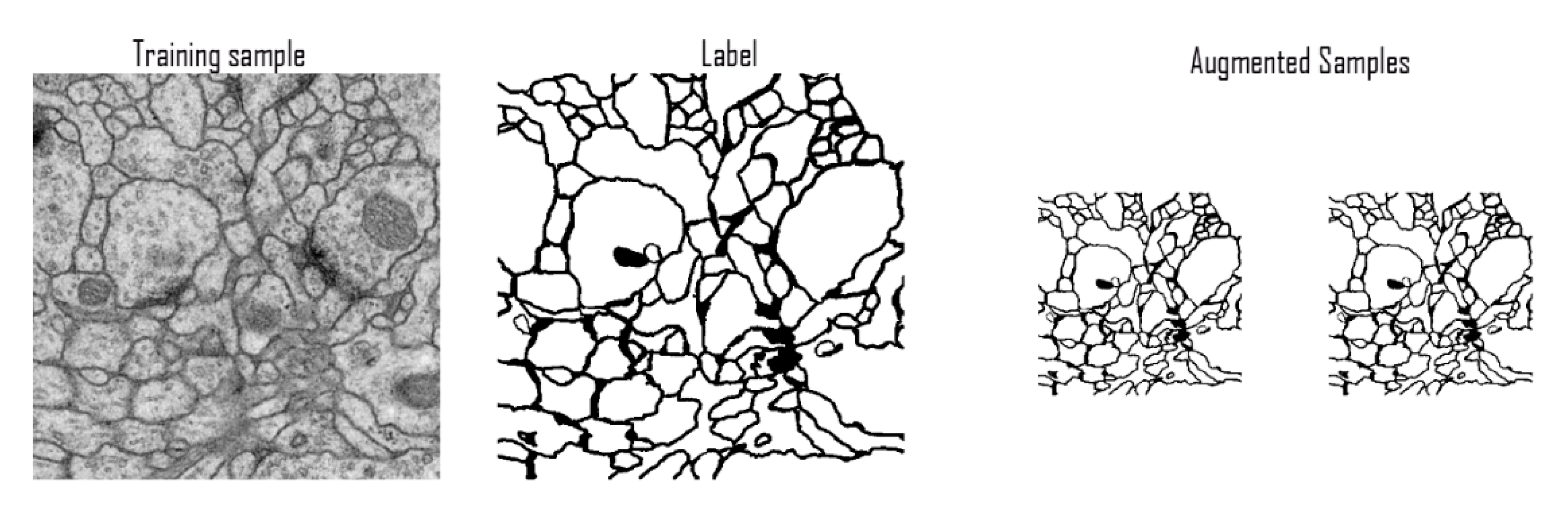
3.4.1. ISBI Challenge Dataset
3.4.2. MSD Challenge Brain Tumor Segmentation (BRATs) Dataset
3.4.3. Data Pre-Processing
3.5. Optimization
3.6. Performance Metrics
3.6.1. Accuracy (AC)
3.6.2. Intersection over Union (IOU)
3.6.3. Sorensen-Dice Co-Efficient (Dice Co-Eff)
3.6.4. Maximal Foreground-Restricted Rand Score (VRand)
3.6.5. Maximal Foreground-Restricted Information Theoretic Score (VInfo)
3.6.6. Floating Point Operations Per Second (FLOPs)
4. Results
4.1. Ablation Study
- U-Net (GN = 32)—Original U-Net architecture with GN only with 32 groups.
- U-Net (depthwise + BN)—U-Net architecture with depthwise separable layers replacing standard convolution layers and BN layers only
- U-Net (depthwise + GN)—U-Net architecture with depthwise separable layers replacing standard convolution layers and GN layers only
- SD-UNet (depthwise + BN + WS)—Proposed SD-UNet based on BN, WS, and depthwise separable convolutions
4.2. Computational Results
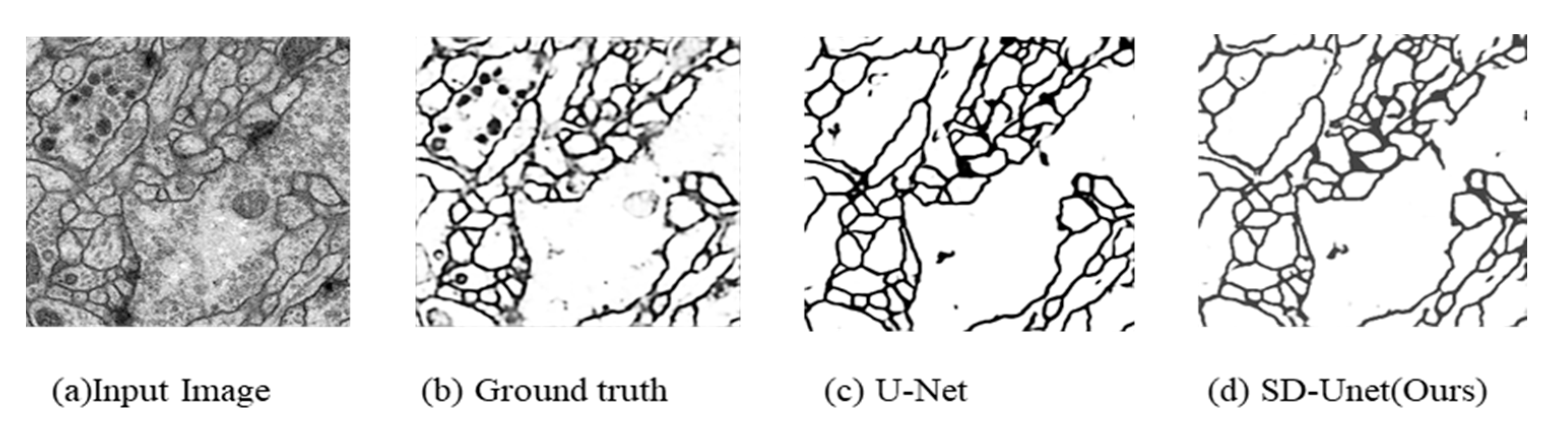
4.3. Results on ISBI Challenge Dataset
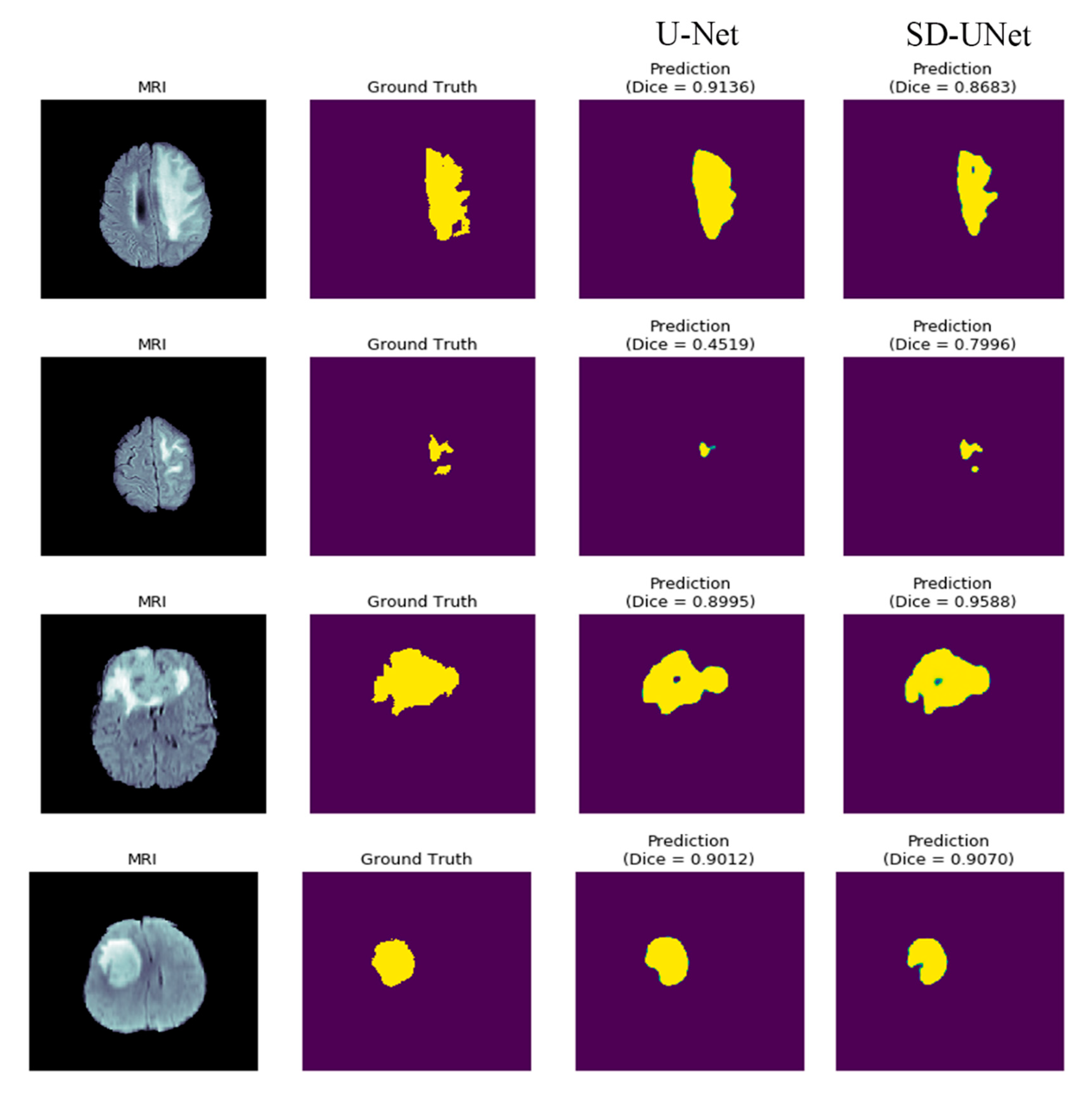
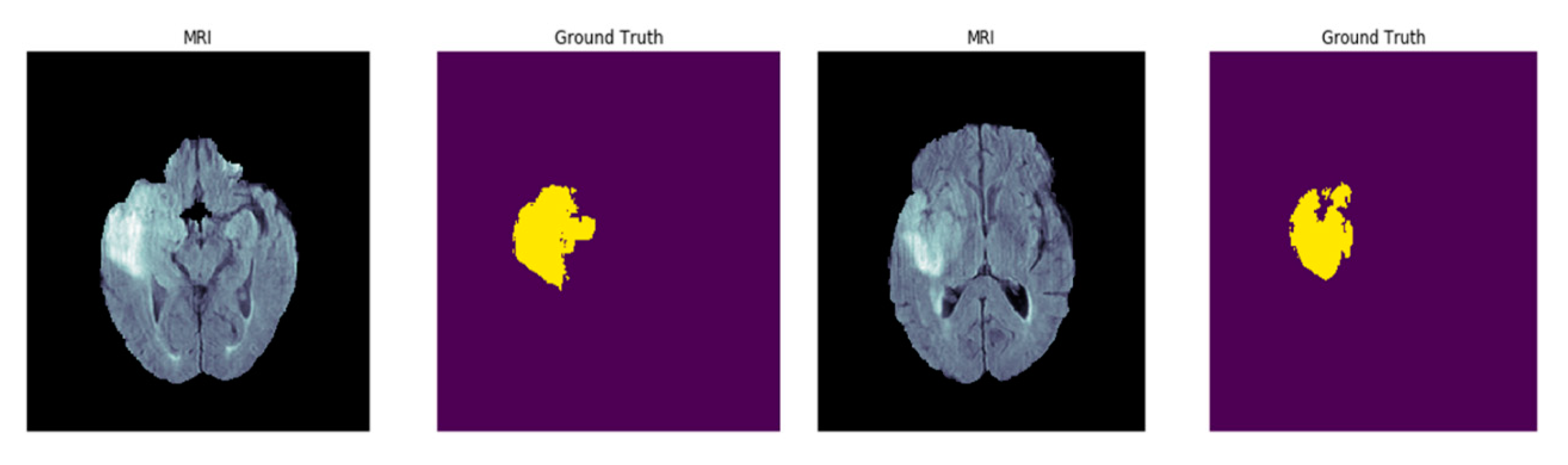
4.4. Results on BRATs Dataset
5. Discussion and Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Wismüller, A.; Vietze, F.; Behrends, J.; Meyer-Baese, A.; Reiser, M.; Ritter, H. Fully automated biomedical image segmentation by self-organized model adaptation. Neural Netw. 2004, 17, 1327–1344. [Google Scholar] [CrossRef]
- Chen, C.; Ozolek, J.A.; Wang, W.; Rohde, G. A General System for Automatic Biomedical Image Segmentation Using Intensity Neighborhoods. Int. J. Biomed. Imaging 2011, 2011, 1–12. [Google Scholar] [CrossRef]
- Aganj, I.; Harisinghani, M.G.; Weissleder, R.; Fischl, B. Unsupervised Medical Image Segmentation Based on the Local Center of Mass. Sci. Rep. 2018, 8, 13012. [Google Scholar] [CrossRef]
- Hesamian, M.H.; Jia, W.; He, X.; Kennedy, P. Deep Learning Techniques for Medical Image Segmentation: Achievements and Challenges. J. Digit. Imaging 2019, 32, 582–596. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Adv. Neural Inf. Process. 2012, 1097–11059. [Google Scholar] [CrossRef]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015; pp. 1–9. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. arXiv 2014, arXiv:1409.1556, 1–14. [Google Scholar]
- Davis, J.W.; Sharma, V. Simultaneous detection and segmentation of pedestrians using top-down and bottom-up processing. In Proceedings of the 2007 IEEE Conference on Computer Vision and Pattern Recognition Processing, Minneapolis, MN, USA, 17–22 June 2007; pp. 1–8. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. Form. Asp. Compon. Softw. 2015, 9351, 234–241. [Google Scholar]
- Shelhamer, E.; Long, J.; Darrell, T. Fully Convolutional Networks for Semantic Segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 640–651. [Google Scholar] [CrossRef]
- Badrinarayanan, V.; Badrinarayanan, V.; Cipolla, R. SegNet: A Deep Convolutional Encoder-Decoder Architecture for Image Segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef]
- Hu, R.; Dollar, P.; He, K.; Darrell, T.; Girshick, R. Learning to Segment Every Thing 2017. Available online: http://openaccess.thecvf.com/content_cvpr_2018/papers/Hu_Learning_to_Segment_CVPR_2018_paper.pdf (accessed on 18 February 2020).
- Simonyan, K.; Zisserman, A. Two-Stream Convolutional Networks for Action Recognition in Videos. Available online: http://papers.nips.cc/paper/5353-two-stream-convolutional (accessed on 18 February 2020).
- Ehsani, K.; Bagherinezhad, H.; Redmon, J.; Mottaghi, R.; Farhadi, A. Who Let the Dogs Out? Modeling Dog Behavior from Visual Data. In Proceedings of the 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition, Salt Lake City, AL, USA, 18–22 June 2018; pp. 4051–4060. [Google Scholar]
- Iqbal, U.; Milan, A.; Gall, J. PoseTrack: Joint Multi-Person Pose Estimation and Tracking. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Colorado Springs, CO, USA, 20–25 June 2011; pp. 2011–2020. [Google Scholar]
- Kehl, W.; Tombari, F.; Ilic, S.; Navab, N. Real-Time 3D Model Tracking in Color and Depth on a Single CPU Core. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), City of Honolulu, HI, USA, 21–26 July 2017; pp. 465–473. [Google Scholar]
- Girshick, R.; Donahue, J.; Darrell, T.; Malik, J.; Malik, J. Rich Feature Hierarchies for Accurate Object Detection and Semantic Segmentation. In Proceedings of the 2014 IEEE Conference on Computer Vision and Pattern Recognition, Washington, DC, USA, 24–27 June 2014; pp. 580–587. [Google Scholar]
- Li, J.; Wu, Y.; Zhao, J.; Guan, L.; Ye, C.; Yang, T. Pedestrian detection with dilated convolution, region proposal network and boosted decision trees. In Proceedings of the 2017 International Joint Conference on Neural Networks (IJCNN), Anchorage, AK, USA, 14–19 May 2017; pp. 4052–4057. [Google Scholar]
- Sun, W.; Wang, R. Fully Convolutional Networks for Semantic Segmentation of Very High Resolution Remotely Sensed Images Combined With DSM. IEEE Geosci. Remote. Sens. Lett. 2018, 15, 474–478. [Google Scholar] [CrossRef]
- Roth, H.R.; Shen, C.; Oda, H.; Oda, M.; Hayashi, Y.; Misawa, K.; Mori, K. Deep learning and its application to medical image segmentation. Med Imaging Technol. 2018, 36, 63–71. [Google Scholar]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; Van Der Laak, J.A.; Van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- Yao, W.; Zeng, Z.; Lian, C.; Tang, H. Pixel-wise regression using U-Net and its application on pansharpening. Neurocomputing 2018, 312, 364–371. [Google Scholar] [CrossRef]
- Iglovikov, V.; Shvets, A. TernausNet: U-Net with VGG11 Encoder Pre-Trained on ImageNet for Image Segmentation 2018. arXiv 2018, arXiv:1801.05746. [Google Scholar]
- Russakovsky, O.; Deng, J.; Su, H.; Krause, J.; Satheesh, S.; Ma, S.; Huang, Z.; Karpathy, A.; Khosla, A.; Bernstein, M.; et al. ImageNet Large Scale Visual Recognition Challenge. Int. J. Comput. Vis. 2015, 115, 211–252. [Google Scholar] [CrossRef]
- Oktay, O.; Schlemper, J.; Le Folgoc, L.; Lee, M.C.H.; Heinrich, M.; Misawa, K.; Mori, K.; McDonagh, S.; Hammerla, N.Y.; Kainz, B.; et al. Attention U-Net: Learning Where to Look for the Pancreas 2018. Available online: https://arxiv.org/abs/1804.03999 (accessed on 18 February 2020).
- Denil, M.; Shakibi, B.; Dinh, L.; Ranzato, M.; de Freitas, N. Predicting Parameters in Deep Learning. Available online: https://papers.nips.cc/paper/5025-predicting-parameters-in-deep-learning.pdf (accessed on 18 February 2020).
- LeCun, Y.; Denker, J.S.; Solla, S.A. Optimal Brain Damage. Adv. Neural Inf. Process. Syst. 1990, 2, 598–605. [Google Scholar]
- Hassibi, B.; Stork, D.G. Second Order Derivaties for Network Prunning: Optimal Brain Surgeon. Available online: https://authors.library.caltech.edu/54983/3/647-second-order-derivatives-for-network-pruning-optimal-brain-surgeon(1).pdf (accessed on 15 February 2020).
- Alvarez, J.M.; Salzmann, M. Compression-aware Training of Deep Networks 2017. Available online: http://papers.nips.cc/paper/6687-compression-aware-training-of-deep-networks (accessed on 17 February 2020).
- Han, S.; Mao, H.; Dally, W.J. Deep Compression: Compressing Deep Neural Networks with Pruning, Trained Quantization, and Huffman Coding. Available online: https://arxiv.org/abs/1510.00149 (accessed on 17 February 2020).
- Howard, A.G.; Zhu, M.; Chen, B.; Kalenichenko, D.; Wang, W.; Weyand, T.; Andreetto, M.; Adam, H. MobileNets: Efficient Convolutional Neural Networks for Mobile Vision Applications. Available online: https://arxiv.org/abs/1704.04861 (accessed on 18 February 2020).
- Zhang, X.; Zhou, X.; Lin, M.; Sun, J. ShuffleNet: An Extremely Efficient Convolutional Neural Network for Mobile Devices. In Proceedings of the 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition 2018, Salt Lake City, AL, USA, 18–22 June 2018; pp. 6848–6856. [Google Scholar]
- Qin, Z.; Zhang, Z.; Chen, X.; Peng, Y. FD-MobileNet: Improved MobileNet with a Fast Downsampling Strategy. 2018. Available online: https://ieeexplore.ieee.org/abstract/document/8451355 (accessed on 17 February 2020).
- Sifre, L. Rigid-Motion Scattering for Image Classification. Ph.D. Thesis, Ecole Polytechnique, Palaiseau, France, 2014. CMAP Rigid-Motion Scattering For Image Classification. Available online: https://www.di.ens.fr/data/publications/papers/phd_sifre.pdf (accessed on 17 February 2020).
- Chollet, F. Xception: Deep Learning with Depthwise Separable Convolutions. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26July 2017; pp. 1800–1807. [Google Scholar]
- Ioffe, S.; Szegedy, C. Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift. Available online: https://arxiv.org/abs/1502.03167 (accessed on 17 February 2020).
- Wu, Y.; He, K. Group Normalization. Formal Asp. Compon. Softw. 2018, 11217 LNCS, 3–19. [Google Scholar]
- Qiao, S.; Wang, H.; Liu, C.; Shen, W.; Yuille, A. Weight Standardization. Available online: https://arxiv.org/abs/1903.10520 (accessed on 18 February 2020).
- Srivastava, N.; Hinton, A.; Sutskever, K.I.; Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 2017, 15, 1929–1958. [Google Scholar]
- Arganda-Carreras, I.; Turaga, S.C.; Berger, D.R.; Cireşan, D.; Giusti, A.; Gambardella, L.M.; Schmidhuber, J.; Laptev, D.; Dwivedi, S.; Buhmann, J.M.; et al. Crowdsourcing the creation of image segmentation algorithms for connectomics. Front. Neuroanat. 2015, 9, 898. [Google Scholar] [CrossRef]
- Cardona, A.; Saalfeld, S.; Preibisch, S.; Schmid, B.; Cheng, A.; Pulokas, J.; Tomancak, P.; Hartenstein, V. An integrated micro- and macroarchitectural analysis of the Drosophila brain by computer-assisted serial section electron microscopy. PLoS Boil. 2010, 8, e1000502. [Google Scholar] [CrossRef] [PubMed]
- Simpson, A.; Antonelli, M.; Bakas, S.; Bilello, M.; Farahani, K.; Van Ginneken, B.; Kopp-Schneider, A.; Landman, B.A.; Litjens, G.; Menze, B.; et al. A Large Annotated Medical Image Dataset for the Development and Evaluation of Segmentation Algorithms. Available online: https://arxiv.org/abs/1902.09063 (accessed on 18 February 2020).
- Bakas, S.; Reyes, M.; Jakab, A.; Bauer, S.; Rempfler, M.; Cromo, A.; Shinohara, R.T.; Berger, C.; Ha, S.M.; Rozycki, M.; et al. Identifying the Best Machine Learning Algorithms for Brain Tumor Segmentation, Progression Assessment, and Overall Survival Prediction in the BRATS Challenge. Available online: https://arxiv.org/abs/1811.02629 (accessed on 17 February 2020).
- Bakas, S.; Akbari, H.; Sotiras, A.; Bilello, M.; Rozycki, M.; Kirby, J.; Freymann, J.B.; Farahani, K.; Davatzikos, C. Advancing The Cancer Genome Atlas glioma MRI collections with expert segmentation labels and radiomic features. Sci. Data 2017, 4, 170117. [Google Scholar] [CrossRef]
- Menze, B.H.; Jakab, A.; Bauer, S.; Kalpathy-Cramer, J.; Farahani, K.; Kirby, J.; Burren, Y.; Porz, N.; Slotboom, J.; Wiest, R.; et al. The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans. Med Imaging 2015, 34, 1993–2024. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. Available online: https://arxiv.org/abs/1412.6980 (accessed on 17 February 2020).






| Model | # Params | # Flops | Size in Memory | Inference (ms) |
|---|---|---|---|---|
| U-Net | 31M | 62.04M | 372.5MB | 188 |
| U-Net (GN = 32) | 26M | 51.9M | 311.7MB | 283 |
| U-Net (depthwise + BN) | 3.9M | 7.8M | 47.0MB | 94 |
| U-Net (depthwise + GN = 32) | 3.9M | 7.7M | 47.1MB | 87 |
| SD-UNet (depthwise +BN + WS) | 3.9M | 7.8M | 15.8MB | 99 |
| SD-UNet (Proposed) | 3.9M | 7.8M | 15.8MB | 107 |
| Model | Loss | Accuracy | Mean IOU | Dice Co-Eff |
|---|---|---|---|---|
| U-Net | 0.0533 | 97.67 | 87.35 | 98.51 |
| U-Net (GN = 32) | 0.0439 | 98.08 | 88.67 | 98.77 |
| U-Net (depthwise + BN ) | 0.0435 | 93.62 | 74.62 | 95.91 |
| U-Net (depthwise + GN = 32) | 0.1393 | 93.94 | 76.54 | 96.13 |
| SD-UNet (depthwise + BN + WS) | 0.1065 | 96.36 | 82.10 | 97.67 |
| SD-UNet (Proposed) | 0.0775 | 96.73 | 83.26 | 97.84 |
| Model | Training Loss | Test Accuracy | Dice Co-Eff | Inference(ms) |
|---|---|---|---|---|
| U-Net | 0.5601 | 98.71 | 80.30 | 91 |
| SD-UNet (Proposed) | 0.0666 | 98.66 | 82.75 | 56 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gadosey, P.K.; Li, Y.; Agyekum, E.A.; Zhang, T.; Liu, Z.; Yamak, P.T.; Essaf, F. SD-UNet: Stripping down U-Net for Segmentation of Biomedical Images on Platforms with Low Computational Budgets. Diagnostics 2020, 10, 110. https://doi.org/10.3390/diagnostics10020110
Gadosey PK, Li Y, Agyekum EA, Zhang T, Liu Z, Yamak PT, Essaf F. SD-UNet: Stripping down U-Net for Segmentation of Biomedical Images on Platforms with Low Computational Budgets. Diagnostics. 2020; 10(2):110. https://doi.org/10.3390/diagnostics10020110
Chicago/Turabian StyleGadosey, Pius Kwao, Yujian Li, Enock Adjei Agyekum, Ting Zhang, Zhaoying Liu, Peter T. Yamak, and Firdaous Essaf. 2020. "SD-UNet: Stripping down U-Net for Segmentation of Biomedical Images on Platforms with Low Computational Budgets" Diagnostics 10, no. 2: 110. https://doi.org/10.3390/diagnostics10020110
APA StyleGadosey, P. K., Li, Y., Agyekum, E. A., Zhang, T., Liu, Z., Yamak, P. T., & Essaf, F. (2020). SD-UNet: Stripping down U-Net for Segmentation of Biomedical Images on Platforms with Low Computational Budgets. Diagnostics, 10(2), 110. https://doi.org/10.3390/diagnostics10020110





