Host-Microbe Interactions in Microgravity: Assessment and Implications
Abstract
:1. Introduction
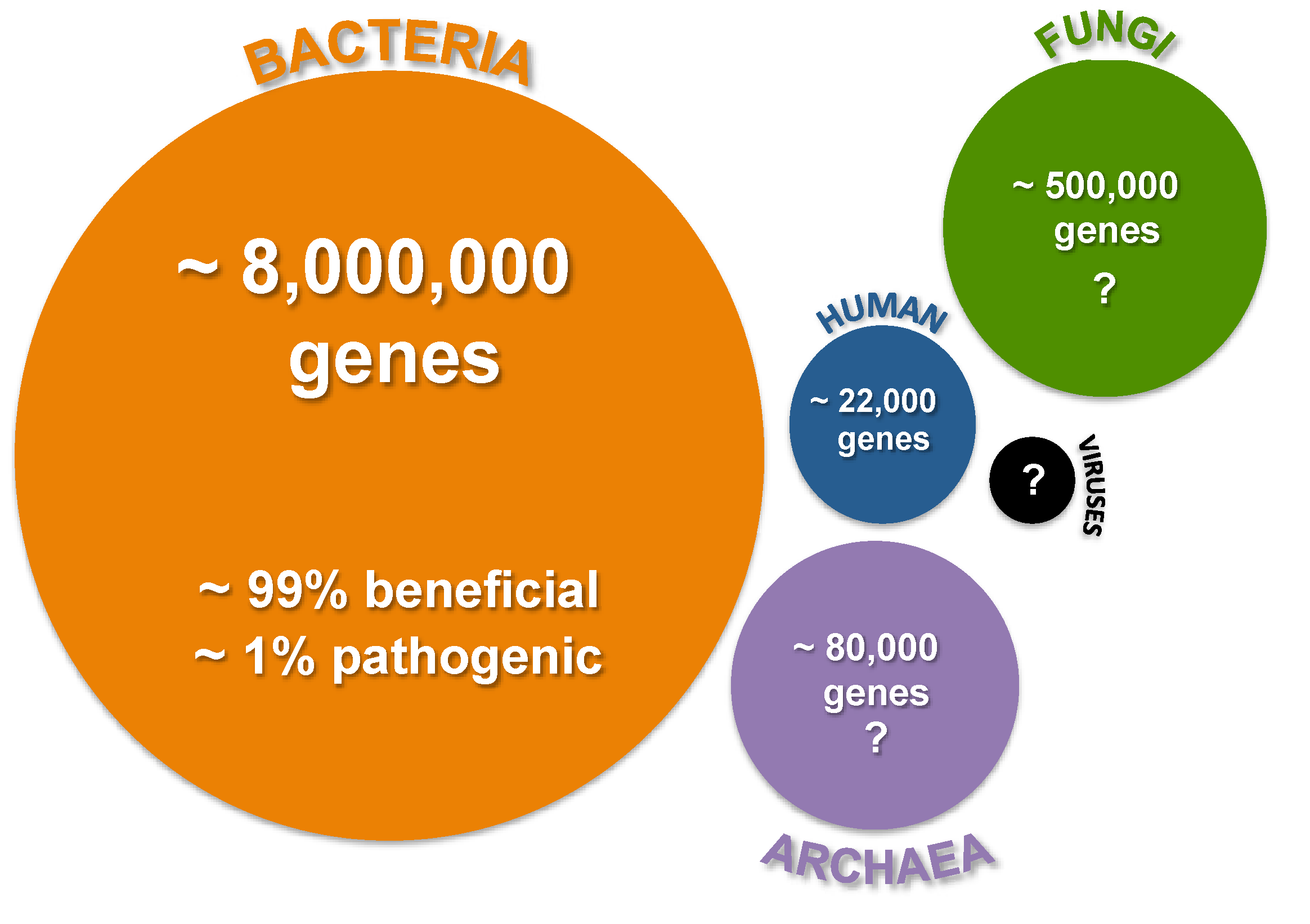
2. Spaceflight and Simulated Microgravity Environments
3. Impact of Microgravity on Animal-Microbe Associations
3.1. Pathogenic Interactions with Animals
3.2. Mutualistic Interactions with Animals
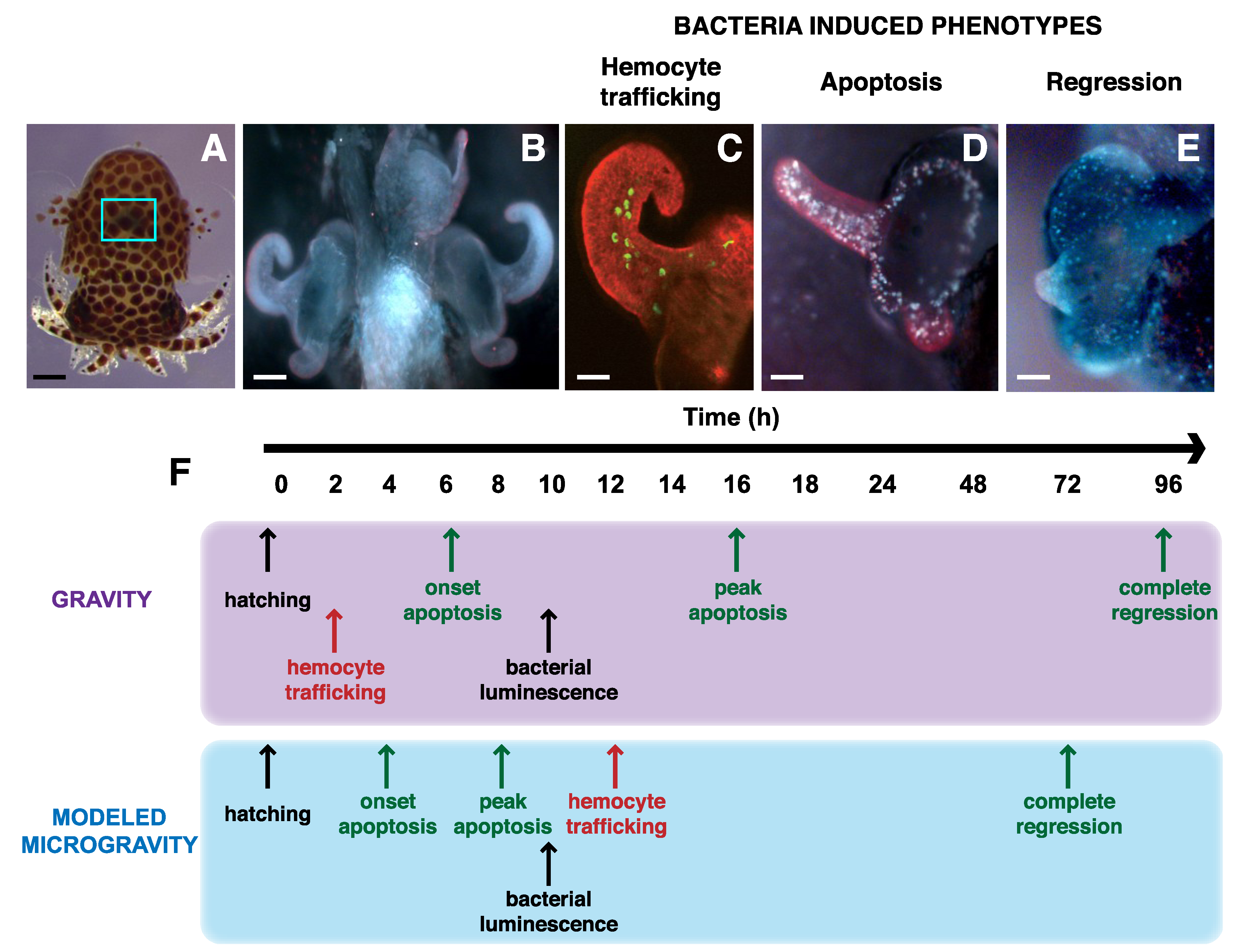
4. Impact of Microgravity on Plant-Microbe Associations
5. Summary
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Nickerson, C.A.; Ott, C.M.; Wilson, J.W.; Ramamurthy, R.; Pierson, D.L. Microbial responses to microgravity and other low-shear environments. Microbiol. Mol. Biol. Rev. 2004, 68, 345–361. [Google Scholar] [CrossRef]
- Chopra, V.; Fadl, A.A.; Sha, J.; Chopra, S.; Galindo, C.L.; Chopra, A.K. Alterations in the virulence potential of enteric pathogens and bacterial-host cell interactions under simulated microgravity conditions. J. Toxicol. Environ. Health A 2006, 69, 1345–1370. [Google Scholar] [CrossRef]
- Klaus, D.M.; Howard, H.N. Antibiotic efficacy and microbial virulence during space flight. Trends Biotechnol. 2006, 24, 131–136. [Google Scholar] [CrossRef]
- Lynch, S.V.; Matin, A. Travails of microgravity: Man and microbes in space. Biologist 2005, 52, 80–92. [Google Scholar]
- Wilson, J.W.; Ott, C.M.; Honer zu Bentrup, K.; Ramamurthy, R.; Quick, L.; Porwollik, S.; Cheng, P.; McClelland, M.; Tsaprailis, G.; et al. Space flight alters bacterial gene expression and virulence and reveals a role for global regulator Hfq. Proc. Natl. Acad. Sci. USA 2007, 104, 16299–16304. [Google Scholar] [CrossRef]
- Mauclaire, L.; Egli, M. Effect of simulated microgravity on growth and production of exopolymeric substances of Micrococcus luteus space and Earth isolates. FEMS Immunol. Med. Microbiol. 2010, 59, 350–356. [Google Scholar]
- Vukanti, R.; Model, M.A.; Leff, L.G. Effect of modeled reduced gravity conditions on bacterial morphology and physiology. BMC Microbiol. 2012, 12. [Google Scholar] [CrossRef]
- Morey-Holton, E.R. The Impact of Gravity on Life. In Evolution on Planet Earth: The Impact of the Physical Environment; Rothschild, L.J., Lister, A., Eds.; Academic Press: New York, NY, USA, 2003; pp. 143–159. [Google Scholar]
- Volkmann, D.; Baluska, F. Gravity: One of the driving forces for evolution. Protoplasma 2006, 229, 143–148. [Google Scholar] [CrossRef]
- Globus, R.K.; Morey-Holton, E.R. Advances in understanding the skeletal biology of spaceflight. Grav. Space Biol. 2009, 22, 3–12. [Google Scholar]
- Stein, T.P. Weight, muscle and bone loss during space flight: Another perspective. Eur. J. Appl. Physiol. 2013, 113, 2171–2181. [Google Scholar] [CrossRef]
- Guéguinou, N.; Huin-Schohn, C.; Bascove, M.; Bueb, J.L.; Tschirhart, E.; Legrand-Frossi, C.; Frippiat, J.P. Could spaceflight-associated immune system weakening preclude the expansion of human presence beyond Earth’s orbit? J. Leukoc. Biol. 2009, 86, 1027–1038. [Google Scholar] [CrossRef]
- Ott, C.M.; Crabbe, A.; Wilson, J.W.; Barrila, J.; Castro, S.L.; Nickerson, C.A. Microbial Stress: Spaceflight-Induced Alterations in Microbial Virulence nad Infectious Disease Risks for the Crew. In Stress Challenges and IMMUNITY in Space; Chouker, A., Ed.; Springer-Verlag: Berlin/Heidelberg, Gemany, 2012; pp. 203–225. [Google Scholar]
- Mermel, L.A. Infection prevention and control during prolonged human space travel. Clin. Infect. Dis. 2013, 56, 123–130. [Google Scholar] [CrossRef]
- Leach, J.E.; Ryba-White, M.; Sun, Q.; Wu, C.J.; Hilaire, E.; Gartner, C.; Nedukha, O.; Kordyum, E.; Keck, M.; Leung, H.; et al. Plants, plant pathogens, and microgravity—A deadly trio. Grav. Space Biol. 2001, 14, 15–23. [Google Scholar]
- Halstead, T.W.; Dutcher, F.R. Plants in space. Annu. Rev. Plant Physiol. 1987, 38, 317–345. [Google Scholar] [CrossRef]
- Henry, R.L.; Green, P.D.; Wong, P.P.; Guikema, J.A. Binding of isolated plant lectin by rhizobia during episodes of reduced gravity obtained by parabolic flight. Plant Physiol. 1990, 92, 262–264. [Google Scholar] [CrossRef]
- Ferl, R.; Wheeler, R.; Levine, H.G.; Paul, A.L. Plants in space. Curr. Opin. Plant Biol. 2002, 5, 258–263. [Google Scholar] [CrossRef]
- Lederberg, J.; McCray, A. The scientis: ‘Ome sweet’ omics—A geneological treasury of words. Scientist 2001, 17, No. 7. [Google Scholar]
- Shade, A.; Hogan, C.S.; Klimowicz, A.K.; Linske, M.; McManus, P.S.; Handelsman, J. Culturing captures members of the soil rare biosphere. Environ. Microbiol. 2012, 14, 2247–2252. [Google Scholar]
- Human Microbiome Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef]
- Dethlefsen, L.; McFall-Ngai, M.; Relman, D.A. An ecological and evolutionary perspective on human-microbe mutualism and disease. Nature 2007, 449, 811–818. [Google Scholar] [CrossRef]
- Mitreva, M. The genome of a blood fluke associated with human cancer. Nat. Genet. 2012, 44, 116–118. [Google Scholar] [CrossRef]
- Cho, I.; Blaser, M.J. The human microbiome: At the interface of health and disease. Nat. Rev. Genet. 2012, 13, 260–270. [Google Scholar]
- Rakoff-Nahoum, S.; Paglino, J.; Eslami-Varzaneh, F.; Edberg, S.; Medzhitov, R. Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell 2004, 118, 229–241. [Google Scholar] [CrossRef]
- Mazmanian, S.K.; Liu, C.H.; Tzianabos, A.O.; Kasper, D.L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 2005, 122, 107–118. [Google Scholar] [CrossRef]
- Wikoff, W.R.; Anfora, A.T.; Liu, J.; Schultz, P.G.; Lesley, S.A.; Peters, E.C.; Siuzdak, G. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc. Natl. Acad. Sci. USA 2009, 106, 3698–3703. [Google Scholar]
- Wen, L.; Ley, R.E.; Volchkov, P.Y.; Stranges, P.B.; Avanesyan, L.; Stonebraker, A.C.; Hu, C.; Wong, F.S.; Szot, G.L.; Bluestone, J.A.; et al. Innate immunity and intestinal microbiota in the development of type 1 diabetes. Nature 2008, 455, 1109–1113. [Google Scholar] [CrossRef]
- Ivanov, I.I.; Atarashi, K.; Manel, N.; Brodie, E.L.; Shima, T.; Karaoz, U.; Wei, D.; Goldfarb, K.C.; Santee, C.A.; Lynch, S.V.; et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 2009, 139, 485–498. [Google Scholar] [CrossRef]
- Brown, C.T.; Davis-Richardson, A.G.; Giongo, A.; Gano, K.A.; Crabb, D.B.; Mukherjee, N.; Casella, G.; Drew, J.C.; Ilonen, J.; Knip, M.; et al. Gut microbiome metagenomics analysis suggests a functional model for the development of autoimmunity for type 1 diabetes. PLoS One 2011, 6, e25792. [Google Scholar] [CrossRef]
- Giongo, A.; Gano, K.A.; Crabb, D.B.; Mukherjee, N.; Novelo, L.L.; Casella, G.; Drew, J.C.; Ilonen, J.; Knip, M.; Hyoty, H.; et al. Toward defining the autoimmune microbiome for type 1 diabetes. ISME J. 2011, 5, 82–91. [Google Scholar] [CrossRef]
- Plottel, C.S.; Blaser, M.J. Microbiome and malignancy. Cell Host Microbe 2011, 10, 324–335. [Google Scholar] [CrossRef]
- Ley, R.E.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Microbial ecology: Human gut microbes associated with obesity. Nature 2006, 444, 1022–1023. [Google Scholar] [CrossRef]
- Kennedy, T.A.; Naeem, S.; Howe, K.M.; Knops, J.M.; Tilman, D.; Reich, P. Biodiversity as a barrier to ecological invasion. Nature 2002, 417, 636–638. [Google Scholar] [CrossRef]
- Taylor, G.R. Recovery of medically important microorganisms from Apollo astronauts. Aerosp. Med. 1974, 45, 824–828. [Google Scholar]
- Pierson, D.L.; Chidambaram, M.; Heath, J.D.; Mallary, L.; Mishra, S.K.; Sharma, B.; Weinstock, G.M. Epidemiology of Staphylococcus aureus during space flight. FEMS Immunol. Med. Microbiol. 1996, 16, 273–281. [Google Scholar] [CrossRef]
- Rosenzweig, J.A.; Abogunde, O.; Thomas, K.; Lawal, A.; Nguyen, Y.; Sodipe, A.; Jejelowo, O. Space flight and modeled microgravity effects on microbial growth and virulence. Appl. Microbiol. Biotechnol. 2010, 85, 885–891. [Google Scholar] [CrossRef]
- Schwarz, R.P.; Goodwin, T.J.; Wolf, D.A. Cell culture for three-dimensional modeling in rotating-wall vessels: An application of simulated microgravity. J. Tissue Cult. Methods 1992, 14, 51–58. [Google Scholar]
- Klaus, D. Clinostats and bioreactors. Grav. Space Biol. 2001, 14, 55–64. [Google Scholar]
- Horneck, G.; Klaus, D.M.; Mancinelli, R.L. Space microbiology. Microbiol. Mol. Biol. Rev. 2010, 74, 121–156. [Google Scholar] [CrossRef]
- Foster, J.S.; Khodadad, C.L.; Ahrendt, S.R.; Parrish, M.L. Impact of simulated microgravity on the normal developmental time line of an animal-bacterial symbiosis. Sci. Rep. 2013, 3. [Google Scholar] [CrossRef]
- Nyholm, S.V.; McFall-Ngai, M.J. The winnowing: Establishing the squid-vibrio symbiosis. Nat. Rev. Microbiol. 2004, 2, 632–642. [Google Scholar] [CrossRef]
- Barrila, J.; Radtke, A.L.; Crabbe, A.; Sarker, S.F.; Herbst-Kralovetz, M.M.; Ott, C.M.; Nickerson, C.A. Organotypic 3D cell culture models: Using the rotating wall vessel to study host-pathogen interactions. Nat. Rev. Microbiol. 2010, 8, 791–801. [Google Scholar] [CrossRef]
- Hammond, T.G.; Stodieck, L.; Birdsall, H.H.; Becker, J.L.; Koenig, P.; Hammond, J.S.; Gunter, M.A.; Allen, P.L. Effects of Microgravity on the Virulence of Listeria monocytogenes, Enterococcus faecalis, Candida albicans, and Methicillin-Resistant Staphylococcus aureus. Astrobiology 2013, 13, 1081–1090. [Google Scholar] [CrossRef]
- Grant, K.A.; Khodadad, C.L.; Foster, J.S. Role of Hfq in an animal-microbe symbiosis under simulated microgravity conditions. Int. J. Astrobiol. 2014, 13, 53–61. [Google Scholar] [CrossRef]
- Fang, A.; Pierson, D.L.; Koenig, D.W.; Mishra, S.K.; Demain, A.L. Effect of simulated microgravity and shear stress on microcin B17 production by Escherichia coli and on its excretion into the medium. Appl. Environ. Microbiol. 1997, 63, 4090–4092. [Google Scholar]
- Nauman, E.A.; Ott, C.M.; Sander, E.; Tucker, D.L.; Pierson, D.; Wilson, J.W.; Nickerson, C.A. Novel quantitative biosystem for modeling physiological fluid shear stress on cells. Appl. Environ. Microbiol. 2007, 73, 699–705. [Google Scholar]
- Nickerson, C.A.; Ott, C.M.; Wilson, J.W.; Ramamurthy, R.; LeBlanc, C.L.; Honer zu Bentrup, K.; Hammond, T.; Pierson, D.L. Low-shear modeled microgravity: A global environmental regulatory signal affecting bacterial gene expression, physiology, and pathogenesis. J. Microbiol. Methods 2003, 54, 1–11. [Google Scholar] [CrossRef]
- Wilson, J.W.; Ott, C.M.; Quick, L.; Davis, R.; Honer zu Bentrup, K.; Crabbe, A.; Richter, E.; Sarker, S.; Barrila, J.; Porwollik, S.; et al. Media ion composition controls regulatory and virulence response of Salmonella in spaceflight. PLoS One 2008, 3, e3923. [Google Scholar] [CrossRef]
- Klaus, D.; Simske, S.; Todd, P.; Stodieck, L. Investigation of space flight effects on Escherichia coli and a proposed model of underlying physical mechanisms. Microbiology 1997, 143, 449–455. [Google Scholar] [CrossRef]
- Kacena, M.A.; Merrell, G.A.; Manfredi, B.; Smith, E.E.; Klaus, D.M.; Todd, P. Bacterial growth in space flight: Logistic growth curve parameters for Eschechia coli and Bacillus subtilus. Appl. Microbiol. Biotechnol. 1999, 51, 229–234. [Google Scholar] [CrossRef]
- Demain, A.L.; Fang, A. Secondary metabolism in simulated microgravity. Chem. Rec. 2001, 1, 333–346. [Google Scholar] [CrossRef]
- Ciferri, O.; Tiboni, O.; di Pasquale, G.; Orlandoni, A.M.; Marchesi, M.L. Effects of microgravity on genetic recombination in Escherichia coli. Naturwissenschaften 1986, 73, 418–421. [Google Scholar] [CrossRef]
- De Boever, P.; Mergeay, M.; Ilyin, V.; van der Auwera, G.; Mahillon, J. Conjugation-mediated plasmid exchange between bacteria grown under space flight conditions. Microgravity Sci. Technol. 2007, 19, 138–144. [Google Scholar] [CrossRef]
- Lynch, S.V.; Mukundakrishnan, K.; Benoit, M.R.; Ayyaswamy, P.S.; Matin, A. Escherichia coli biofilms formed under low-shear modeled microgravity in a ground-based system. Appl. Environ. Microbiol. 2006, 72, 7701–7710. [Google Scholar] [CrossRef]
- Searles, S.C.; Woolley, C.M.; Petersen, R.A.; Hyman, L.E.; Nielsen-Preiss, S.M. Modeled microgravity increases filamentation, biofilm formation, phenotypic switching, and antimicrobial resistance in Candida albicans. Astrobiology 2011, 11, 825–836. [Google Scholar] [CrossRef]
- Wilson, J.W.; Ott, C.M.; Ramamurthy, R.; Porwollik, S.; McClelland, M.; Pierson, D.L.; Nickerson, C.A. Low-Shear modeled microgravity alters the Salmonella enterica serovar typhimurium stress response in an RpoS-independent manner. Appl. Environ. Microbiol. 2002, 68, 5408–5416. [Google Scholar] [CrossRef]
- Kaur, I.; Simons, E.R.; Kapadia, A.S.; Ott, C.M.; Pierson, D.L. Effect of spaceflight on ability of monocytes to respond to endotoxins of gram-negative bacteria. Clin. Vaccine Immunol. 2008, 15, 1523–1528. [Google Scholar] [CrossRef]
- Crabbé, A.; Schurr, M.J.; Monsieurs, P.; Morici, L.; Schurr, J.; Wilson, J.W.; Ott, C.M.; Tsaprailis, G.; Pierson, D.L.; Stefanyshyn-Piper, H.; et al. Transcriptional and proteomic responses of Pseudomonas aeruginosa PAO1 to spaceflight conditions involve Hfq regulation and reveal a role for oxygen. Appl. Environ. Microbiol. 2011, 77, 1221–1230. [Google Scholar] [CrossRef]
- Arunasri, K.; Adil, M.; Venu Charan, K.; Suvro, C.; Himabindu Reddy, S.; Shivaji, S. Effect of simulated microgravity on E. coli K12 MG1655 growth and gene expression. PLoS One 2013, 8, e57860. [Google Scholar]
- Wilson, J.W.; Ramamurthy, R.; Porwollik, S.; McClelland, M.; Hammond, T.; Allen, P.; Ott, C.M.; Pierson, D.L.; Nickerson, C.A. Microarray analysis identifies Salmonella genes belonging to the low-shear modeled microgravity regulon. Proc. Natl. Acad. Sci. USA 2002, 99, 13807–13812. [Google Scholar]
- Castro, S.L.; Nelman-Gonzalez, M.; Nickerson, C.A.; Ott, C.M. Induction of attachment-independent biofilm formation and repression of Hfq expression by low-fluid-shear culture of Staphylococcus aureus. Appl. Environ. Microbiol. 2011, 77, 6368–6378. [Google Scholar] [CrossRef]
- Valentin-Hansen, P.; Eriksen, M.; Udesen, C. The bacterial Sm-like protein Hfq: A key player in RNA transactions. Mol. Microbiol. 2004, 51, 1525–1533. [Google Scholar] [CrossRef]
- Sittka, A.; Pfeiffer, V.; Tedin, K.; Vogel, J. The RNA chaperone Hfq is essential for the virulence of Salmonella typhimurium. Mol. Microbiol. 2007, 63, 193–217. [Google Scholar] [CrossRef]
- Robertson, G.T.; Roop, R.M., Jr. The Brucella abortus host factor I (HF-I) protein contributes to stress resistance during stationary phase and is a major determinant of virulence in mice. Mol. Microbiol. 1999, 34, 690–700. [Google Scholar] [CrossRef]
- Ding, Y.; Davis, B.M.; Waldor, M.K. Hfq is essential for Vibrio cholerae virulence and downregulates sigma expression. Mol. Microbiol. 2004, 53, 345–354. [Google Scholar] [CrossRef]
- Hammond, T.G.; Hammond, J.M. Optimized suspension culture: The rotating-wall vessel. Am. J. Physiol. 2001, 281, F12–F25. [Google Scholar]
- McFall-Ngai, M.J.; Ruby, E.G. Symbiont recognition and subsequent morphogenesis as early events in an animal-bacterial mutualism. Science 1991, 254, 1491–1494. [Google Scholar]
- McFall-Ngai, M.; Nyholm, S.V.; Castillo, M.G. The role of the immune system in the initiation and persistence of the Euprymna scolopes—Vibrio fischeri symbiosis. Semin. Immunol. 2010, 22, 48–53. [Google Scholar] [CrossRef]
- Altura, M.A.; Stabb, E.; Goldman, W.; Apicella, M.; McFall-Ngai, M.J. Attenuation of host No production by MAMPs potentiates development of the host in the squid-vibrio symbiosis. Cell. Microbiol. 2011, 13, 527–537. [Google Scholar] [CrossRef]
- McFall-Ngai, M.; Heath-Heckman, E.A.; Gillette, A.A.; Peyer, S.M.; Harvie, E.A. The secret languages of coevolved symbioses: Insights from the Euprymna scolopes-Vibrio fischeri symbiosis. Semin. Immunol. 2012, 24, 3–8. [Google Scholar] [CrossRef]
- Buchon, N.; Broderick, N.A.; Lemaitre, B. Gut homeostasis in a microbial world: Insights from Drosophila melanogaster. Nat. Rev. Microbiol. 2013, 11, 615–626. [Google Scholar] [CrossRef]
- Nyholm, S.V.; Stabb, E.V.; Ruby, E.G.; McFall-Ngai, M.J. Establishment of an animal-bacterial association: Recruiting symbiotic vibrios from the environment. Proc. Natl. Acad. Sci. USA 2000, 97, 10231–10235. [Google Scholar]
- Foster, J.S.; Apicella, M.A.; McFall-Ngai, M.J. Vibrio fischeri lipopolysaccharide induces developmental apoptosis, but not complete morphogenesis, of the Euprymna scolopes symbiotic light organ. Dev. Biol. 2000, 226, 242–254. [Google Scholar] [CrossRef]
- Koropatnick, T.A.; Engle, J.T.; Apicella, M.A.; Stabb, E.V.; Goldman, W.E.; McFall-Ngai, M.J. Microbial factor-mediated development in a host-bacterial mutualism. Science 2004, 306, 1186–1188. [Google Scholar] [CrossRef]
- Montgomery, M.K.; McFall-Ngai, M. Bacterial symbionts induce host organ morphogenesis during early postembryonic development of the squid Euprymna scolopes. Development 1994, 120, 1719–1729. [Google Scholar]
- Doino, J.A.; McFall-Ngai, M. Transient exposures to competent bacteria initiates symbiosis-specific squid light organ morphogenesis. Biol. Bull. 1995, 189, 347–355. [Google Scholar] [CrossRef]
- Foster, J.S.; McFall-Ngai, M.J. Induction of apoptosis by cooperative bacteria in the morphogenesis of host epithelial tissues. Dev. Genes Evol. 1998, 208, 295–303. [Google Scholar] [CrossRef]
- Crucian, B.E.; Stowe, R.P.; Pierson, D.L.; Sams, C.F. Immune system dysregulation following short- vs long-duration space flight. Aviat. Space Environ. Med. 2008, 79, 835–843. [Google Scholar] [CrossRef]
- Baqai, F.P.; Gridley, D.S.; Slater, J.M.; Luo-Owen, X.; Stodieck, L.S.; Ferguson, V.; Chapes, S.K.; Pecaut, M.J. Effects of spaceflight on innate immune function and antioxidant gene expression. J. Appl. Physiol. 2009, 106, 1935–1942. [Google Scholar] [CrossRef]
- Read, D.J.; Perez-Moreno, J. Mycorrhizas and nutrient cycling in ecosystems—A journy towards relevance? New Phytol. 2003, 157, 475–492. [Google Scholar] [CrossRef]
- Harrison, M.J. Signaling in the arbuscular mycorrhizal symbiosis. Annu. Rev. Microbiol. 2005, 59, 19–42. [Google Scholar] [CrossRef]
- Marschner, H.; Kirkby, E.A.; Cakmak, I. Effect of mineral nutritional status on shoot-root partitioning of photoassimilates and cycling of mineral nutrients. J. Exp. Bot. 1996, 47, 1255–1263. [Google Scholar] [CrossRef]
- Mendes, R.; Garbeva, P.; Raaijmakers, J.M. The rhizosphere microbiome: Significance of plant beneficial, plant pathogenic, and human pathogenic microorganisms. FEMS Microbiol. Rev. 2013, 37, 634–663. [Google Scholar] [CrossRef]
- Mendes, R.; Kruijt, M.; de Bruijn, I.; Dekkers, E.; van der Voort, M.; Schneider, J.H.; Piceno, Y.M.; DeSantis, T.Z.; Andersen, G.L.; Bakker, P.A.; et al. Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 2011, 332, 1097–1100. [Google Scholar]
- Berendsen, R.L.; Pieterse, C.M.; Bakker, P.A. The rhizosphere microbiome and plant health. Trends Plant Sci. 2012, 17, 478–486. [Google Scholar] [CrossRef]
- Zamioudis, C.; Pieterse, C.M. Modulation of host immunity by beneficial microbes. Mol. Plant Microbe Interact. 2012, 25, 139–150. [Google Scholar] [CrossRef]
- Millet, Y.A.; Danna, C.H.; Clay, N.K.; Songnuan, W.; Simon, M.D.; Werck-Reichhart, D.; Ausubel, F.M. Innate immune responses activated in Arabidopsis roots by microbe-associated molecular patterns. Plant Cell 2010, 22, 973–990. [Google Scholar] [CrossRef]
- Van der Ent, S.; van Wees, S.C.; Pieterse, C.M. Jasmonate signaling in plant interactions with resistance-inducing beneficial microbes. Phytochemistry 2009, 70, 1581–1588. [Google Scholar] [CrossRef]
- Porterfield, D.M. Astroculture root metabolism and cytochemical analysis. Adv. Space Res. 2000, 26, 315–318. [Google Scholar] [CrossRef]
- Heinse, R.; Jones, S.B.; Steinberg, S.L.; Tuller, M.; Or, D. Measurements and modeling of variable gravity effects on water distribution and flow in unsaturated ourous media. Vadose Zone J. 2007, 6, 713–724. [Google Scholar] [CrossRef]
- Stutte, G.W.; Monje, O.; Goins, G.D.; Tripathy, B.C. Microgravity effects on thylakoid, single leaf, and whole canopy photosynthesis of dwarf wheat. Planta 2005, 223, 46–56. [Google Scholar] [CrossRef]
- Paul, A.L.; Wheeler, R.M.; Levine, H.G.; Ferl, R.J. Fundamental plant biology enabled by the space shuttle. Am. J. Bot. 2013, 100, 226–234. [Google Scholar] [CrossRef]
- Croxdale, J.; Cook, M.; Tibbitts, T.W.; Brown, C.S.; Wheeler, R.M. Structure of potato tubers formed during spaceflight. J. Exp. Bot. 1997, 48, 2037–2043. [Google Scholar] [CrossRef]
- Paul, A.L.; Daugherty, C.J.; Bihn, E.A.; Chapman, D.K.; Norwood, K.L.; Ferl, R.J. Transgene expression patterns indicate that spaceflight affects stress signal perception and transduction in arabidopsis. Plant Physiol. 2001, 126, 613–621. [Google Scholar] [CrossRef]
- Ryba-White, M.; Nedukha, O.; Hilaire, E.; Guikema, J.A.; Kordyum, E.; Leach, J.E. Growth in microgravity increases susceptibility of soybean to a fungal pathogen. Plant Cell Phyiol. 2001, 42, 657–664. [Google Scholar] [CrossRef]
- Bingham, G.E.; Podolsky, I.G.; Topham, T.S.; Mullholland, J.M. Lada: The ISS Plant Substrate Microgravity Testbed; SAE Technical Paper, No. 2002-01-187; Society of Automotive Engineers (SAE): Warrendale, PA, USA, 2002. [Google Scholar]
- Hummerick, M.E.; Garland, J.; Bingham, G.E.; Sychev, V.N.; Podolsky, I.G. Microbiological analysis of Lada vegetable units (VPU) to define critical control points and procedures to endure the safety of space grown vegetables. In Proceedings of the Internaitonal Conference on Environmental Systems, Barcelona, Spain, 11–15 July 2010. Volume AIAA-2010–6253.
- Wheeler, R.M.; Mackowiak, C.L.; Yorio, N.C.; Sager, J.C. Effects of CO2 on stomatal conductance: Do stomata open at very high CO2 concentrations? Ann. Bot. 1999, 83, 234–251. [Google Scholar]
- Kaplan, F.; Zhao, W.; Richards, J.T.; Wheeler, R.M.; Guy, C.L.; Levine, L.H. Transcriptional and metabolic insights into the differential physiological responses of Arabidopsis to optimal and supraoptimal atmospheric CO2. PLoS One 2012, 7, e43583. [Google Scholar]
- Bishop, D.L.; Levine, H.G.; Kropp, B.R.; Anderson, A.J. Seedborne fungal contamination: Consequences in space-grown wheat. Phytopathology 1997, 87, 1125–1133. [Google Scholar] [CrossRef]
- Nedukha, E.M. Possible mechanisms of plant cell wall changes at microgravity. Adv. Space Res. 1996, 17, 37–45. [Google Scholar] [CrossRef]
- Urban, J.E.; Gerren, R.; Zoelle, J. Effets of microgravity on the binding of acetylsalicylic acid by Rhizobium leguminosarum bv. trifolii. Acta Astronaut. 1995, 36, 129–133. [Google Scholar] [CrossRef]
- Oke, V.; Long, S.R. Bacterial genes induced within the nodule during the Rhizobium-legume symbiosis. Mol. Microbiol. 1999, 32, 837–849. [Google Scholar] [CrossRef]
- National Research Council. Recapturing a Future for Space Exploration; Life and Physical Sciences Research for a New Era; Council, N.R., Ed.; The National Academies Press: Washington, DC, USA, 2011. [Google Scholar]
- Bosch, T.C.G.; McFall-Ngai, M.J. Metaorganisms as the new frontier. Zoology 2011, 114, 185–190. [Google Scholar] [CrossRef]
- Liu, L.; Chen, X.; Skogerbo, G.; Zhang, P.; Chen, R.; He, S.; Huang, D.W. The human microbiome: A hot spot of microbial horizontal gene transfer. Genomics 2012, 100, 265–270. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Foster, J.S.; Wheeler, R.M.; Pamphile, R. Host-Microbe Interactions in Microgravity: Assessment and Implications. Life 2014, 4, 250-266. https://doi.org/10.3390/life4020250
Foster JS, Wheeler RM, Pamphile R. Host-Microbe Interactions in Microgravity: Assessment and Implications. Life. 2014; 4(2):250-266. https://doi.org/10.3390/life4020250
Chicago/Turabian StyleFoster, Jamie S., Raymond M. Wheeler, and Regine Pamphile. 2014. "Host-Microbe Interactions in Microgravity: Assessment and Implications" Life 4, no. 2: 250-266. https://doi.org/10.3390/life4020250
APA StyleFoster, J. S., Wheeler, R. M., & Pamphile, R. (2014). Host-Microbe Interactions in Microgravity: Assessment and Implications. Life, 4(2), 250-266. https://doi.org/10.3390/life4020250



