Air Pollution and microRNAs: The Role of Association in Airway Inflammation
Abstract
1. Introduction
2. miRNA: General Consideration
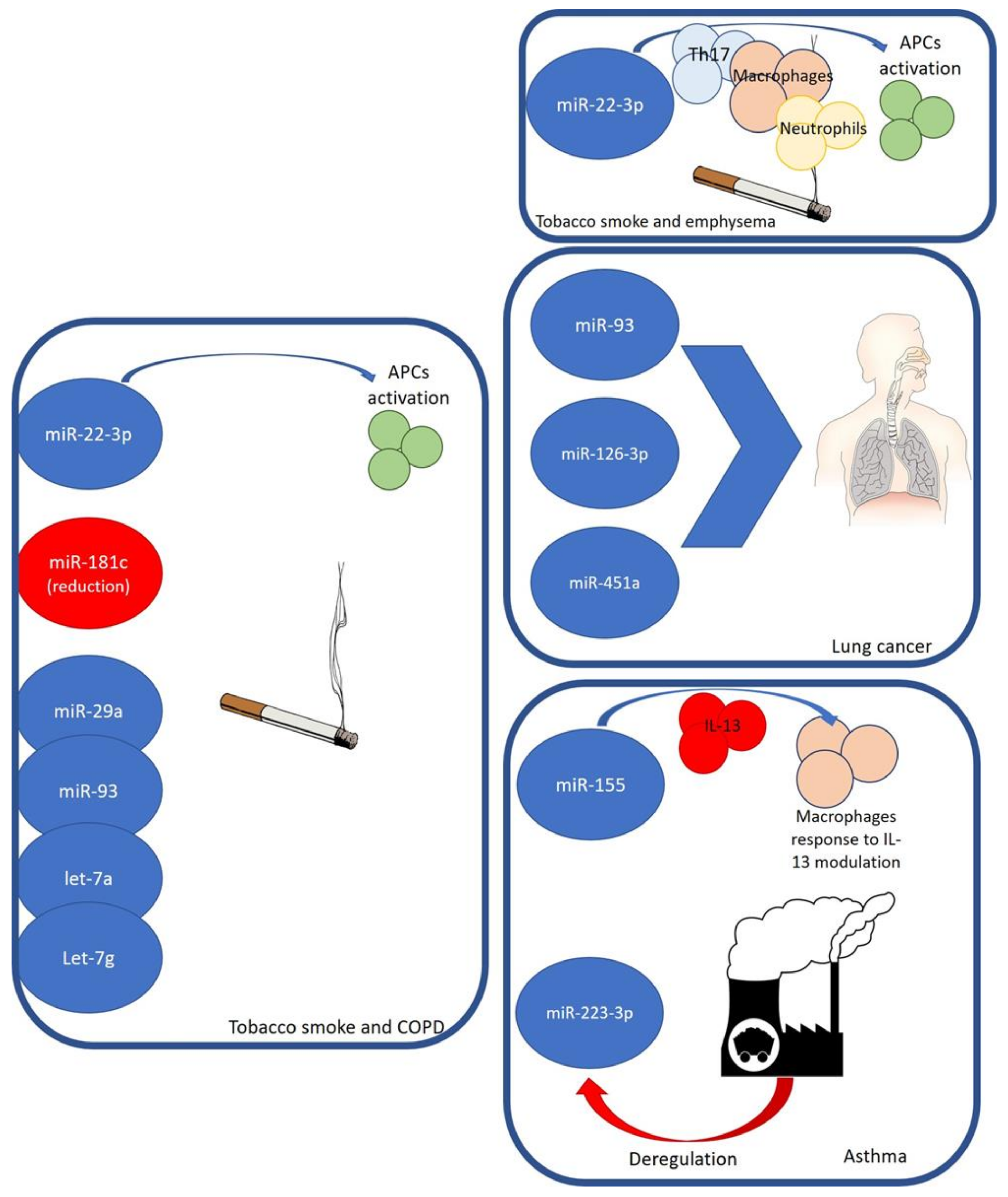
3. Tobacco Smoke and miRNAs in Airway Diseases
4. miRNA, Air Pollutants, and Airways Diseases
5. The Use of miRNA as Therapeutic Strategy in Lung Disease
6. Future Perspectives
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Couzin, J. MicroRNAs make big impression in disease after disease. Science 2008, 319, 1782–1784. [Google Scholar] [CrossRef]
- Heffler, E.; Allegra, A.; Pioggia, G.; Picardi, G.; Musolino, C.; Gangemi, S. MicroRNA Profiling in Asthma: Potential Biomarkers and Therapeutic Targets. Am. J. Respir. Cell Mol. Biol. 2017, 57, 642–650. [Google Scholar] [CrossRef] [PubMed]
- Vrijens, K.; Bollati, V.; Nawrot, T.S. MicroRNAs as Potential Signatures of Environmental Exposure or Effect: A Systematic Review. Environ. Health Perspect. 2015, 123, 399–411. [Google Scholar] [CrossRef] [PubMed]
- Wild, C. Complementing the genome with an “exposome”: The outstanding challenge of environmental exposure measurement in molecular epidemiology. Cancer Epidemiol. Biomark. Prev. 2005, 14, 1847–1850. [Google Scholar] [CrossRef] [PubMed]
- Eguiluz-Gracia, I.; Mathioudakis, A.G.; Bartel, S.; Vijverberg, S.J.H.; Fuertes, E.; Comberiati, P.; Cai, Y.S.; Tomazic, P.V.; Diamant, Z.; Vestbo, J.; et al. The need for clean air: The way air pollution and climate change affect allergic rhinitis and asthma. Allergy 2020, 75, 2170–2184. [Google Scholar] [CrossRef] [PubMed]
- Del Giacco, S.R.; Bakirtas, A.; Bel, E.; Custovic, A.; Diamant, Z.; Hamelmann, E.; Heffler, E.; Kalayci, Ö.; Saglani, S.; Sergejeva, S.; et al. Allergy in severe asthma. Allergy 2017, 72, 207–220. [Google Scholar] [CrossRef]
- Rondon, C.; Bogas, G.; Barrionuevo, E.; Blanca, M.; Torres, M.J.; Campo, P. Nonallergic rhinitis and lower airway disease. Allergy 2017, 72, 24–34. [Google Scholar] [CrossRef]
- Tomari, Y.; Zamore, P.D. MicroRNA Biogenesis: Drosha Can’t Cut It without a Partner. Curr. Biol. 2005, 15, R61–R644. [Google Scholar] [CrossRef]
- Michlewski, G.; Cáceres, J.F. Post-Transcriptional Control of MiRNA Biogenesis. RNA 2019, 25, 1–16. [Google Scholar] [CrossRef]
- Rupani, H.; Sanchez-Elsner, T.; Howarth, P. MicroRNAs and respiratory diseases. Eur. Respir. J. 2013, 41, 695–705. [Google Scholar] [CrossRef]
- Breving, K.; Esquela-Kerscher, A. The complexities of microRNA regulation: Mirandering around the rules. Int. J. Biochem. Cell Biol. 2010, 42, 1316–1329. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.C.; Wentzel, E.A.; Kent, O.A.; Ramachandran, K.; Mullendore, M.; Lee, K.H.; Feldmann, G.; Yamakuchi, M.; Ferlito, M.; Lowenstein, C.J.; et al. Transactivation of miR34a by p53 broadly influences gene expression and promotes apoptosis. Mol. Cell 2007, 26, 745–752. [Google Scholar] [CrossRef]
- He, L.; He, X.; Lim, L.P.; de Stanchina, E.; Xuan, Z.; Liang, Y.; Xue, W.; Zender, L.; Magnus, J.; Ridzon, D.; et al. A microRNA component of the p53 tumour suppressor network. Nature 2007, 447, 1130–1134. [Google Scholar] [CrossRef]
- Raver-Shapira, N.; Marciano, E.; Meiri, E.; Spector, Y.; Rosenfeld, N.; Moskovits, N.; Bentwich, Z.; Oren, M. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol. Cell 2007, 26, 731–743. [Google Scholar] [CrossRef] [PubMed]
- Brueckner, B.; Stresemann, C.; Kuner, R.; Mund, C.; Musch, T.; Meister, M.; Sültmann, H.; Lyko, F. The human let-7a-3 locus contains an epigenetically regulated microRNA gene with oncogenic function. Cancer Res. 2007, 67, 1419–1423. [Google Scholar] [CrossRef]
- Williams, A.E.; Moschos, S.A.; Perry, M.M.; Barnes, P.J.; Lindsay, M.A. Maternally imprinted microRNAs are differentially expressed during mouse and human lung development. Dev. Dyn. 2007, 236, 572–580. [Google Scholar] [CrossRef] [PubMed]
- Williams, A.E.; Perry, M.M.; Moschos, S.A.; Lindsay, M.A. MicroRNA expression in the aging mouse lung. BMC Genom. 2007, 8, 172. [Google Scholar] [CrossRef] [PubMed]
- Ventura, A.; Young, A.G.; Winslow, M.M.; Lintault, L.; Meissner, A.; Erkeland, S.J.; Newman, J.; Bronson, R.T.; Crowley, D.; Stone, J.R.; et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell 2008, 132, 875–886. [Google Scholar] [CrossRef]
- Rodriguez, A.; Vigorito, E.; Clare, S.; Warren, M.V.; Couttet, P.; Soond, D.R.; van Dongen, S.; Grocock, R.J.; Das, P.P.; Miska, E.A.; et al. Requirement of bic/microRNA-155 for normal immune function. Science 2007, 316, 608–611. [Google Scholar] [CrossRef]
- Hulin, M.; Simoni, M.; Viegi, G.; Annesi-Maesano, I. Respiratory health and indoor air pollutants based on quantitative exposure assessments. Eur. Respir. J. 2012, 40, 1033–1045. [Google Scholar] [CrossRef]
- Vardavas, C.I.; Hohmann, C.; Patelarou, E.; Martinez, D.; Henderson, A.J.; Granell, R.; Sunyer, J.; Torrent, M.; Fantini, M.P.; Gori, D.; et al. The independent role of prenatal and postnatal exposure to active and passive smoking on the development of early wheeze in children. Eur. Respir. J. 2016, 48, 115–124. [Google Scholar] [CrossRef] [PubMed]
- Burke, H.; Leonardi-Bee, J.; Hashim, A.; Pine-Abata, H.; Chen, Y.; Cook, D.G.; Britton, J.R.; McKeever, T.M. Prenatal and passive smoke exposure and incidence of asthma and wheeze: Systematic review and meta-analysis. Pediatrics 2012, 129, 735–744. [Google Scholar] [CrossRef] [PubMed]
- Melén, E.; Barouki, R.; Barry, M.; Boezen, H.M.; Hoffmann, B.; Krauss-Etschmann, S.; Koppelman, G.H.; Forsberg, B. Promoting respiratory public health through epigenetics research: An ERS Environment Health Committee workshop report. Eur. Respir. J. 2018, 51, 1702410. [Google Scholar] [CrossRef]
- Hwang, S.H.; Hwang, J.H.; Moon, J.S.; Lee, D.H. Environmental tobacco smoke and children’s health. Korean J. Pediatr. 2012, 55, 35–41. [Google Scholar] [CrossRef] [PubMed]
- Birzele, L.T.; Depner, M.; Ege, M.J.; Engel, M.; Kublik, S.; Bernau, C.; Loss, G.J.; Genuneit, J.; Horak, E.; Schloter, M.; et al. Environmental and mucosal microbiota and their role in childhood asthma. Allergy 2017, 72, 109–119. [Google Scholar] [CrossRef]
- Marsland, B.J.; Gollwitzer, E.S. Host-microorganism interactions in lung diseases. Nat. Rev. Immunol. 2014, 14, 827–835. [Google Scholar] [CrossRef]
- Kulkarni, R.; Antala, S.; Wang, A.; Amaral, F.E.; Rampersaud, R.; Larussa, S.J.; Planet, P.J.; Ratner, A.J. Cigarette smoke increases Staphylococcus aureus biofilm formation via oxidative stress. Infect. Immun. 2012, 80, 3804–3811. [Google Scholar] [CrossRef]
- Izzotti, A.; Calin, G.A.; Arrigo, P.; Steele, V.E.; Croce, C.M.; De Flora, S. Downregulation of microRNA expression in the lungs of rats exposed to cigarette smoke. FASEB J. 2009, 23, 806–812. [Google Scholar] [CrossRef]
- Schembri, F.; Sridhar, S.; Perdomo, C.; Gustafson, A.M.; Zhang, X.; Ergun, A.; Lu, J.; Liu, G.; Zhang, X.; Bowers, J.; et al. MicroRNAs as modulators of smoking-induced gene expression changes in human airway epithelium. Proc. Natl. Acad. Sci. USA 2009, 106, 2319–2324. [Google Scholar] [CrossRef]
- Izzotti, A.; Calin, G.A.; Steele, V.E.; Croce, C.M.; De Flora, S. Relationships of microRNA expression in mouse lung with age and exposure to cigarette smoke and light. FASEB J. 2009, 23, 3243–3250. [Google Scholar] [CrossRef]
- Takahashi, K.; Yokota, S.; Tatsumi, N.; Fukami, T.; Yokoi, T.; Nakajima, M. Cigarette smoking substantially alters plasma microRNA profiles in healthy subjects. Toxicol. Appl. Pharmacol. 2013, 272, 154–160. [Google Scholar] [CrossRef]
- Goldberg, A.D.; Allis, C.D.; Bernstein, E. Epigenetics: A landscape takes shape. Cell 2007, 128, 635–638. [Google Scholar] [CrossRef]
- Gluckman, P.D.; Hanson, M.A.; Cooper, C.; Thornburg, K.L. Effect of in utero and early-life conditions on adult health and disease. N. Engl. J. Med. 2008, 359, 61–73. [Google Scholar] [CrossRef]
- Herberth, G.; Bauer, M.; Gasch, M.; Hinz, D.; Röder, S.; Olek, S.; Kohajda, T.; Rolle-Kampczyk, U.; von Bergen, M.; Sack, U.; et al. Lifestyle and Environmental Factors and Their Influence on Newborns Allergy Risk study group. Maternal and cord blood miR-223 expression associates with prenatal tobacco smoke exposure and low regulatory T-cell numbers. J. Allergy Clin. Immunol. 2014, 133, 543–550. [Google Scholar] [CrossRef]
- Tsamou, M.; Vrijens, K.; Madhloum, N.; Lefebvre, W.; Vanpoucke, C.; Nawrot, T.S. Air pollution-induced placental epigenetic alterations in early life: A candidate miRNA approach. Epigenetics 2018, 13, 135–146. [Google Scholar] [CrossRef]
- Velasco-Torres, Y.; Ruiz, V.; Montaño, M.; Pérez-Padilla, R.; Falfán-Valencia, R.; Pérez-Ramos, J.; Pérez-Bautista, O.; Ramos, C. Participation of the MiR-22-HDAC4-DLCO Axis in Patients with COPD by Tobacco and Biomass. Biomolecules 2019, 9, 837. [Google Scholar] [CrossRef]
- Du, Y.; Ding, Y.; Chen, X.; Mei, Z.; Ding, H.; Wu, Y.; Jie, Z. MicroRNA-181c Inhibits Cigarette Smoke–Induced Chronic Obstructive Pulmonary Disease by Regulating CCN1 Expression. Respir. Res. 2017, 18, 155. [Google Scholar] [CrossRef]
- Khan, A.; Thatcher, T.H.; Woeller, C.F.; Sime, P.J.; Phipps, R.P.; Hopke, P.K.; Utell, M.J.; Krahl, P.L.; Mallon, T.M.; Thakar, J. Machine Learning Approach for Predicting Past Environmental Exposures from Molecular Profiling of Post-Exposure Human Serum Samples. J. Occup. Environ. Med. 2019, 61 (Suppl. S12), S55–S64. [Google Scholar] [CrossRef]
- Tan, W.; Shen, H.; Wong, W. Dysregulated autophagy in COPD: A pathogenic process to be deciphered. Pharmacol. Res. 2019, 144, 1–7. [Google Scholar] [CrossRef]
- Lu, W.; You, R.; Yuan, X.; Yang, T.; Samuel, E.L.G.; Marcan, D.C.; Sikkema, W.K.A.; Tour, J.M.; Rodriguez, A.; Kheradmand, F.; et al. The microRNA miR-22 inhibits the histone deacetylase HDAC4 to promote T(H)17 cell-dependent emphysema. Nat. Immunol. 2015, 16, 1185–1194. [Google Scholar] [CrossRef]
- Gao, Y.; Deng, K.; Liu, X.; Dai, M.; Chen, X.; Chen, J.; Chen, J.; Huang, Y.; Dai, S.; Chen, J. Molecular Mechanism and Role of MicroRNA-93 in Human Cancers: A Study Based on Bioinformatics Analysis, Meta-analysis, and Quantitative Polymerase Chain Reaction Validation. J. Cell Biochem. 2019, 120, 6370–6383. [Google Scholar] [CrossRef]
- Micolucci, L.; Akhtar, M.M.; Olivieri, F.; Rippo, M.R.; Procopio, A.D. Diagnostic Value of MicroRNAs in Asbestos Exposure and Malignant Mesothelioma: Systematic Review and Qualitative Meta-Analysis. Oncotarget 2016, 7, 58606–58637. [Google Scholar] [CrossRef]
- Chen, Q.; Hu, H.; Jiao, D.; Yan, J.; Xu, W.; Tang, X.; Chen, J.; Wang, J. MiR-126-3p and MiR-451a Correlate with Clinicopathological Features of Lung Adenocarcinoma: The Underlying Molecular Mechanisms. Oncol. Rep. 2016, 36, 909–917. [Google Scholar] [CrossRef]
- Gulei, D.; Raduly, L.; Broseghini, E.; Ferracin, M.; Berindan-Neagoe, I. The Extensive Role of MiR-155 in Malignant and Non-Malignant Diseases. Mol. Asp. Med. 2019, 70, 33–56. [Google Scholar] [CrossRef]
- Malmhall, C.; Johansson, K.; Winkler, C.; Alawieh, S.; Ekerljung, L.; Radinger, M. Altered miR-155 Expression in Allergic Asthmatic Airways. Scand. J. Immunol. 2017, 85, 300–307. [Google Scholar] [CrossRef]
- Varikuti, S.; Verma, C.; Natarajan, G.; Oghumu, S.; Satoskar, A.R. MicroRNA155 Plays a Critical Role in the Pathogenesis of Cutaneous Leishmania major Infection by Promoting a Th2 Response and Attenuating Dendritic Cell Activity. Am. J. Pathol. 2021, 191, 809–816. [Google Scholar] [CrossRef]
- Zech, A.; Ayata, C.K.; Pankratz, F.; Meyer, A.; Baudiss, K.; Cicko, S.; Yegutkin, G.G.; Grundmann, S.; Idzko, M. MicroRNA-155 modulates P2R signaling and Th2 priming of dendritic cells during allergic airway inflammation in mice. Allergy 2015, 70, 1121–1129. [Google Scholar] [CrossRef]
- Jakiela, B.; Gielicz, A.; Plutecka, H.; Hubalewska-Mazgaj, M.; Mastalerz, L.; Bochenek, G.; Soja, J.; Januszek, R.; Aab, A.; Musial, J.; et al. Th2-type cytokine induced mucus metaplasia decreases susceptibility of human bronchial epithelium to rhinovirus infection. Am. J. Respir. Cell Mol. Biol. 2014, 51, 229–241. [Google Scholar] [CrossRef]
- Gomez, J.L.; Chen, A.; Diaz, M.P.; Zirn, N.; Gupta, A.; Britto, C.; Sauler, M.; Yan, X.; Stewart, E.; Santerian, K.; et al. A Network of Sputum MicroRNAs Is Associated with Neutrophilic Airway Inflammation in Asthma. Am. J. Respir. Crit. Care Med. 2020, 202, 51–64. [Google Scholar] [CrossRef]
- Maio, S.; Fasola, S.; Marcon, A.; Angino, A.; Baldacci, S.; Bilò, M.B.; Bono, R.; La Grutta, S.; Marchetti, P.; Sarno, G.; et al. Relationship of long-term air pollution exposure with asthma and rhinitis in Italy: An innovative multipollutant approach. Environ. Res. 2023, 224, 115455. [Google Scholar] [CrossRef]
- Martinez-Nunez, R.T.; Louafi, F.; Sanchez-Elsner, T. The interleukin 13 (IL-13) pathway in human macrophages is modulated by microRNA-155 via direct targeting of interleukin 13 receptor a1 (IL13Ra1). J. Biol. Chem. 2011, 286, 1786–1794. [Google Scholar] [CrossRef]
- Chiba, Y.; Nakazawa, S.; Todoroki, M.; Shinozaki, K.; Sakai, H.; Misawa, M. Interleukin-13 augments bronchial smooth muscle contractility with an upregulation of RhoA protein. Am. J. Respir. Cell Mol. Biol. 2009, 40, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Chiba, Y.; Todoroki, M.; Nishida, Y.; Tanabe, M.; Misawa, M. A novel STAT6 inhibitor AS1517499 ameliorates antigen-induced bronchial hypercontractility in mice. Am. J. Respir. Cell Mol. Biol. 2009, 41, 516–524. [Google Scholar] [CrossRef]
- Chiba, Y.; Sakai, H.; Misawa, M. Augmented acetylcholine-induced translocation of RhoA in bronchial smooth muscle from antigen induced airway hyperresponsive rats. Br. J. Pharmacol. 2001, 133, 886–890. [Google Scholar] [CrossRef]
- Chiba, Y.; Ueno, A.; Shinozaki, K.; Takeyama, H.; Nakazawa, S.; Sakai, H.; Misawa, M. Involvement of RhoA mediated Ca2+ sensitization in antigen-induced bronchial smooth muscle hyperresponsiveness in mice. Respir. Res. 2005, 6, 4. [Google Scholar] [CrossRef]
- Lu, T.X.; Munitz, A.; Rothenberg, M.E. MicroRNA-21 is up-regulated in allergic airway inflammation and regulates IL-12p35 expression. J. Immunol. 2009, 182, 4994–5002. [Google Scholar] [CrossRef]
- McCarthy, J.J.; Esser, K.A. MicroRNA-1 and microRNA-133a expression are decreased during skeletal muscle hypertrophy. J. Appl. Physiol. 2007, 102, 306–313. [Google Scholar] [CrossRef]
- Luo, X.; Lin, H.; Pan, Z.; Xiao, J.; Zhang, Y.; Lu, Y.; Yang, B.; Wang, Z. Down-regulation of miR-1/miR-133 contributes to re-expression of pacemaker channel genes HCN2 and HCN4 in hypertrophic heart. J. Biol. Chem. 2011, 286, 28656. [Google Scholar] [CrossRef]
- Mattes, J.; Collison, A.; Plank, M.; Phipps, S.; Foster, P.S. Antagonism of microRNA126 suppresses the effector function of TH2 cells and the development of allergic airways disease. Proc. Natl. Acad. Sci. USA 2009, 106, 18704–18709. [Google Scholar] [CrossRef]
- Larner-Svensson, H.M.; Williams, A.E.; Tsitsiou, E.; Perry, M.M.; Jiang, X.; Chung, K.F.; Lindsay, M.A. Pharmacological studies of the mechanism and function of interleukin-1b-induced miRNA-146a expression in primary human airway smooth muscle. Respir. Res. 2010, 11, 68. [Google Scholar] [CrossRef]
- Perry, M.M.; Moschos, S.A.; Williams, A.E.; Williams, A.E.; Shepherd, N.J.; Larner-Svensson, H.M.; Lindsay, M.A. Rapid changes in microRNA-146a expression negatively regulate the IL-1binduced inflammatory response in human lung alveolar epithelial cells. J. Immunol. 2008, 180, 5689–5698. [Google Scholar] [CrossRef]
- Williams, A.E.; Larner-Svensson, H.; Perry, M.M.; Campbell, G.A.; Herrick, S.E.; Adcock, I.M.; Erjefalt, J.S.; Chung, K.F.; Lindsay, M.A. MicroRNA expression profiling in mild asthmatic human airways and effect of corticosteroid therapy. PLoS ONE 2009, 4, e5889. [Google Scholar] [CrossRef]
- Antognelli, C.; Gambelunghe, A.; Muzi, G.; Talesa, V.N. Glyoxalase I drives epithelial-to-mesenchymal transition via argpy-rimidine-modified Hsp70, miR-21 and SMAD signalling in human bronchial cells BEAS-2B chronically exposed to crystalline silica Min-U-Sil 5: Transformation into a neoplastic-like phenotype. Free Radic. Biol. Med. 2016, 92, 110–125. [Google Scholar] [CrossRef]
- Antognelli, C.; Gambelunghe, A.; Muzi, G.; Talesa, V.N. Peroxynitrite-mediated glyoxalase I epigenetic inhibition drives apoptosis in airway epithelial cells exposed to crystalline silica via a novel mechanism involving argpyrimidine-modified Hsp70, JNK, and NF-κB. Free Radic. Biol. Med. 2015, 84, 128–141. [Google Scholar] [CrossRef]
- Kumar, M.S.; Erkeland, S.J.; Pester, R.E.; Chen, C.Y.; Ebert, M.S.; Sharp, P.A.; Jacks, T. Suppression of nonsmall cell lung tumor development by the let-7 microRNA family. Proc. Natl. Acad. Sci. USA 2008, 105, 3903–3908. [Google Scholar] [CrossRef]
- Trang, P.; Medina, P.P.; Wiggins, J.F.; Ruffino, L.; Kelnar, L.; Omotola, M.; Homer, R.; Brown, D.; Bader, A.G.; Weidhaas, J.B.; et al. Regression of murine lung tumors by the let-7 microRNA. Oncogene 2010, 29, 1580–1587. [Google Scholar] [CrossRef]
- Oglesby, I.K.; McElvaney, N.G.; Greene, C.M. MicroRNAs in inflammatory lung disease--master regulators or target practice? Respir. Res. 2010, 11, 148. [Google Scholar] [CrossRef]
- Fujita, Y.; Takeshita, F.; Mizutani, T.; Ohgi, T.; Kuwano, K.; Ochiya, T. A novel platform to enable inhaled naked RNAi medicine for lung cancer. Sci. Rep. 2013, 3, 3325. [Google Scholar] [CrossRef]
- Fujita, Y.; Takeshita, F.; Kuwano, K.; Ochiya, T. RNAi Therapeutic Platforms for Lung Diseases. Pharmaceuticals 2013, 6, 223–250. [Google Scholar] [CrossRef]
- Fujita, Y.; Kosaka, N.; Araya, J.; Kuwano, K.; Ochiya, T. Extracellular vesicles in lung microenvironment and pathogenesis. Trends Mol. Med. 2015; Epub ahead of print. [Google Scholar] [CrossRef]
- Fujita, Y.; Yoshioka, Y.; Ito, S.; Araya, J.; Kuwano, K.; Ochiyaet, T. Intercellular communication by extracellular vesicles and their microRNAs in asthma. Clin. Ther. 2014, 36, 873–881. [Google Scholar] [CrossRef]
- Chen, L.L. The biogenesis and emerging roles of circular RNAs. Nat. Rev. Mol. Cell Biol. 2016, 17, 205–211. [Google Scholar] [CrossRef]
- Liu, C.X.; Li, X.; Nan, F.; Jiang, S.; Gao, X.; Guo, S.K.; Xue, W.; Cui, Y.; Dong, K.; Ding, H.; et al. Structure and degradation of circular RNAs regulate PKR activation in innate immunity. Nat. Rev. Mol. Cell Biol. 2016, 17, 205–211. [Google Scholar] [CrossRef]
- Allegra, A.; Cicero, N.; Tonacci, A.; Musolino, C.; Gangemi, S. Circular RNA as a Novel Biomarker for Diagnosis and Prognosis and Potential Therapeutic Targets in Multiple Myeloma. Cancers 2022, 14, 1700. [Google Scholar] [CrossRef]
- Shen, S.; Wu, Y.; Chen, J.; Xie, Z.; Huang, K.; Wang, G.; Yang, Y.; Ni, W.; Chen, Z.; Shi, P.; et al. CircSERPINE2 protects against osteoarthritis by targeting miR-1271 and ETS-related gene. Ann. Rheum. Dis. 2019, 78, 826–836. [Google Scholar] [CrossRef]
- Wesselhoeft, R.A.; Kowalski, P.S.; Anderson, D.G. Engineering circular RNA for potent and stable translation in eukaryotic cells. Nat. Commun. 2018, 9, 2629. [Google Scholar] [CrossRef] [PubMed]
- Meng, Q.; Wang, J.; Cui, J.; Li, B.; Wu, S.; Yun, J.; Aschner, M.; Wang, C.; Zhang, L.; Li, X.; et al. Prediction of COPD acute exacerbation in response to air pollution using exosomal circRNA profile and Machine learning. Environ. Int. 2022, 168, 107469. [Google Scholar] [CrossRef]
- Zhao, J.; Xia, H.; Wu, Y.; Lu, L.; Cheng, C.; Sun, J.; Xiang, Q.; Bian, T.; Liu, Q. CircRNA_0026344 via miR-21 is involved in cigarette smoke-induced autophagy and apoptosis of alveolar epithelial cells in emphysema. Cell Biol. Toxicol. 2021. online ahead of print. [Google Scholar] [CrossRef]
- Li, Z.; Ma, J.; Shen, J.; Chan, M.T.V.; Wu, W.K.K.; Wu, Z. Differentially expressed circular RNAs in air pollution-exposed rat embryos. Environ. Sci. Pollut. Res. Int. 2019, 26, 34421–34429. [Google Scholar] [CrossRef] [PubMed]
- Ağaç, D.; Gill, M.A.; Farrar, J.D. Adrenergic Signaling at the Interface of Allergic Asthma and Viral Infections. Front. Immunol. 2018, 9, 736. [Google Scholar] [CrossRef]
- Guttman, M.; Amit, I.; Garber, M.; French, C.; Lin, M.F.; Feldser, D. Chromatin signature reveals over a thousand highly conserved large non- coding RNAs in mammals. Nature 2009, 458, 223–227. [Google Scholar] [CrossRef]
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martin, D.; Merkel, A.; Knowles, D.G.; et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 2012, 22, 1775–1789. [Google Scholar] [CrossRef] [PubMed]
- Batista, P.J.; Chang, H.Y. Long noncoding RNAs: Cellular address codes in development and disease. Cell 2013, 152, 1298–1307. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Allegra, A.; Mania, M.; D’Ascola, A.; Oteri, G.; Siniscalchi, E.N.; Avenoso, A.; Innao, V.; Scuruchi, M.; Allegra, A.G.; Musolino, C.; et al. Altered Long Noncoding RNA Expression Profile in Multiple Myeloma Patients with Bisphosphonate-Induced Osteonecrosis of the Jaw. Biomed. Res. Int. 2020, 2020, 9879876. [Google Scholar] [CrossRef]
- He, F.; Wang, N.; Yu, X.; Zheng, Y.; Liu, Q.; Chen, Q.; Pu, J.; Li, N.; Zou, W.; Li, B.; et al. GATA3/long noncoding RNA MHC-R regulates the immune activity of dendritic cells in chronic obstructive pulmonary disease induced by air pollution particulate matter. J. Hazard Mater. 2022, 438, 129459. [Google Scholar] [CrossRef]
- Lin, H.; Zhang, X.; Feng, N.; Wang, R.; Zhang, W.; Deng, X.; Wang, Y.; Yu, X.; Ye, X.; Li, L.; et al. LncRNA LCPAT1 mediates smoking/ particulate matter 2.5-induced cell autophagy and epithelial-mesenchymal transition in lung cancer cells via RCC2. Cell Physiol. Biochem. 2018, 47, 1244–1258. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Furci, F.; Allegra, A.; Tonacci, A.; Isola, S.; Senna, G.; Pioggia, G.; Gangemi, S. Air Pollution and microRNAs: The Role of Association in Airway Inflammation. Life 2023, 13, 1375. https://doi.org/10.3390/life13061375
Furci F, Allegra A, Tonacci A, Isola S, Senna G, Pioggia G, Gangemi S. Air Pollution and microRNAs: The Role of Association in Airway Inflammation. Life. 2023; 13(6):1375. https://doi.org/10.3390/life13061375
Chicago/Turabian StyleFurci, Fabiana, Alessandro Allegra, Alessandro Tonacci, Stefania Isola, Gianenrico Senna, Giovanni Pioggia, and Sebastiano Gangemi. 2023. "Air Pollution and microRNAs: The Role of Association in Airway Inflammation" Life 13, no. 6: 1375. https://doi.org/10.3390/life13061375
APA StyleFurci, F., Allegra, A., Tonacci, A., Isola, S., Senna, G., Pioggia, G., & Gangemi, S. (2023). Air Pollution and microRNAs: The Role of Association in Airway Inflammation. Life, 13(6), 1375. https://doi.org/10.3390/life13061375








