Abstract
The current digitization of industrial processes is leading to the development of smart machines and smart applications in the field of engineering technologies. The basis is an advanced sensor system that monitors selected characteristic values of the machine. The obtained data need to be further analysed, correctly interpreted, and visualized by the machine operator. Thus the machine operator can gain a sixth sense for keeping the machine and the production process in a suitable condition. This has a positive effect on reducing the stress load on the operator in the production of expensive components and in monitoring the safe condition of the machine. The key element here is the use of a suitable classification model for data evaluation of the monitored machine parameters. The article deals with the comparison of the success rate of classification models from the MATLAB Classification Learner App. Classification models will compare data from the frequency and time domain, the data source is the same. Both data samples are from real measurements on the CNC vertical machining center (CNC-Computer Numerical Control). Three basic states representing machine tool damage are recognized. The data are then processed and reduced for the use of the MATLAB Classification Learner app, which creates a model for recognizing faults. The article aims to compare the success rate of classification models when the data source is a dataset in time or frequency domain and combination.
1. Introduction
The current development of Industry 4.0 brings, in addition to the digitization of production, the collection of large amounts of data, which are suitable for the use of artificial intelligence methods. Particularly in the field of technical diagnostics, this opens possibilities for the early detection state of an object (machine) based on the measured data, which are obtained from different standard diagnostic methods (e.g., vibrodiagnostics, thermodiagnostics or electrodiagnostics) and it is possible to use special methods of technical diagnostics such as measuring the magnetic field of the machine [1]. It is therefore a multi-parameter diagnostic that can more accurately detect, identify, and localize the emerging fault, thanks to a combination of input data from different areas of diagnostics.
Vibrations are an important carrier of information about the condition of rotating equipment and this fact is commonly used in the field of vibrodiagnostics. This article focuses on the machining center and here it is possible to use the vibrodiagnostic system to detect other information such as machining quality or tool damage. Thus, unexpected states can be detected by a vibrodiagnostic signal, which may indicate a fault or a safety risk. However, these signals are unexpected and a simple evaluation mechanism cannot be designed, for example, with the detection of frequent failures of rotary machines such as misalignment, unbalance, or bearing damage. In these cases, it is appropriate to use self-learning artificial intelligence methods, i.e., they can learn to recognize a fault condition from historical data for a specific machine or specific machining operations.
In the development process of smart machine tools capable of operating within the framework of industry 4.0, it is necessary to implement risk analysis already during the machine design stage not only in terms of potential damage to the health of the machine operator (according to ISO 12100 [2]), but also in terms of quality management and monitoring of the capability of the manufacturing process on the machine tool (according to ISO 9001 [3]). We can apply the good technical practice as described in IEC 31010 [4] and then implement the necessary diagnostic tools into the machine design or modify this machine design so that the measurement uncertainty of the sensor system (according to ISO 3534-1 [5]) would not affect the interpretation of the measured data and the subsequent decision-making processes in a negative way. A typical example could be early detection of tool damage and subsequent bringing the machine to a safe state. Breaking and ejecting the carbide insert from a tool holder can cause damage to the transparent guard of the machine’s working area, and thus lead to a subsequent loss of its safety function, or it may cause irreversible damage to the machined surface. Therefore, by early detection of a machine fault, we can protect the health of the operator as well as reduce the financial losses caused by poor production or increased wear of the smart machine’s spindle.
This article will, therefore, compare classification methods that can detect a specific fault from real data, namely a missing insert on a machining tool, the absence of which could have occurred due to tool damage. A method will be used where predictors for machine learning will be evaluated in advance and their choice is therefore up to the expert. The correct choice of predictors has a major impact on the classification success rate. The authors of the work have already dealt with the issue of evaluating vibrodiagnostic signals in [6,7,8]. The idea of using artificial intelligence methods in diagnostics is described in other publications such as in [9,10,11,12], where artificial intelligence methods are used for the recognition of gearbox faults. Also, in the paper [13] authors focus on the classification using neural networks in the planetary gear. The article [14] uses artificial intelligence methods to recognize the shape of a rotary machine shaft’s orbit; it is a picture classification according to the shape of shaft orbit. The aim of this research is to determine a fast method (real-time evaluation) for the timely evaluation of a failure that would reduce the safety of the machine. Thus, the primary goal is not to diagnose the condition of the machine, such as a bearing or gearbox failure, and to predict residual life, but to detect failures related to machine safety and to warn the operator immediately.
This article has several goals and they are:
- Verify that the missing insert on the tool is detectable on the vibrodiagnostic signal measured on the machine spindle.
- Select and compare predictors from time and frequency domains.
- Effect spindle speeds on the classification success rate.
- Select and compare the most successful classification methods
- Evaluate detection success rate of fault-free state
It is assumed that the process of data collection and evaluation takes place on an IPC (Industrial PC) computer near the machine (for example IPC SIMATIC PC-based, where one IPC will be prepared for each CNC machine), and the learning and model generated process will take place on a server containing commercial software such as MATLAB (server will be used for many CNC machines). Learning and exporting the model is an important operation and will take place under strict supervision. The created MATLAB model will be exported for the use of individual IPCs without MATLAB.
The Python environment also includes Machine Learning tools such as scikit [15] or Artap used in [16], however in this case the ambitions are to be used in a safety environment and the authors there rely on the professional Matlab environment. In the future, the above-mentioned toolboxes will be used and tested to evaluate the condition of the machine.
Python environment is used for signal processing, the MATLAB and Classification Learner App [17] are used for data classification.
2. Experimental Procedure
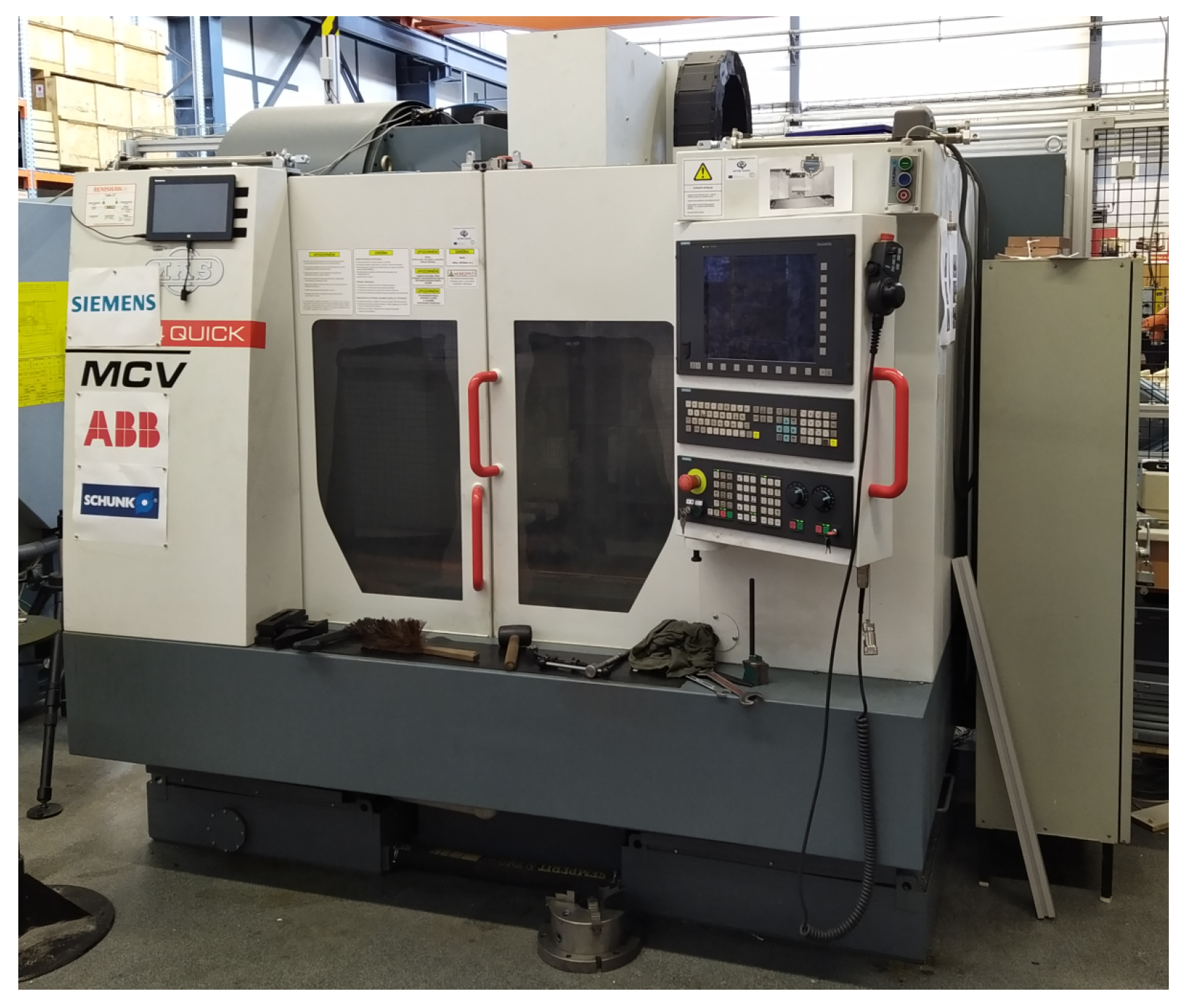
The data were obtained from the machine MCV 754 QUICK (Figure 1), it is a three-axis CNC (Computer Numerical Control) machining centre, more information can be obtained on the manufacturer’s website [18]. The obtained data are from the measuring instrument Microlog CMXA48 and the measurement parameters are in the Table 1.
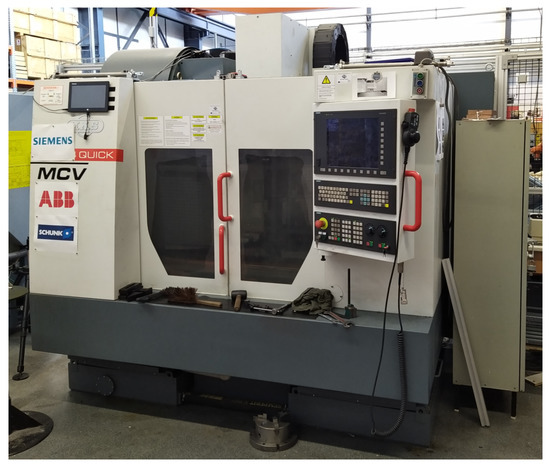

Figure 1.
CNC vertical machining center MCV 754.

Table 1.
Measurement parameters of Microlog CMXA48.
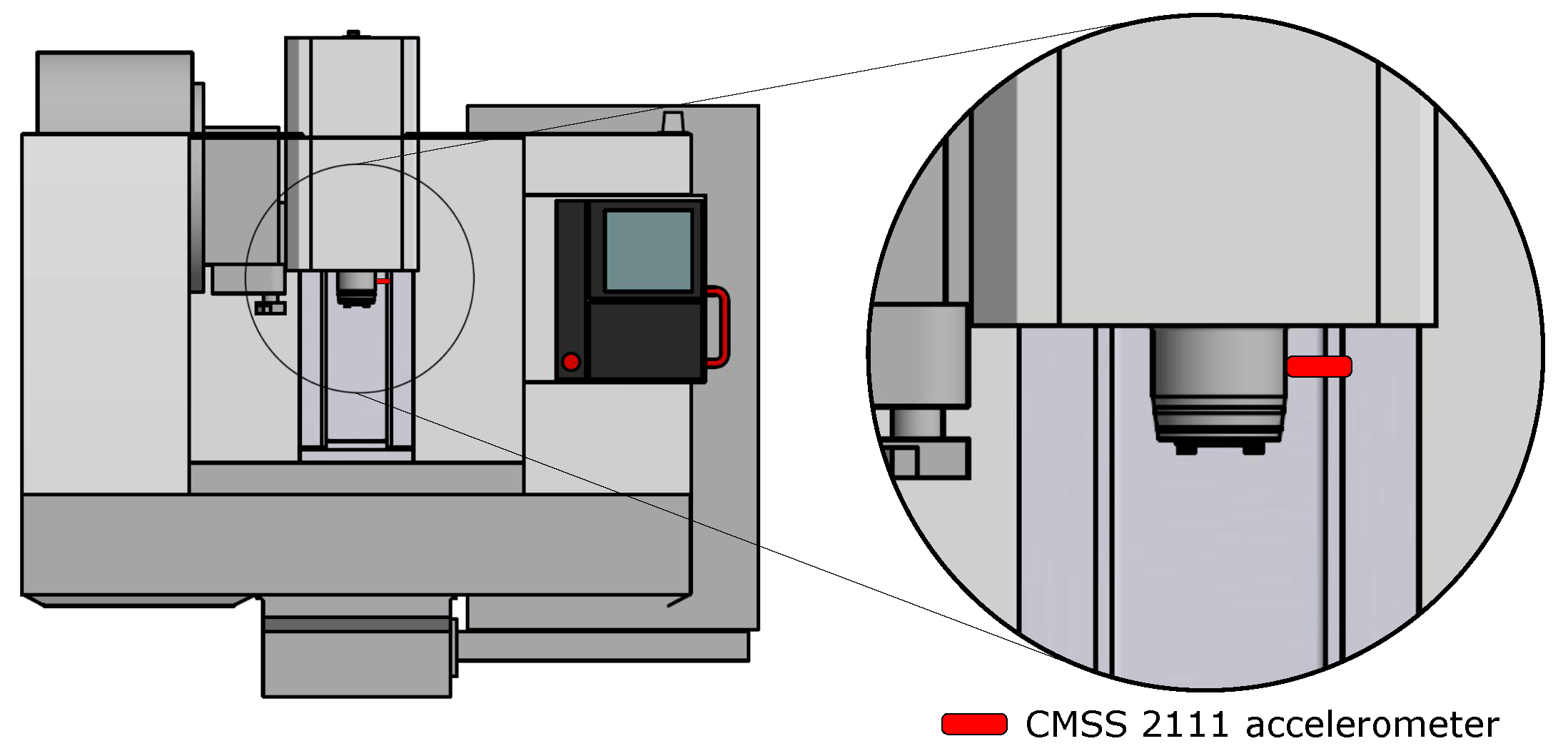
The instrument Microlog uses a CMSS 2111 accelerometer sensor that has been attached to the spindle with a magnet. The position of the sensor on the machine is shown in Figure 2. Used tool H490 F90AX D040-4-16-12 has the possibility of changing the teeth (inserts), which are fixed with a screw, as shown in Figure 3. Three states/classes were chosen for the experiment, namely:
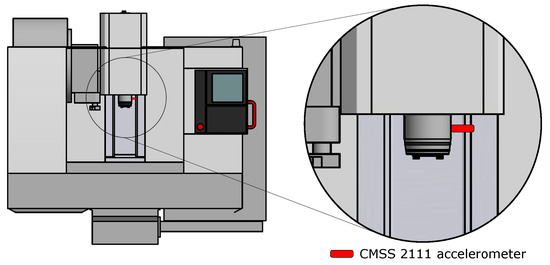
Figure 2.
Placement of the CMSS 2111 accelerometer to the machine MCV 754.
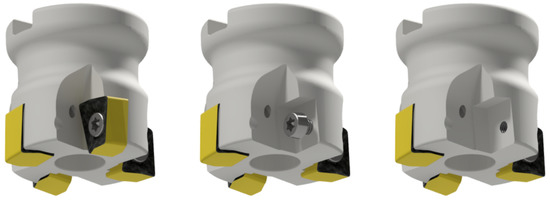

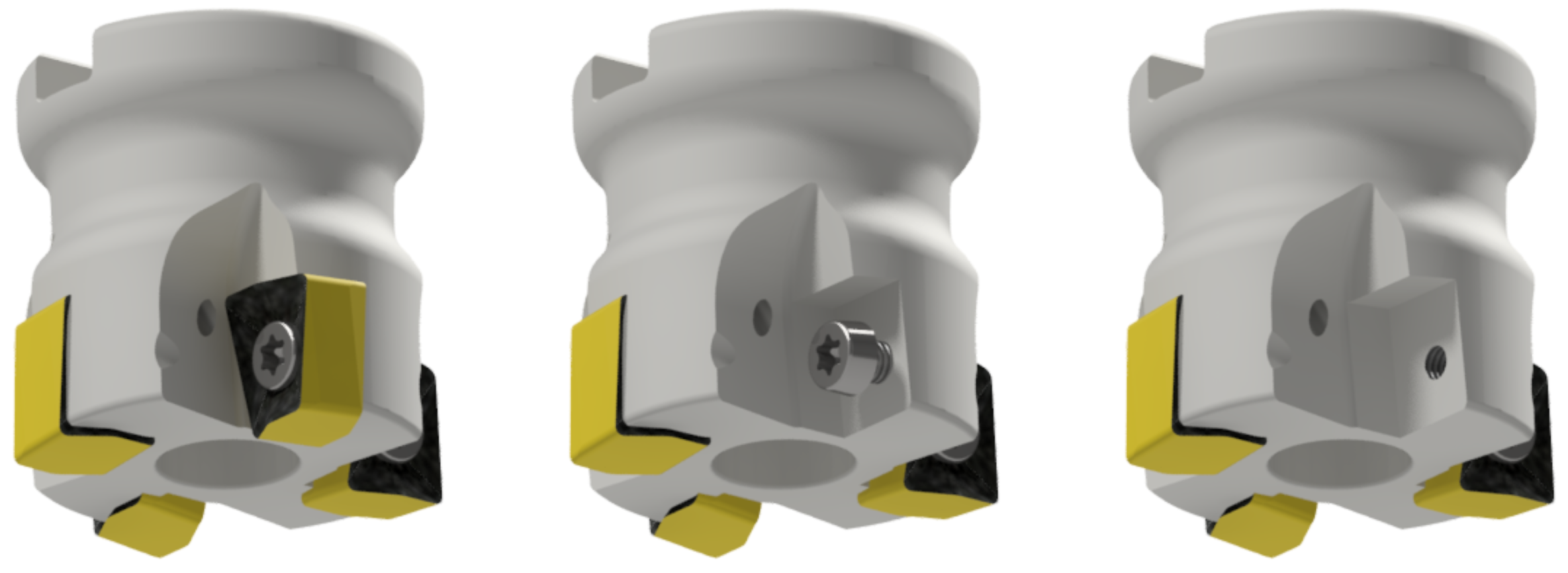
Figure 3.
The tool H490 F90AX D040-4-16-12 in fault-free state-Class 0 (left), without the insert Class 1 (middle) and without the insert and without the screw Class 2 (right).
Expected measurable changes are caused by the unbalance of the tool when this vibration is transmitted to the spindle. The failure of unbalance is characterized by an increase in vibrations in the region of the first harmonic frequency. The unbalance is described in the [19,20,21] and is divided into a static unbalance and a dynamic unbalance.
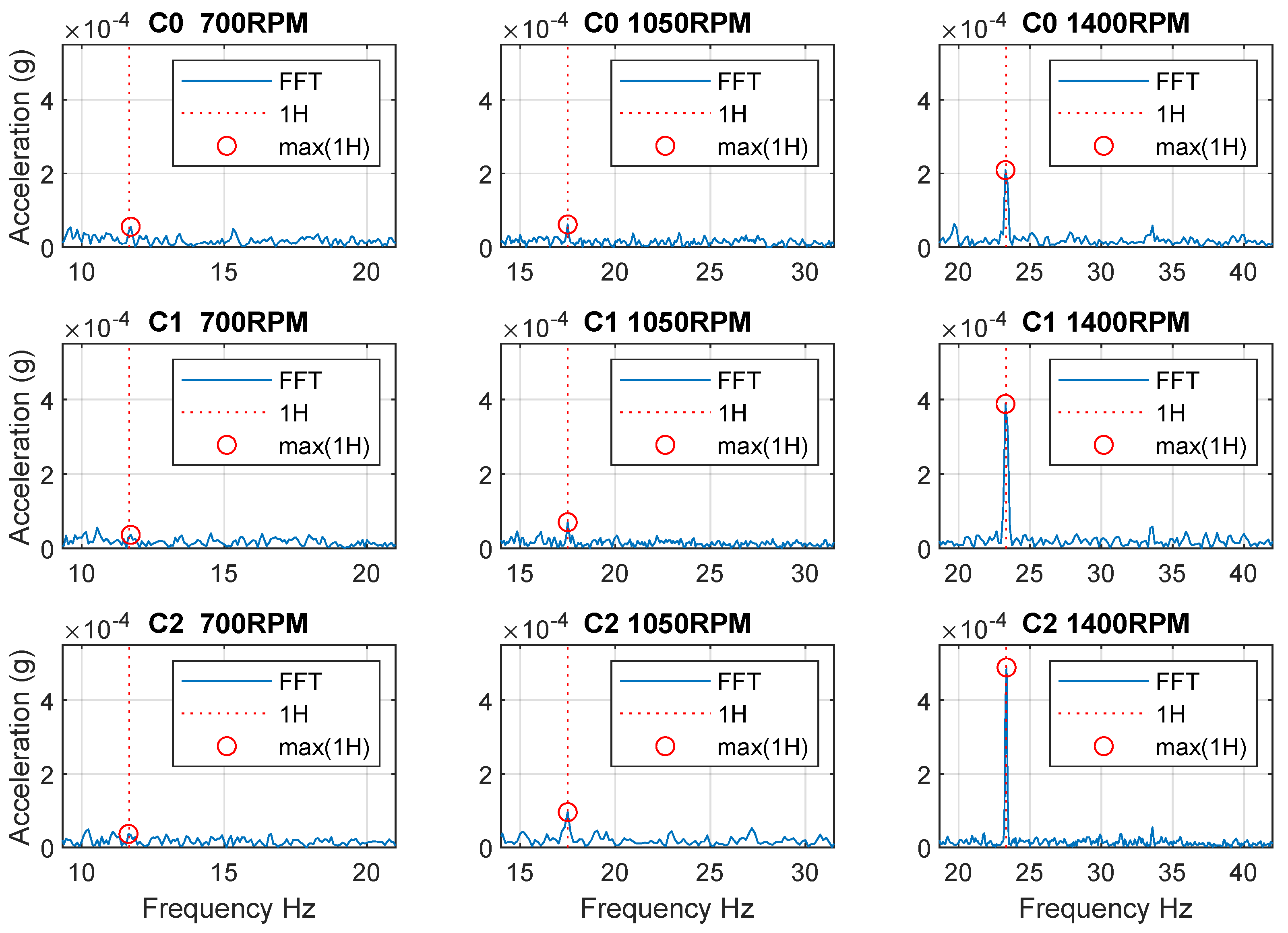
The first experiment verified that the missing insert was reflected in spindle vibrations at 700, 1050, and 1400 RPM. The results are shown in Figure 4, which shows the area around the first harmonic frequency. The aim of this experiment was to determine at which spindle speeds the manifestations are visible in the frequency spectrum and which speeds are suitable for data collection in a machine learning dataset. From the result, it is evident that at the speed 700 RPM the effect of the unbalances in the frequency domain is insignificant, and therefore undetectable. For this reason, speeds were selected to obtain a dataset from 1000 to 1500 RPM.
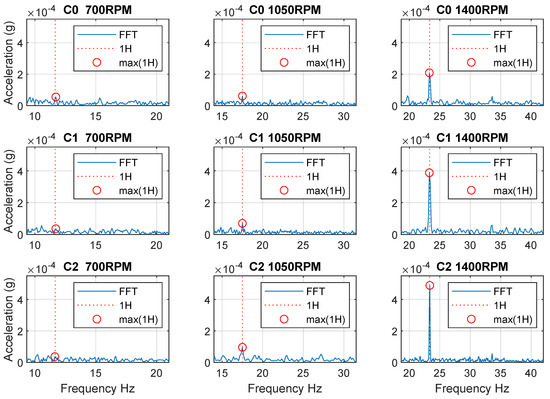
Figure 4.
The first test of the effect of the fault in frequency domain (C0−C2 are Class 0−Class 2, 1H—first harmonic frequency, max(1H)—maximum of the first harmonic frequency).
2.1. Dataset Obtain
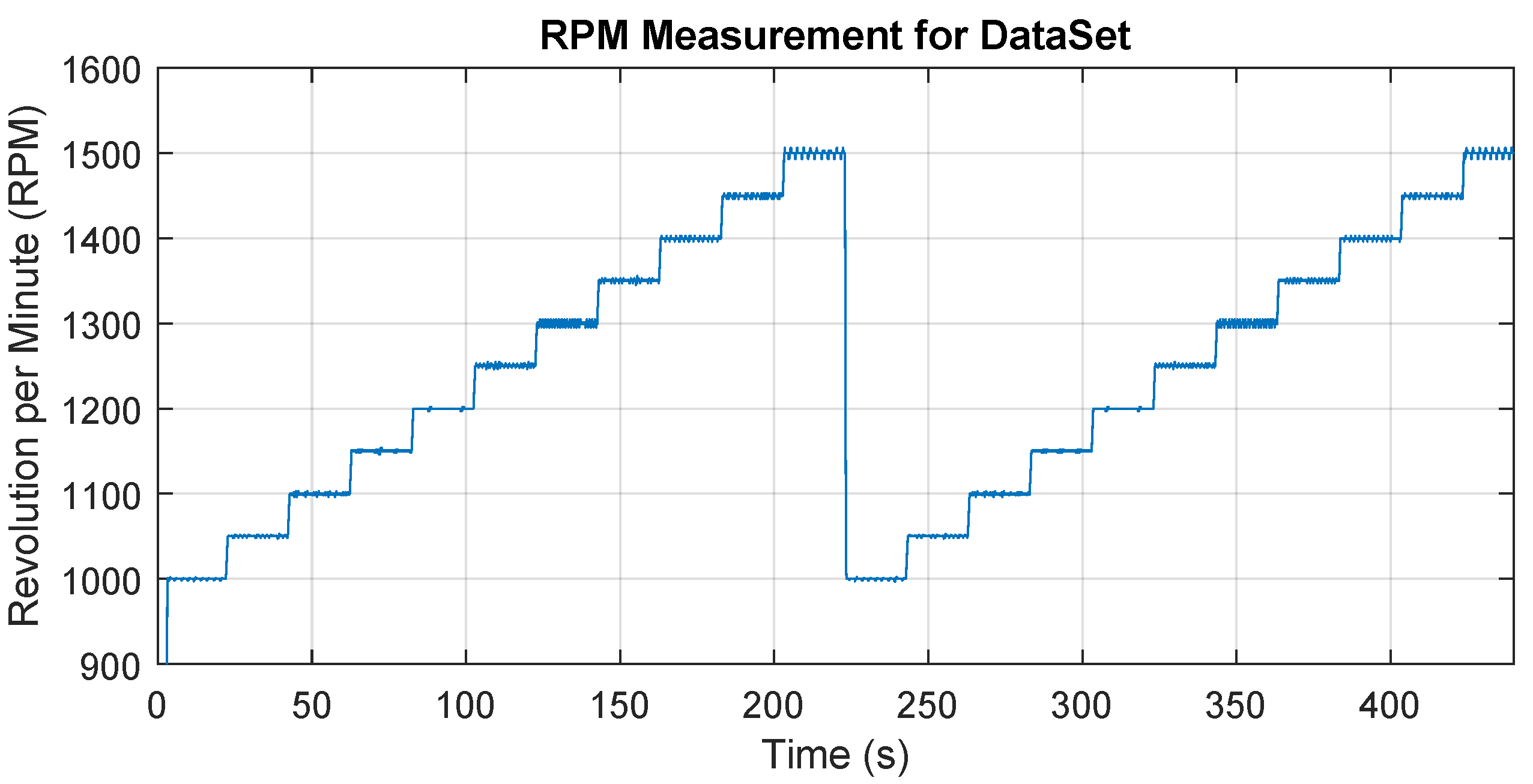
For each state/class, a series of data was measured for different operating speeds that are within the recommended speed range for the tool. Specifically, the speed was from 1000 RPM to 1500 RPM after the step of 50 RPM in two repetitions and 20 s is maintained at each speed level. The total time is therefore 20 × 11 × 2 = approx. 440 s for one state (class). The speed setting was programmed and started for each state in the same way, Figure 5 shows the time record of the speed change over time. The aim is to obtain data at a wide range of operating speeds. The data were captured in a recorder mode, i.e., one long time record was obtained, which was later processed into individual samples, which formed a dataset for machine learning.
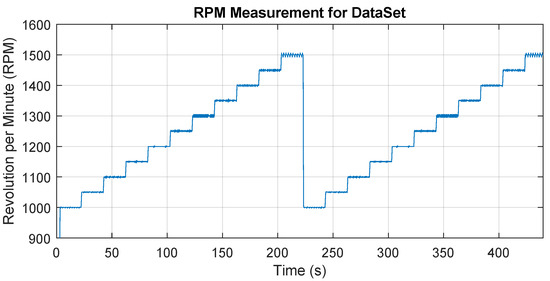
Figure 5.
The variation of the spindle speed over time during the dataset obtain.
2.2. Predictors
Predictors are arbitrary parameters, and their values (or combinations thereof) determine the properties of the class being searched. If these parameters did not adequately reflect the properties of the class, the data classification would not be successful. An example of the use of a predictor is the known area for image data processing. The input is information about individual pixels and at full HD resolution () it is more than pixels and each pixel contains information about three colours (RGB). This amount of data is disproportionately large and contains a lot of useless information, so there is a need to reduce data such as reducing the number of colours, edge detection, and more. The outputs of this reduction are predictors, which are the input for artificial intelligence, and based on their values (and their combinations) it is possible to classify individual classes. The properties of predictors should be:
- Clearly describe the class properties (state)-improved achievement classification
- Low computational complexity-speeds up the predictor calculation
- Low number of predictors-speeds up the learning and evaluation process
The input data in vibrodiagnostics are time domain signals. In this particular case, the sampling frequency is 2560 Hz (sampling period is approximately 0.4 ms), which means 2560 values every second of recording for one axis. It is, therefore, necessary to reduce this data flow and evaluate suitable predictors from it, which will be the input for artificial intelligence. The aim is to select appropriate predictors and compare their impact on the success of data classification.
2.3. Data Processing and Selection of Predictors
In this article, a method is chosen that classifies data into individual classes based on predictors. The selection and calculation of predictors are performed before the classification of data and the correct choice of these predictors has a significant impact on the results of the classification and it is necessary to know the issue and have experience with similar experiments. The author of the article has already dealt with this issue in [7,8] and verified which predictors are applicable for a similar type of problem. This article will compare the classification successes for time and frequency domain predictors and their combinations. It was also necessary to choose the length of the time signal and divide the resulting samples. The times of 0.25 s and 0.5 s were chosen and again the advantages and disadvantages of these lengths were compared. The following predictors were used.
Predictors from the time domain:
- Speed-Current RPM during one sample
- RMS-Root Mean Square of acceleration amplitudes from one sample
- STD-The standard deviation of the acceleration amplitudes from a single sample
- PCA-Principal Component Analysis realized with scikit learn library R2 (with parameters n_components = 1 and explained_variance_ = 0) [22]
The result is a vector that contains 4 predictors for each sample (RPM, RMS, STD and PCA); the values in this vector are further labelled as “T predictors”.
Frequency domain predictors are based on FFT (Fast Fourier Transform) frequency analysis performed using the NumPy library “Standard FFTs” [23] and a range of 4 to 100 Hz is used. The minimum value of 4 Hz is determined by the minimum measuring range of the sensor used. The maximum value of 100 Hz is the maximum value of the occurrence of the first harmonic in all operating speeds of the tool and any tooth frequency. The maximum operating speed is 1432 RPM, the maximum tooth frequency is therefore (1432 × 4)/60 = 95.47 Hz, therefore the maximum frequency of 100 Hz was chosen.
- Speed-Current RPM during one sample
- Spectrum energy (eF)-calculated as for the 4–100 Hz range, where are amplitudes of spectral lines.
- Af(i)-the sequence of amplitudes of individual spectral lines for frequencies 4–100 Hz
The result is a vector whose size varies according to the length of the signals used, i.e., division into individual samples. For example, if the signal is 0.25 s long and has a sampling frequency of 2560 Hz, the situation is as follows:
For range 4–100 Hz:
For a 0.25 s signal (t1), the output will be a vector containing 26 (24 + 1 + 1) predictors for each data sample. The disadvantage of such a short signal is the frequency resolution () of 4 Hz, where this value may be insufficient. Furthermore, signals with a length of 0.5 s will be compared. There is the following situation
For range 4–100 Hz:
For a 0.5 s signal (t2), the output will be a vector containing 50 (48 + 1 + 1) predictors for each data sample. With this signal distribution, we get a frequency resolution () of 2 Hz, which is more suitable than the previous case, but we get only half of the samples for machine learning from the limited recording.
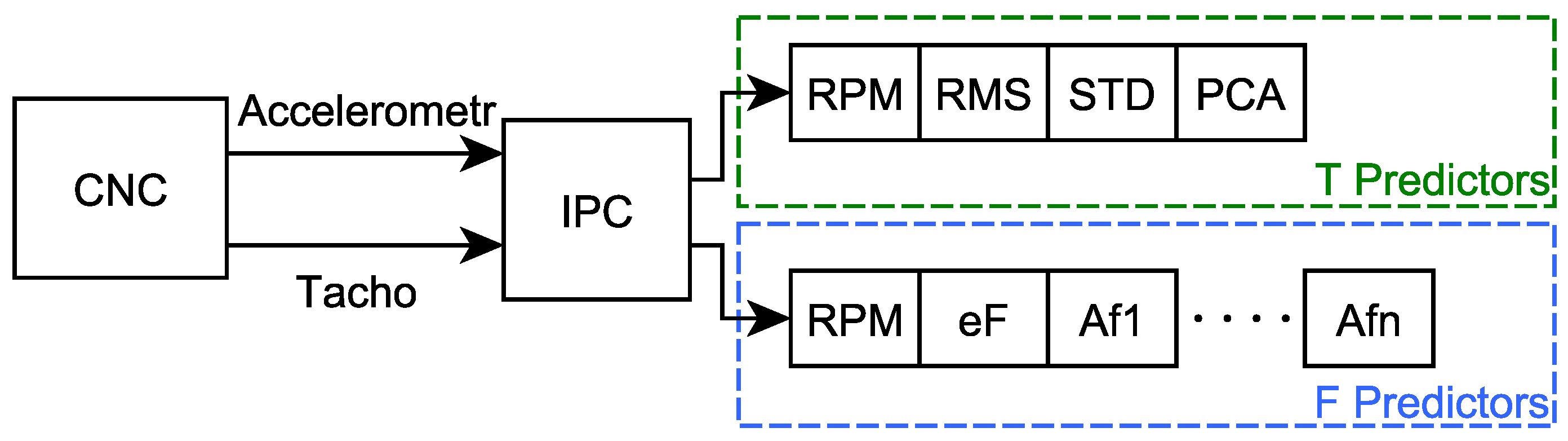
The resulting vector for frequency domain predictors contains the measured RPM value, the calculated Spectrum energy value (labeled as eF) and the sequence of amplitudes of individual spectral lines (Af1 to Af24 for t1 and Af1 to Af48 for t2), calculated using the FFT function, see Figure 6. The values in this vector are further labelled as “F predictors”.
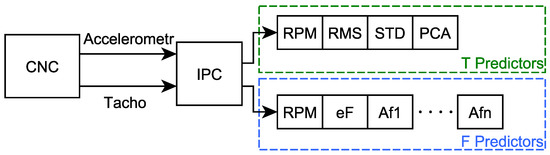
Figure 6.
Scheme of predictors calculation using IPC (“eF”—Spectrum energy, “Af1” to “Afn”—the sequence of amplitudes of individual spectral lines).
Both variants will be compared with each other.
The detailed data processing procedure is as follows:
Data processing (outside MATLAB)
- Combination of vibration channel and tacho channel data
- Splitting the signal according to the selected time (t1 = 0.25 s or t2 = 0.5 s)
- Checking whether the change is not greater than 10 RPM during the duration (t1 or t2) (elimination of transients)
- Sorting the signal into individual classes
- Predictor calculation, for each sample (T-time domain, F-Frequency domain or FT-all predictors)
- Saving predictors to a file according to individual classes (preparation for MATLAB)
Processed in MATLAB:
- Random mixing and merging of data of individual classes into one dataset
- Evaluating in the environment “Classification learner app”
- Generating and saving the most successful model
Verification of the generated model:
- Randomly selected n samples from each class (selected n = 100 per class)
- Accuracy and TPR (True positive rate) and FNR (False negative rate) evaluated
- Results displayed and saved as the Confusion matrix
3. Classification Methods
Classification Learner App (CL) compared different classification methods to determine which methods are appropriate for the problem. This application allows you to compare different methods and then select and create a model containing the selected method (chosen were the most successful). The individual methods are described in more detail in [24]. Datasets were tested on all methods that CL allows and their success is in a Table 2, Table 3 and Table 4. The number of repetitions of the learning process was 10 for each model, and the interval estimate was calculated using “The t Confidence interval” [25] with alpha = 0.05. Furthermore, the hyperparametric optimization provided by Matlab was verified. Each winning model was subjected to hyperparametric optimization with the same dataset, and these results are shown in Table 5. Due to the time consuming, only one repetition was performed, the average improvement of the prediction accuracy with hyperparametric optimization is only 0.55%. Models without hyperparametric optimization were used for further validation.

Table 2.
Resulting success rate of classification of individual methods for experiments with sample time t1 (0.25 s). The best results are marked in bold.

Table 3.
Resulting success rate of classification of individual methods for experiments with sample time t2 (0.5 s). The best results are marked in bold. Note: F-model failed (one or more of the classes have singular covariance matrices).

Table 4.
Resulting success rate of classification of individual methods for experiments with combined predictors. The best results are marked in bold. Note: F-model failed (one or more of the classes have singular covariance matrices).

Table 5.
Results of prediction accuracy after hyperparametric optimization, “Hyperparameter opt”-best model with hyperparameter optimization, “Difference”-Difference between result with and without hyperparameter optimization, “Avr. difference”-average difference from all experiments.
Briefly, these are the following classification methods:
3.1. Classification Trees
Binary decision trees for multiclass learning [26]. This method is easy to interpret and fast for data prediction, has low memory requirements, but may not achieve sufficient prediction accuracy. Decision making takes place through a tree structure from the beginning (root) to the final class (leaves). The method was probably first published by J. Ross Quinlan in 1975. The tested variants are:
- Fine tree
- Medium tree
- Coarse tree
3.2. Discriminant Analysis
Regularized linear and quadratic discriminant analysis [27]. This method is easy to interpret and fast for data prediction for large datasets. Using “Quadratic Discriminant” increases memory requirements. It is one of the methods of N-dimensional statistical analysis (multivariate statistical analysis), where, based on the decision rule, objects are divided into groups according to probability densities. The method was probably first published by Ronald Fisher in 1936. The tested variants are:
- Linear discriminant
- Quadratic discriminant
3.3. Naive Bayes
Naive Bayes model with Gaussian, multinomial, or kernel predictors [28]. This method is easy to interpret and achieves good results for multi-class classification. Prediction speed is slow (medium for Gaussian), memory usage is a medium (small for Gaussian). The method is based on Bayes’ theorem on conditional probabilities and class affiliation testing. Bayes’ Theorem was published by Thomas Bayes in 1736 and the classifier was probably first published in 1960. The tested variants are:
- Gaussian Naive Bayes
- Kernel Naive Bayes
3.4. Support Vector Machine Classification
Support vector machines (SVM) for binary or multiclass classification [29].
This method is difficult to interpret (not for the linear variant). If there are more than two classification classes, then the problem must be divided into several partial binary problems, therefore, it has high memory requirements and is therefore slow in multi-class classification. The principle of the method is to find a hyperplane that separates individual objects based on class affiliation. The method was probably first published by Vladimir N. Vapnik and Alexey Ya. Chervonenkis in 1963. The tested variants are:
- Linear SVM
- Quadratic SVM
- Cubic SVM
- Fine Gaussian SVM
- Medium Gaussian SVM
- Coarse Gaussian SVM
3.5. Nearest Neighbors
k nearest neighbours (KNN) classification using Kd-tree search [30]. This method is difficult to interpret and the success of the method decreases with the growth of classes. Prediction speed and memory requirements are media. The principle of the method consists of finding the nearest neighbours of objects in N-dimensional space with the properties of the class. The method was probably first published by Evelyn Fix and Joseph Hodges in 1951. The tested variants are:
- Fine KNN
- Medium KNN
- Coarse KNN
- Cosine KNN
- Cubic KNN
- Weighted KNN
3.6. Classification Ensembles
Boosting, random forest, bagging, random subspace, and ECOC (Error-Correcting Output Codes) ensembles for multiclass learning [31]. This method combines several simple methods into one powerful model. Interpretability is difficult, however, the prediction speed is from fast to medium (depending on the combination used) and memory usage is low (medium for Subspace KNN). The tested variants are:
- Boosted Trees
- Bagged Trees
- Subspace Discriminant
- Subspace KNN
- RUSBoosted Trees
4. Comparison of Classification Results
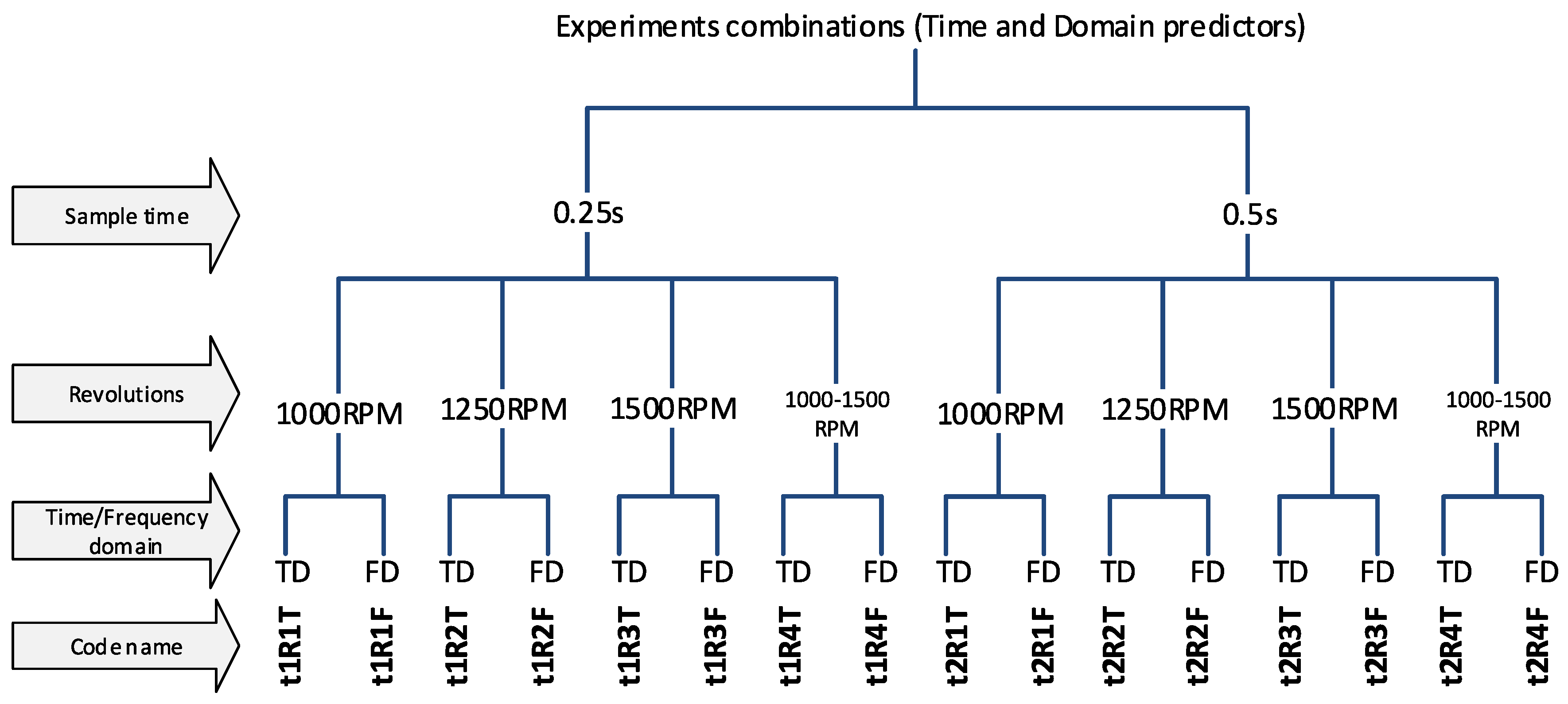
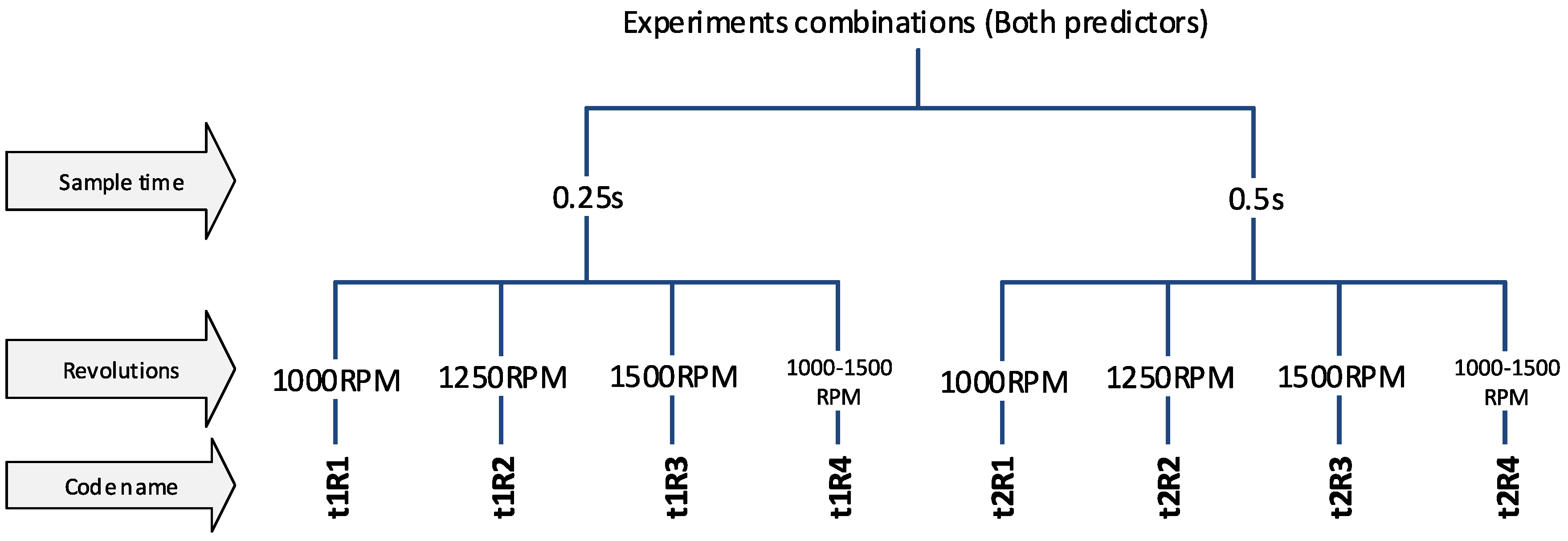
The experiments were divided into several variants, according to the properties of the input data, to be able to compare results of the classification and determine which parameters/predictors and under what conditions they affect the success of the classification. The first step is to use a sample length of 0.25 s and 0.5 s (t1 and t2). The choice of this parameter affects the number of predictors-(frequency spectrum resolution) and also the number of samples for the dataset because the measured data have a limited length (for more details, see Section 2 in Figure 5). Another division will focus on determining the effect of spindle speed variability. Datasets labeled R1-R3 have a constant speed (1000, 1250, 1500 RPM) and experiments labelled R4 have a dataset with combinations of all speeds, so it is a random mixing of sample R1 to R3 regardless of speed. And the last experiments contain a combination of all predictors, ie. from the time and frequency domain (without T or F marking). A unique code name was chosen for each experiment so that it was possible to refer to a specific variant in the text. The overall division of the experiment is shown in Figure 7 and Figure 8.
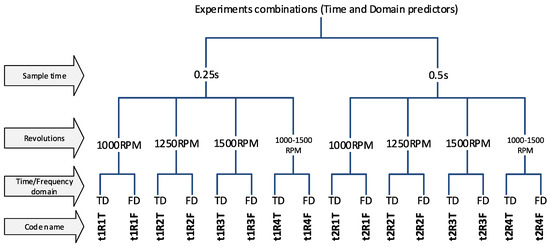
Figure 7.
The code names of experiments.
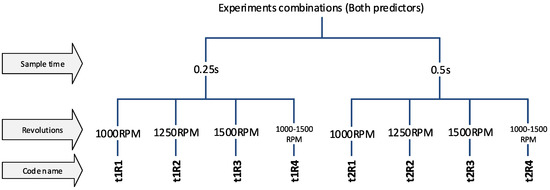
Figure 8.
The code names of experiments (the combined predictors).
5. Verification of Resulting Models
The following section compares the success of each method for all variants of experiments. The goal was to find out what methods are suitable for the given predictors, and how the success of the classification differs with the spindle speed, and how successful the time and frequency domain predictors are.
The accuracy of the models is compared and is determined as follows. In the case of selecting the most successful model, an evaluation method was used, which the Matlab Learner app calls “Accuracy” (in the generated code as “validationAccuracy”)
where:
—success rate of classification (ideal value is 100%)
—True prediction from all class
—All (true and false) prediction from all class
To verify the model (Figures 10 and 11) the method used by Matlab was used, namely “True positive rate”-TPR, where each class was evaluated separately; the remaining fields on the lines in CM (Confusion matrix) are False negative rate-FNR for the class. The line amount is therefore 100%.
where:
—True positiv rate (ideal value is 100%)
—True (correct) prediction per class
—All (true + false) prediction per class
where:
-False positiv rate (ideal value is 0%)
—Predicted class (incorrect) for True class
—All (true + false) prediction per class
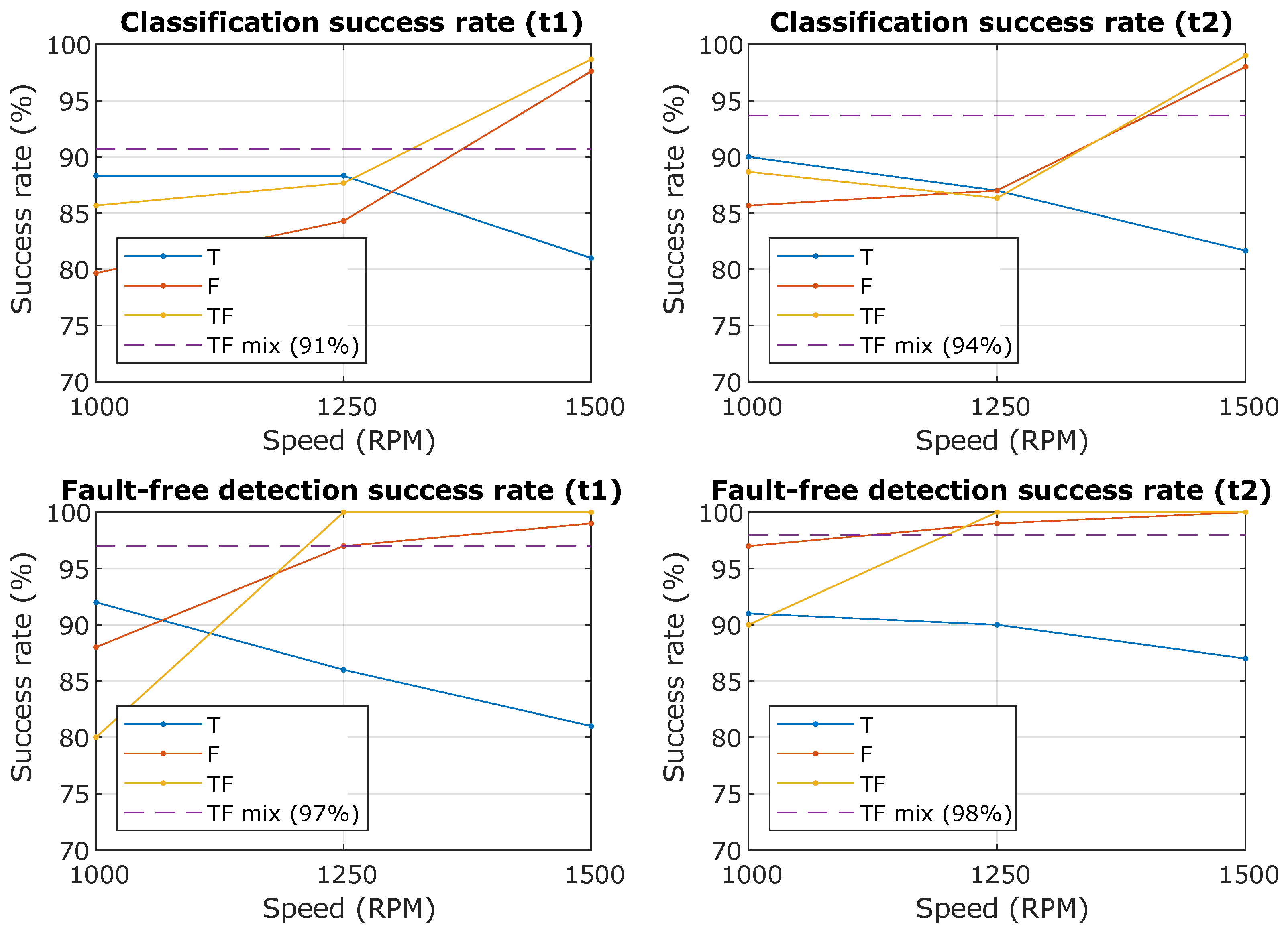
In Table 2, Table 3 and Table 4 are the results of individual methods and it is clear that some methods are more successful for predictors from the time domain than predictors from the frequency domain. More interesting is the graphic display in Figure 9, where the success rate of the classification depends on the spindle speed. Descriptions in the graph mean “Classification success rate” is the Accuracy () for the created models at the number of a random selection of 100 samples per class and “Fault-free detection success rate” is the TPR (True positive rate) for class 0, i.e., faultless condition.
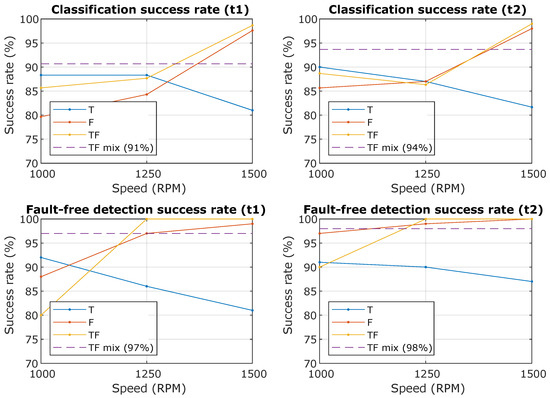
Figure 9.
The resulting successes rate of the classification and the success rate of the prediction of the fault-free state. The waveforms from the graph are T—Time domain predictors only, F—Frequency domain predictor only, TF—All predictors together, TF mix—All predictors together for all RPM (from 1000 to 1500 with step 50).
In the case of predictors from the frequency domain, the success rate increases with spindle speed, because the effect of the unbalances fault is more noticeable with increasing spindle speed, but in the case of the predictor from the time domain it the course is the opposite, see Figure 9.
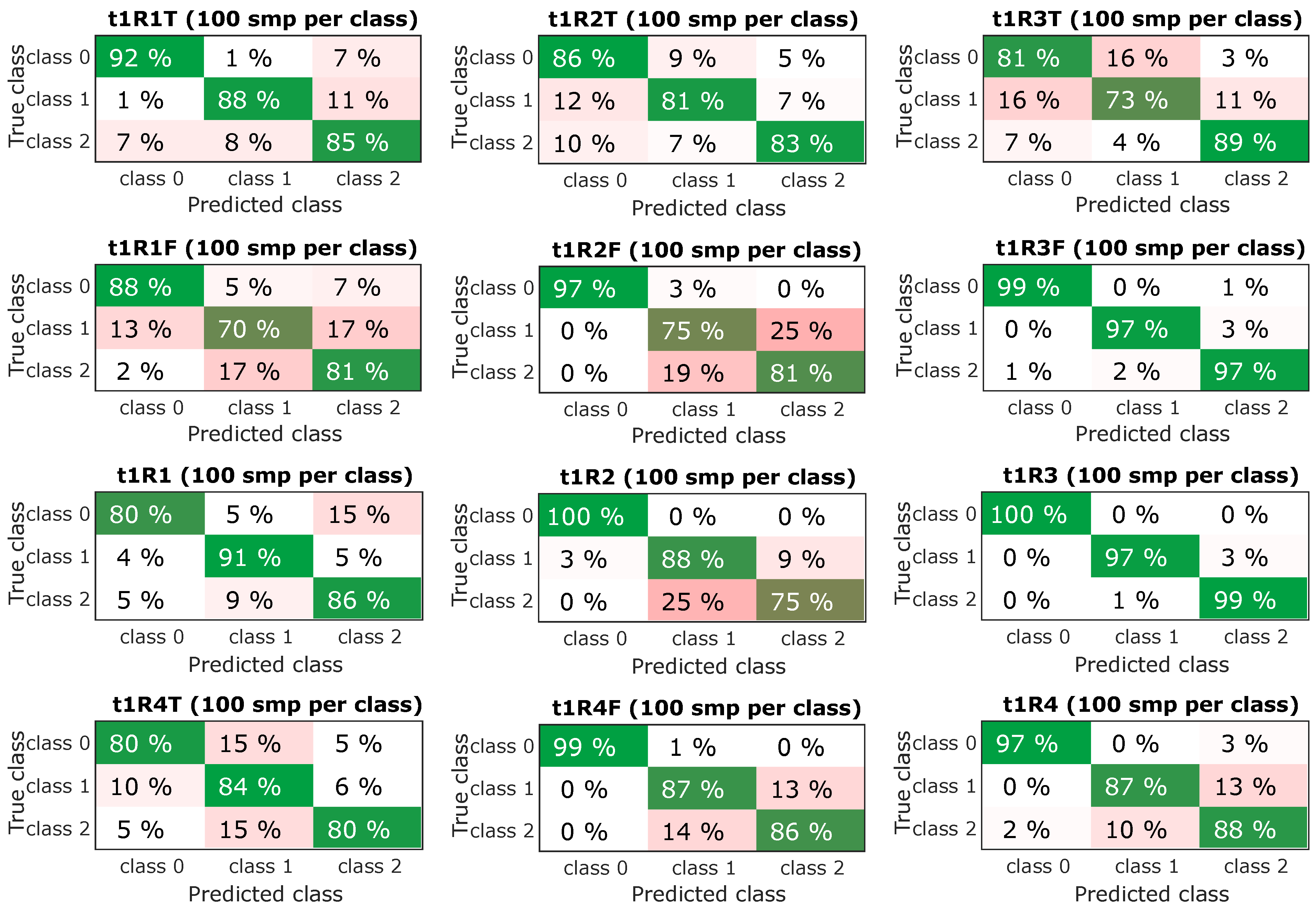
The next step was to select the most successful models and subject them to further verification. From the confusing matrix in Figure 10 and Figure 11 is clear that the models had a problem recognizing classes 1 and 2, it is a variant without an insert with a screw and a variant without an insert and a screw, i.e., a defect was determined, but incorrectly classified. However, the classification of the fault-free state is more important, it determines the success with which the model classifies the fault-free state-class 0, this information is the most important and is graphically shown in Figure 9-bottom. The experiment, which combined all predictors and contained datasets from all speeds, achieved a success rate of the classification of the fault-free state (classification of class 0-marked “TF mix”) equally 97% for the dataset for lengths of 0.25 s (t1) and 98% for the dataset for lengths of 0.5 s (t2). Experiment with speed 1500 RPM and combined predictors achieved a success rate of 100% for both datasets for lengths of 0.25 s (t1) and 0.5 s (t2).
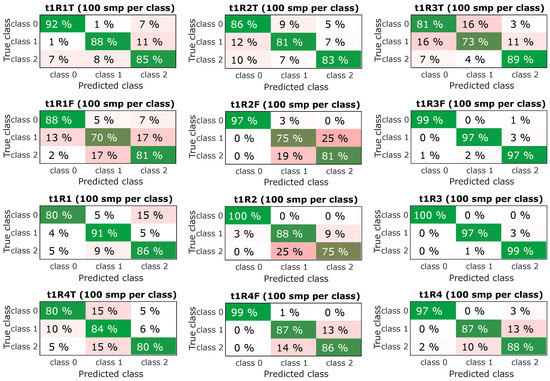
Figure 10.
Resulting confusion matrix for experiments with sample time t1 (0.25 s).
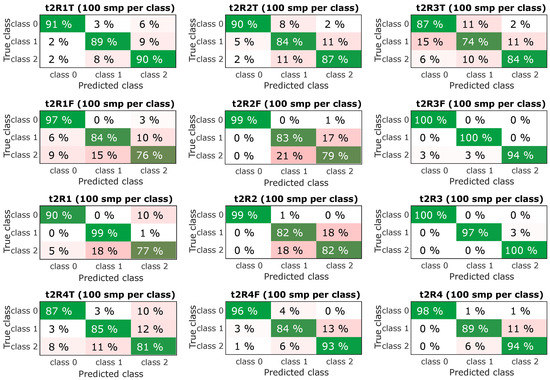
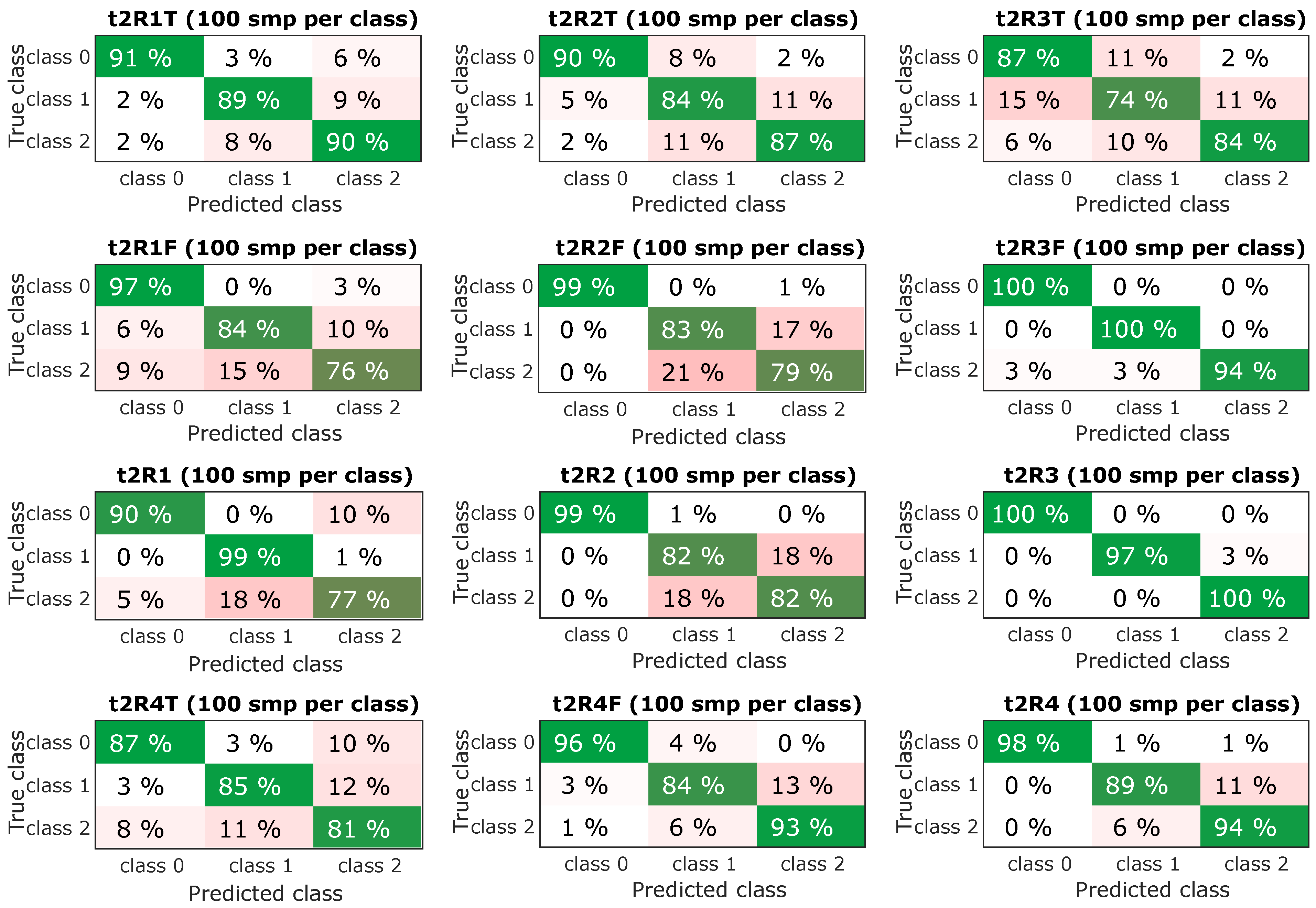
Figure 11.
Resulting confusion matrix for experiments with sample time t2 (0.5 s).
Time Consumption of Prediction
The time consumption of the prediction is highly dependent on the performance of the computing unit that will perform the prediction. In general, the time-consuming process is the learning of the model, but this operation is only performed once when the systems are deployed or when a new dataset is available for learning. Thus, the learning process can be efficiently scheduled and is not critical to the running of the system.
More important is the prediction time and the possible reaction of the system to a possible problem. The prediction time consists of two steps, namely data preprocessing, i.e., extraction of predictors, and then the prediction itself. These times were tested on a personal computer with Intel(R) Core(TM) i7-2600 CPU @ 3.40 GHz, 16 GB RAM and a similar configuration is possible for the current IPC. Time was measured for 100 repetitions each time to eliminate random variations in performance. Measured time includes only computation time without loading or saving data from recording media. Winning models for the most challenging variants containing all predictors (time and frequency domain) were used. For experiment T1R4, the winning model was “Bagged Trees” and the results are as follows. The data processing time for 100 repetitions is 1.13 s and the prediction time for 100 repetitions is 5.12 s, that is, the average total time is (1.13 + 5.12)/100 = 0.0625 s i.e., 62.5 ms. For T2R4 experiment, the winning model was “Cubic SVM” and the results are as follows. The data processing time for 100 repetitions is 2.04 s and the prediction time for 100 repetitions is 1.43 s, that is, the average total time is (2.04 + 1.43)/100 = 0.0347 s i.e., 34.7 ms. The measured times are only approximate and depend on many circumstances, also there is a possibility for optimizing the classifier selection according to its prediction time. For our purposes, the achieved times are sufficient.
6. Implementation of the Trained Model
The procedure for implementing a trained model is as follows. The model can be generated as a function of MATLAB, or as source code in C language, then it is easy to compile it for any platform. Its function (ability to classify) does not change over time because there is no learning process. However, provided the conditions have changed or a new dataset is available, it is possible to start the learning process again and train the model for the new data.
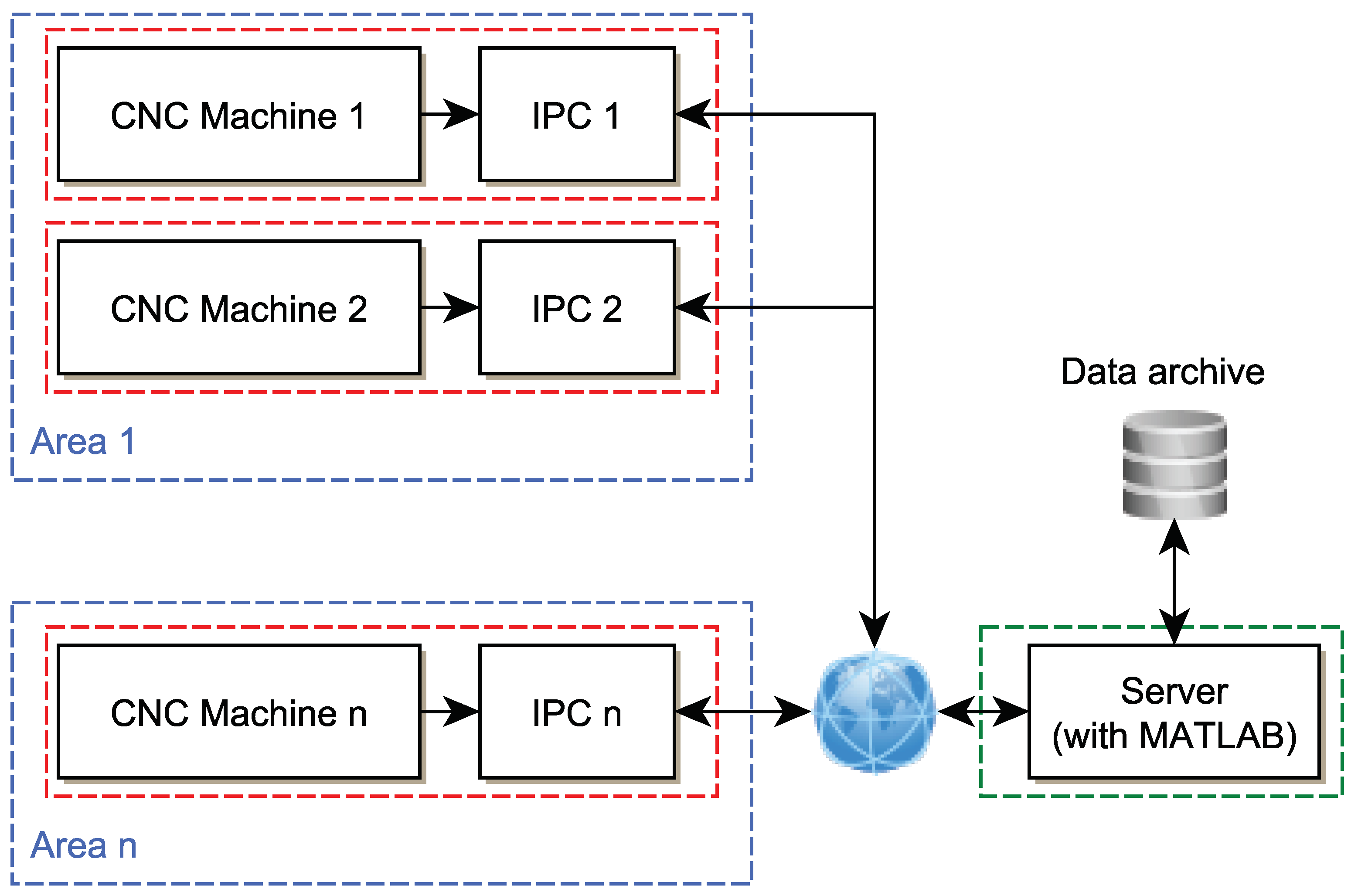
Figure 12 schematically shows the possible connection of the server with individual clients. Each IPC performs data collection and especially timely evaluation based on a trained model that was learned in the MATLAB environment on the server and then exported. The process is as follows:
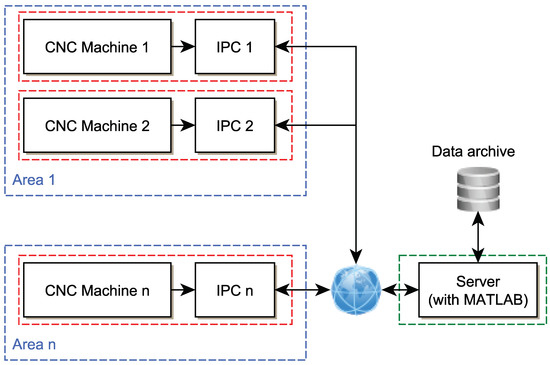
Figure 12.
IPC and server location topology.
- (a)
- Data acquisition and learning process
- The server requests data from individual IPCs and creates a dataset for creating a classification model (Data is predictors-it evaluates IPCs, only predictors are transmitted).
- If there is a sufficient amount of data (or conditions have changed, etc.), then MATLAB performs learning and creates a trained classification model for a specific IPC and exports it to the C language (this MATLAB allows).
- The trained model is sent to a specific IPC.
Individual machines and their IPCs can be distributed anywhere in different places and there will be only one server with MATLAB, which will generate a model for a specific machine and send it to a specific IPC. If necessary, the MATLAB SW can be replaced by another system, however, the functionality will be maintained and the change will be made on only one server.
- (b)
- Evaluation process
- IPC performs data collection and predictor calculation-this is a simple operation and can be performed in any programming language, for simplicity and availability Python was chosen, which performs statistical operations and frequency analysis.
- Performs classification using MATLAB created and trained model, which was exported to C language (Python allows to run code in C language).
Such a topology and partitioning allows for easy changes on both the IPC and server-side, including changing the programming environment.
As for the specific deployment for monitoring the machine, it is important to note that nowadays it is not easy to implement elements of artificial intelligence in the safety segment, where it is necessary to certify the software and verify its functions. However, it is possible to deploy a similar system only as an informative element that would alert the operator, but not interfere with the operation of the machine. The use of artificial intelligence methods is particularly advantageous in the field of technical diagnostics for the evaluation of multiple criteria and operator alerts.
7. Results and Discussions
Figure 9 summarizes the results in graphical form and expresses the dependence of the predictors on the spindle speed. For predictors from the frequency domain, an increase in success with increasing speed is evident due to a more pronounced manifestation of an unbalance fault in the frequency spectrum (increasing the first harmonic). In contrast, time domain predictors achieve a higher success rate at lower speeds than frequency domain predictors. Thus, using all predictors (time and frequency) increases the success rate at lower speeds and does not decrease at higher speeds. The figure also focuses on the detection of a fault-free state, i.e., assuming that we modify the classes only to the fault-free state (class0) and any fault state (class 1 or 2), then it is obvious that the success of fault-free detection will increase because we do not include misclassification between class 1 and class 2, which are similar. This is important if we only want to detect a fault-free condition and in any other case, an operator warning is issued.
Important results are shown in the Table 2, Table 3 and Table 4, which compare the successes of individual classification methods. Furthermore, Figure 10 and Figure 11 compare the successes of the classification of individual classes and the most important is the comparison in Figure 9, which shows the success of the classification and the success of fault-free state detection, wherewith variant R4 (TF mix-the combination of all speeds), 97% for t1 and 98% for t2 success rate is achieved.
8. Conclusions
The article aimed to verify the possibilities of artificial intelligence for the classification of machine tool failure based on an experiment from a real machine. Early detection of the disorder is an important factor in protecting the health and eliminating damage. The use of artificial intelligence methods brings the possibility of creating universal algorithms, which, thanks to the ability to learn, adapt to a specific machine, and a specific phase of machining.
The described method approaches the possibility of using artificial intelligence for data classification, in this case for the classification of various machine states. A similar method can be used to classify any classes, where only a change of predictors that describe the problem sought is needed.
The choice of predictors used is an important factor which, together with the operating conditions, influences the success of the classification. Future experiments will focus on the possibility of detecting a fault condition of a tool during machining and will focus on obtaining a larger dataset with different machining conditions.
Author Contributions
Resources, Software, Writing—original draft, D.Z.; Writing—original draft, P.B.; Validation, Visualization, T.M.; Resources, J.T.; Writing—review and editing, R.H.; Validation, K.M.; Resources, V.F. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by the Ministry of Education, Youth and Sports of the Czech Republic and The European Union-European Structural and Investments Funds in the frame of Operational Programme Research Development and Education-Project: Machine Tools and Precision Engineering (Project No. CZ.02.1.01/0.0/0.0/16_026/0008404).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Oravec, M.; Pacaiova, H.; Izarikova, G.; Hovanec, M. Magnetic Field Image-Source of Information for Action Causality. In Proceedings of the 2019 IEEE 17th World Symposium on Applied Machine Intelligence and Informatics (SAMI), Herl’any, Slovakia, 24–26 January 2019; pp. 101–105, ISBN 978-1-7281-0250-4. [Google Scholar]
- ISO 12100:2010. Safety of Machinery—General Principles for Design—Risk Assessment and Risk Reduction; ISO: Genova, Switzerland, 2010. [Google Scholar]
- ISO 9001:2008. Quality Management Systems; ISO: Genova, Switzerland, 2008. [Google Scholar]
- ISO 31010:2019. Risk Management—Risk Assessment Techniques; IEC: Genova, Switzerland, 2019. [Google Scholar]
- ISO 3534-1:2006. Statistics—Vocabulary and Symbols—Part 1: General Statistical Terms and Terms Used in Probability; ISO: Genova, Switzerland, 2006. [Google Scholar]
- Zuth, D.; Marada, T. Using artificial intelligence to determine the type of rotary machine fault. Mendel 2018, 24, 49–54. [Google Scholar] [CrossRef]
- Zuth, D.; Marada, T. Utilization of machine learning in vibrodiagnostics. Adv. Intell. Syst. Comput. 2018. [Google Scholar] [CrossRef]
- Zuth, D.; Marada, T. Comparison of Faults Classification in Vibrodiagnostics from Time and Frequency Domain Data. In Proceedings of the 2018 18th International Conference on Mechatronics-Mechatronika, Brno, Czech Republic, 5–7 December 2018; Volume 18, pp. 482–487, ISBN 978-802145544-3. [Google Scholar]
- Ympa, A. Learning Methods for Machine Vibration Analysis and Health Monitoring. Ph.D. Dissertation, Delft University of Technology, Delft, The Netherlands, 2001. [Google Scholar]
- Maluf, D.A.; Daneshmend, L. Application of machine learning for machine monitoring and diagnosis. Fla. Artif. Intell. Res. Symp. 1997, 10, 232–236. [Google Scholar]
- Li, C.; Sánchez, R.V.; Zurita, G.; Cerrada, M.; Cabrera, D. Fault Diagnosis for Rotating Machinery Using Vibration Measurement Deep Statistical Feature Learning. Sensors 2016, 16, 895. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Z.; Chen, X.; Li, C.; Sanchez, R.V.; Qin, H. Vibration-based gearbox fault diagnosis using deep neural networks. J. Vibroeng. 2017, 19, 2475–2496. [Google Scholar] [CrossRef]
- Popiołek, K. Diagnosing the technical condition of planetary gearbox using the artificial neural network based on analysis of non-stationary signals. Diagnostyka 2016, 17, 57–64. [Google Scholar]
- Haedong, J.; Park, S.; Woo, S.; Lee, S. Rotating Machinery Diagnostics Using Deep Learning on Orbit Plot Images. Procedia Manuf. 2016, 5, 1107–1118. [Google Scholar] [CrossRef] [Green Version]
- Scikit-Learn: Machine Learning in Python. Available online: https://scikit-learn.org/stable/index.html (accessed on 6 January 2018).
- Pánek, D.; Orosz, T.; Karban, P. Artap: Robust Design Optimization Framework for Engineering Applications. In Proceedings of the 2019 Third International Conference on Intelligent Computing in Data Sciences (ICDS), Marrakech, Morocco, 28–30 October 2019; pp. 1–6. [Google Scholar] [CrossRef] [Green Version]
- Train Models to Classify Data Using Supervised Machine Learning. MathWorks-Makers of MATLAB and Simulink. Available online: https://www.mathworks.com/help/stats/classificationlearner-app.html (accessed on 20 April 2018).
- MCV 750 | KOVOSVIT MAS. Machine tools, CNC machines, CNC lathes | KOVOSVIT MAS [online]. Copyright© KOVOSVIT MAS 2016. Available online: https://www.kovosvit.com/mcv-750-p7.html (accessed on 20 April 2018).
- Broch, J.T. Mechanical Vibration and Shock Measurements, 2nd ed.; Bruel & Kjaer: Nellum, Denmark, 1980; ISBN 978-8787355346. [Google Scholar]
- Blata, J.; Juraszek, J. Metody Technické Diagnostiky, Teorie a Praxe. Metody Diagnostyki Technicznej, Teoria i Praktika; Repronis: Ostrava, Czech Republic, 2013; ISBN 978-80-248-2997-5. [Google Scholar]
- Zuth, D.; Vdoleček, F. Měření vibrací ve vibrodiagnostice. In Automa; FCC Public: Praha, Czech Republic, 2010; pp. 32–36. ISSN 1210-9592. [Google Scholar]
- Scikit-Learn 0.23.1 Documentation. Scikit-Learn: Machine learning in Python—Scikit-learn 0.16.1 Documentation. Available online: https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html (accessed on 20 April 2018).
- Discrete Fourier Transform (numpy.fft)—NumPy v1.19 Manual. NumPy. Available online: https://numpy.org/doc/stable/reference/routines.fft.html (accessed on 20 April 2018).
- Classification, MathWorks-Makers of MATLAB and Simulink-MATLAB & Simulink. Available online: https://www.mathworks.com/help/stats/classification.html (accessed on 20 April 2018).
- Montgomery, D. Applied Statistics and Probability for Engineers; John Wiley: New York, NY, USA, 2003; ISBN 0-471-20454-4. [Google Scholar]
- Decision Trees, MATLAB and Simulink. Available online: https://www.mathworks.com/help/stats/decision-trees.html (accessed on 20 April 2018).
- Discriminant Analysis Classification, MATLAB and Simulink. Available online: https://www.mathworks.com/help/stats/discriminant-analysis.html (accessed on 20 April 2018).
- Naive Bayes Classification, MATLAB and Simulink. Available online: https://www.mathworks.com/help/stats/naive-bayes-classification.html (accessed on 20 April 2018).
- Support Vector Machines for Binary Classification, MATLAB and Simulink. Available online: https://www.mathworks.com/help/stats/support-vector-machines-for-binary-classification.html (accessed on 20 April 2018).
- Classification Using Nearest Neighbors, MATLAB and Simulink. Available online: https://www.mathworks.com/help/stats/classification-using-nearest-neighbors.html (accessed on 20 April 2018).
- Framework for Ensemble Learning, MATLAB and Simulink. Available online: https://www.mathworks.com/help/stats/framework-for-ensemble-learning.html (accessed on 20 April 2018).
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).