Mapping Tropical Forest Biomass by Combining ALOS-2, Landsat 8, and Field Plots Data
Abstract
:1. Introduction
2. Study Area and Dataset
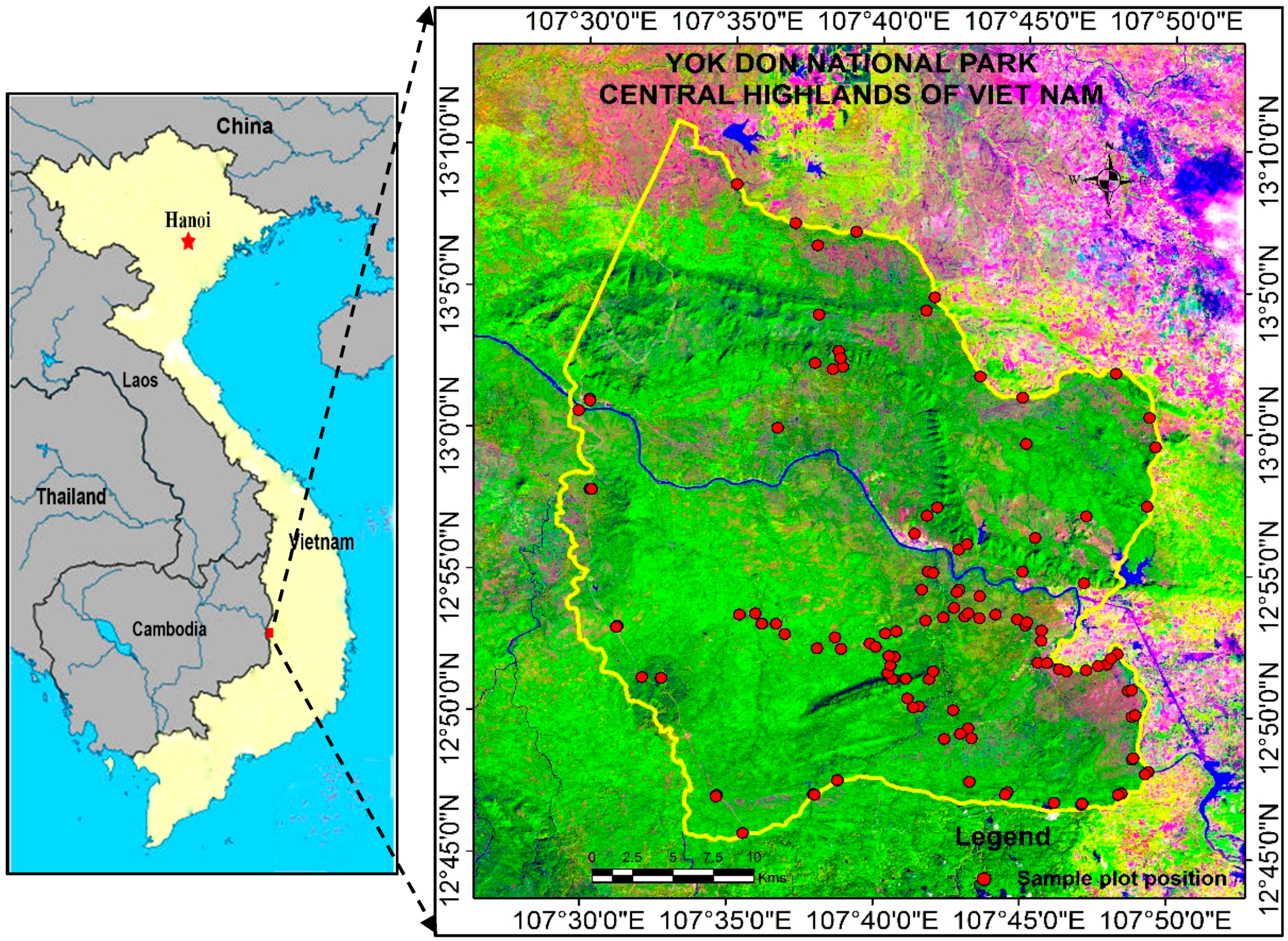
2.1. Study Area
2.2. Satellite Dataset
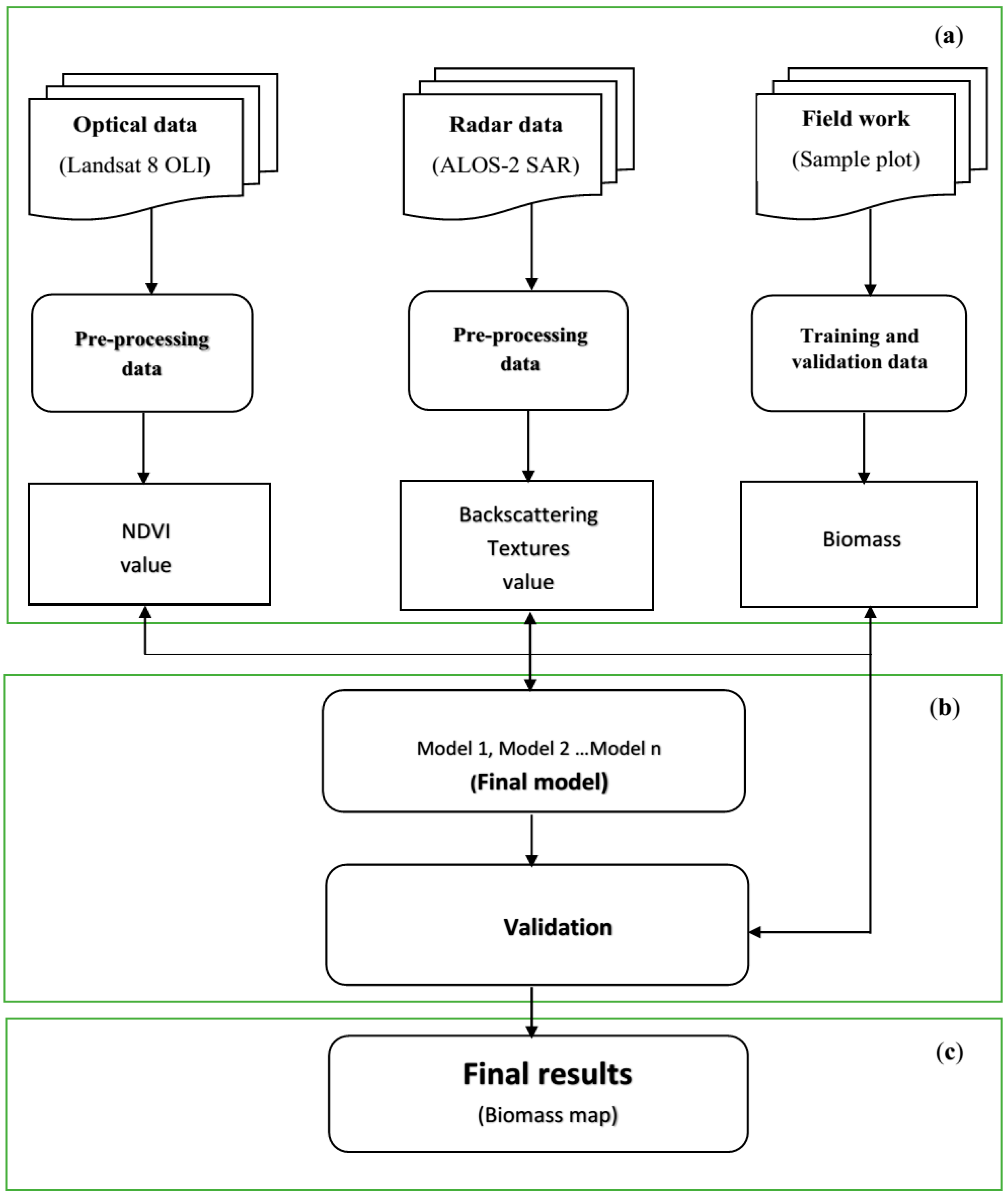
3. Methodology
3.1. Field Biomass Estimation
3.2. Processing of Satellite Data
3.3. Accuracy Analysis
4. Results and Discussion
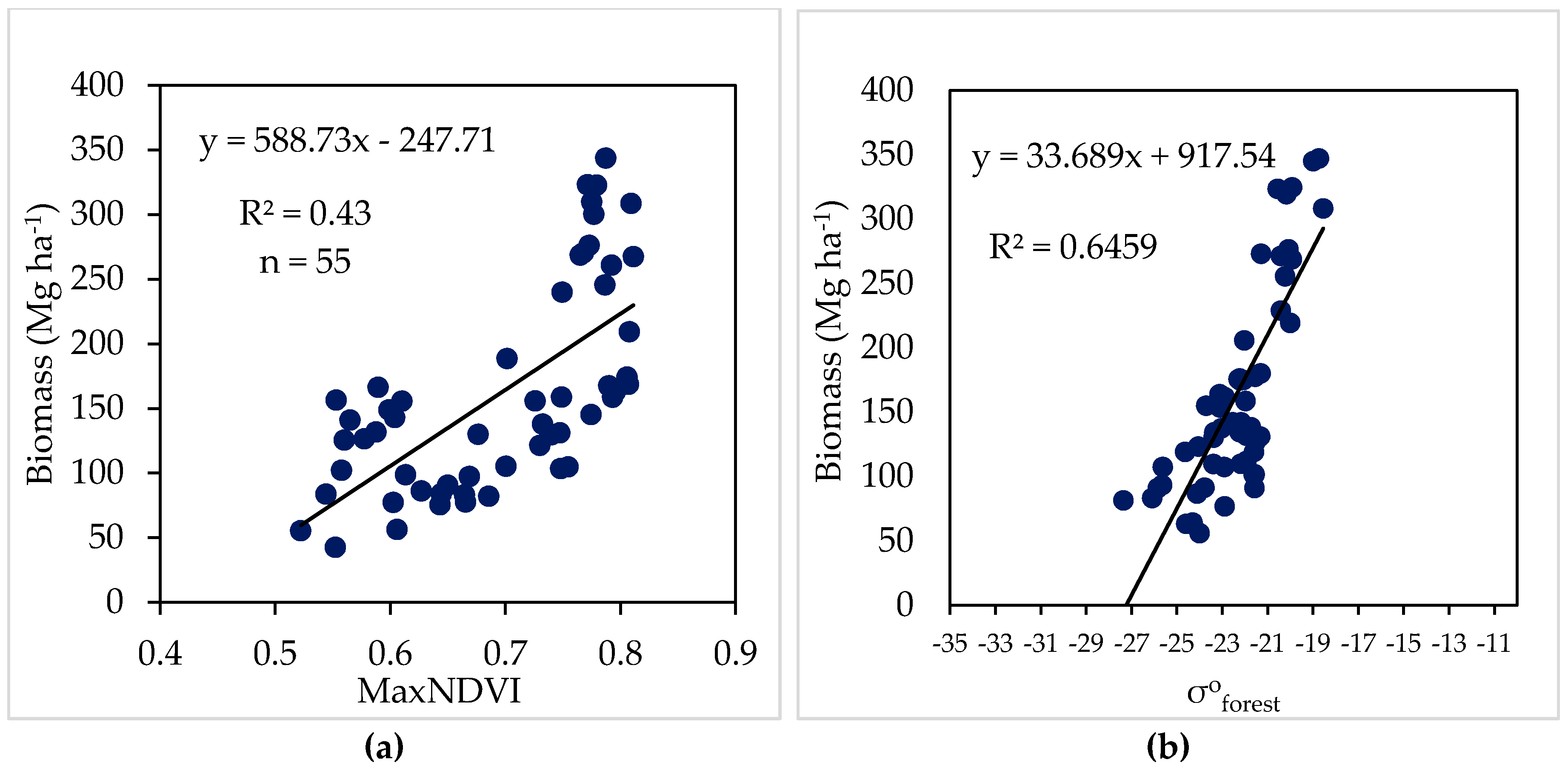
4.1. Estimates Using Single Sensor
4.2. Estimates Using Both Sensors
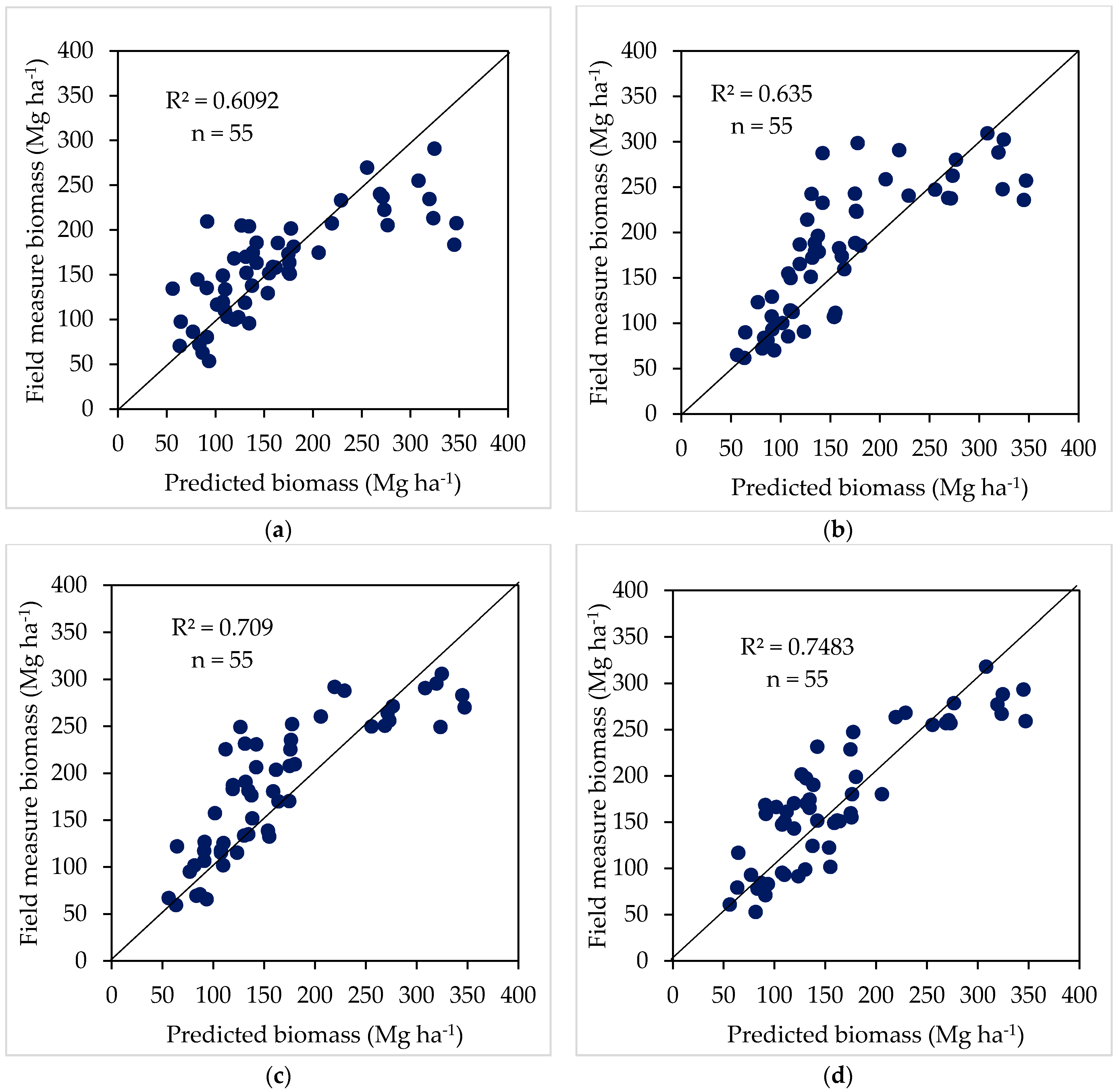
4.3. Validation of Biomass Models
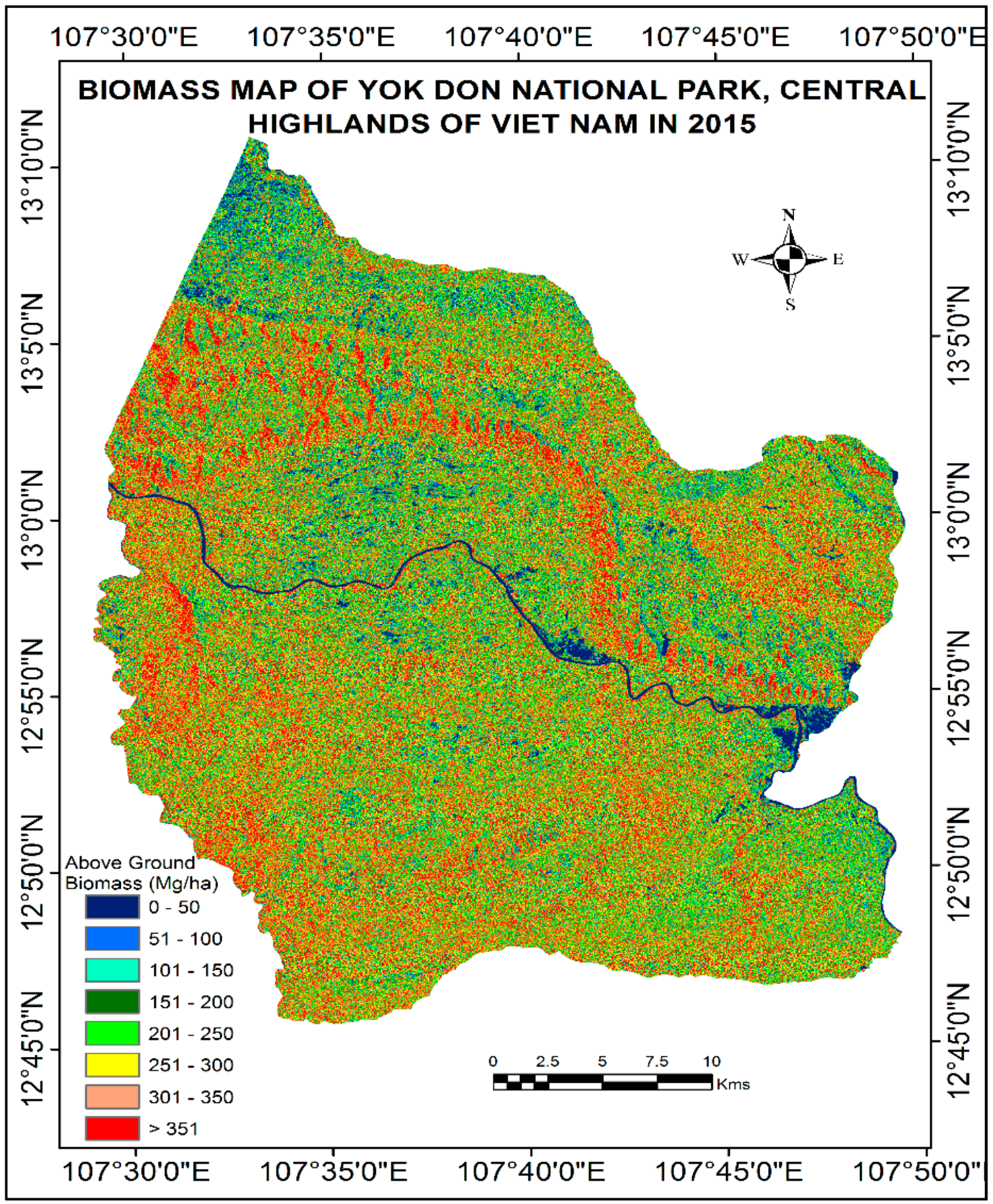
4.4. Mapping of Aboveground Biomass
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- FAO. Global Forest Resources Assessment 2015: How Are The World’s Forests Changing? FAO: Rome, Italy, 2015. [Google Scholar]
- Keenan, R.J.; Reams, G.A.; Achard, F.; de Freitas, J.V.; Grainger, A.; Lindquist, E. Dynamics of global forest area: Results from the FAO global forest resources assessment 2015. For. Ecol. Manag. 2015, 352, 9–20. [Google Scholar] [CrossRef]
- Busch, J.; Engelmann, J. The Future of Forests: Emissions from Tropical Deforestation with and without a Carbon Price, 2016–2050: Working Paper 411; CGD: Washington, DC, USA, 2015. [Google Scholar]
- Lu, D. The potential and challenge of remote sensing-based biomass estimation. Int. J. Remote Sens. 2006, 27, 1297–1328. [Google Scholar] [CrossRef]
- Bombelli, A.; Avitabile, V.; Balzter, H.; Marchesini, L.B.; Bernoux, M.; Brady, M. Biomass—Assessment of the Status of the Development of the Standards for the Terrestrial Essential Climate Variables; Food and Agriculture Organization-Global Terrestrial Observation System: Rome, Italy, 2009. [Google Scholar]
- Ridder, R.M. Options and Recommendations for a Global Remote Sensing Survey of Forests: Global Forest Resources Assessment 2010. Available online: http://www.citeulike.org/group/7954/article/3997800 (accessed on 1 August 2015).
- Foody, G.M.; Boyd, D.S.; Cutler, M.E. Predictive relations of tropical forest biomass from Landsat TM data and their transferability between regions. Remote Sens. Environ. 2003, 85, 463–474. [Google Scholar] [CrossRef]
- Hall, R.J.; Skakun, R.S.; Arsenault, E.J.; Case, B.S. Modeling forest stand structure attributes using Landsat ETM+ data: Application to mapping of aboveground biomass and stand volume. For. Ecol. Manag. 2006, 225, 378–390. [Google Scholar] [CrossRef]
- Meng, Q.; Cieszewski, C.J.; Madden, M.; Borders, B. A linear mixed-effects model of biomass and volume of trees using Landsat ETM+ images. For. Ecol. Manag. 2007, 244, 93–101. [Google Scholar] [CrossRef]
- Powell, S.L.; Cohen, W.B.; Healey, S.P.; Kennedy, R.E.; Moisen, G.G.; Pierce, K.B.; Ohmann, J.L. Quantification of live aboveground forest biomass dynamics with Landsat time-series and field inventory data: A comparison of empirical modeling approaches. Remote Sens. Environ. 2010, 114, 1053–1068. [Google Scholar] [CrossRef]
- Sharma, R.C.; Kajiwara, K.; Honda, Y. Automated extraction of canopy shadow fraction using unmanned helicopter-based color vegetation indices. Trees 2013. [Google Scholar] [CrossRef]
- Mutanga, O.; Skidmore, A.K. Narrow band vegetation indices overcome the saturation problem in biomass estimation. Int. J. Remote Sens. 2004, 25, 3999–4014. [Google Scholar] [CrossRef]
- Sinha, S.; Jeganathan, C.; Sharma, L.K.; Nathawat, M.S. A review of radar remote sensing for biomass estimation. Int. J. Environ. Sci. Technol. 2015, 12, 1779–1792. [Google Scholar] [CrossRef]
- Kumar, L.; Sinha, P.; Taylor, S.; Alqurashi, A.F. Review of the use of remote sensing for biomass estimation to support renewable energy generation. J. Appl. Remote Sens. 2015. [Google Scholar] [CrossRef]
- Ulaby, F.T.; Moore, R.K.; Fung, A.K. Microwave Remote Sensing: Active and Passive; Addison-Wesley: Reading, MA, USA, 1981. [Google Scholar]
- Le Toan, T.; Beaudoin, A.; Riom, J.; Guyon, D. Relating forest biomass to SAR data. IEEE Trans. Geosci. Remote Sens. 1992, 30, 403–411. [Google Scholar] [CrossRef]
- Jensen, J.R. Remote Sensing of the Environment: An Earth Resource Perspective; Pearson Education India: Delhi, India, 2000. [Google Scholar]
- Richards, J.; Sun, G.Q.; Simonett, D.S. L-band radar backscatter modeling of forest stands. IEEE Trans. Geosci. Remote Sens. 1987, 25, 487–498. [Google Scholar] [CrossRef]
- Freeman, A.; Durden, S.L. A three-component scattering model for polarimetric SAR data. IEEE Trans. Geosci. Remote Sens. 1998, 36, 963–973. [Google Scholar] [CrossRef]
- Ordonez, F.Y.; Ruiz, S.J.; Leblon, B. Forest Inventory Using Optical and Radar Remote Sensing; INTECH: Rijeka, Yugoslavia, 2008. [Google Scholar]
- Poolla, S.B. Polarimetric Scattering Model for Biophysical Characterization of Multilayer Vegetation Using Space Borne Polsar Data. Master’s Thesis, University of Twente, Enschede, The Netherlands, 2013. [Google Scholar]
- Kim, C. Quantitative analysis of relationship between ALOS PALSAR backscatter and forest stand volume. J. Mar. Sci. Technl. 2012, 20, 624–628. [Google Scholar]
- Park, S.E.; Kim, J.S.; Moon, W.M.; Boerner, W.M. Inversion of Surface Parameters from NASA/JPL AIRSAR Polarimetric SAR Data. In Proceedings of the 3rd International Workshop on Science and Applications of SAR Polarimetry and Polarimetric Interferometry, Frascati, Italy, 22–26 January 2007.
- Park, S.E.; Moon, W.M.; Pottier, E. Assessment of scattering mechanism of polarimetric SAR signal from mountainous forest areas. IEEE Trans. Geosci. Remote Sens. 2012, 50, 4711–4719. [Google Scholar] [CrossRef]
- Panciera, R.; Tanase, M.; Lowell, K.; Walker, J.P. Evaluation of IEM, Dubois, and Oh radar backscatter models using airborne L-Band SAR. IEEE Trans. Geosci. Remote Sens. 2014, 52, 4966–4979. [Google Scholar] [CrossRef]
- Mermoz, S.; Réjou-Méchain, M.; Villard, L.; Le Toan, T.; Rossi, V.; Gourlet-Fleury, S. Decrease of L-band SAR backscatter with biomass of dense forests. Remote Sens. Environ. 2015, 159, 307–317. [Google Scholar] [CrossRef]
- Ranson, K.J.; Sun, G. Mapping biomass of a northern forest using multifrequency SAR data. IEEE Trans. Geosci. Remote Sens. 1994, 32, 388–396. [Google Scholar] [CrossRef]
- Ranson, K.J.; Sun, G. Effect of environmental temperatures on SAR forest biomass estimates. In Proceedings of the 1997 IEEE International Geoscience and Remote Sensing, IGARSS ’97. Remote Sensing—A Scientific Vision for Sustainable Development, Greenbelt, MD, USA, 3–8 August 1997; pp. 1722–1724.
- Shi, J.; Wang, J.; Hsu, A.Y.; O’Neill, P.E.; Engman, E.T. Estimation of bare surface soil moisture and surface roughness parameter using L-band SAR image data. IEEE Trans. Geosci. Remote Sens. 1997, 35, 1254–1266. [Google Scholar]
- Bindlish, R.; Barros, A.P. Parameterization of vegetation backscatter in radar-based, soil moisture estimation. Remote Sens. Environ. 2001, 76, 130–137. [Google Scholar] [CrossRef]
- Kasischke, E.S.; Bourgeau-Chavez, L.L.; Rober, A.R.; Wyatt, K.H.; Waddington, J.M.; Turetsky, M.R. Effects of soil moisture and water depth on ERS SAR back scatter measurements from an Alaskan wetland complex. Remote Sens. Environ. 2009, 113, 1868–1873. [Google Scholar] [CrossRef]
- Huang, W.; Sun, G.; Zhang, Z.; Ni, W. Sensitivity of multi-source SAR backscatter to changes of forest aboveground biomass. Remote Sens. 2015, 7, 9587–9609. [Google Scholar] [CrossRef]
- Le Toan, T.; Mermoz, S.; Bouvet, A.; Villard, L.K.C. Forest Cover Change and Biomass Mapping; K&C Initiative Report on an International Science Collaboration Led by JAXA: Tokyo, Japan, 2013. [Google Scholar]
- Iizuka, K.; Tateishi, R. Estimation of CO2 sequestration by the forests in Japan by discriminating precise tree age category using remote sensing techniques. Remote Sens. 2015, 7, 15082–15113. [Google Scholar] [CrossRef]
- Luckman, A.; Baker, J.; Kuplich, M.T.; Yanasse, F.C.C.; Frery, C.A. A study of the relationship between radar backscatter and regenerating tropical forest biomass for spaceborne SAR instruments. Remote Sens. Environ. 1997, 60, 1–13. [Google Scholar] [CrossRef]
- Kurvonen, L.; Pulliainen, J.; Hallikainen, M. Retrieval of biomass in boreal forests from multitemporal ERS-1 and JERS-1 SAR images. IEEE Trans. Geosci. Remote Sens. 1999, 37, 198–205. [Google Scholar] [CrossRef]
- Hajnsek, I.; Kugler, F.; Lee, S.K.; Papathanassiou, K.P. Tropical-forest-parameter estimation by means of Pol-InSAR: The INDREX-II campaign. IEEE Trans. Geosci. Remote Sens. 2009, 47, 481–493. [Google Scholar] [CrossRef]
- Mika, K.; Ulla, P.; Karila, K.; Juha, H. Forest biomass estimation using ALOS PALSAR images in challenging natural forest area in Finland. In Proceedings of the ALOS PI 2008 Symposium, ESA Special Publication. Island of Rhodes, Greece, 3–7 November 2008.
- Lucas, R.; Armston, J.; Fairfax, R.; Fensham, R.; Accad, A.; Carreiras, J.; Metcalfe, D. An evaluation of the ALOS PALSAR L-band backscatter—Above ground biomass relationship Queensland, Australia: Impacts of surface moisture condition and vegetation structure. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2010, 3, 576–593. [Google Scholar] [CrossRef]
- Morel, A.C.; Saatchi, S.S.; Malhi, Y.; Berry, N.J.; Banin, L.; Burslem, D.; Ong, R.C. Estimating aboveground biomass in forest and oil palm plantation in Sabah, Malaysian Borneo using ALOS PALSAR data. For. Ecol. Manag. 2011, 262, 1786–1798. [Google Scholar] [CrossRef]
- Englhart, S.; Keuck, V.; Siegert, F. Aboveground biomass retrieval in tropical forests–The potential of combined X- and L-band SAR data use. Remote Sens. Environ. 2011, 115, 1260–1271. [Google Scholar] [CrossRef]
- Hame, T.; Rauste, Y.; Antropov, O.; Ahola, H.A.; Kilpi, J. Improved mapping of tropical forests with optical and SAR imagery, Part II: Above ground biomass estimation. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2013, 6, 92–101. [Google Scholar] [CrossRef]
- Carreiras, J.M.B.; Melo, J.B.; Vasconcelos, M.J. Estimating the above-ground biomass in Miombo Savanna woodlands (Mozambique, East Africa) using L-band synthetic aperture radar data. Remote Sens. 2013, 5, 1524–1548. [Google Scholar] [CrossRef]
- Peregon, A.; Yamagata, Y. The use of ALOS/PALSAR backscatter to estimate above-ground forest biomass: A case study in Western Siberia. Remote Sens. Environ. 2013, 137, 139–146. [Google Scholar] [CrossRef]
- Rahman, M.M.; Tetuko Sri Sumantyo, J. Retrieval of tropical forest biomass information from ALOS PALSAR data. Geocarto Int. 2013, 28, 382–403. [Google Scholar] [CrossRef]
- Nezry, E.; Mougin, E.; Lopes, A.; Gastellu-Etchegorry, J.P.; Laumonier, Y. Tropical vegetation mapping with combined visible and SAR spaceborne data. Remote Sens. 1993, 14, 2165–2184. [Google Scholar] [CrossRef]
- Andersen, H.E.; Strunk, J.; Temesgen, H.; Atwood, D.; Winterberger, K. Using multilevel remote sensing and ground data to estimate forest biomass resources in remote regions: A case study in the boreal forests of interior Alaska. Can. J. Remote Sens. 2012, 37, 596–611. [Google Scholar] [CrossRef]
- Cutler, M.E.J.; Boyd, D.S.; Foody, G.M.; Vetrivel, A. Estimating tropical forest biomass with a combination of SAR image texture and Landsat TM data: An assessment of predictions between regions. ISPRS J. Photogramm. Remote Sens. 2012, 70, 66–77. [Google Scholar] [CrossRef]
- Goh, J.; Miettinen, J.; Chia, A.S.; Chew, P.T.; Liew, S.C. Biomass estimation in humid tropical forest using a combination of ALOS PALSAR and SPOT 5 satellite imagery. Asian J. Geoinform. 2014, 13, 59–72. [Google Scholar]
- Hamdan, O.; Hasmadi, I.M.; Aziz, H.K. Combination of SPOT-5 and ALOS PALSAR images in estimating aboveground biomass of lowland Dipterocarpus forest. IOP Conf. Ser. Earth Environ. Sci. 2014. [Google Scholar] [CrossRef]
- Luong, N.V.; Tateishi, R.; Thanh Nguyen, H.C.; Sharma, R.; Trong To, T.; Mai Le, S. Estimation of Tropical Forest Structural Characteristics Using ALOS-2 SAR Data. Adv. Remote Sens. 2016, 5, 131–144. [Google Scholar]
- Champion, I.; Dubois-Fernandez, P.; Guyon, D.; Cottrel, D. Radar SAR images texture as a function of forest stand age. Int. J. Remote Sens. 2008, 29, 1795–1800. [Google Scholar] [CrossRef]
- Thapa, R.B.; Watanabe, M.; Shimada, M.; Motohka, T. Examining high-resolution PiSAR-L2 textures for estimating tropical forest carbon stocks. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2016, 9, 3202–3209. [Google Scholar] [CrossRef]
- Le Toan, T.; Quegan, S.; Davidson, M.W.J.; Balzter, H.; Paillou, P.; Papathanassiou, K.; Ulander, L. The BIOMASS mission: Mapping global forest biomass to better understand the terrestrial carbon cycle. Remote Sens. Environ. 2011, 115, 2850–2860. [Google Scholar] [CrossRef]
- Minh, D.H.T.; Tebaldini, S.; Rocca, F.; Le Toan, T.; Villard, L.; Dubois-Fernandez, P.C. Capabilities of BIOMASS tomography for investigating tropical forests. IEEE Trans. Geosci. Remote Sens. 2015, 53, 965–975. [Google Scholar] [CrossRef]
- Trung, T.V. Chapter IV. In The Tropical Forest Ecosystem in Vietnam; Science and Technics Publishing House: Hanoi, Vietnam, 1998. [Google Scholar]
- Lan, P.N.; Hong, P.N.; Hung, T.V.; Thin, N.N.; Chan, L.T. Natural Forest Ecosystems Viet Nam; Ministry of Agriculture and Rural Development of Vietnam: Hanoi, Vietnam, 2006.
- Thin, N.N.; Anh Nguyen, D.; Tai, V.A.; Thanh, N.T.K.; Thi Nguyen, D. Update Information of Flora and Plant’s Resources of Yok Don National Park, Daclak Province in 2008. Available online: http://www.botanyvn.com/cnt.asp?param=news&newsid=46&lg=en (accessed on 9 June 2008).
- Nguyen, T.T. Modelling Growth and Yield of Dipterocarpa Forests in Central Highlands of Vietnam. Ph.D. Thesis, Technische Universität München, München, Germany, 2009. [Google Scholar]
- MARD. Soil Map in Highlands’s Region in 2010; Scale 1:250.000; National Institute for Agricultural Planning–Remote Sensing Center, Ministry of Agriculture and Rural Development of Vietnam: Hanoi, Vietnam, 2010.
- Canh, N.X.; Quynh, H.Q.; Anh, L.T.; Luong, N.V. Report on Conservation Planning and Sustainable Development of Yok Don National Park in 2010–2020; Ministry of Agriculture and Rural Development of Vietnam: Hanoi, Vietnam, 2009.
- MONRE. Survey Data Base of Hydrometeorology Sector, Has Been Archived at Center of National Hydro-Meteorological Service; Ministry of Natural Resources and Environment: Hanoi, Vietnam, 2015.
- Phuong, V.T.; Xuan, N.V.; Trieu, D.T.; Trung, P.D.; Giap, N.X.; Thanh, P.N. Tree Allometric Equations in Evergreen Broadleaf, Deciduous, and Bamboo Forest in Central Highland Region, Vietnam; UN-REDD Vietnam Programme: Hanoi, Vietnam, 2013. [Google Scholar]
- IPCC. Good Practice Guidance for Land Use, Land-Use Change and Forestry; IPCC National Greenhouse Gas Inventories Programme Technical Support Unit: Hayama, Japan, 2003. [Google Scholar]
- JAXA. ALOS-2/Calibration Result of JAXA Standard Products; Japan Aerospace Exploration Agency, Earth Observation Research Center: Tsukuba, Japan, 2014. [Google Scholar]
- Attema, E.P.W.; Ulaby, F.T. Vegetation modeled as a water cloud. Radio Sci. 1978, 13, 357–364. [Google Scholar] [CrossRef]
- Yamaguchi, Y.; Yajima, Y.; Yamada, H. A four-component decomposition of POLSAR images based on the coherency matrix. IEEE Geosci. Remote Sens. Lett. 2006, 3, 292–296. [Google Scholar] [CrossRef]
- Haralick, R.M.; Shanmugam, K.; Dinstein, I.H. Textural features for image classification. IEEE Trans. Syst. Man Cybern. 1973, 6, 610–621. [Google Scholar] [CrossRef]
- Anys, H.; Bannari, A.; He, D.C.; Morin, D. Texture analysis for the mapping of urban areas using airborne MEIS-II images. In Proceedings of the First International Airborne Remote Sensing Conference and Exhibition, Strasbourg, France, 12–15 September 1994; pp. 231–245.
- Rouse, J., Jr.; Haas, R.H.; Schell, J.A.; Deering, D.W. Monitoring vegetation systems in the Great Plains with ERTS. NASA Spec. Publ. 1974, 351, 309. [Google Scholar]
- Anderson, G.L.; Hanson, J.D.; Haas, R.H. Evaluating Landsat Thematic Mapper derived vegetation indices for estimating above-ground biomass on semiarid rangelands. Remote Sens. Environ. 1993, 45, 165–175. [Google Scholar] [CrossRef]
- Hunt, E.R., Jr. Relationship between woody biomass and PAR conversion efficiency for estimating net primary production from NDVI. Int. J. Remote Sens. 1994, 15, 1725–1729. [Google Scholar] [CrossRef]
- Dong, J.; Kaufmann, R.K.; Myneni, R.B.; Tucker, C.J.; Kauppi, P.E.; Liski, J.; Hughes, M.K. Remote sensing estimates of boreal and temperate forest woody biomass: carbon pools, sources, and sinks. Remote Sens. Environ. 2003, 84, 393–410. [Google Scholar] [CrossRef]
- Saatchi, S.S.; Harris, N.L.; Brown, S.; Lefsky, M.; Mitchard, E.T.; Salas, W.; Petrova, S. Benchmark map of forest carbon stocks in tropical regions across three continents. Proc. Natl. Acad. Sci. USA 2011, 108, 9899–9904. [Google Scholar] [CrossRef] [PubMed]
| No. | Scene ID | Observation Date | Observation Angle | Polarizations | Season |
|---|---|---|---|---|---|
| 1 | ALOS2040600240-150222-FBDR2.1GUA | 22 February 2015 | 32.9° | HH, HV | Dry |
| 2 | ALOS2040600250-150222-FBDR2.1GUA | 22 February 2015 | 32.9° | HH, HV | Dry |
| No | ID | Observation Date | Path/ Row | Bands Used | Season |
|---|---|---|---|---|---|
| 1 | LC81240512015289LGN00 | 16 October 2015 | 124/051 | B6, B5, B4, B8 | Rainy |
| Month/Year | Daily Rainfall (mm) | Total Rainfall (mm) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| February 2015 | Days | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | 0.0 |
| Rainfall (mm) | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ||
| October 2015 | Days | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 40.5 |
| Rainfall (mm) | 0.0 | 0.0 | 12.0 | 27.7 | 0.8 | 0.0 | 0.0 | 0.0 | ||
| Parameter | Deciduous Forest | Evergreen Forest | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Minimum | Maximum | Mean | Standard Deviation | Minimum | Maximum | Mean | Standard Deviation | ||
| Diameter (m) | 8.14 | 38.52 | 18.29 | 6.73 | 16.87 | 48.74 | 19.39 | 13.33 | |
| Height (m) | 6.33 | 38.52 | 10.97 | 2.93 | 10.01 | 18.23 | 12.18 | 3.14 | |
| Biomass (Mg·ha−1) | 42.46 | 350.18 | 134.82 | 73.10 | 167.49 | 347.98 | 309.59 | 73.84 | |
| Parameter | Deciduous Forest | Evergreen Forest | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Minimum | Maximum | Mean | Standard Deviation | Minimum | Maximum | Mean | Standard Deviation | ||
| Diameter (m) | 9.89 | 39.41 | 16.74 | 6.17 | 12.51 | 44.82 | 20 | 15.11 | |
| Height (m) | 6.06 | 17.67 | 10.635 | 2.73 | 6.17 | 17.67 | 12.95 | 4.57 | |
| Biomass (Mg·ha−1) | 63.45 | 324.75 | 137.99 | 69.8 | 56.19 | 347.21 | 308.34 | 135.07 | |
| Model | Parameter used | R2 | RMSE |
|---|---|---|---|
| Model 1 | Max NDVI | 0.43 | 60.45 |
| Model 2 | σ°forest | 0.64 | 48.04 |
| Model 3 | SAR textures * | 0.36 | 64.06 |
| Model 4 | HV, HH | 0.57 | 52.51 |
| Model | Parameter Used | Training Model | Validation Model | Significance F | ||
|---|---|---|---|---|---|---|
| R2 | RMSE | R2 | RMSE | |||
| Model 5 | Max NDVI and SAR textures * | 0.62 | 49.36 | 0.60 | 55.13 | 3 × 10−7 |
| Model 6 | HV and SAR textures * | 0.66 | 46.69 | 0.63 | 44.64 | 3 × 10−8 |
| Model 7 | Max NDVI, HV and SAR textures * | 0.73 | 41.60 | 0.70 | 39.10 | 7 × 10−1 |
| Model 8 | Max NDVI, σ°forest, and SAR textures * | 0.75 | 35.18 | 0.74 | 35.88 | 3 × 10−1 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Viet Nguyen, L.; Tateishi, R.; Kondoh, A.; Sharma, R.C.; Thanh Nguyen, H.; Trong To, T.; Ho Tong Minh, D. Mapping Tropical Forest Biomass by Combining ALOS-2, Landsat 8, and Field Plots Data. Land 2016, 5, 31. https://doi.org/10.3390/land5040031
Viet Nguyen L, Tateishi R, Kondoh A, Sharma RC, Thanh Nguyen H, Trong To T, Ho Tong Minh D. Mapping Tropical Forest Biomass by Combining ALOS-2, Landsat 8, and Field Plots Data. Land. 2016; 5(4):31. https://doi.org/10.3390/land5040031
Chicago/Turabian StyleViet Nguyen, Luong, Ryutaro Tateishi, Akihiko Kondoh, Ram C. Sharma, Hoan Thanh Nguyen, Tu Trong To, and Dinh Ho Tong Minh. 2016. "Mapping Tropical Forest Biomass by Combining ALOS-2, Landsat 8, and Field Plots Data" Land 5, no. 4: 31. https://doi.org/10.3390/land5040031
APA StyleViet Nguyen, L., Tateishi, R., Kondoh, A., Sharma, R. C., Thanh Nguyen, H., Trong To, T., & Ho Tong Minh, D. (2016). Mapping Tropical Forest Biomass by Combining ALOS-2, Landsat 8, and Field Plots Data. Land, 5(4), 31. https://doi.org/10.3390/land5040031







