Characteristics of Bacterial Communities in Cyanobacteria-Blooming Aquaculture Wastewater Influenced by the Phytoremediation with Water Hyacinth
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strains and Culture Conditions
2.2. Co-Existence Experiments
2.3. General Enumeration of Bacteria
2.4. DNA Extraction and High-Throughput Sequencing
2.5. Statistical Analysis
3. Results
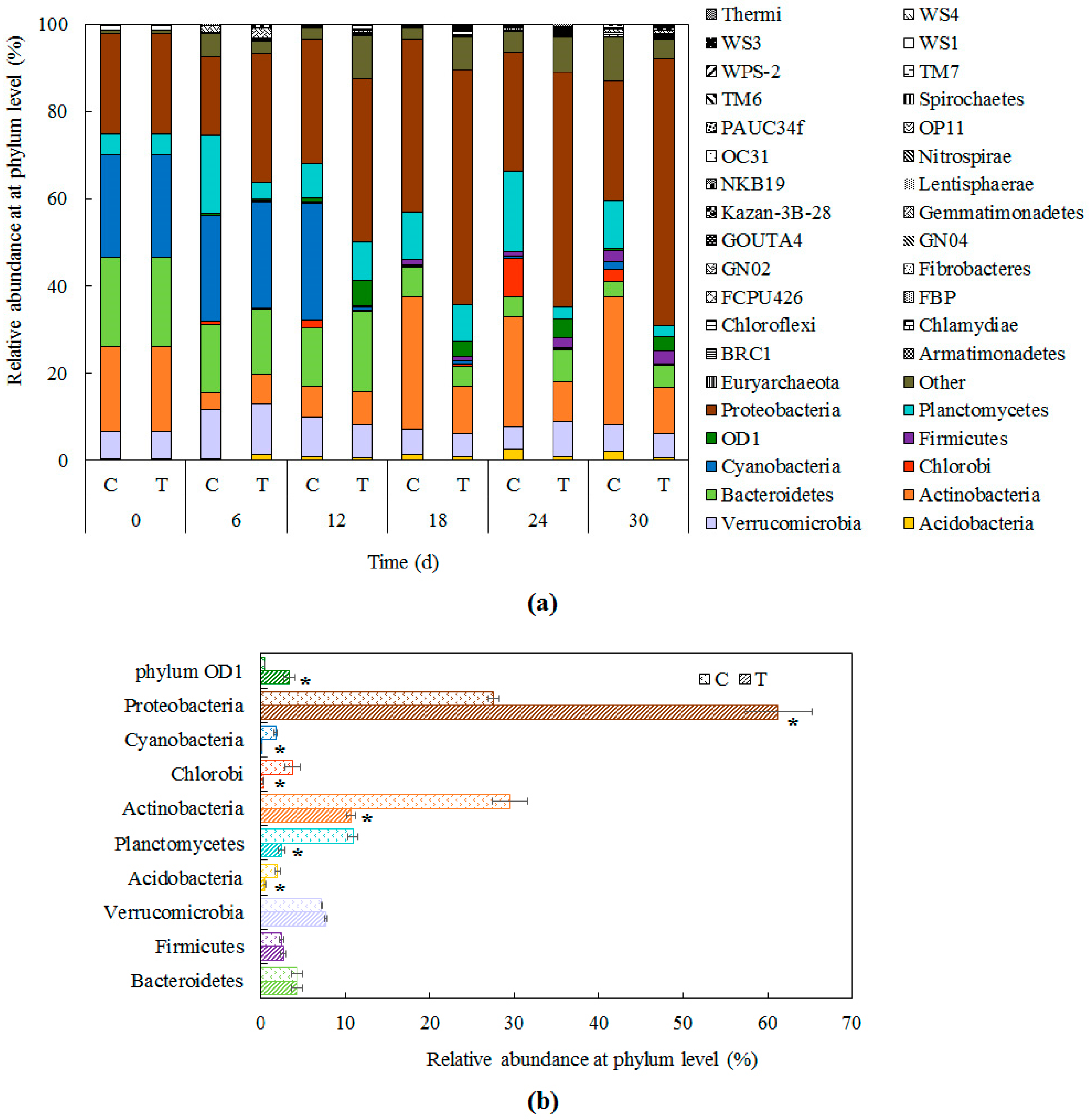
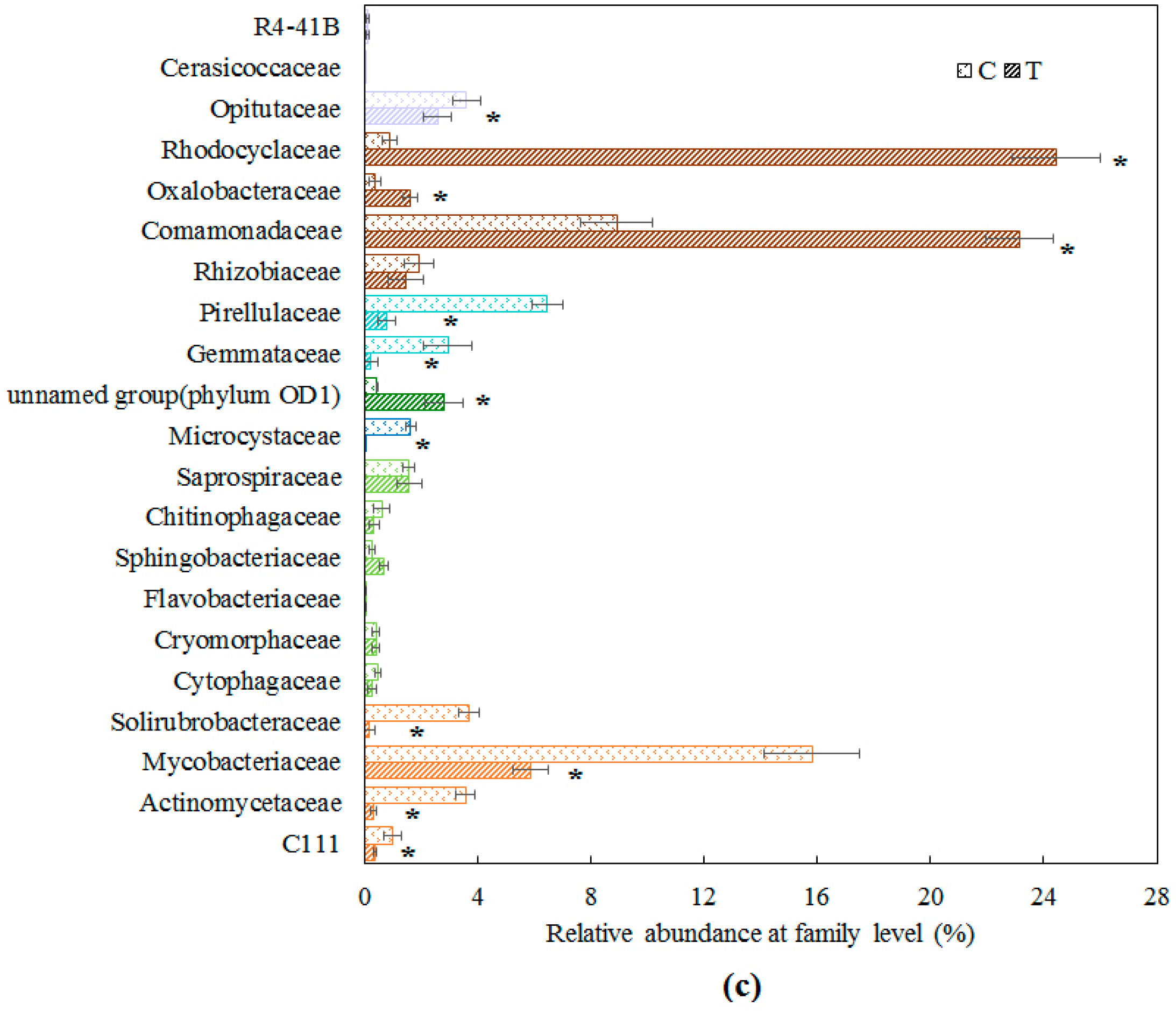
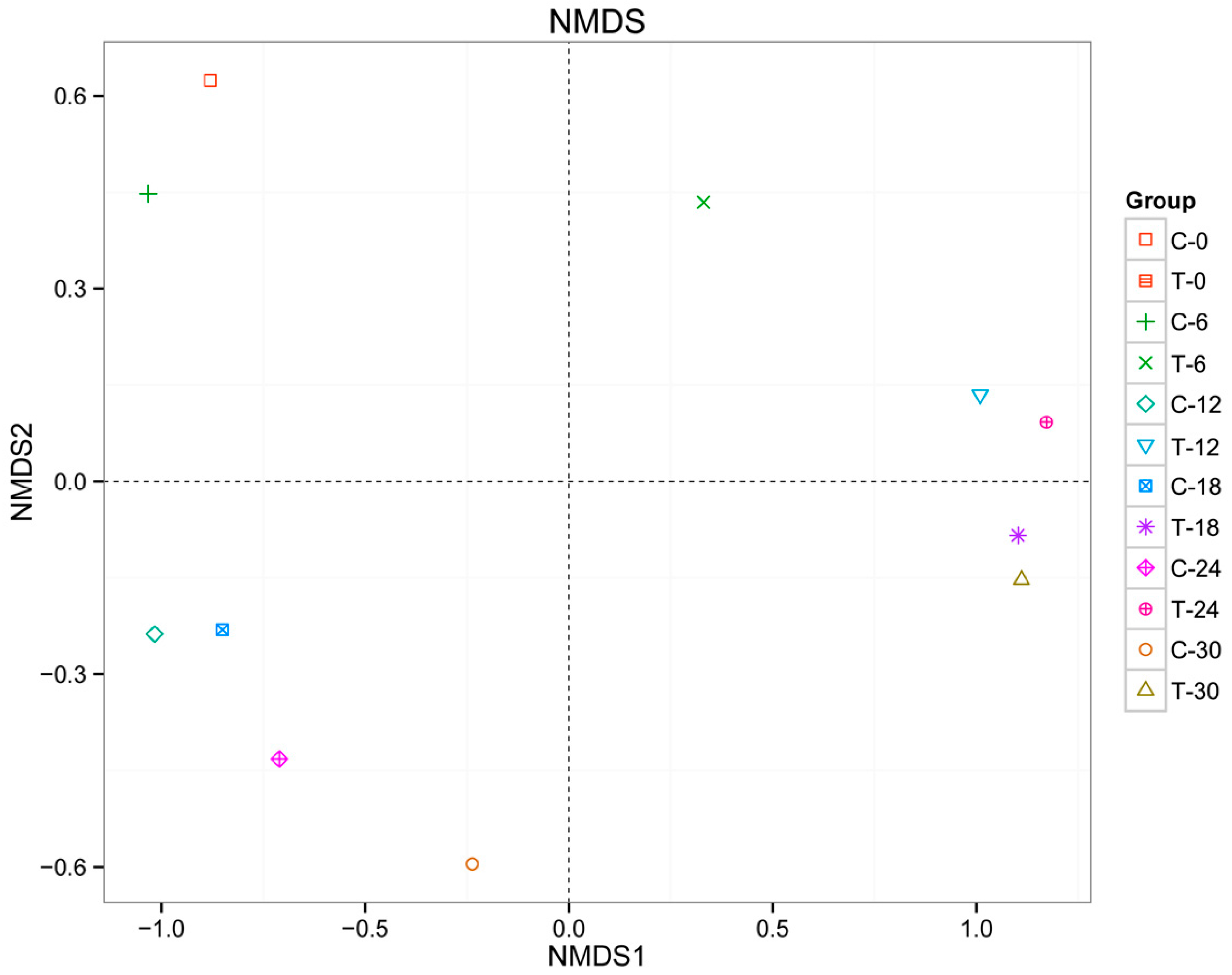
3.1. Bacterial Community Structure in Cyanobacteria-Blooming Aquaculture Wastewater Influenced by Water Hyacinth
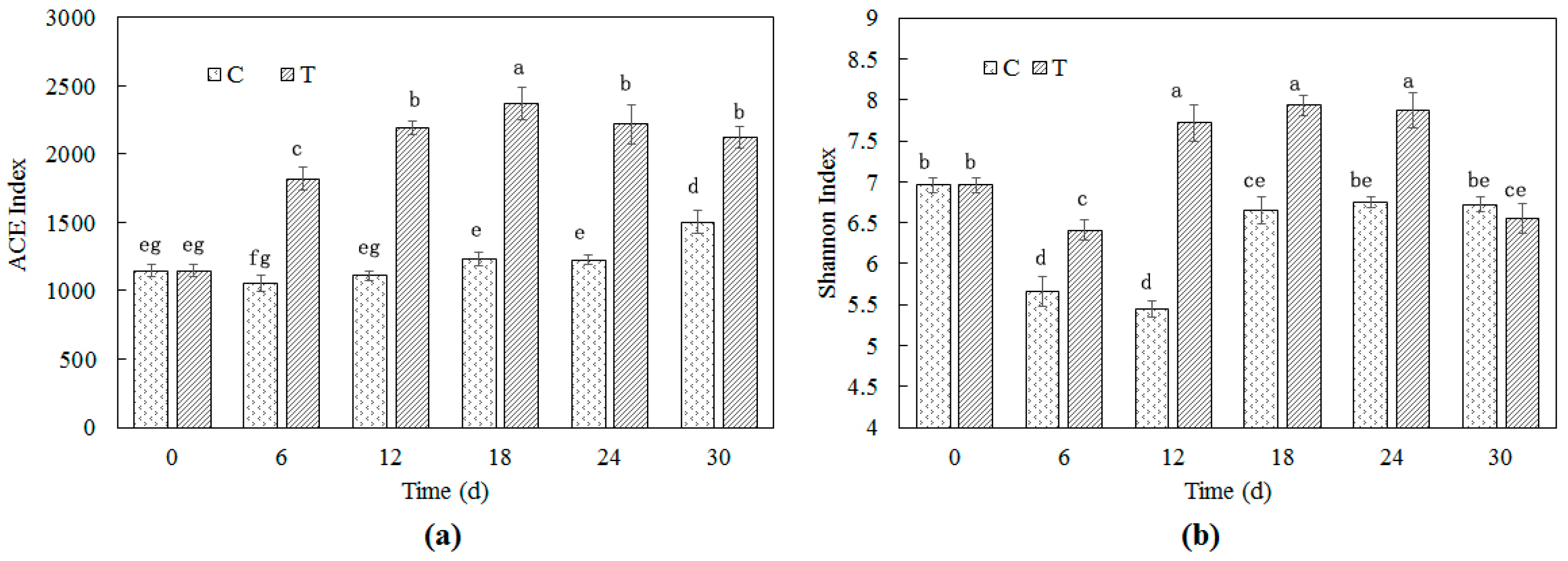
3.2. Bacterial Richness and Diversity in Cyanobacteria-Blooming Aquaculture Wastewater Influenced by Water Hyacinth
3.3. Specific Bacteria and Disease-Related Genes in Cyanobacteria-Blooming Aquaculture Wastewater Influenced by Water Hyacinth
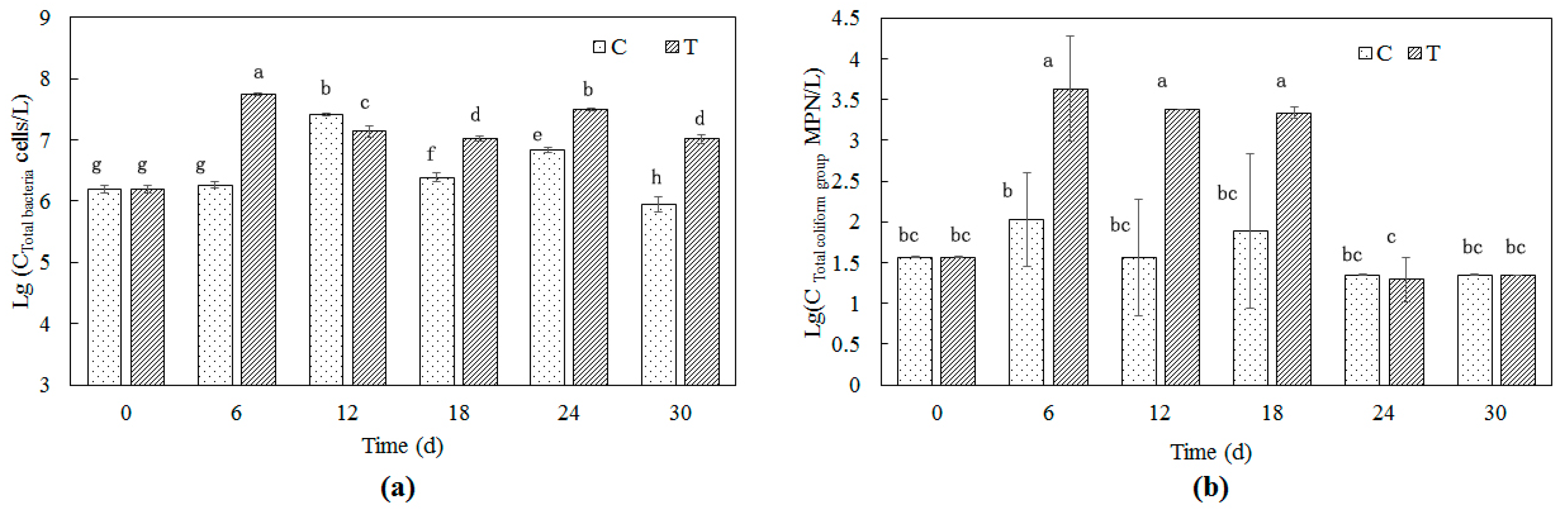
3.3.1. Total Bacteria
3.3.2. Total Coliform Group
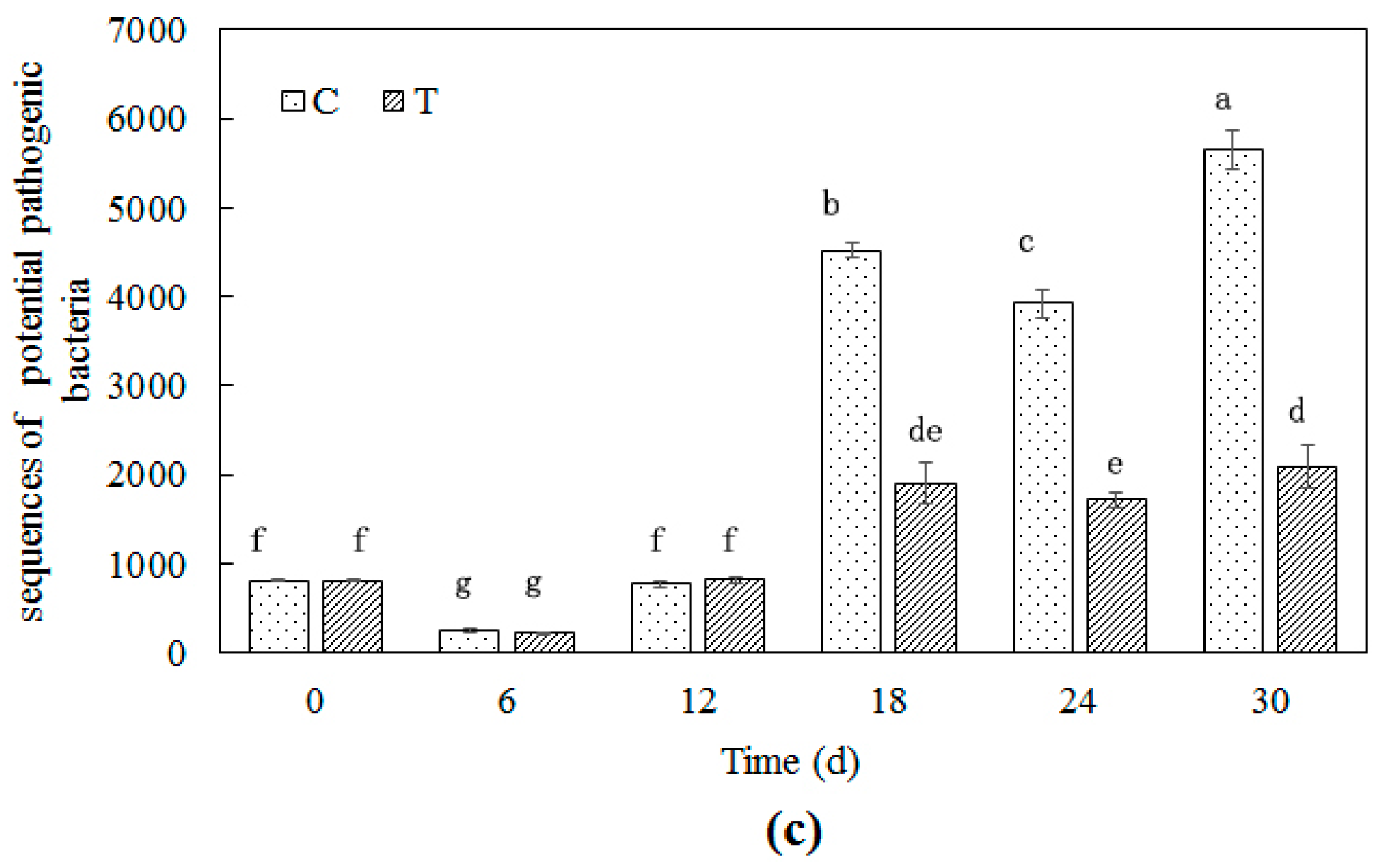
3.3.3. Potential Pathogenic Bacteria
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Yi, Q. Aquaculture industry exports nearly 300 million cubic meters of wastewater per year in our country. Mod. Prop. Manag. 2014, Z1, 93. [Google Scholar]
- Malik, A. Environmental challenge vis a vis opportunity: The case of water hyacinth. Environ. Int. 2007, 33, 122–138. [Google Scholar] [CrossRef] [PubMed]
- Akinbile, C.O.; Yusoff, M.S. Assessing water hyacinth (Eichhornia crassopes) and lettuce (Pistia stratiotes) effectiveness in aquaculture wastewater treatment. Int. J. Phytoremediat. 2012, 14, 201–211. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.X.; Zhang, L.M.; Chua, H.; Li, X.D.; Xia, M.F.; Pu, P.M. A mosaic community of macrophytes for the ecological remediation of eutrophic shallow lakes. Ecol. Eng. 2009, 35, 582–590. [Google Scholar] [CrossRef]
- Muyodi, F.J.; Rubindamayugi, M.S.; Semesi, A.K. Effect of water hyacinth on distribution of sulphate-reducing bacteria in sediments of Lake Victoria. Water SA 2004, 30, 421–425. [Google Scholar] [CrossRef]
- Yi, N.; Gao, Y.; Long, X.; Zhang, Z.; Guo, J.; Shao, H.; Zhang, Z.; Yan, S. Eichhornia crassipes cleans wetlands by enhancing the nitrogen removal and modulating denitrifying bacteria community. Clean Soil Air Water 2014, 42, 664–673. [Google Scholar] [CrossRef]
- Wei, B.; Yu, X.; Zhang, S.; Gu, L. Comparison of the community structures of ammonia-oxidizing bacteria and archaea in rhizoplanes of floating aquatic macrophytes. Microbiol. Res. 2011, 166, 468–474. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Li, B.; Sun, S.; Wang, L.; Zhang, M.; Zhu, J. Phytoplankton community structure and its relationship to water quality parameters in a compound aquaculture system. J. Hydroecol. 2015, 36, 81–88. [Google Scholar]
- Valdor, R.; Aboal, M. Effects of living cyanobacteria, cyanobacterial extracts and pure microcystins on growth and ultrastructure of microalgae and bacteria. Toxicon 2007, 49, 769–779. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Kong, F. The determination of in situ growth rates of the bloomed Microcystis in Meiliang Bay, Lake Taihu. China Environ. Sci. 2008, 28, 552–555. [Google Scholar]
- Deng, J. Studies on the algicidal effects and mechanisms of an algae-lysing bacterial culture agaist Microcystis aeruginosa. In Chengdu Institute of Biology; Chinese Academy of Sciences: Chengdu, China, 2009. [Google Scholar]
- De Oliveira, L.F.V.; Margis, R. The source of the river as a nursery for microbial diversity. PLoS ONE 2015, 10, e120608. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Li, G.; Wang, C.; Jing, Y.; Zhu, Y.; Zhang, S.; Liu, Y. Community dynamics of prokaryotic and eukaryotic microbes in an estuary reservoir. Sci. Rep. UK 2014, 4, 6966. [Google Scholar] [CrossRef] [PubMed]
- Wilhelm, S.W.; Farnsley, S.E.; LeCleir, G.R.; Layton, A.C.; Satchwell, M.F.; DeBruyn, J.M.; Boyer, G.L.; Zhu, G.; Paerl, H.W. The relationships between nutrients, cyanobacterial toxins and the microbial community in Taihu (Lake Tai), China. Harmful Algae 2011, 10, 207–215. [Google Scholar] [CrossRef]
- Shen, P.; Fan, X.; Li, G. Microbiology Experiment, 3rd ed.; Higher Education Press: Beijing, China, 2001; p. 93. [Google Scholar]
- SEPAC (State Environmental Protection Administration of China). HJ/T 347 Water Quality-Determination of Fecal Coliform-Manifold Zymotechnics and Filter Membrane; China Environmental Science Press: Beijing, China, 2007.
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Met. 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microb. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Bibby, K.; Viau, E.; Peccia, J. Pyrosequencing of the 16S rRNA gene to reveal bacterial pathogen diversity in biosolids. Water Res. 2010, 44, 4252–4260. [Google Scholar] [CrossRef] [PubMed]
- Ye, L.; Zhang, T. Pathogenic bacteria in sewage treatment plants as revealed by 454 pyrosequencing. Environ. Sci. Technol. 2011, 45, 7173–7179. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Li, Z. Insitu nutrient removal from aquaculture wasterwater by aquatic vegetable Ipomoea aquatica on floating beds. Water Sci. Technol. 2009, 59, 1937–1943. [Google Scholar] [CrossRef] [PubMed]
- Eiler, A.; Bertilsson, S. Composition of freshwater bacterial communities associated with cyanobacterial blooms in four Swedish lakes. Environ. Microbiol. 2004, 6, 1228–1243. [Google Scholar] [CrossRef] [PubMed]
- Xiong, W.; Sun, Y.; Zhang, T.; Ding, X.; Li, Y.; Wang, M.; Zeng, Z. Antibiotics, antibiotic resistance genes, and bacterial community composition in fresh water aquaculture environment in China. Microb. Ecol. 2015, 70, 425–432. [Google Scholar] [CrossRef] [PubMed]
- Zhong, F.; Wu, J.; Dai, Y.; Yang, L.; Zhang, Z.; Cheng, S.; Zhang, Q. Bacterial community analysis by PCR-DGGE and 454-pyrosequencing of horizontal subsurface flow constructed wetlands with front aeration. Appl. Microbiol. Biotechnol. 2015, 99, 1499–1512. [Google Scholar] [CrossRef] [PubMed]
- Ansola, G.; Arroyo, P.; Sáenz de Miera, L.E. Characterisation of the soil bacterial community structure and composition of natural and constructed wetlands. Sci. Total Environ. 2014, 473–474, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Ofek, M.; Hadar, Y.; Minz, D. Ecology of Root Colonizing Massilia (Oxalobacteraceae). PLoS ONE 2012, 7, e40117. [Google Scholar] [CrossRef] [PubMed]
- Weissbrodt, D.; Lochmatter, S.; Neu, T.R.; Holliger, C. Significance of Rhodocyclaceae for the formation of aerobic granular sludge biofilms and nutrient removal from wastewater. In Proceedings of the IWA Biofilm Conference, Shanghai, China, 27–30 October 2011; pp. 106–107. [Google Scholar]
- Ge, H.; Batstone, D.J.; Keller, J. Biological phosphorus removal from abattoir wastewater at very short sludge ages mediated by novel PAO clade Comamonadaceae. Water Res. 2015, 69, 173–182. [Google Scholar] [CrossRef] [PubMed]
- Rejmankova, E. The role of macrophytes in wetland ecosystems. J. Ecol. Field Biol. 2011, 34, 333–344. [Google Scholar] [CrossRef]
- Lu, S.; Zhang, C.; Xu, J. Root exudates of wetland plants and the influence on the microbial community in constructed wetlands. Ecol. Environ. Sci. 2011, 20, 676–680. [Google Scholar]
- Zhou, Q.; Han, S.; Yan, S.; Guo, J.; Song, W.; Liu, G. Impacts of Eichhornia crassipes (Mart.) Solms stress on the physiological characteristics, microcystin production and release of Microcystis aeruginosa. Biochem. Syst. Ecol. 2014, 55, 148–155. [Google Scholar] [CrossRef]
- Ramirez, E.; Robles, E.; Martincz, B. Free-living amoebae isolated from water-hyacinth root (Eichhornia crassipes). Exp. Parasitol. 2010, 126, 42–44. [Google Scholar] [CrossRef] [PubMed]
- El-Deeb, B.; Gherbawy, Y.; Hassan, S. Molecular charaterization of endophytic bacteria from metal hyperaccumulator aquatic plant (Eichhornia crassipes) and its role in heavy metal removal. Geomicrobiol. J. 2012, 29, 906–915. [Google Scholar] [CrossRef]
- Orta de Velásquez, M.T.; Rojas-Valencia, M.N.; Ayala, A. Wastewater disinfection using ozone to remove free-living, highly pathogenic bacteria and amoebae. Ozone Sci. Eng. 2008, 30, 367–375. [Google Scholar] [CrossRef]
- Sotiriou, G.A.; Pratsinis, S.E. Antibacterial activity of nanosilver ions and particles. Environ. Sci. Technol. 2010, 44, 5649–5654. [Google Scholar] [CrossRef] [PubMed]
- Bomo, A.M.; Husby, A.; Steik, T.K. Removal of fish pathogenic bacteria in biological sand filters. Water Res. 2003, 37, 2618–2626. [Google Scholar] [CrossRef]
| Sample | Valid Sequences | Sample | Valid Sequences |
|---|---|---|---|
| C0 | 40,114 ± 1224 | T0 | 40,114 ± 1224 |
| C6 | 41,332 ± 2231 | T6 | 38,333 ± 2264 |
| C12 | 42,015 ± 1989 | T12 | 38,563 ± 2005 |
| C18 | 41,142 ± 2388 | T18 | 37,364 ± 1908 |
| C24 | 40,805 ± 1868 | T24 | 38,579 ± 2179 |
| C30 | 38,876 ± 2121 | T30 | 39,961 ± 1645 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhou, Q.; Chen, T.; Han, S. Characteristics of Bacterial Communities in Cyanobacteria-Blooming Aquaculture Wastewater Influenced by the Phytoremediation with Water Hyacinth. Water 2017, 9, 956. https://doi.org/10.3390/w9120956
Zhou Q, Chen T, Han S. Characteristics of Bacterial Communities in Cyanobacteria-Blooming Aquaculture Wastewater Influenced by the Phytoremediation with Water Hyacinth. Water. 2017; 9(12):956. https://doi.org/10.3390/w9120956
Chicago/Turabian StyleZhou, Qing, Ting Chen, and Shiqun Han. 2017. "Characteristics of Bacterial Communities in Cyanobacteria-Blooming Aquaculture Wastewater Influenced by the Phytoremediation with Water Hyacinth" Water 9, no. 12: 956. https://doi.org/10.3390/w9120956
APA StyleZhou, Q., Chen, T., & Han, S. (2017). Characteristics of Bacterial Communities in Cyanobacteria-Blooming Aquaculture Wastewater Influenced by the Phytoremediation with Water Hyacinth. Water, 9(12), 956. https://doi.org/10.3390/w9120956




