1. Introduction
Packaging is an integral part of logistics systems, continuously accompanying, supporting, and complementing the processes that occur in time and space. Throughout logistics processes, the packaged product is exposed to various forms, intensities, and combinations of external influences, such as stresses, which may result in quantitative and/or qualitative damage to the product. In the case of food, any decrease in quality is of enormous importance, as it can impact not only the product’s sensory and nutritional properties but also its safety. The distribution of goods subjects them to several types of mechanical stresses, such as stationary sinusoidal vibration, stationary random vibration, non-stationary vibration, fall, overturn, rocking, tumbling, and constant acceleration. Sustained shaking stresses, including stationary random vibration, are unavoidable during the distribution process [
1]. The shock stress on product packaging systems is caused by a combination of track unevenness, unbalanced rotational masses in vehicles, the vehicle’s suspension system, and the properties of the track support [
2,
3,
4]. The movements measured on the load surface are random, meaning that both the frequency and amplitude of the vibration strongly vary over time. The shaking involves six degrees of freedom, including surge, heave, sway, roll, yaw, and pitch [
5], of which the vertical jolt is the most intense in reality [
6].
The purpose of conducting a laboratory vibration test is to replicate the random vibration environment that occurs during real transportation. This broad-spectrum random vibration is generated by an electrodynamic shaker system in the frequency range of 1–200 Hz [
7]. The specific frequency range was chosen because it encompasses the most intense vibration responses of the vehicle’s components, including the suspension, wheels, and structure. This is often referred to as the power spectral density (PSD) of random vibrations, which is calculated as the fast Fourier transform (FFT) of random and non-stationary acceleration signals (g). The PSD provides information on the characteristic of motion based on the power of motion, expressed in g
2/Hz, as a function of frequency (Hz) [
6].
In Hungary, there are few foods whose per capita consumption is increasing as rapidly as bottled natural mineral water. In the 1980s and early 1990s, the per capita consumption was relatively stable at around 3 L/person per year. However, in 1993, consumption started to grow dynamically at a rate of 20–30% per year, and by 2019, it reached 131 L/person per year. Although the amount of mineral water consumption per person decreased in 2020, Hungary is still among the top five countries in the European Union [
8].
Consumers worldwide tend to believe that bottled mineral water is safer to drink than tap water [
9,
10,
11,
12]. Despite the limitations of microbial development, such as the lack of organic nutrients or high mineral content, mineral waters are characterized by a specific microbiota that is unique to the water source [
13,
14,
15,
16]. This microbiota, which is a complex ecosystem with phenotypic and genotypic diversity [
10,
17,
18], can be broadly divided into two groups: allochthonous and autochthonous. The allochthonous microbiota includes microorganisms introduced from the environment, whose survival in bottled water is low due to the low nutrient content. However, several authors have isolated pathogenic bacteria and viruses from natural mineral water a few weeks after bottling [
17,
19,
20,
21,
22]. The majority of microorganisms living in groundwater are heterotrophic bacteria, among which pathogens occur, mainly coliforms such as
Escherichia,
Enterobacter,
Citrobacter, and
Klebsiella spp., as well as opportunistic pathogens [
23].
Due to the limited availability of nutrients in the water base, the bacteria that make up the natural microbiota of springs and mineral waters are often in a state of starvation. As a result, their morphology can change [
24], and the shape of the cells can become deformed; they can take on an egg shape, and their diameter can decrease. These cells may pass through the usual 0.45 µm membrane filter and even a 0.2 µm membrane filter [
14,
25]. Therefore, it may happen that the membrane filtration used in everyday routine is ineffective for their detection. A study conducted by França et al. [
26] has revealed that the number of microbes that can be detected using traditional culture methods is only 0.0002% of the actual microbial content.
Studies by other authors show that initial low microbial counts of approximately 10
1 CFU/cm
3 a few days after bottling [
10,
14,
27] can increase to 10
4–10
5 CFU/cm
3 following 1–3 weeks of storage [
9,
28,
29]. This increased colony count then becomes permanent for a longer period, and after up to 2 years of storage, the total microbial content of bottled mineral water can still be 10
3 CFU/cm
3 [
30]. Directive 2009/54/EC [
31] establishes water quality requirements at the time of bottling. After bottling, the total colony count of water samples at the source must not exceed 100 CFU/cm
3 when incubated at 20 to 22 °C for 72 h on agar-agar or an agar-gelatin mixture, and 20 CFU/cm
3 if incubated at 37 °C for 24 h on agar-agar. The total colony count must be measured within 12 h of bottling, with the water maintained at 4 ± 1 °C during this period. The traditional culture method is required for carrying out microbiological tests for natural mineral water, as indicated by the wording of the directive. However, there are no regulations specifying a maximum allowable cell count during storage.
Bacteria tend to grow more slowly in glass bottles than in plastic ones, such as polyethylene terephthalate (PET) or polyvinyl chloride (PVC) [
30]. Extremophilic bacteria present in heterotrophic plate count (HPC) are viable for a long time in bottled water due to a number of survival mechanisms. In the presence of trace elements, they are even capable of degrading PET [
32]. Gautam et al. [
33] found a close correlation between the type of bottle and HPC, coliform, and
Pseudomonas counts. They also found that 40% more substandard-quality water was present in damaged bottles than in intact ones. A dent on the bottle promotes the formation of biofilm by lipase-positive
Pseudomonas spp., which subsequently causes molecular changes in the surface of the plastic, indicating the initial stage of plastic degradation [
34].
The primary reasons for choosing bottled water are its perceived quality and safety. While acute illnesses from bottled water are rare indeed, some outbreaks have been linked to the consumption of bottled mineral water [
35,
36,
37]. High levels of heterotrophic bacteria in bottled water can lead to illness, particularly in vulnerable populations, such as infants, the elderly, pregnant women, immunosuppressed individuals, like those with HIV/AIDS, and people suffering from COVID-19, tuberculosis, or malnourishment [
33,
38,
39,
40].
Transportation-induced vibration can cause various physicochemical changes in different types of food, such as beer, wine, and milk [
41,
42,
43,
44,
45,
46,
47,
48]. In a previous study [
49], low-intensity mechanical agitation was found to increase the specific growth rate of both autochthonous and allochthonous microbes in freshly bottled natural mineral water. When examining the microbiological status of freshly bottled mineral water using a traditional method, high-intensity shaking resulted in a significant increase in the specific growth rate of allochthonous species compared to autochthonous ones, which could pose a potential health risk.
The objective of this research was to investigate the potential correlation between the intensity of mechanical agitation and the number of detectable microorganisms in bottled mineral water.
2. Materials and Methods
2.1. Mineral Water Samples
In this study, two model matrices were used as follows: (1) freshly bottled (F) non-carbonated natural mineral water sourced from a bottling plant located in north-western Hungary, and (2) three different types of non-carbonated natural mineral water brands (A, B, C) purchased from various bottling locations and times within Hungary. The hypothesis was that subjecting these different mineral waters, which have varying content properties, to a certain intensity of time-accelerated mechanical impact would not result in significant differences in their microbiome.
All mineral water samples were filled into 0.5 L PET bottles, with six bottles comprising each packaging unit. For each vibration test, 15 packaging units (i.e., 15 × 6 bottles) were shaken. New units were always used for microbiological testing. The main chemical components of the natural mineral waters examined are listed in
Table 1 and
Table 2. These data are mandatory information on mineral water bottles. The size and shape of the PET bottles varied across different bottlers. The dimensions of each bottle type were as follows: F: 8 cm diameter × 21 cm height, A: 7 cm diameter × 21.7 cm height, B: 7 cm diameter × 23 cm height, and C: 6 cm diameter × 23.3 cm height.
2.2. Vibration Test
Dynamic mechanical vibration was simulated at various performance levels using both time-accelerated and real-time tests, with each level being tested three times. The PSD curve in the ASTM D4169-16 standard of the American Society of Testing Materials [
50] was utilized during the research. This standard replicates the conditions of the shaking motion experienced during semi-trailer transportation with leaf springs, which is most typical for bottled products. The standard proposes three curves characterized by different performance levels (i.e., low, medium, and high). The test parameters are shown in
Table 3. In this study, the vertical vibration relationship was reproduced.
In the laboratory, the shaking stress was simulated on a shaking table operating according to the servo-hydraulic principle. The shaking table performs vibrations of different waveforms in the frequency range of 1–200 Hz, with a maximum amplitude of 200 mm. For the laboratory simulation of vibration and climatic stress, the electrodynamic shaking device (TIRA TV59355, TIRA GmbH, Schalkau, Germany) and VR Research 1100 controller, as well as a connected climate cabinet (Angelantoni AV600C, Angelantoni S.p.a., Massa Martana, Italy) were used. The latter ensured a constant temperature of 22 ± 1 °C during the shaking test. After shaking, the samples were stored in an air-conditioning cabinet (ESPEC PR-3ST, Espec, Osaka, Japan) at 5 ± 3 °C to avoid changes in their microbiological status. In this test, a test duration of 60 min can be understood as 480 km.
2.3. Sampling
The samples collected from the electrodynamic shaker were kept at a temperature of 5 ± 3 °C until the microbiological tests were conducted. During transport to the laboratory, a Styrofoam thermal box (Gasztronagyker Ltd., Debrecen, Hungary) and an ice battery were used to maintain the same temperature. Samples (3 × 1 bottle, 0.5 L each) were taken before and after each shaking, and then at 12 h intervals for time-accelerated tests and 24 h intervals for real-time shaking tests. A control sample of non-shaken natural mineral water, sourced from the same bottling location and time as the other samples, was included in all cases. This control sample was stored at 22 ± 1 °C, with the same bottling time as the shaken samples.
2.4. Molecular Biological Studies
Tihanyi-Kovács et al. [
49] isolated
Acidovorax temperans from freshly bottled natural mineral water. The genus
Acidovorax belongs to the class of Betaproteobacteria. So far, all described species have been divided into two major groups, namely, plant pathogens and species occupying aquatic and soil habitats.
The linear order of nucleic acid bases in the DNA was determined by capillary-based, Sanger-type sequencing [
51] with the assistance of a service provider (Macrogen, Amsterdam, The Netherlands). In addition to
A. temperans, sequences of the bacterial species/strains listed in
Table 4 were used to design primers. Since the microbial composition of mineral water is specific to each water source, degenerate oligonucleotides were designed to quantitatively determine the presence of bacteria relevant to the food industry. The oligonucleotides showed a significant match to the reference sequences listed in
Table 4.
The changes in microbial counts during agitation were monitored using the qPCR method, which employed specific primers. The primers used in the qPCR tests to detect bacteria relevant to the food industry were as follows:
These primers were slightly modified sequences used generally in metagenome analyses that apply new generation sequencing of the V4 region of the 16S rRNA coding gene.
2.4.1. DNA Purification from Mineral Water
After homogenizing the water samples, 1 mL was measured and transferred into a 1.5 mL sterile Eppendorf tube (Greiner Bio-One Hungary Ltd., Mosonmagyaróvár, Hungary). The samples were then centrifuged using a BioSan Microspin 12 centrifuge (Biocenter Ltd., Szeged, Hungary) for 6 min at 11,700× g. The supernatant was removed and 100 μL of a 10% Chelex-100 solution (Bio-Rad Hungary Ltd., Budapest, Hungary) was added to the pellet. The pellet–Chelex mix was incubated at 100 °C for 10 min with continuous shaking using a Thermo-Shaker (PHMT; Grant Bio, Shepreth, UK). After incubation, the samples were centrifuged at 13,000× g for 1 min. The purified DNA samples were stored at −20 ± 1 °C until the qPCR tests were performed.
It should be noted that the Chelex-100 treatment of the bacterial pellet at 100 °C does not isolate DNA, but rather purifies it by binding and depleting PCR inhibitors that can interfere with subsequent amplification steps [
52,
53].
2.4.2. Quantitative PCR (qPCR)
To detect bacterial DNA in the water using the Piko Real PCR system (Thermo Fisher Scientific, Waltham, MA, USA), we used the Bioline SensiFAST SYBR No-Rox kit (Izinta Ltd., Budapest, Hungary) as recommended by the manufacturer. The PCR cycling conditions are shown in
Table 5.
The frozen, purified DNA samples were thawed at room temperature and then centrifuged at 13,000× g for 1 min. We used 9 μL of master mix and 1 μL of supernatant of the purified DNA in each PCR reaction.
2.4.3. Determination of Total Microbial Count
To ensure that the test is appropriate for determining the changes in total microbial counts, we prepared a decimal dilution series from the suspension of
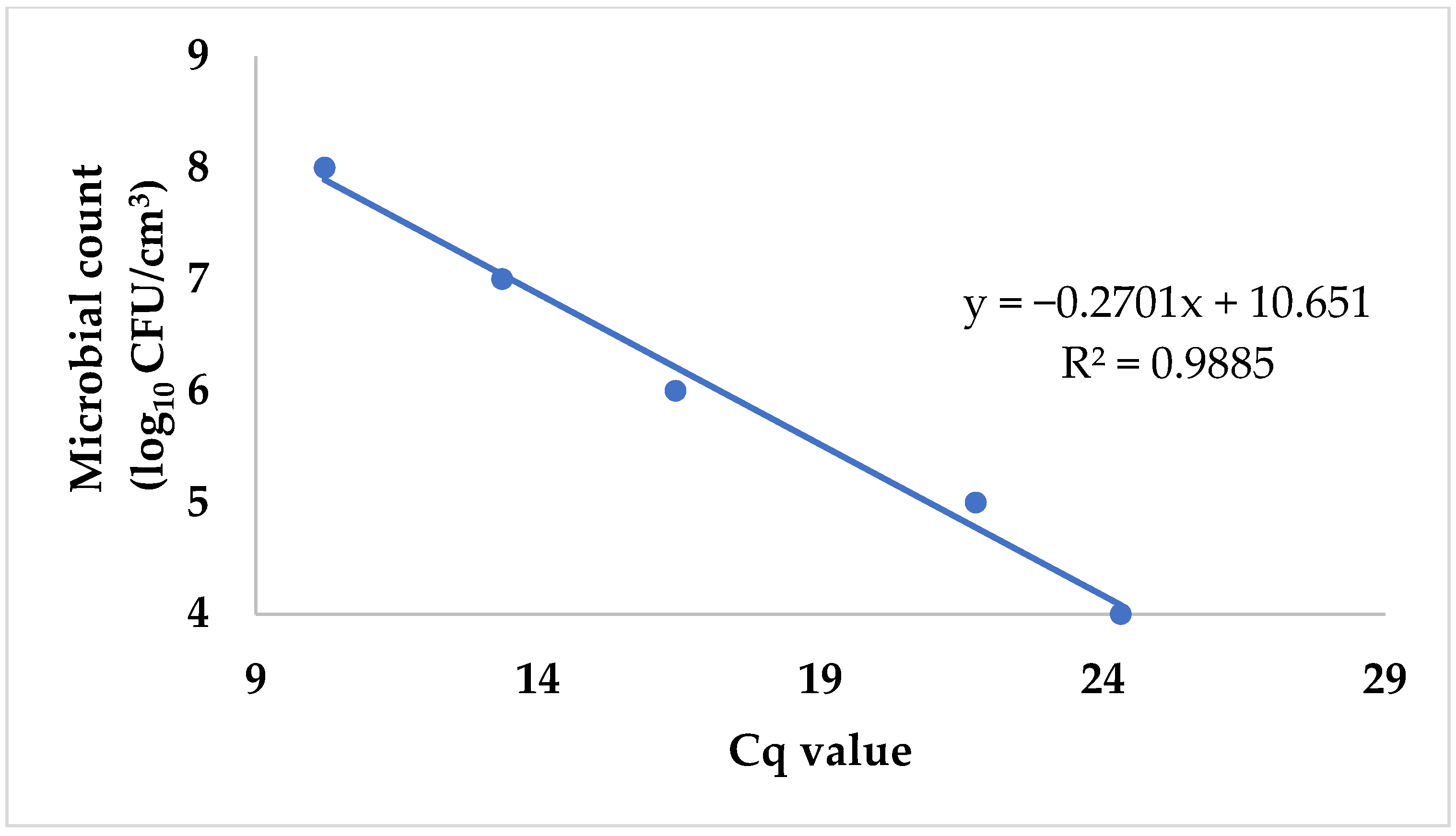
Escherichia coli ATCC 8739. Next, we purified the DNA from each sample of the dilution series and ran them following the qPCR protocol previously described. In addition, we used the pour plate technique with ChromoBio Coliform agar (Biolab Ltd., Budapest, Hungary) to determine the total plate count of the original solution. The Petri dishes were then incubated at 36 ± 1 °C for 21 ± 3 h, and the colony count was performed. We assigned the logarithmically transformed cell counts to the corresponding Cq values obtained during qPCR runs. Plotting the data allowed us to create a standard curve that could subsequently be used to convert Cq values into logarithmic cell counts (
Figure 1).
2.5. Statistical Analysis
Microsoft Excel 365 (Microsoft Corporation, Redmond, WA, USA) was used to process and statistically evaluate the results. Differences between shakes performed at different intensities were evaluated using the F-test and Student’s
t-test at a 95% significance level (
p < 0.05). To supplement these calculations, the Kruskal–Wallis test extended with the Mann–Whitney U test was used [
54,
55,
56], also at a 95% significance level (
p < 0.05). A Bonferroni correction was applied during the comparison. To analyze the relationship between time-accelerated and real-time simulations, a regression analysis was performed.
3. Results and Discussion
3.1. Real-Time Shaking Tests
As a general rule, the microbial growth in control samples that were not subjected to shaking was slower (μcontrol = 0.005 ± 0.001 1/h) than in samples that were exposed to mechanical agitation (μlow = 0.013 ± 0.001 1/h, μmedium = 0.006 ± 0.001 1/h, μhigh = 0.008 ± 0.001 1/h), with a significant difference (p < 0.05) being observed at both low and high intensities. It is worth noting that the specific growth rate (μ) represents the rate at which the microbial population increases over time. During the exponential phase, the number of microbes obtained was half an order of magnitude higher at low-intensity agitation and one order of magnitude higher at high-intensity agitation compared to the initial cell count. However, medium-intensity shaking had no effect (p > 0.05) on the number of microbes reached at the end of the exponential phase.
There was no difference (p > 0.05) in the maximum cell counts achieved during the entire study period when compared to the control (5.49 ± 0.01 log10 CFU/cm3). The maximum colony count values obtained from the shaken samples ranged from 5.44 to 5.51 log10 CFU/cm3. Medium-intensity agitation resulted in the lowest count, whereas high-intensity agitation produced the highest count.
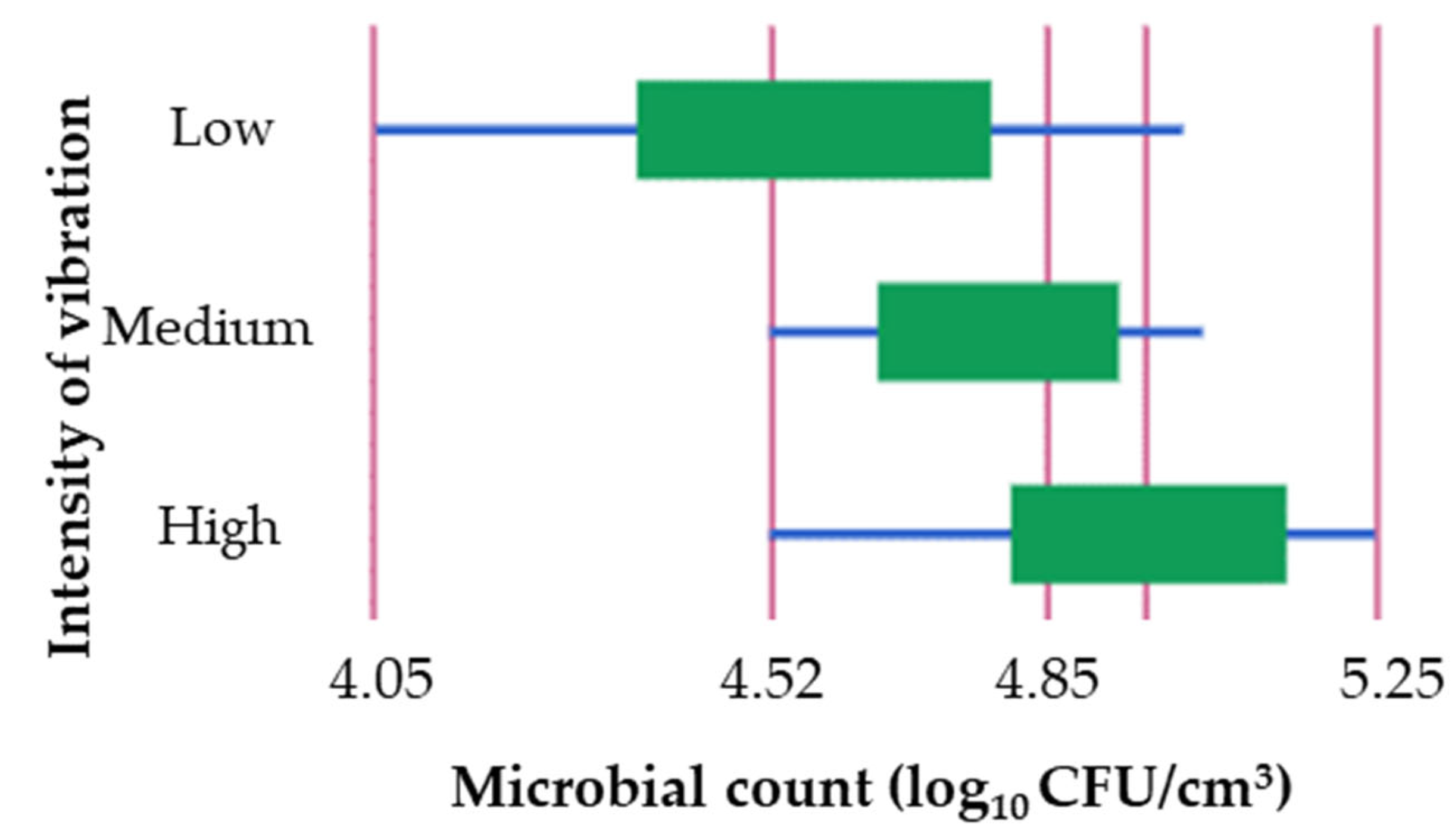
To investigate whether there were differences in microbial counts obtained during the exponential phase of growth when shaken with different intensities, a study was conducted. The results showed a significant difference (
p < 0.05) between low and high intensity levels (
Table 6). The highest mean cell count of 4.95 log
10 CFU/cm
3 was observed at high intensity during the exponential phase of growth (
Figure 2).
Comparing the specific growth rates of microorganisms at all three levels of vibration intensity, a significant difference (
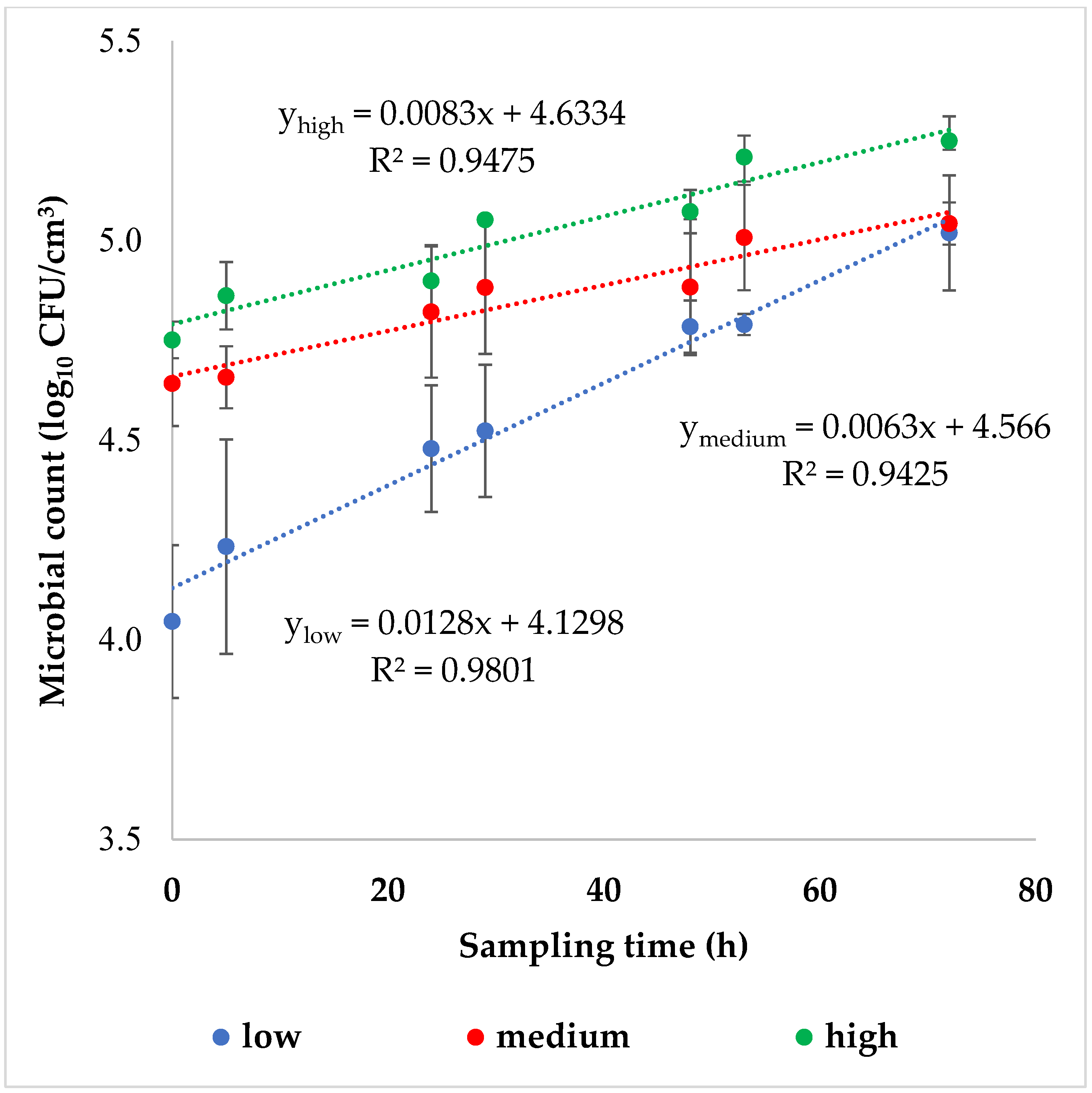
p < 0.05) was found. The generation time (tg), which is the time it takes for the microbial population to double in size, was greatly reduced in the samples shaken at low intensity (tg = 76.9 ± 6.0 h), compared to those shaken at medium (tg = 166.7 ± 28.6 h) or high (tg = 125.0 ± 15.9 h) intensities. Despite the initial cell counts being considerably different, the final cell counts were similar across all three agitation intensities tested, as shown in
Figure 3.
3.2. Time-Accelerated Shaking
As shown in
Table 7, significant differences (
p < 0.05) were observed in the specific growth rates of microbial populations in bottled natural mineral water samples subjected to various shaking intensities specified in the ASTM D-4169-16 standard [
50]. It is noteworthy that the generation times in freshly bottled water (F) were reduced by approximately 70–80% compared to the control group (i.e., tg
control F = 200.0 ± 41.5 h), with the lowest generation time being 41.7 ± 3.2 h for high-intensity shaking, followed by 45.5 ± 9.9 h and 58.8 ± 3.5 h for medium and low shaking intensities, respectively.
The microbiome of samples A, B, and C exhibited a significant increase (
p < 0.05) in growth rate due to shaking, compared to their respective controls, except for sample A under low intensity and sample B under high intensity. However, for sample C, the high-intensity mechanical impact resulted in a reduction of the specific growth rate (
Table 7).
The effect of low-intensity mechanical vibration on microbial growth varied across the samples, resulting in different cell counts in the exponential growth phase (data not shown). Samples B and C demonstrated the highest mean cell counts during this phase (5.36 and 5.55 log10 CFU/cm3, respectively), whereas sample A had the lowest mean cell count (4.87 log10 CFU/cm3). Statistical analysis revealed significant differences (p < 0.05) between several sample pairs, including F–A, A–B, and A–C.
For samples F, B, and C, there was no difference (p > 0.05) in the cell count achieved during the study, with maximum log10 CFU/cm3 values ranging from 5.98 to 6.09. In contrast, sample A had a significantly lower (p < 0.05) maximum cell count of 5.19 log10 CFU/cm3.
As shown in
Table 7, the specific growth rates of microbial populations in each sample were affected differently by shaking at low intensity. Sample C showed the highest rate of microbial population growth, whereas sample A demonstrated the lowest growth rate.
For all samples, medium-intensity vibration resulted in an extension of the exponential phase of growth, which was observed to last between 40 h and 70 h, as opposed to the previous duration of 25 h. In sample A, microbial multiplication began after the second shaking, while in sample B, after the first. Despite the initially low microbial count of 4.70 log10 CFU/cm3, with medium mechanical vibration, the number of CFUs in the exponential phase of growth for sample A increased by more than one order of magnitude (1.64 Δlog10 CFU/cm3), compared to low-intensity agitation (0.27 Δlog10 CFU/cm3). For samples F, B, and C, the changes in log10 CFU/cm3 values during the exponential phase of growth with medium-intensity agitation were 0.77, 0.83, and 0.91, respectively.
There was no significant difference (
p > 0.05) in the specific growth rates at medium intensity, as indicated in
Table 7. However, when comparing the cell numbers of the sample pairs, a significant difference (
p < 0.05) was observed for both the F–B and F–C pairs during medium-intensity vibration. The maximum cell numbers ranged from 6.2 to 6.7 log
10 CFU/cm
3, higher than those observed during low-intensity shaking.
As for high-intensity mechanical agitation, a latent period of microbial growth was only observed for the C water sample. The time of rapid microbial growth in sample A was 13 h, whereas it was 37 h in the other three samples. In all cases, except for the F–C sample pair, a significant difference (
p < 0.05) was observed in the specific growth rates, as shown in
Table 7. For samples F, A, B, and C, the changes in cell numbers (i.e., Δlog
10 CFU/cm
3 values) during the exponential phase of growth with high-intensity agitation were 0.91, 0.46, 0.53, and 0.77 log
10 cycles, respectively. Moreover, when comparing the viable cell counts of sample pairs, a significant difference (
p < 0.05) was found for F–A, F–B, F–C, and A–B during high-intensity shaking.
The correlation between shaking intensity and microbial growth was also investigated in each sample. Results showed a significant difference (
p < 0.05) in the specific growth rate at each intensity in almost all cases (
Table 7), except for the low–medium pairwise comparison of the freshly bottled samples. Sample C was the only one that exhibited a decrease in specific growth rate with increasing agitation intensity. In contrast, sample A showed an increase in specific growth rate with increasing vibration intensity.
The generation times calculated from the specific growth rates determined during the exponential growth phase are presented in
Table 8. The results show that all samples, except for sample C at high intensity, experienced a decrease in generation time due to vibration. The most significant changes were observed in freshly bottled mineral water.
3.3. Comparison of Real-Time and Time-Accelerated Test Results of Freshly Bottled Natural Mineral Water
The time-accelerated tests conducted at all three intensities of the ASTM D-4169-16 standard [
50] resulted in significantly higher growth rates (
Table 9) and lower generation times (
Table 10) compared to real-time shaking.
The rate of increase in cell counts observed at both medium and high vibration intensities was similar for real-time and time-accelerated test methods (
Table 11). In contrast, low-intensity time-accelerated shaking induced half a log
10 cycle higher increase in microbial counts compared to low-intensity real-time shaking.
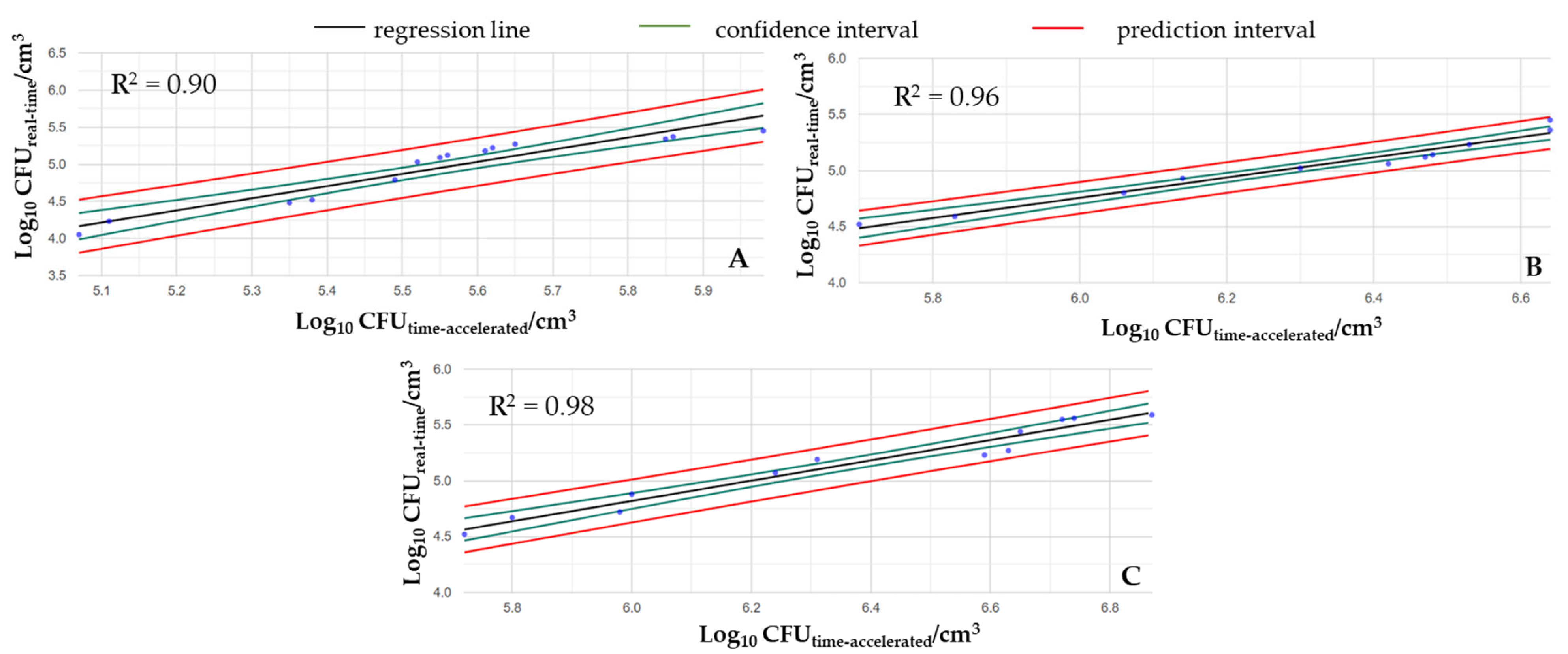
Figure 4 shows that the changes in cell counts caused by real-time mechanical shocks can be approximated well by time-accelerated mechanical agitation. As the intensity of shaking increased, the regression of the examined variable also increased, which is illustrated by the R
2 values of the fitted curves (i.e., R
2low = 0.90, R
2medium = 0.96, and R
2high = 0.98).
To our knowledge, there are no publications on changes in the microbiome of natural mineral waters due to mechanical agitation or tests conducted using the ASTM D4169-16 standard [
50]. Therefore, we were only able to compare our results to those from publications that used somewhat different methodologies.
The time lapse between individual tests, which ranged from several weeks to several months, may have impacted the microbial composition of the catchment sites. Carraturo et al. [
57] also observed similar changes in water samples collected from the same location during different seasons. These changes may have influenced the initial plate counts through microbial interactions, which could have affected the test results. The intensity of mechanical agitation affected the samples from various bottling locations and times differently. This may be due to the presence of distinct genotypic and phenotypic microbial populations at each water catchment site, as reported by several studies [
10,
13,
14,
15,
16,
17,
18,
26], as well as the time elapsed since bottling. The microbial composition can change with time [
58], which may account for the differences in the observed growth rates among the various natural mineral waters. There was no significant difference in microbial count among the individual samples tested.
Using the qPCR method, we detected a maximum microbial load of 10
5–10
7 CFU/cm
3, which is one to two orders of magnitude higher than the findings of other researchers [
9,
14,
27,
28,
29]. In these studies, however, microbial counts were determined using conventional culturing techniques. Moreover, it is important to note that live cells were not distinguished from dead cells in the present study. The qPCR method and the primers designed were shown to be appropriate for accurately quantifying total microbial DNA in natural mineral water samples, regardless of whether the cells were live or dead. PCR is a rapid and sensitive technique for detecting microbes in food [
59]. However, it cannot differentiate between viable and dead cells [
60], resulting in an overestimation of the number of viable cells in samples [
61,
62]. While this may not be crucial for the study’s objectives, which involved evaluating the correlation between mechanical agitation and microbial counts, future improvements to the method may benefit from distinguishing between live and dead cells. The qPCR-PMA (propidium monoazide) method is a potential solution for such differentiation [
63,
64,
65].
The time-accelerated tests were performed in accordance with the ASTM D4169-16 standard [
50], and it was observed that microbial growth during the exponential phase was significantly faster at all intensities than during real-time agitation. The change in cell numbers over time between the two methods exhibited a close correlation. These findings suggest that the time-accelerated method is a suitable approach for simulating the impact of real-time mechanical agitation on microbiome changes at low, medium, and high vibration intensities.
It would be worthwhile to extend this investigation to monitor the microbiological status of other food matrices, such as beer, wine, and milk, which are susceptible to physicochemical changes during transportation that can affect product quality.