Isolation and Identification of Bacterial Communities in Neutral Mine Drainage in Central Slovakian Neovolcanites (Slovakia)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sampling and Storage
2.2. Cultivation, Gram Staining
2.3. Preparation of Samples for MALDI TOF MS Profiling
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Mine Drainage. Available online: https://www.usgs.gov/mission-areas/water-resources/science/mine-drainage?qt-science_center_objects=0#qt-science_center_objects (accessed on 16 January 2022).
- Koděra, P.; Lexa, J. Classic localities in Central Slovakia Volcanic Field: Gold, silver and base metal mineralizations and minig history at Banská Štiavnica and Kremnica. Acta Mineral.-Petrogr. 2010, 29, 19. [Google Scholar]
- Števko, M.; Sejkora, J.; Malíková, R. New data on supergene minerals from the Banská Štiavnica deposit (Slovak Republic). Bull. Mineral. Petrol. 2018, 26, 90–101. [Google Scholar]
- Čiastkový Monitorovací Systém Geologických Faktorov Životného Prostredia Slovenskej Republiky—Vplyv Ťažby na Životné Prostredie Subsystém 04 Správa za rok 2010. Available online: https://dionysos.geology.sk/cmsgf/files/Hodn_monitor_2010/04_Vplyv_tazby_2010/Hodnotenie_monitorovania_za_rok_2010_v_subsysteme_04_TAZBA.pdf (accessed on 29 October 2022).
- Podsystém 04 Vplyv Ťažby Nerastov na Životné Prostredie—Správa za rok 2020. Available online: https://dionysos.geology.sk/cmsgf/files/Hodn_monitor_2020/04_Vplyv_tazby_2020.pdf (accessed on 29 October 2022).
- Bajtoš, P.; Záhorová, Ľ.; Rapant, S.; Pramuka, S. Monitoring geologických faktorov vplyvu ťažby nerastov na životné prostredie v rizikových oblastiach na Slovensku v rokoch 2007–2011. Miner. Slovaca 2012, 44, 375–392. [Google Scholar]
- Nosáľová, L.; Maliničová, L.; Kisková, J.; Timková, I.; Sedláková-Kaduková, J.; Pristaš, P. Cultivable Microbiota Associated with Gold Ore from the Rozália Gold Mine, Hodruša-Hámre, Slovakia. Geomicrobiol. J. 2021, 38, 415–425. [Google Scholar] [CrossRef]
- Kisková, J.; Perháčová, Z.; Vlčko, L.; Sedláková, J.; Kvasnová, S.; Pristaš, P. The Bacterial Population of Neutral Mine Drainage Water of Elizabeth’s Shaft (Slovinky, Slovakia). Curr. Microbiol. 2018, 75, 988–996. [Google Scholar] [CrossRef]
- Šipošová, N.; Liptáková, V.; Kvasnová, S.; Kosorínová, P.; Pristaš, P. Genetic diversity of Acinetobacter spp. adapted to heavy metal polluted environments. Nova Biotechnol. Chim. 2017, 16, 42–47. [Google Scholar] [CrossRef]
- Serial Dilution Problem Help. Available online: https://www.uvm.edu/~btessman/calc/serhelp.html (accessed on 17 January 2022).
- Kožurková, M. Cvičenia z Biochémie Mikroorganizmov, 1st ed.; Univerzita Pavla Jozefa Šafárika v Košiciach: Košice, Slovakia, 2011; pp. 74–88. [Google Scholar]
- Gram Staining: Principle, Procedure, Interpretation, Examples and Animation. Available online: https://microbiologyinfo.com/gram-staining-principle-procedure-interpretation-examples-and-animation/ (accessed on 20 January 2022).
- Seng, P.; Drancourt, M.; Gouriet, F.; La Scola, B.; Fournier, P.E.; Rolain, J.M.; Raoult, D. Ongoing revolution in bacteriology: Routine identification of bacteria by matrix-assisted laser desorption ionization time-of-flight mass spectrometry. Clin. Infect. Dis 2009, 49, 543–551. [Google Scholar] [CrossRef]
- Singhal, N.; Kumar, M.; Kanaujia, P.K.; Virdi, J.S. MALDI-TOF mass spectrometry: An emerging technology for microbial identification and diagnosis. Front. Microbiol. 2015, 6, 791. [Google Scholar] [CrossRef] [Green Version]
- Murray, P.R. 16—The Clinician and the Microbiology Laboratory. In Mandell, Douglas, and Bennett’s Principles and Practice of Infectious Diseases, 8th ed.; Bennett, J.E., Dolin, R., Blaser, M.J., Eds.; W.B. Saunders: Philadelphia, PA, USA, 2015; pp. 191–223. [Google Scholar] [CrossRef]
- Prepilková, V.; Poništ, J. Analysis of Environmental Impacts of Mine Water in the Area of Central Slovak Neovolcanites. In Proceedings of the Science of Youth 2021, Nitra, Slovakia, 21 May 2021. [Google Scholar]
- Kuang, J.L.; Huang, L.N.; Chen, L.X.; Hua, Z.S.; Li, S.J.; Hu, M.; Li, J.T.; Shu, W.S. Contemporary environmental variation determines microbial diversity patterns in acid mine drainage. ISME J. 2013, 7, 1038–1050. [Google Scholar] [CrossRef] [Green Version]
- Kadnikov, V.V.; Ivasenko, D.A.; Beletsky, A.V.; Mardanov, V.; Danilova, E.V.; Pimenov, N.V.; Karnachuk, O.V.; Ravin, N.V. Effect of metal concentration on the microbial community in acid mine drainage of a polysulfide ore deposit. Microbiology 2016, 85, 745–751. [Google Scholar] [CrossRef]
- Travisany, D.; Di Genova, A.; Sepulveda, A.; Bobadilla-Fazzini, R.A.; Parada, P.; Maass, A. Draft genome sequence of the Sulfobacillus thermosulfidooxidans Cutipay strain, an indigenous bacterium isolated from a naturally extreme mining environment in Northern Chile. J. Bacteriol. 2012, 194, 6327–6328. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pereira, L.B.; Vicentini, R.; Ottoboni, L.M.M. Changes in the Bacterial Community of soil from a Neutral Mine Drainage Channel. PLoS ONE 2014, 9, e96605. [Google Scholar] [CrossRef] [PubMed]
- Pereira, L.B.; Vicentini, R.; Ottoboni, L.M.M. Characterization of the core microbiota of the drainage and surrounding soil of a Brazilian copper mine. Genet. Mol. Biol. 2015, 38, 484–489. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kisková, J.; Stramová, Z.; Javorský, P.; Sedláková-Kaduková, J.; Pristaš, P. Analysis of the bacterial community from high alkaline (pH > 13) drainage water at a brown mud disposal site near Žiar nad Hronom (Banská Bystrica region, Slovakia) using 454 pyrosequencing. Folia Microbiol. 2019, 64, 83–90. [Google Scholar] [CrossRef]
- Majzlan, J.; Lalinska, B.; Chovan, M.; Blass, U.; Brecht, B.; Gottlicher, J.; Gescher, J. A mineralogical, geochemical, and microbiogical assessment of the antimony- and arsenic-rich neutral mine drainage tailings near Pezinok, Slovakia. Am. Mineral. 2010, 96, 1–13. [Google Scholar] [CrossRef]
- Lopes, A.R.; Madureira, D.; Diaz, A.; Santos, S.; Vila, M.C.; Nunes, O.C. Characterisation of bacterial communities from an active mining site and assessment of its potential metal solubilising activity. J. Environ. Chem. Eng. 2020, 8, 104495. [Google Scholar] [CrossRef]
- Lukhele, T.; Selvarajan, R.; Nyoni, H.; Mamba, B.B.; Makudali Msagati, T.A. Affiliations expand Diversity and functional profile of bacterial communities at Lancaster acid mine drainage dam, South Africa as revealed by 16S rRNA gene high-throughput sequencing analysis. Extremophiles 2019, 23, 719–734. [Google Scholar] [CrossRef]
- Oyetibo, G.O.; Enahoro, J.A.; Ikwubuzo, C.A.; Ukwuoma, C.S. Microbiome of highly polluted coal mine drainage from Onyeama, Nigeria, and its potential for sequestrating toxic heavy metals. Sci. Rep. 2021, 11, 17496. [Google Scholar] [CrossRef]
- Sedláková-Kaduková, J.; Kisková, J.; Maliničová, L.; Timková, I.; Jeleň, S.; Pristaš, P. The first evidence of Acidithiobacillus albertensis in weathered ore samples from active gold mine Hodruša-Hámre (Slovakia). Nova Biotechnol. Chim. 2019, 18, 18–24. [Google Scholar] [CrossRef] [Green Version]
- Perháčová, Z.; Pristaš, P.; Trnková, K.; Prepilková, V.; Schwarz, M.; Jankech, A.; Kvasnová, S.; Kisková, J. Dynamics of the Galionella spp. and Leptothrix spp. Communities of Polluted Mine Drainage Water in Slovinky and Markušovce (Slovakia). Geomicrobiol. J. 2022, 39, 597–605. [Google Scholar] [CrossRef]
- Liu, J.; Li, C.; Jing, J.; Zhao, P.; Luo, Z.; Cao, M.; Ma, Z.; Jia, T.; Chai, B. Ecological patterns and adaptability of bacterial communities in alkaline copper mine drainage. Water Res. 2018, 133, 99–109. [Google Scholar] [CrossRef] [PubMed]
- Zucoloto da Costa, B.; Drumond Rodrigues, V.; Maia de Oliveira, V.; Mariscal Ottoboni, L.M.; Marsaioli, A.J. Enzymatic potential of heterothophic bacteria from a neutral copper mine drainage. Braz. J. Microbiol. 2016, 47, 846–852. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chaudhary, P.; Beniwal, V.; Umar, A.; Kumar, R.; Sharma, P.; Kumar, A.; Al-Hadeethi, Y.; Chhokar, V. In vitro microcosm of Co-cultured bacteria for the removal of hexavalent Cr and tannic acid: A mechanistic approach to study the impact of operational parameters. Ecotoxicol. Environ. Saf. 2021, 208, 111484. [Google Scholar] [CrossRef] [PubMed]
- Colak, F.; Atar, N.; Yazicioglu, D.; Olgun, A. Biosorption of lead from aqueous solutions by Bacillus strains possessing heavy-metal resistance. Chem. Eng. J. 2011, 173, 422–428. [Google Scholar] [CrossRef]
- Al Disi, Z.; Al-Ghouti, M.A.; Zouari, N. Investigating the simultaneous removal of hydrocarbons and heavy metals by highly adapted Bacillus and Pseudomonas strains. Environ. Technol. Innov. 2022, 27, 102513. [Google Scholar] [CrossRef]
- Sharma, R.; Jasrotia, T.; Umar, A.; Sharma, M.; Sharma, S.; Kumar, R.; Alkhanjaf, A.A.M.; Vats, R.; Beniwal, V.; Kumar, R.; et al. Effective removal of Pb(II) and Ni(II) ions by Bacillus cereus and Bacillus pumilus: An experimental and mechanistic approach. Environ. Res. 2022, 212, 113337. [Google Scholar] [CrossRef]
- Ayangbenro, A.S.; Babalola, O.O. Genomic analysis of Bacillus cereus NWUAB01 and its heavy metal removal from polluted soil. Sci. Rep. 2020, 10, 19660. [Google Scholar] [CrossRef]
- Raja, C.E.; Omine, K. Characterization of boron resistant and accumulating bacteria Lysinibacillus fusiformis M1, Bacillus cereus M2, Bacillus cereus M3, Bacillus pumilus M4 isolated from former mining site, Hokkaido, Japan. J. Environ. Sci. Health 2012, 47, 1341–1349. [Google Scholar] [CrossRef]
- Njoku, K.L.; Akinyede, O.R.; Obidi, O.F. Microbial Remediation of Heavy Metals Contaminated Media by Bacillus megaterium and Rhizopus stolonifer. Sci. Afr. 2020, 10, e00545. [Google Scholar] [CrossRef]
- Chellaiah, E.R. Cadmium (heavy metals) bioremediation by Pseudomonas aeruginosa: A minireview. Appl. Water Sci. 2018, 8, 154. [Google Scholar] [CrossRef] [Green Version]
- Escamilla-Rodríguez, A.; Carlos-Hernández, S.; Díaz-Jiménez, L. Evidence of Resistance of Heavy Metals from Bacteria Isolated from Natural Waters of a Mining Area in Mexico. Water 2021, 13, 2766. [Google Scholar] [CrossRef]
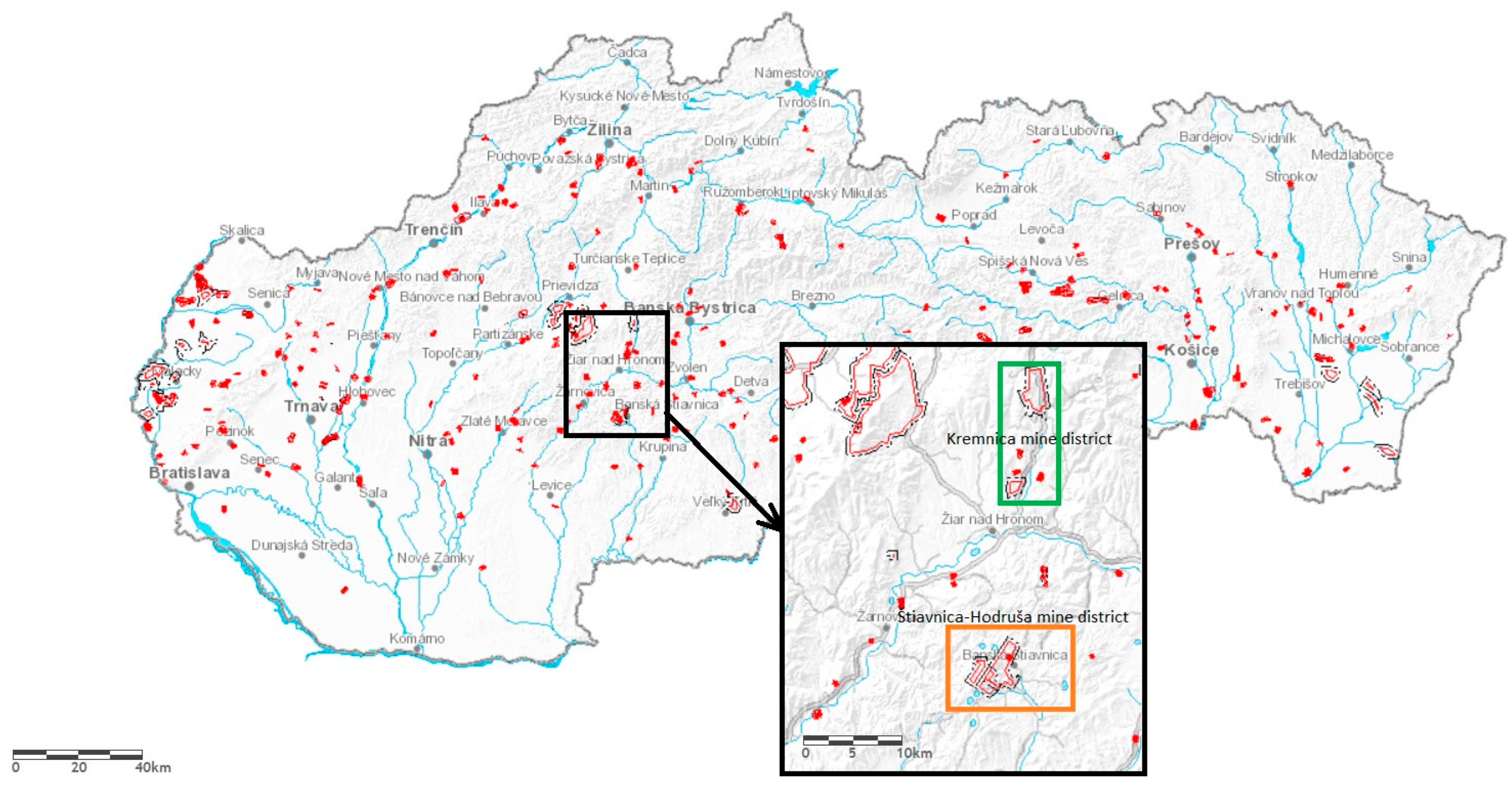

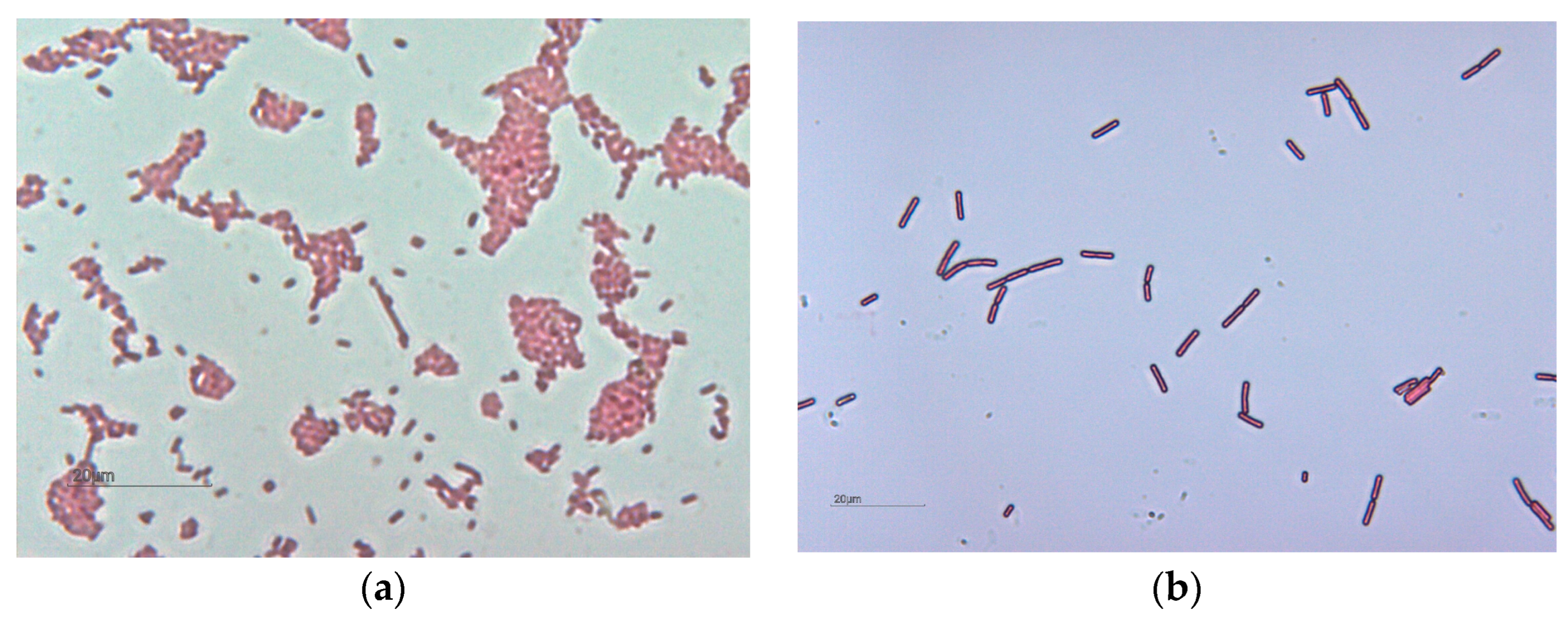
| Kremnica Ore District | pH | Hodruša-Štiavnica Ore District | pH |
|---|---|---|---|
| Main inheritance adit 48°35′4.2″ N 18°51′46.44″ E | 7.35 | Voznická inheritance adit 48° 27′ 51.87″ N 18°42′ 28.38″ E | 7.18 |
| Horná Ves adit 48°40′40.8″ N 18°54’23.04″ E | 5.63 | Tailings pond Hodruša-Hámre 48°27′52.01″ N 18°45′17.84″ E | 7.74 |
| Deep inheritance adit 48°41′9.24″ N 18°55′5.16″ E | 6.92 | Zlatý stôl adit 48°28′3.1″ N 18°50′24.38″ E | 7.25 |
| Isolate | Direct Transfer (ICP) | Score | Extraction (PEP) | Score |
|---|---|---|---|---|
| 1.1. | not reliable identification | 1.596 | Providencia rettgeri | 1.860 |
| 1.3 | not reliable identification | 1.412 | not reliable identification | 1.697 |
| 1.4 | not reliable identification | 1.478 | Morganella morganii | 2.175 |
| 1.5 | Pseudomonas monteilii | 1.710 | Pseudomonas monteilii | 1.980 |
| 2.2 | not reliable identification | 1.550 | not reliable identification | 1.505 |
| 2.3 | Aeromonas hydrophila | 1.719 | Aeromonas hydrophila | 1.809 |
| 2.4 | Lysinibacillus fusiformis | 1.998 | Lysinibacillus fusiformis | 2.395 |
| 2.5 | Bacillus cereus | 1.847 | Bacillus cereus | 1.951 |
| 3.2 | not reliable identification | 1.558 | Bacillus cereus | 2.076 |
| 3.3 | Bacillus cereus | 1.712 | Bacillus cereus | 1.985 |
| 4.1 | not reliable identification | 1.271 | Bacillus pumilus | 1.843 |
| 4.2 | not reliable identification | 1.460 | Serratia liquefaciens | 1.719 |
| 4.3 | not reliable identification | 1.113 | Serratia liquefaciens | 1.903 |
| 4.4 | not reliable identification | 1.292 | not reliable identification | 1.435 |
| 4.5 | not reliable identification | 1.544 | Bacillus altitudinis | 1.956 |
| 4.6 | not reliable identification | 1.170 | not reliable identification | 1.364 |
| 5.1 | not reliable identification | 1.222 | Bacillus cereus | 2.151 |
| 5.2 | not reliable identification | 1.485 | Bacillus cereus | 2.700 |
| 5.5 | Pseudomonas fragi | 1.805 | not reliable identification | 1.391 |
| 5.6 | not reliable identification | 1.568 | Pseudomonas fluorescens | 1.774 |
| 6.2 | not reliable identification | 1.387 | not reliable identification | 1.496 |
| 6.3 | not reliable identification | 1.263 | not reliable identification | 1.414 |
| 6.4 | not reliable identification | 1.178 | Bacillus pumilus | 1.946 |
| Isolate | MALDI TOF | Gram Staining | BLAST |
|---|---|---|---|
| 1.1 | Providencia rettgeri | G− | Providencia rettgeri |
| 1.3 | not reliable identification 1.697 | G− | Serratia plymuthica, S. quinivorans, S. grimesii |
| 1.4 | Morganella morganii | G− | Microbacterim oxydans, M. maritypicum, M. algeriense |
| 1.5 | Pseudomonas monteilii | G− | Bacillus pumilus, B. zhangzhouensis |
| 2.3 | Aeromonas hydrophila | G− | Proteus hauseri, P. vulgaris |
| 2.5 | Bacillus cereus | G+ | Bacillus cereus, Bacillus thuringiensis |
| 3.2 | Bacillus cereus | G+ | Bacillus cereus |
| 3.3 | Bacillus cereus | G+ | Bacillus cereus, Bacillus thuringiensis |
| 4.1 | Bacillus pumilus | G+ | Bacillus safensis, Bacillus pumilus |
| 4.2 | Serratia liquefaciens | G− | Serratia liquefaciens |
| 4.3 | Serratia liquefaciens | G− | Serratia liquefaciens |
| 4.4 | Pseudomonas cedrina, P. fluorescens 1.435 | G− | Pseudomonas fluorescens, P. gessardii |
| 4.5 | Bacillus altitudinis | G+ | Bacillus pumilus, B. zhangzhouensis |
| 4.6 | 1.364 Staphylococcus arlettae | G+ | Pseudomonas fluorescens, P. gessardii |
| 5.1 | Bacillus cereus | G+ | Bacillus cereus, B. thuringiensis |
| 5.2 | Bacillus cereus | G+ | Bacillus velezensis, B. cereus |
| 5.5 | Pseudomonas fragi | G− | Pseudomonas fragi |
| 5.6 | Pseudomonas fluorescens | G− | Pseudomonas veronii, P. fluorescens |
| 6.2 | not reliable identification 1.496 | G+ | Proteus hauseri, P. vulgaris |
| 6.3 | not reliable identification 1.414 | G+ | Bacillus safensis, B. pumilus |
| 6.4 | Bacillus pumilus | G+ | Bacillus pumilus |
| Site | pH | Bacterial Communities | References | ||
|---|---|---|---|---|---|
| Slovakia neovolcanites | 7–8 | Proteobacteria | Bacteroidetes | Actinobacteria | our results |
| Žiar nad Hronom, Slovakia | 13.2 | Proteobacteria | Firmicutes | Bacteroidetes | [22] |
| Slovinky, Slovakia | 6.8 | Proteobacteria | Chloroflexi | Actinobacteria | [8] |
| Pezinok, Slovakia | 6.5–8.0 | Proteobacteria | Actinobacteria | Chloroflexi | [23] |
| Neves-Corvo, Portugal | 4.6–8.7 | Thiobacillus | Halothiobacillus | Thiomicrospira | [24] |
| Lancaster, South Africa | 2.3–2.9 | Acidobacteria | Actinobacteria | Chloroflexi | [25] |
| Onyeama, Nigeria | 3.1 | Proteobacteria | Bacteroidetes | Acidobacteria | [26] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Prepilková, V.; Perháčová, Z.; Schwarz, M.; Trnková, K. Isolation and Identification of Bacterial Communities in Neutral Mine Drainage in Central Slovakian Neovolcanites (Slovakia). Water 2023, 15, 951. https://doi.org/10.3390/w15050951
Prepilková V, Perháčová Z, Schwarz M, Trnková K. Isolation and Identification of Bacterial Communities in Neutral Mine Drainage in Central Slovakian Neovolcanites (Slovakia). Water. 2023; 15(5):951. https://doi.org/10.3390/w15050951
Chicago/Turabian StylePrepilková, Veronika, Zuzana Perháčová, Marián Schwarz, and Katarína Trnková. 2023. "Isolation and Identification of Bacterial Communities in Neutral Mine Drainage in Central Slovakian Neovolcanites (Slovakia)" Water 15, no. 5: 951. https://doi.org/10.3390/w15050951
APA StylePrepilková, V., Perháčová, Z., Schwarz, M., & Trnková, K. (2023). Isolation and Identification of Bacterial Communities in Neutral Mine Drainage in Central Slovakian Neovolcanites (Slovakia). Water, 15(5), 951. https://doi.org/10.3390/w15050951






