How to Statistically Disentangle the Effects of Environmental Factors and Human Disturbances: A Review
Abstract
:1. Introduction
2. Modeling Based on Stratified Randomized Survey
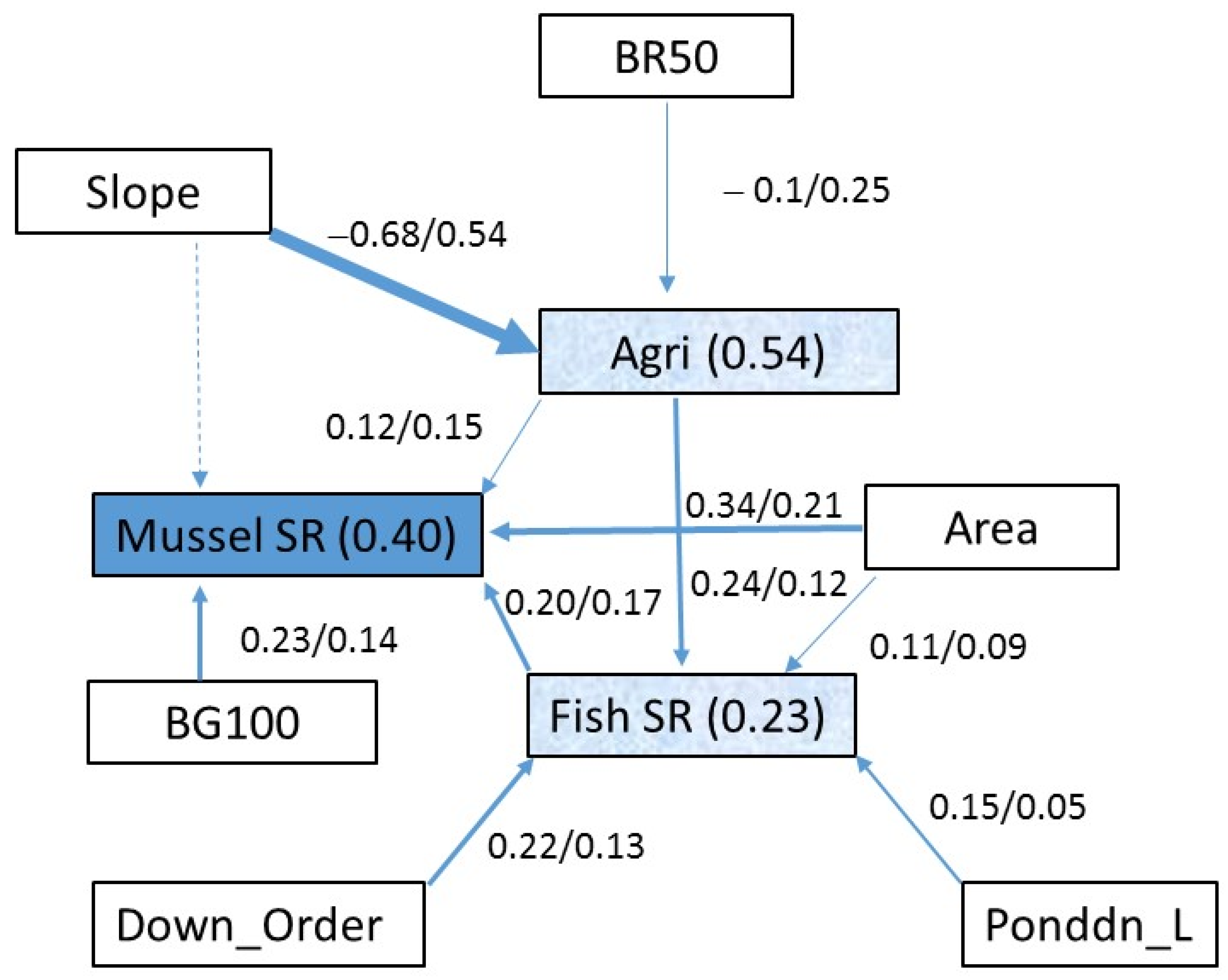
3. Structured Equation Model (SEM)
4. Propensity Scores (PS)
5. Hierarchical Partitioning (HP)
6. Commonality Analysis (CA)
7. Sums of AIC Weight (SW)
8. Tree-Based Approaches: Random Forest (RF) and Boosted Regression Tree (BRT)
9. Assessing the Observation against the Expectation (O/E)
10. Ordination-Based Variance Partitioning for Multivariate Responses
11. Summary and Remarks
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hurlbert, S.H. Pseudoreplication and the design of ecological field experiments. Ecol. Monogr. 1984, 54, 187–211. [Google Scholar] [CrossRef]
- Quinn, G.P.; Keough, M.J. Experimental Design and Data Analysis for Biologist; Cambridge University Press: New York, NY, USA, 2002. [Google Scholar]
- Tredennick, A.T.; Hooker, G.; Ellner, S.P.; Adler, P.B. A practical guide to selecting models for exploration, inference, and prediction in ecology. Ecology 2021, 102, e03336. [Google Scholar] [CrossRef]
- Cao, Y.; Hawkins, C.P.; Olson, J.R.; Kosterman, M.A. Modeling natural environmental gradients improves the accuracy and precision of diatom-based indicators for Idaho streams. J. N. Am. Benthol. Soc. 2007, 26, 566–585. [Google Scholar] [CrossRef]
- Cao, Y.; Hinz, L.; Taylor, C.; Metzke, B.; Cummings, K. Species richness of mussel assemblages and trait guilds in relation to environment and fish diversity in streams of Illinois, the U.S.A. Hydrobiologia 2022, 849, 2193–2208. [Google Scholar] [CrossRef]
- Piggott, J.J.; Lange, K.; Townsend, C.R.; Matthaei, C.D. Multiple stressors in agricultural streams: A mesocosm study of interactions among raised water temperature, sediment addition and nutrient enrichment. PLoS ONE 2012, 7, e49873. [Google Scholar] [CrossRef]
- Dormann, C.F.; Elith, J.; Bacher, S.; Buchmann, C.; Carl, G.; Carré, G.; Marquéz, J.R.G.; Gruber, B.; Lafourcade, B.; Leitão, P.J.; et al. Collinearity: A review of methods to deal with it and a simulation study evaluating their performance. Ecography 2013, 36, 27–46. [Google Scholar] [CrossRef]
- Ormerod, S.J.; Dobson, M.; Hildrew, A.G.; Towsend, C.R. Multiple stressors in freshwater ecosystems. Freshwater Biology 2010, 55 (Suppl. 1), 1–4. [Google Scholar] [CrossRef]
- USEPA. A Practitioner’s Guide to the Biological Condition Gradient: A Framework to Describe Incremental Change in Aquatic Ecosystems; EPA-842-R-16-001; U.S. Environmental Protection Agency: Washington, DC, USA, 2016.
- Clements, W.H.; Kashian, D.R.; Kiffney, P.M.; Zuellig, R.E. Perspectives on the context-dependency of stream community responses to contaminants. Freshw. Biol. 2016, 61, 2162–2170. [Google Scholar] [CrossRef]
- Statzner, B.; Bêche, L.A. Can biological invertebrate traits resolve effects of multiple stressors on running water ecosystems? Freshwatewer Biol. 2010, 55, 80–119. [Google Scholar] [CrossRef]
- Pyne, M.I.; Russell, B.; Christensen, W.F. Predicting local biological characteristics in streams: A comparison of landscape classifications. Freshwater Biol. 2007, 52, 1302–1321. [Google Scholar] [CrossRef]
- McManmay, R.A.; Christopher, R.D. Data descriptor: A stream classification system for the conterminous United States. Sci. Data 2019, 6, 190017. [Google Scholar] [CrossRef] [PubMed]
- Wright, J.F.; Sutcliffe, D.W.; Furse, M.T. Assessing the Biological Quality of Fresh Waters: RIVPACS and Other Techniques; Freshwater Biological Association: Ambleside, UK, 2000. [Google Scholar]
- Friberg, N.; Bonada, N.; Bradley, D.C.; Dunbar, M.J.; Edwards, F.K.; Grey, J.; Hayes, R.B.; Hildrew, A.G.; Lamouroux, N.; Trimmer, M.; et al. Biomonitoring of Human Impacts in Freshwater Ecosystems: The Good, the Bad and the Ugly. In Advances in Ecological Research; Academic press: Cambridge, MA, USA, 2011; Volume 44, pp. 1–68. [Google Scholar]
- Kuhn, M.; Johnson, K. Appled Preditive Modeling; Springer: New York, NY, USA, 2016. [Google Scholar]
- Schartel, T.; Cao, Y.; Hennings, B.; Feng, E.; Hinz, L. Modeling and predicting freshwater mussel distributions in the Midwestern United States. Aquatic Conservation: Freshw. Mar. Ecosyst. 2021, 31, 3370–3385. [Google Scholar] [CrossRef]
- SAS, Inc. Visual Data Mining and Machine Learning. 2022. Available online: https://documentation.sas.com/doc/en/vdmmlcdc/v_014/vdmmlref/n12jcjwia3hb21n1104tdpkl9d1v.htm (accessed on 8 February 2023).
- Mac Nally, R. Regression and model-building in conservation biology, biogeography and ecology: The distinction between-and reconciliation of -’predictive’ and ‘explanatory’ models. Biodivers. Conserv. 2000, 9, 655–671. [Google Scholar] [CrossRef]
- Smith, A.C.; Koper, N.; Francis, C.M.; Fahrig, L. Confronting collinearity: Comparing methods for disentangling the effects of habitat loss and fragmentation. Landsc. Ecol. 2009, 24, 1271–1285. [Google Scholar] [CrossRef]
- Freckleton, R.P. On the misuse of residuals in ecology: Regression of residuals vs. multiple regression. J. Anim. Ecol. 2002, 71, 542–545. [Google Scholar] [CrossRef]
- Redlich, S.; Zhang, J.; Benjamin, C.; Dhillon, M.S.; Englmeier, J.; Ewald, J.; Fricke, U.; Ganuza, C.; Haensel, M.; Hovestadt, T.; et al. Disentangling effects of climate and land use on biodiversity and ecosystem services—A multi-scale experimental design. Methods Ecol. Evol. 2021, 13, 514–527. [Google Scholar] [CrossRef]
- Fricke, U.; Redlich, S.; Zhang, J.; Tobisch, C.; Rojas-Botero, S.; Benjamin, C.S.; Englmeier, J.; Ganuza, C.; Riebl, R.; Uhler, J.; et al. Plant richness, land use and temperature differently shape invertebrate leaf-chewing herbivory on plant functional groups. Oecologia 2022, 199, 407–417. [Google Scholar] [CrossRef]
- Ganuza, C.; Redlich, S.; Uhler, J.; Tobisch, C.; Rojas-Botero, S.; Peters, M.K.; Zhang, J.; Benjamin, C.S.; Englmeier, J.; Ewald, J.; et al. Interactive effects of climate and land use on pollinator diversity differ among taxa and scales. Sci. Adv. 2022, 8, eabm9359. [Google Scholar] [CrossRef] [PubMed]
- Englmeier, J.; von Hoermann, C.; Rieker, D.; Benbow, M.E.; Benjamin, C.; Fricke, U.; Ganuza, C.; Haensel, M.; Lackner, T.; Mitesser, O.; et al. Dung-visiting beetle diversity is mainly affected by land use, while community specialization is driven by climate. Ecol. Evol. 2022, 12, e9386. [Google Scholar] [CrossRef]
- Hynes, H.B.N. The Ecology of Running Waters; University of Toronto Press: Toronto, ON, Canada, 1970. [Google Scholar]
- Wang, L.; Lyons, J.; Rasmussen, P.; Seelbach, P.; Simon, T.; Wiley, M.; Kanehl, P.; Baker, E.; Niemela, S.; Stewart, P.M. Watershed, reach, and riparian influences on stream fish assemblages in the Northern Lakes and Forest Ecoregion, U.S.A. Can. J. Fish. Aquat. Sci. 2003, 60, 491–505. [Google Scholar] [CrossRef]
- Werner, C.; Schermelleh-Engel, K. Structural Equation Modeling: Advantages, Challenges, and Problems. In Introduction to Structural Equation Modeling with LISREL; Goethe University: Frankfurt, Germany, 2009. [Google Scholar]
- Vaughn, C.C.; Taylor, C.M. Macroecology of a host-parasite relationship: Distribution patterns of mussels and fishes. Ecography 2000, 23, 11–20. [Google Scholar] [CrossRef]
- Leitão, R.P.; Zuanon, J.; Mouillot, D.; Leal, C.G.; Hughes, R.M.; Kaufmann, P.R.; Villéger, S.; Pompeu, P.S.; Kasper, D.; de Paula, F.R.; et al. Disentangling the pathways of land use impacts on the functional structure of fish assemblages in Amazon streams. Ecography 2018, 41, 219–232. [Google Scholar] [CrossRef]
- Lefcheck, J.S. PIECEWISESEM: Piecewise structural equationmodelling in R for ecology, evolution, and systematics. Methods Ecol. Evol. 2016, 7, 573–579. [Google Scholar] [CrossRef]
- Schmidt, T.S.; Van Metre, P.C.; Carlisle, D.M. Linking the agricultural landscape of the Midwest to stream health with Structural Equation Modeling. Environ. Sci. Technol. 2019, 53, 452–462. [Google Scholar] [CrossRef]
- Alvarenga, L.R.P.; Pompeu, P.S.; Leal, C.G.; Hughes, R.M.; Fagundes, D.C.; Leitão, R.P. Land-use changes affect the functional structure of stream fish assemblages in the Brazilian Savanna. Neotrop. Ichthyol. 2021, 19, e210035. [Google Scholar] [CrossRef]
- Mao, Z.; Cao, Y.; Gu, X.; Zeng, Q.; Chen, H.; Jeppesen, E. Response of zooplankton to nutrient reduction and enhanced fish predation in a shallow eutrophic lake. Ecol. Appl. 2023, 33, e2750. [Google Scholar] [CrossRef] [PubMed]
- Rosenbaum, P.R.; Rubin, D.B. The central role of the propensity score in observational studies for causal effects. Biometrika 1983, 70, 41–55. [Google Scholar] [CrossRef]
- Ramsey, D.S.L.; Forsyth, D.M.; Wright, E.; McKay, M.; Westbrooke, I. Using propensity scores for causal inference in ecology: Options, considerations, and a case study. Methods Ecol. Evol. 2019, 10, 320–331. [Google Scholar] [CrossRef]
- Keller, B.; Tipton, E. Propensity score analysis in R: A software review. J. Educ. Behav. Stat. 2016, 41, 326–348. [Google Scholar] [CrossRef]
- Yuan, L.L. Estimating the effects of excess nutrients on stream invertebrates from observational data. Ecol. Appl. 2010, 20, 110–125. [Google Scholar] [CrossRef] [PubMed]
- Pearson, C.E.; Ormerod, S.J.; Symondson, W.O.; Vaughan, I.P. Resolving large-scale pressures on species and ecosystems: Propensity modelling identifies agricultural effects on streams. J. Appl. Ecol. 2016, 53, 408–417. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Breiman, L.; Ffriedman, J.H.; Olshen, R.A.; Stone, C.G. Classification and Regression Trees; Wadsworth Statistics/Probability Series; Chapman and Hall: New York, NY, USA, 1984. [Google Scholar]
- Shimose, S.; Tanaka, M.; Iwamoto, H.; Niizeki, T.; Shirono, T.; Aino, H.; Noda, Y.; Kamachi, N.; Okamura, S.; Nakano, M.; et al. Prognostic impact of transcatheter arterial chemoembolization (TACE) combined with radiofrequency ablation in patients with unresectable hepatocellular carcinoma: Comparison with TACE alone using decision-tree analysis after propensity score matching. Hepatol. Res. 2019, 49, 919–928. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Levine, R.A.; Fan, J.J. Causal effect random forest of interaction trees for learning individualized treatment regimes with multiple treatments in observational studies. Stat 2022, 11, e457. [Google Scholar] [CrossRef]
- Chevan, A.; Sutherland, M. Hierarchical partitioning. Am. Stat. 1991, 45, 90–96. [Google Scholar]
- South, E.J.; DeWalt, R.E.; Cao, Y. Relative importance of Conservation Reserve programs to aquatic insects in an agricultural landscape. Hydrobiologia 2018, 829, 327–340. [Google Scholar]
- Walsh, C.J.; Papas, P.J.; Crowther, D.; Sim, P.T.; Yoo, J. Stormwater drainage pipes as a threat to a streamdwelling amphipod of conservation significance, Austrogammarus australis, in southeastern Australia. Biodivers. Conserv. 2004, 13, 781–793. [Google Scholar] [CrossRef]
- Lai, J.-S.; Zou, Y.; Zhang, J.-L.; Peres-Neto, P.R. Generalizing hierarchical and variation partitioning in multiple regression and canonical analyses using the rdacca.hp R package. Methods Ecol. Evol. 2022, 13, 782–788. [Google Scholar] [CrossRef]
- Lai, J.-S.; Zou, Y.; Zhang, S.; Zhang, X.-G.; Mao, L.-F. glmm.hp: An R package for computing individual effect of predictors in generalized linear mixed models. J. Plant Ecol. 2022, 15, 1302–1307. [Google Scholar] [CrossRef]
- Olea, P.P.; Mateo-Tomas, P.; de Frutos, A. Estimating and modelling bias of the hierarchical partitioning public-domain software: Implications in environmental management and conservation. PLoS ONE 2010, 5, e11698. [Google Scholar] [CrossRef]
- Warton, D.I. Eco-Stats: Data Analysis in Ecology from t-Tests to Multivariate Abundances; Springer Nature: Cham, Switzerland, 2022. [Google Scholar]
- Newton, R.G.; Spurell, D.J. Examples of the use of elements for classifying regression analysis. Appl. Stat. 1967, 16, 165–172. [Google Scholar] [CrossRef]
- Nimon, K.; Reio, T. Regression commonality analysis: A technique for quantitative theory building. Hum. Resour. Dev. 2011, 10, 329–340. [Google Scholar] [CrossRef]
- Nimon, K.; Oswald, F.L.; Roberts, J.K. Yhat: Interpreting Regression Effects. R Package Version 2.0–3. 2022. Available online: https://cran.r-project.org/web/packages/yhat/yhat.pdf (accessed on 8 February 2023).
- Ray-Mukherjee, J.; Nimon, K.; Mukherjee, S.; Morris, D.W.; Slotow, R.; Hamer, M. Using commonality analysis in multiple regressions: A tool to decompose regression effects in the face of multicollinearity. Methods Ecol. Evol. 2014, 5, 320–328. [Google Scholar] [CrossRef]
- Prunier, J.G.; Dubut, V.; Loot, G.; Tudesque, L.; Blanchet, S. The relative contribution of river network structure and anthropogenic stressors to spatial patterns of genetic diversity in two freshwater fishes: A multiple-stressors approach. Freshw. Biol. 2018, 63, 6–21. [Google Scholar] [CrossRef]
- Alahuhta, J.; Lindholm, M.; Bove, C.P.; Chappuis, E.; Clayton, J.; de Winton, M.; Feldmann, T.; Ecke, F.; Gacia, E.; Grillas, P.; et al. Global patterns in the metacommunity structuring of lake macrophytes: Regional variations and driving factors. Oecologia 2018, 188, 1167–1182. [Google Scholar] [CrossRef]
- Schneider, W.J. Playing statistical ouija board with commonality analysis: Good questions, wrong assumptions. Appl. Neuropschol. 2008, 15, 44–53. [Google Scholar] [CrossRef]
- Anderson, D.R.; Burnham, K.P. Avoiding pitfalls when using information-theoretic methods. J. Wildl. Manag. 2002, 66, 912–918. [Google Scholar] [CrossRef]
- Galipaud, M.; Gillingham, M.A.F.; Dechaume-Moncharmont, F.-X. A farewell to the sum of Akaike weights: The benefits of alternative metrics for variable importance estimations in model selection. Methods Ecol. Evol. 2017, 8, 1668–1678. [Google Scholar] [CrossRef]
- Burnham, K.P.; Anderson, D.R.; Huyvaert, K.P. AIC model selection and multimodel inference in behavioral ecology: Some background, observations, and comparisons. Behav. Ecol. Sociobiol. 2011, 65, 23–35. [Google Scholar] [CrossRef]
- Murray, K.; Conner, M.M. Methods to quantify variable importance: Implications for the analysis of noisy ecological data. Ecology 2009, 90, 348–355. [Google Scholar] [CrossRef]
- Galipaud, M.; Gillingham, M.A.F.; David, M.; Dechaume-Moncharmont, F.-X. Ecologists overestimate the importance of predictor variables in model averaging: A plea for cautious interpretations. Methods Ecol. Evol. 2014, 5, 983–991. [Google Scholar] [CrossRef]
- Giam, X.-L.; Olden, J.D. Quantifying variable importance in a multimodel inference framework. Methods Ecol. Evol. 2016, 7, 388–397. [Google Scholar] [CrossRef]
- Li, W.Q.; Kou, X.J. WiBB: An integrated method for quantifying the relative importance of predictive variables. Ecography 2022, 44, 1557–1567. [Google Scholar] [CrossRef]
- Wright, M.N.; Ziegler, A.; Köning, I.R. Do little interactions get lost in dark random forests? B.M.C. Bioinform. 2016, 17, 145. [Google Scholar] [CrossRef]
- Strobl, C.; Boulesteix, A.-L.; Zeileis, A.; Hothorn, T. Bias in random forest variable importance measures: Illustrations, sources and a solution. B.M.C. Bioinform. 2017, 8, 25. [Google Scholar] [CrossRef]
- Probst, P.; Wright, M.N.; Boulesteix, A.-L. Hyperparameters and tuning strategies for random forest. Wires Data Min. Knowl. Discov. 2019, 9, e1301. [Google Scholar] [CrossRef]
- Elith, J.; Leathwick, J.R.; Hastie, T. A working guide to boosted regression trees. J. Anim. Ecol. 2008, 77, 803–813. [Google Scholar] [CrossRef]
- Paumier, A.; Drouineau, H.; Boutry, S.; Sillero, N.; Lambert, P. Assessing the relative importance of temperature, discharge, and day length on the reproduction of an anadromous fish (Alosa alosa). Freshw. Biol. 2020, 65, 253–263. [Google Scholar] [CrossRef]
- Waite, I. Agricultural disturbance response models for invertebrate and algal metrics from streams at two spatial scales within the U.S. Hydrobiologia 2014, 726, 285–303. [Google Scholar] [CrossRef]
- Stoddard, J.L.; Larsen, D.P.; Hawkins, C.P.; Johnson, R.K.; Norris, R.H. Setting expectations for the ecological condition of running waters: The concept of reference conditions. Ecol. Appl. 2006, 16, 1267–1276. [Google Scholar] [CrossRef]
- Clarke, R.T.; Wright, J.F.; Furse, M.T. RIVPACS models for predicting the expected macroinvertebrate fauna and assessing the ecological quality of rivers. Ecol. Model. 2003, 160, 219–233. [Google Scholar] [CrossRef]
- Hawkins, C.P. Maintaining and restoring the ecological integrity of freshwater ecosystems: Refining biological assessments. Ecol. Appl. 2006, 16, 1249–1250. [Google Scholar] [CrossRef]
- Van Sickle, J. An index of compositional dissimilarity between observed and expected assemblages. J. N. Am. Benthol. Soc. 2008, 27, 227–235. [Google Scholar] [CrossRef]
- Cao, Y.; Hinz, L.; Cummings, K.; Douglass, S.; Price, A.; Holtrop, A. Reconstructing historic distributions of mussel species and diversity patterns in Illinois streams. Freshw. Sci. 2017, 36, 669–682. [Google Scholar] [CrossRef]
- Pont, D.; Hugueny, B.; Beier, U.; Goffaux, D.; Melcher, A.; Noble, R.; Rogers, C.; Roset, N.; Schmutz, S. Assessing river biotic condition at a continental scale: A European approach using functional metrics and fish assemblages. J. Appl. Ecol. 2006, 43, 70–80. [Google Scholar] [CrossRef]
- Hawkins, C.P.; Cao, Y.; Roper, R. Method of predicting reference conditions affects the performance and interpretation of ecological indices. Freshw. Biol. 2010, 55, 1066–1085. [Google Scholar] [CrossRef]
- Carlisle, D.M.; Falcone, J.; Wolock, D.M.; Meador, M.R.; Norris, R.H. Predicting the natural flow regime: Models for assessing hydrological alterantion in streams. River Res. Appl. 2010, 26, 118–136. [Google Scholar] [CrossRef]
- Kaufmann, P.R.; Hughes, R.M.; Paulsen, S.G.; Peck, D.V.; Seeliger, C.W.; Kincaid, T.; Mitchell, R.M. Physical habitat in conterminous U.S. streams and Rivers, part 2: A quantitative assessment of habitat condition. Ecol. Indic. 2022, 141, 109047. [Google Scholar] [CrossRef]
- Hawkins, C.P.; Olson, J.R.; Hill, R.A. The reference condition: Predicting benchmarks for ecological and water-quality assessments. J. N. Am. Benthol. Soc. 2010, 29, 312–343. [Google Scholar] [CrossRef]
- Elith, J.; Phillips, S.J.; Hastie, T.; Dudík, M.; Chee, Y.E.; Yates, C.J. A statistical explanation of MaxEnt for ecologists. Diversity and Distributions 2011, 17, 43–57. [Google Scholar] [CrossRef]
- Hastie, T.; Fithian, W. Inference from presence-only data: The ongoing controversy. Ecography 2013, 36, 864–867. [Google Scholar] [CrossRef] [PubMed]
- Merow, C.; Smith, M.J.; Silander, J.A., Jr. A practical guide to MaxEnt for modeling species’ distributions: What it does, and why inputs and settings matter. Ecography 2013, 36, 1058–1069. [Google Scholar] [CrossRef]
- Legendre, P.; Legendre, L. Numerical Ecology, 3rd ed.; Elsevier: New York, NY, USA, 2012. [Google Scholar]
- Borcard, D.; Lengendre, P.; Drapeau, P. Partitioning out the spatial component of ecological variation. Ecology 1992, 73, 1045–1055. [Google Scholar] [CrossRef]
- Weigel, B.M.; Wang, L.; Rasmussen, P.W.; Butcher, J.T.; Stewart, P.M.; Simon, T.P.; Wiley, M.J. Relative influence of variables at multiple spatial scales on stream macroinvertebrates in the Northern Lakes and Forest ecoregion, U.S.A. Freshw. Biol. 2003, 48, 1440–1461. [Google Scholar] [CrossRef]
- Meißner, T.; Sures, B.; Feld, C.K. Multiple stressors and the role of hydrology on benthic invertebrates in mountainous streams. Sci. Total Environ. 2019, 663, 841–851. [Google Scholar] [CrossRef] [PubMed]
- Morales-Molino, C.; Steffen, M.; Samartin, S.; van Leeuwen, J.F.N.; Hürlimann, D.; Vescovi, E.; Tinner, W. Long-term responses of mediterranean mountain forests to climate change, fire and human activities in the Northern Apennines (Italy). Ecosystems 2021, 24, 1361–1377. [Google Scholar] [CrossRef]
- Hastie, T.; Tibsherian, R.; Wainright, M. Statistical Leaning with Sparsity: Lasso and Generations; CRC Press: Boca Raton, FL, USA, 2016. [Google Scholar]
| Lat | long | Slope | Agri | Forest | BG100 | BR50 | Temp | Precip | |
|---|---|---|---|---|---|---|---|---|---|
| Long | 0.09 | ||||||||
| Slope | −0.43 | −0.33 | |||||||
| Agri | 0.34 | 0.06 | −0.73 | ||||||
| Forest | −0.61 | −0.13 | 0.87 | −0.83 | |||||
| BG100 | 0.42 | 0.37 | −0.44 | 0.36 | −0.43 | ||||
| BR50 | −0.48 | −0.36 | 0.59 | −0.50 | 0.57 | −0.83 | |||
| Temp | −0.99 | −0.08 | 0.39 | −0.30 | 0.57 | −0.40 | 0.44 | ||
| Precip | −0.90 | 0.08 | 0.57 | −0.52 | 0.74 | −0.40 | 0.50 | 0.87 | |
| Perm | 0.28 | 0.16 | −0.09 | 0.05 | −0.11 | 0.17 | −0.20 | −0.28 | −0.16 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cao, Y.; Wang, L. How to Statistically Disentangle the Effects of Environmental Factors and Human Disturbances: A Review. Water 2023, 15, 734. https://doi.org/10.3390/w15040734
Cao Y, Wang L. How to Statistically Disentangle the Effects of Environmental Factors and Human Disturbances: A Review. Water. 2023; 15(4):734. https://doi.org/10.3390/w15040734
Chicago/Turabian StyleCao, Yong, and Lizhu Wang. 2023. "How to Statistically Disentangle the Effects of Environmental Factors and Human Disturbances: A Review" Water 15, no. 4: 734. https://doi.org/10.3390/w15040734
APA StyleCao, Y., & Wang, L. (2023). How to Statistically Disentangle the Effects of Environmental Factors and Human Disturbances: A Review. Water, 15(4), 734. https://doi.org/10.3390/w15040734






