Insights into Increasing Selenate Reductase Enzyme Activity in the Presence of Nitrogen-Doped Graphite Electrodes for Selenium Effluent Treatment
Abstract
:1. Introduction
2. Materials and methods
2.1. Chemicals and Materials
2.2. Recombinant Expression and Affinity Enrichment of TsSer-α, β, and γ Subunits
2.3. Validation of TsSer-αβγ Heterotrimeric Complex Formation
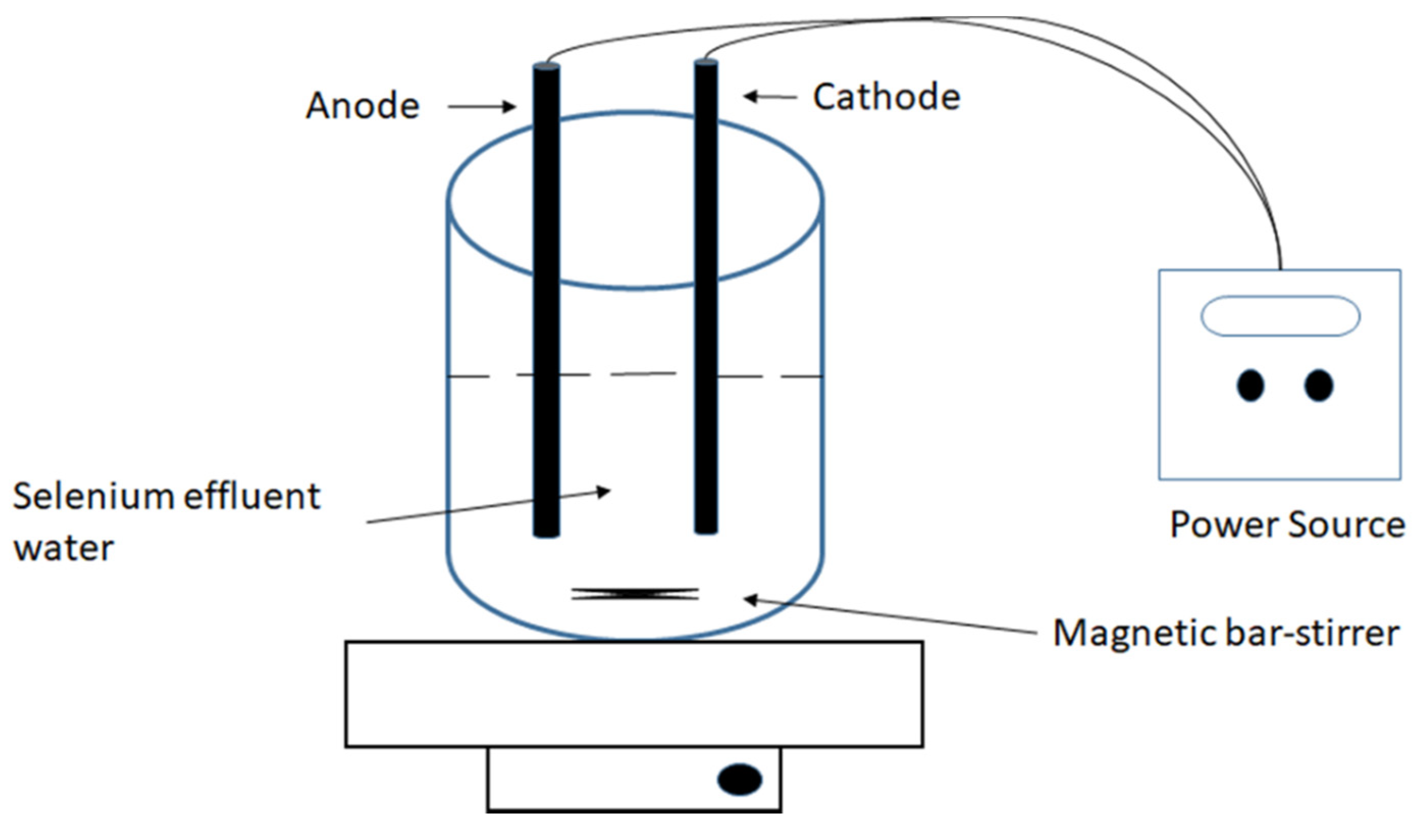
2.4. Selenate Reduction by the TsSer-αβγ Heterotrimeric Complex
3. Results and Discussion
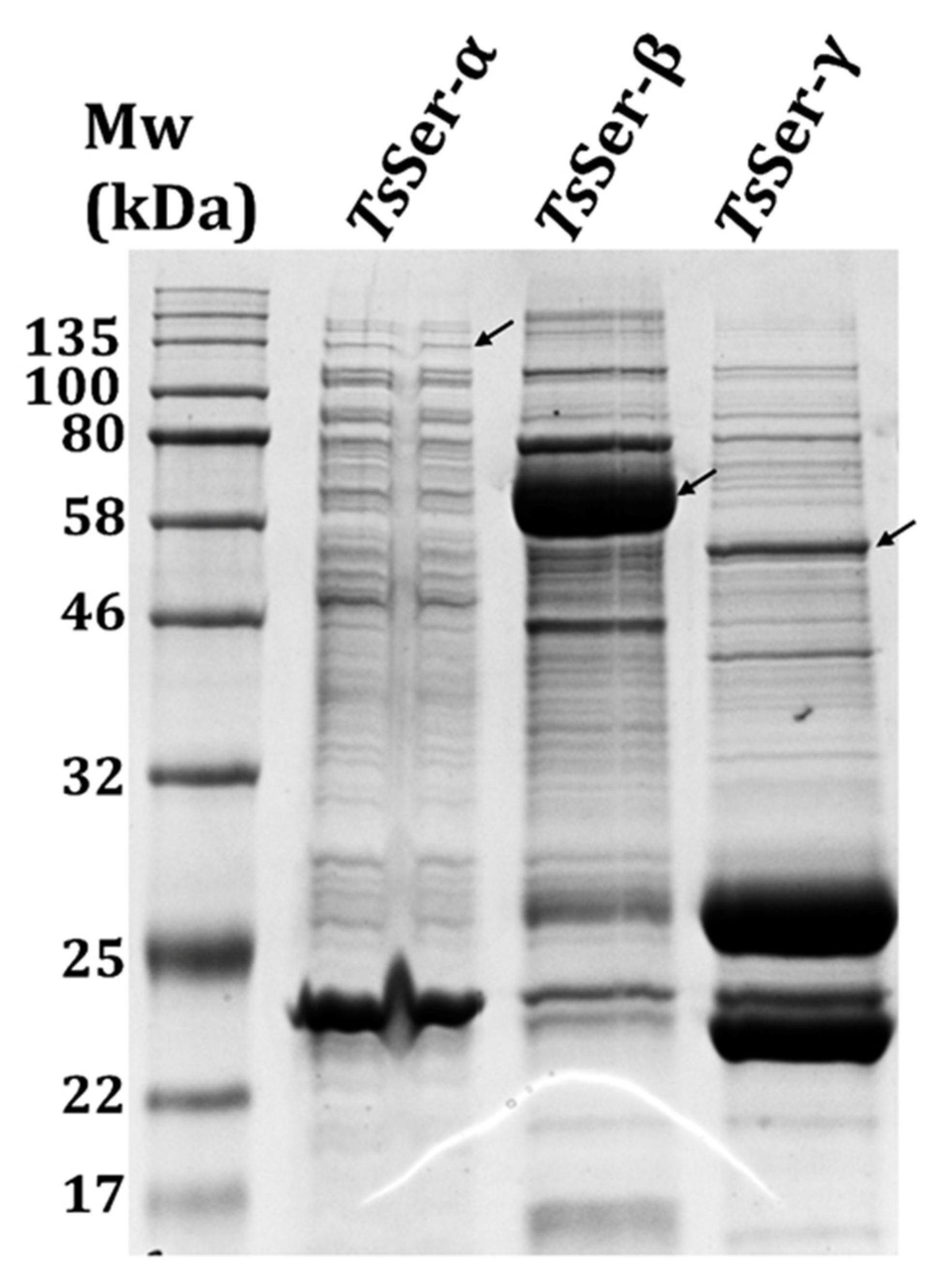
3.1. Expression and Purification of the Individual TsSer-α, -β, and -γ Subunits
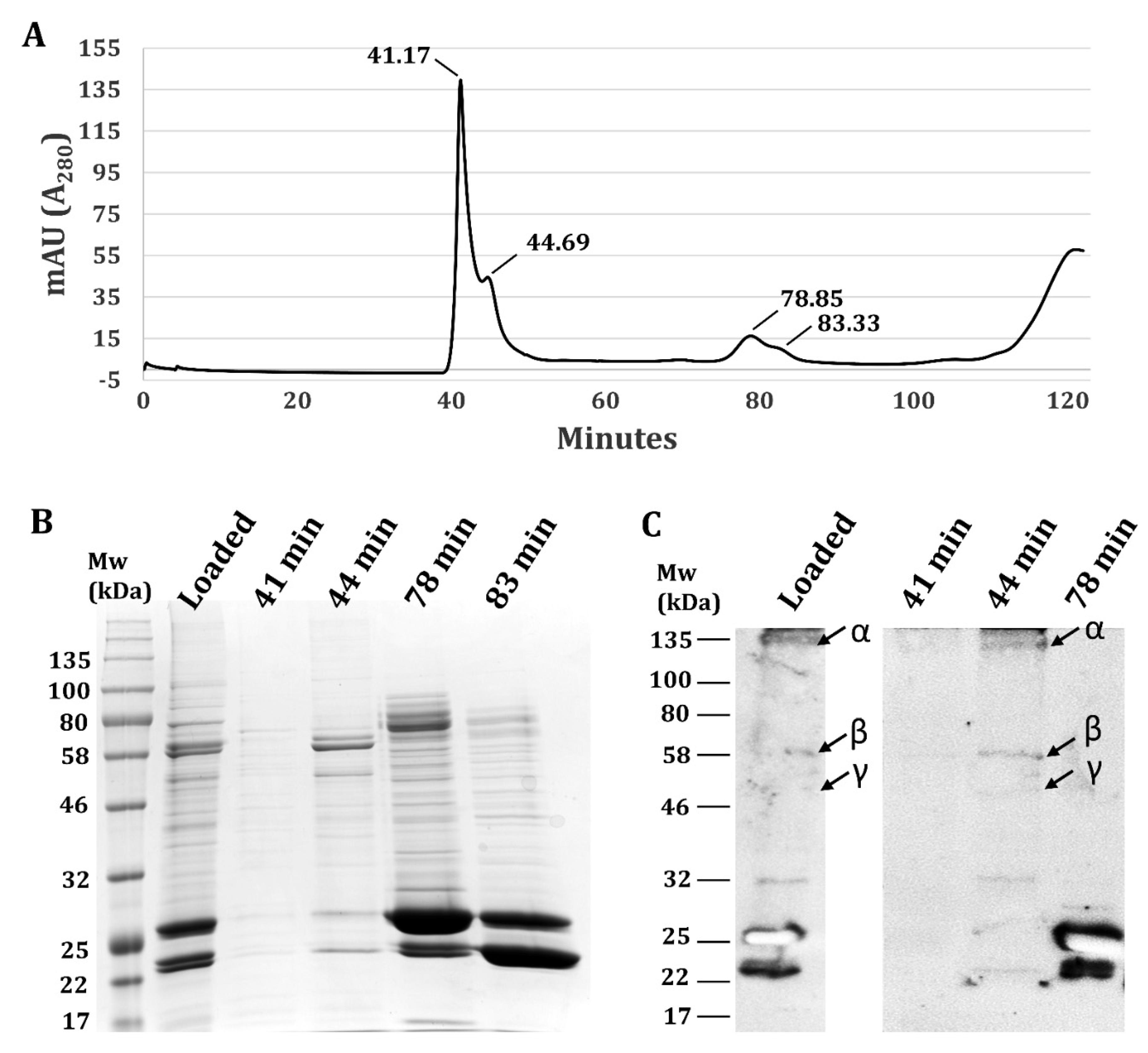
3.2. Validation Heterotrimeric TsSer-αβγ Complex Formation
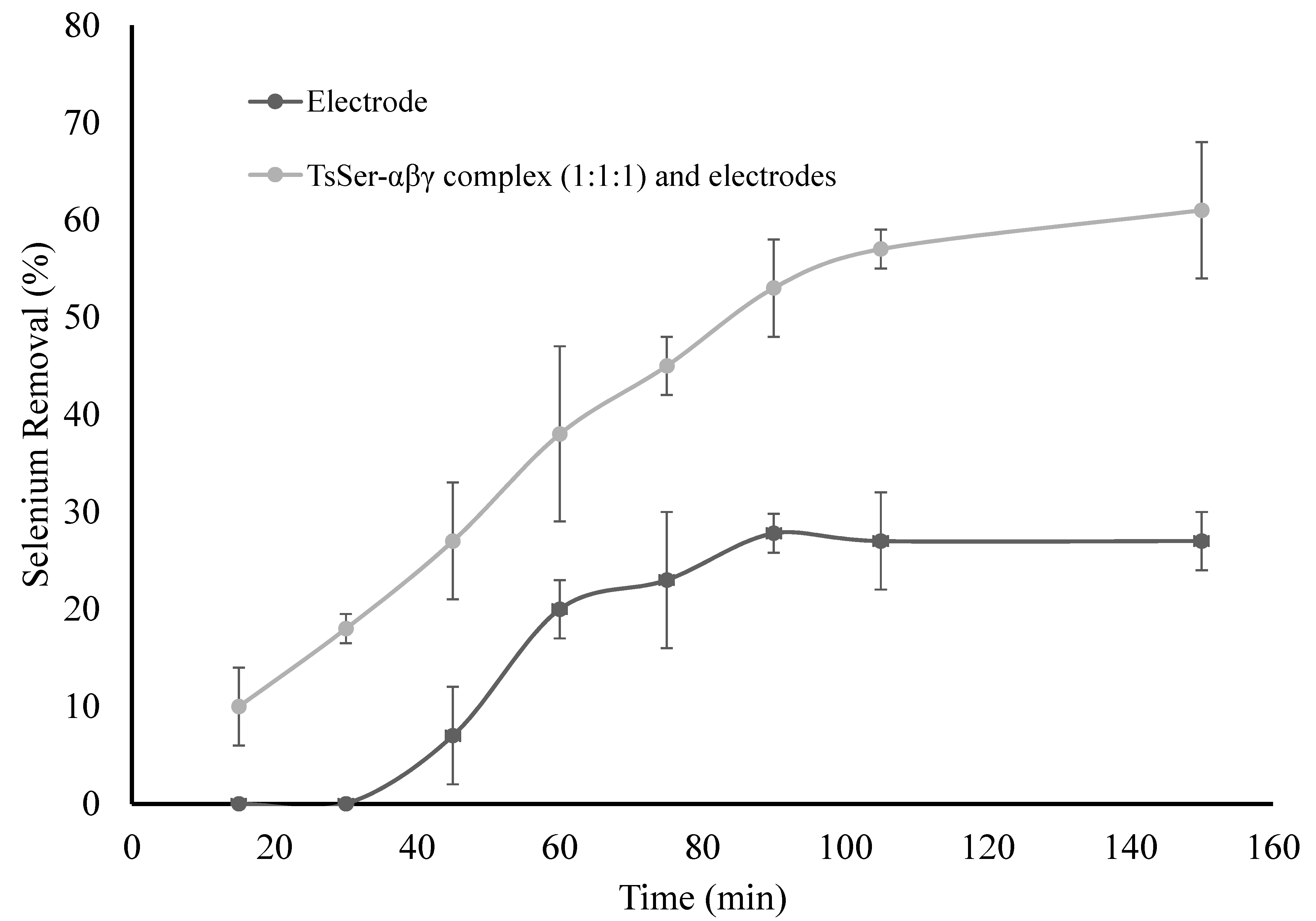
3.3. Enzymatic Reduction of Selenium by Recombinantly Produced TsSer-αβγ Enzyme Complex from Effluent Water
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Martínez, F.G.; Moreno-Martin, G.; Pescuma, M.; Madrid-Albarrán, Y.; Mozzi, F. Biotransformation of selenium by lactic acid bacteria: Formation of seleno-nanoparticles and seleno-amino acids. Front. Bioeng. Biotechnol. 2020, 8, 506. [Google Scholar] [CrossRef] [PubMed]
- Sabuda, M.C.; Rosenfeld, C.E.; DeJournett, T.D.; Schroeder, K.; Wuolo-Journey, K.; Santelli, C.M. Fungal Bioremediation of selenium-contaminated industrial and municipal wastewaters. Front. Microbiol. 2020, 11, 2105. [Google Scholar] [CrossRef] [PubMed]
- Debieux, C.M.; Dridge, E.J.; Mueller, C.M.; Splatt, P.; Paszkiewicz, K.; Knight, I.; Florance, H.; Love, J.; Titball, R.W.; Lewis, R.J.; et al. A bacterial process for selenium nanosphere assembly. Proc. Natl. Acad. Sci. USA 2011, 108, 13480–13485. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, L.; Yang, H.; Shin, H.D.; Chen, R.R.; Li, J.; Du, G.; Chen, J. How to achieve high-level expression of microbial enzymes: Strategies and perspectives. Bioengineered 2013, 4, 212–223. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rehm, F.B.H.; Chen, S.; Rehm, B.H.A. Enzyme engineering for in situ immobilization. Molecules 2016, 21, 1370. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singh, R.K.; Tiwari, M.K.; Singh, R.; Lee, J.K. From protein engineering to immobilization: Promising strategies for the upgrade of industrial enzymes. Int. J. Mol. Sci. 2013, 14, 1232–1277. [Google Scholar] [CrossRef] [PubMed]
- Connelly, K.R.S.; Stevenson, C.; Kneuper, H.; Sargent, F. Biosynthesis of selenate reductase in Salmonella enterica: Critical roles for the signal peptide and DmsD. Microbiology 2016, 162, 2136–2146. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wells, M.; McGarry, J.; Gaye, M.M.; Basu, P.; Oremland, R.S.; Stolz, J.F. Respiratory selenite reductase from Bacillus selenitireducens strain MLS10. J. Bacteriol. 2019, 201, e00614–e00618. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yanke, L.; Bryant, R.; Laishley, E. Hydrogenase I of Clostridium pasteurianum functions as a novel selenite reductase. Anaerobe 1995, 1, 61–67. [Google Scholar] [CrossRef]
- Maher, M.J.; Macy, J.M. Biological Crystallography Crystallization and preliminary X-ray analysis of the selenate reductase from Thauera selenatis. Acta Crystallogr. Sect. D Biol. Crystallogr. 2002, 58, 706–708. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, M.; Yamashita, M.; Miwa, E.; Imao, K.; Fujimoto, N.; Ono, H.; Nagano, K.; Sei, K.; Ike, M. Molecular cloning and characterization of the srdBCA operon, encoding the respiratory selenate reductase complex, from the selenate-reducing Bacterium Bacillus selenatarsenatis SF-1. J. Bacteriol. 2011, 193, 2141–2148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guymer, D.; Maillard, J.; Sargent, F. A genetic analysis of in vivo selenate reduction by Salmonella enterica serovar Typhimurium LT2 and Escherichia coli K12. Arch. Microbiol. 2009, 191, 519–528. [Google Scholar] [CrossRef] [PubMed]
- Yee, N.; Choi, J.; Porter, A.W.; Carey, S.; Rauschenbach, I.; Harel, A. Selenate reductase activity in Escherichia coli requires Isc iron-sulfur cluster biosynthesis genes, FEMS. Microbiol. Lett. 2014, 361, 138–143. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Recht, S.A.; Macy, A.J.M. The terminal reductases for selenate and nitrate respiration in Thauera selenatis are two distinct enzymes. J. Bacteriol. 1992, 174, 7316–7320. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schröder, I.; Rech, S.; Krafft, T.; Macy, J.M. Purification and characterization of the selenate reductase from Thauera selenatis. J. Biol. Chem. 1997, 272, 23765–23768. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krafft, T.; Bowen, A.; Theis, F. Cloning and sequencing of the genes encoding the periplasmic-cytochrome B-containing selenate reductase of Thauera selenatis. DNA Seq. 2000, 10, 365–377. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, V.K.; Park, Y.; Yu, J.; Lee, T. Microbial selenite reduction with organic carbon and electrode as sole electron donor by a bacterium isolated from domestic wastewater. Bioresour. Technol. 2016, 212, 182–189. [Google Scholar] [CrossRef] [PubMed]
- Gates, A.J.; Marritt, S.J.; Bradley, J.M.; Shi, L.; McMillan, D.G.; Jeuken, L.J.; Richardson, D.J.; Butt, J.N. Electrode assemblies composed of redox cascades from microbial respiratory electron transfer chains. Biochem. Soc. Trans. 2013, 41, 1249–1253. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mohapatra, D.P.; Robinson, K.A.; Huang, F.; Kirpalani, D.; Loewen, M.C. Insights into Increasing Selenate Reductase Enzyme Activity in the Presence of Nitrogen-Doped Graphite Electrodes for Selenium Effluent Treatment. Water 2022, 14, 931. https://doi.org/10.3390/w14060931
Mohapatra DP, Robinson KA, Huang F, Kirpalani D, Loewen MC. Insights into Increasing Selenate Reductase Enzyme Activity in the Presence of Nitrogen-Doped Graphite Electrodes for Selenium Effluent Treatment. Water. 2022; 14(6):931. https://doi.org/10.3390/w14060931
Chicago/Turabian StyleMohapatra, Dipti Prakash, Kelly Ann Robinson, Fang Huang, Deepak Kirpalani, and Michele Christine Loewen. 2022. "Insights into Increasing Selenate Reductase Enzyme Activity in the Presence of Nitrogen-Doped Graphite Electrodes for Selenium Effluent Treatment" Water 14, no. 6: 931. https://doi.org/10.3390/w14060931
APA StyleMohapatra, D. P., Robinson, K. A., Huang, F., Kirpalani, D., & Loewen, M. C. (2022). Insights into Increasing Selenate Reductase Enzyme Activity in the Presence of Nitrogen-Doped Graphite Electrodes for Selenium Effluent Treatment. Water, 14(6), 931. https://doi.org/10.3390/w14060931






