Salinity Shapes the Microbial Communities in Surface Sediments of Salt Lakes on the Tibetan Plateau, China
Abstract
1. Introduction
2. Materials and Methods
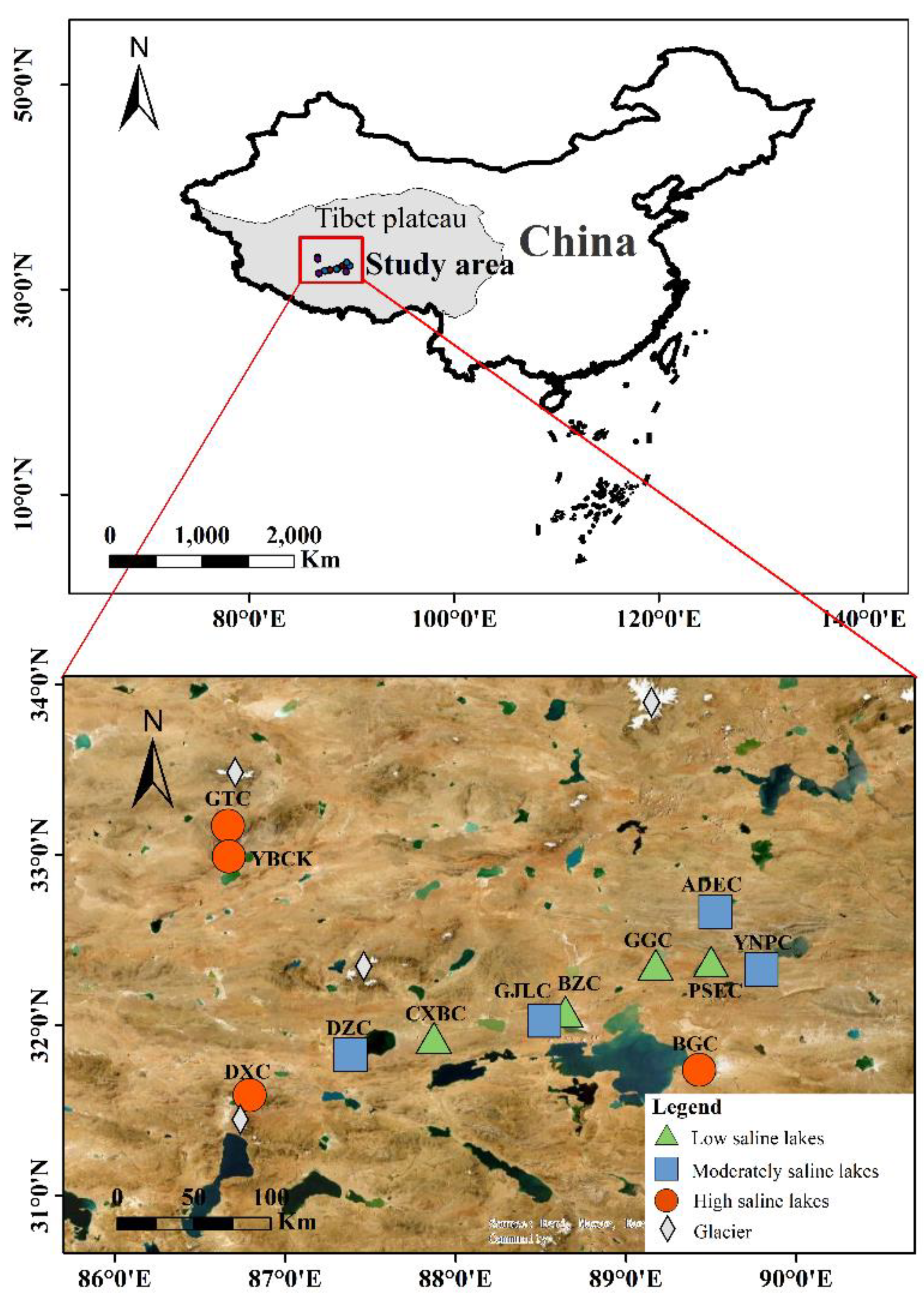
2.1. Study Site and Sampling
2.2. Determination of Environmental Parameters
2.3. DNA Extraction, High-throughput Sequencing and Sequence Analysis
2.4. Data Analysis
3. Results
3.1. Physical and Chemical Parameters of Lake Sediment
3.2. Comparison of Alpha Diversity among All Lakes
3.3. Microbial Community Composition of the 12 Lakes
3.4. Effect of Physicochemical Factors on Microbial Composition of Lakes
4. Discussion
4.1. Distribution of Microbial Community Composition along a Salinity Gradient
4.2. Effects of Environmental and Spatial Factors on Microbial Communities in these Lakes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Williams, W.D. What Future for Saline Lakes? Environ. Sci. Policy Sustain. Dev. 1996, 38, 12–39. [Google Scholar] [CrossRef]
- Bai, J. Situation of Study on Global Salt Lakes and Paleo- Climatic and Environmental Evolution of the Typical Region. Master’s Thesis, Hunan Normal University, Changsha, China, 2010. [Google Scholar]
- Zhang, G.; Xie, H.; Kang, S.; Yi, D.; Ackley, S.F. Monitoring lake level changes on the Tibetan Plateau using ICESat altimetry data (2003–2009). Remote Sens. Environ. 2011, 115, 1733–1742. [Google Scholar] [CrossRef]
- Zhang, G.; Yao, T.; Xie, H.; Zhang, K.; Zhu, F. Lakes’ state and abundance across the Tibetan Plateau. Chin. Sci. Bull. 2014, 59, 3010–3021. [Google Scholar] [CrossRef]
- Liu, Y.; Priscu, J.C.; Yao, T.; Vick-Majors, T.J.; Michaud, A.B.; Jiao, N.; Hou, J.; Tian, L.; Hu, A.; Chen, Z.Q. A comparison of pelagic, littoral, and riverine bacterial assemblages in Lake Bangongco, Tibetan Plateau. FEMS Microbiol. Ecol. 2014, 89, 211–221. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Zhu, L.; Wang, J.; Ju, J.; Ma, Q.; Qiao, B.; Wang, Y.; Xu, T.; Chen, H.; Kou, Q.; et al. In-situ water quality investigation of the lakes on the Tibetan Plateau. Sci. Bull. 2021, 66, 1727–1730. [Google Scholar] [CrossRef]
- Dong, G.; Yi, C.; Chen, L. An Introduction to the Physical Geography of the Qiangtang Plateau: A Frontier for Future Geoscience Research on the Tibetan Plateau. Phys. Geogr. 2010, 31, 475–492. [Google Scholar] [CrossRef]
- Li, H.; Zhang, D.; Hao, Y.; Zhao, H. Analysis of microbial diversity of root microecosystem of Orinus kokonorica. J. Shenyang Norm. Univ. 2016, 34, 227–233. [Google Scholar] [CrossRef]
- Boggs, D.; Eliot, I.; Knott, B. Salt lakes of the northern agricultural region, Western Australia. Hydrobiologia 2007, 576, 49–59. [Google Scholar] [CrossRef]
- Eder, W.; Jahnke, L.L.; Schmidt, M.; Huber, R. Microbial diversity of the brine-seawater interface of the Kebrit Deep, Red Sea, studied via 16S rRNA gene sequences and cultivation methods. Appl. Environ. Microbiol. 2001, 67, 3077–3085. [Google Scholar] [CrossRef]
- Yang, J.; Ma, L.; Jiang, H.; Wu, G.; Dong, H. Salinity shapes microbial diversity and community structure in surface sediments of the Qinghai-Tibetan Lakes. Sci. Rep. 2016, 6, 25078. [Google Scholar] [CrossRef]
- Ren, M.; Li, L.; Chen, L.; Xing, T.; Liu, Y.; Dong, X. Methanogen communities and predominant methanogenic pathways in three saline-alkaline lakes on the Tibetan Plateau. Acta Microbiol. Sin. 2020, 60, 161–171. [Google Scholar] [CrossRef]
- Xiong, J.; Liu, Y.; Lin, X.; Zhang, H.; Zeng, J.; Hou, J.; Yang, Y.; Yao, T.; Knight, R.; Chu, H. Geographic distance and pH drive bacterial distribution in alkaline lake sediments across Tibetan Plateau. Environ. Microbiol. 2012, 14, 2457–2466. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Zhang, G.; Jiang, H.; Yu, B.; Chapman, L.; Lucas, C.; Fields, M. Microbial Diversity in Sediments of Saline Qinghai Lake, China: Linking Geochemical Controls to Microbial Ecology. Microb. Ecol. 2006, 51, 65–82. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Zhang, X.; Shen, G.; Feng, X.; Long, Q.; Zhu, D. Microbial Community Structure and Diversity of Xiaochaidan Salt Lake on the Tibetan Plateau. J. Hydroecology 2017, 38, 55–64. [Google Scholar] [CrossRef]
- Zhang, X.; Liu, J.; Shen, G.; Long, Q.; Han, R.; Zhu, D. Illumina-based sequencing analysis of microbial community composition in Chaka Salt Lake in Qinghai-Tibet Plateau. Microbiol. China 2017, 44, 1834–1846. [Google Scholar]
- Zhu, D.; Liu, J.; Han, R.; Shen, G.; Yang, F.; Long, Q.; Liu, D. Population diversity and phylogeny of halophiles in the Qinghai Lake. Biodivers. Sci. 2012, 20, 495–504. [Google Scholar]
- Li, G.; Xue, L.; Mo, T.; Yang, R.; Li, W. New Progress of Research Methoneds for Microbial Diversity in Lake Sediment. J. Lanzhou Jiaotong Univ. 2015, 34, 12–16. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Huntley, J.; Fierer, N.; Owens, S.M.; Betley, J.; Fraser, L.; Bauer, M.; et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012, 6, 1621–1624. [Google Scholar] [CrossRef]
- Liao, W.; Tong, D.; Li, Z.; Nie, X.; Liu, Y.; Ran, F.; Liao, S. Characteristics of microbial community composition and its relationship with carbon, nitrogen and sulfur in sediments. Sci. Total Environ. 2021, 795, 148848. [Google Scholar] [CrossRef]
- Li, M.; Zhang, K.; Yan, Z.; Liu, L.; Kang, E.; Kang, X. Soil Water Content Shapes Microbial Community Along Gradients of Wetland Degradation on the Tibetan Plateau. Front. Microbiol. 2022, 13, 824267. [Google Scholar] [CrossRef]
- Moreau, M.M.; Eades, S.C.; Reinemeyer, C.R.; Fugaro, M.N.; Onishi, J.C. Illumina sequencing of the V4 hypervariable region 16S rRNA gene reveals extensive changes in bacterial communities in the cecum following carbohydrate oral infusion and development of early-stage acute laminitis in the horse. Vet. Microbiol. 2014, 168, 436–441. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Chen, L.; Liu, S.; Li, F.; Zhang, X.; Chen, H.; Liu, Y. Studying safe storage time of orange peel (Citrus reticulata) using high-throughput sequencing and conventional pure culture. Food Sci. Nutr. 2018, 6, 2545–2552. [Google Scholar] [CrossRef] [PubMed]
- Frade, P.R.; Glasl, B.; Matthews, S.A.; Mellin, C.; Serrão, E.A.; Wolfe, K.; Mumby, P.J.; Webster, N.S.; Bourne, D.G. Spatial patterns of microbial communities across surface waters of the Great Barrier Reef. Commun. Biol. 2020, 3, 442. [Google Scholar] [CrossRef]
- Gao, Q.; Wan, Y.; Li, Y.; Sheng, W.; Jiang, C.; Wang, B.; Li, W. Trends of grassland NPP and its response to human activity in Northern Tibet. Acta Ecol. Sin. 2007, 27, 4612–4619. [Google Scholar]
- Duan, M.; Gao, Q.; Wan, Y.; Li, Y.; Guo, Y.; Dan, J.; Luo, S. Effect of grazing on community characteristics and species diversity of Stipa purpurea alpine grassland in Northern Tibet. Acta Ecol. Sin. 2010, 30, 3892–3900. [Google Scholar]
- Niu, L.; Liu, Y.; Li, Y.; Ou, Y. Microbial community structure of the alpine meadow under different grzing styles in Naqu prefecture of Tibet. Chin. J. Appl. Ecol. 2015, 26, 2298–2306. [Google Scholar] [CrossRef]
- Deng, Y.; Liu, Y.; Dumont, M.; Conrad, R. Salinity Affects the Composition of the Aerobic Methanotroph Community in Alkaline Lake Sediments from the Tibetan Plateau. Microb. Ecol. 2017, 73, 101–110. [Google Scholar] [CrossRef]
- Lin, H.; Su, Y.; Zhuang, Y.; Zhong, H.; Lin, J.; Jin, H. A Comparision of Chlorophyll–a in the Sediment of Two Reservoirs in Fujian Province. J. Subtrop. Resour. Environ. 2011, 6, 42–48. [Google Scholar] [CrossRef]
- Kushner, D.J. Growth and nutrition of halophilic bacteria. In The Biology of Halophilic Bacteria; CRC Press: Boca Raton, FL, USA, 1993; pp. 87–103. [Google Scholar]
- Wang, Y.; Zheng, M.; Yan, L.; Bu, L.; Qi, W. Influence of the regional climate variations on lake changes of Zabuye, Dangqiong Co and Bankog Co salt lakes in Tibet. J. Geogr. Sci. 2019, 29, 1895–1907. [Google Scholar] [CrossRef]
- Ju, J.; Zhu, L.; Huang, L.; Yang, R.; Ma, Q.; Hu, X.; Wang, Y.; Zhen, X. Ranwu Lake, a proglacial lake with the potential to reflect glacial activity in SE Tibet. Chin. Sci. Bull. 2015, 60, 16–26. [Google Scholar]
- Liu, K.; Liu, Y.; Han, B.P.; Xu, B.; Zhu, L.; Ju, J.; Jiao, N.; Xiong, J. Bacterial community changes in a glacial-fed Tibetan lake are correlated with glacial melting. Sci. Total Environ. 2019, 651, 2059–2067. [Google Scholar] [CrossRef] [PubMed]
- Ben Abdallah, M.; Karray, F.; Kallel, N.; Armougom, F.; Mhiri, N.; Quemeneur, M.; Cayol, J.L.; Erauso, G.; Sayadi, S. Abundance and diversity of prokaryotes in ephemeral hypersaline lake Chott El Jerid using Illumina Miseq sequencing, DGGE and qPCR assays. Extremophiles 2018, 22, 811–823. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y. Bacterial Diversity of Haloalkaline Lakes in Desert Hunshandake in Inner Mongolia. Master’s Thesis, Inner Mongolia Agricultural University, Hohhot, China, 2011. [Google Scholar]
- Liu, Y.; Yao, T.; Jiao, N.; Zhu, L.; Hu, A.; Liu, X.; Gao, J.; Chen, Z. Salinity Impact on Bacterial Community Composition in Five High-Altitude Lakes from the Tibetan Plateau, Western China. Geomicrobiol. J. 2013, 30, 462–469. [Google Scholar] [CrossRef]
- Ventosa, A.; Fernandez, A.; Leon, M.; Sanchez-Porro, C.; Rodriguez-Valera, F. The Santa Pola saltern as a model for studying the microbiota of hypersaline environments. Extremophiles 2014, 18, 811–824. [Google Scholar] [CrossRef]
- Zhang, J.; Yang, Y.; Zhao, L.; Li, Y.; Xie, S.; Liu, Y. Distribution of sediment bacterial and archaeal communities in plateau freshwater lakes. Appl. Microbiol. Biotechnol. 2015, 99, 3291–3302. [Google Scholar] [CrossRef]
- Wu, Q.; Zwart, G.; Schauer, M.; Kamst-van Agterveld, M.P.; Hahn, M.W. Bacterioplankton community composition along a salinity gradient of sixteen high-mountain lakes located on the Tibetan Plateau, China. Appl. Environ. Microbiol. 2006, 72, 5478–5485. [Google Scholar] [CrossRef] [PubMed]
- Herlemann, D.P.R.; Lundin, D.; Andersson, A.F.; Labrenz, M.; Jurgens, K. Phylogenetic Signals of Salinity and Season in Bacterial Community Composition Across the Salinity Gradient of the Baltic Sea. Front. Microbiol. 2016, 7, 1883. [Google Scholar] [CrossRef] [PubMed]
- Bouvier, T.C.; del Giorgio, P.A. Compositional changes in free-living bacterial communities along a salinity gradient in two temperate estuaries. Limnol. Oceanogr. 2002, 47, 453–470. [Google Scholar] [CrossRef]
- Dupont, C.L.; Larsson, J.; Yooseph, S.; Ininbergs, K.; Goll, J.; Asplund-Samuelsson, J.; McCrow, J.P.; Celepli, N.; Allen, L.Z.; Ekman, M.; et al. Functional Tradeoffs Underpin Salinity-Driven Divergence in Microbial Community Composition. PLoS ONE 2014, 9, e89549. [Google Scholar] [CrossRef]
- Etesami, H.; Beattie, G.A. Mining Halophytes for Plant Growth-Promoting Halotolerant Bacteria to Enhance the Salinity Tolerance of Non-halophytic Crops. Frontiers in Microbiology 2018, 9, 148. [Google Scholar] [CrossRef]
- Humayoun, S.B.; Bano, N.; Hollibaugh, J.T. Depth distribution of microbial diversity in Mono Lake, a meromictic soda lake in California. Appl. Environ. Microbiol. 2003, 69, 1030–1042. [Google Scholar] [CrossRef] [PubMed]
- Saum, S.H.; Muller, V. Regulation of osmoadaptation in the moderate halophile Halobacillus halophilus: Chloride, glutamate and switching osmolyte strategies. Saline Syst. 2008, 4, 4. [Google Scholar] [CrossRef] [PubMed]
- Sleator, R.D.; Hill, C. Bacterial osmoadaptation: The role of osmolytes in bacterial stress and virulence. Fems Microbiol. Rev. 2002, 26, 49–71. [Google Scholar] [CrossRef] [PubMed]
- Oren, A. Microbial life at high salt concentrations: Phylogenetic and metabolic diversity. Saline Syst. 2008, 4, 2. [Google Scholar] [CrossRef] [PubMed]
- Mujakic, I.; Piwosz, K.; Koblizek, M. Phylum Gemmatimonadota and Its Role in the Environment. Microorg. 2022, 10, 151. [Google Scholar] [CrossRef] [PubMed]
- Huber, K.J.; Overmann, J. Vicinamibacteraceae fam. nov., the first described family within the subdivision 6 Acidobacteria. Int. J. Syst. Evol. Microbiol. 2018, 68, 2331–2334. [Google Scholar] [CrossRef]
- Wang, X.T.; Wang, X.M.; Zheng, W.S.; Zhang, X.K.; Du, Z.J. Aquiflexum aquatile sp. Nov., isolated from lake water. Int. J. Syst. Evol. Microbiol. 2019, 69, 1947–1952. [Google Scholar] [CrossRef]
- Pan, K.L.; Gao, J.F.; Li, D.C.; Fan, X.Y. The dominance of non-halophilic archaea in autotrophic ammonia oxidation of activated sludge under salt stress: A DNA-based stable isotope probing study. Bioresour. Technol. 2019, 291, 121914. [Google Scholar] [CrossRef]
- Banda, J.F.; Zhang, Q.; Ma, L.; Pei, L.; Du, Z.; Hao, C.; Dong, H. Both pH and salinity shape the microbial communities of the lakes in Badain Jaran Desert, NW China. Sci. Total Environ. 2021, 791, 148108. [Google Scholar] [CrossRef]
- Kaempfer, P.; Rekha, P.D.; Busse, H.J.; Arun, A.B.; Priyanka, P.; Glaeser, S.P. Halomonas malpeensis sp nov., isolated from rhizosphere sand of a coastal sand dune plant. Int. J. Syst. Evol. Microbiol. 2018, 68, 1037–1046. [Google Scholar] [CrossRef]
- Yang, H.; Gao, M.; Han, X.; Duan, H.; Wang, Z.; Sui, L. Planktonic Composition and Microbial Biodiversity in Five Salt Lakes in Tibet. J. Tianjin Univ. Sci. Technol. 2022, 37, 30–37. [Google Scholar] [CrossRef]
- Wu, H.; Wang, Z.; Yang, L. The Selective Enrichment and Diversity Analysis of Halophilic Bacteria in Sedimental Sample from Daban Salt Lake in Xinjiang. Microbiol. China 2010, 37, 956–961. [Google Scholar]
- Wang, S.; Hou, X.; Su, H. Exploration of the relationship between biogas production and microbial community under high salinity conditions. Sci. Rep. 2017, 7, 1149. [Google Scholar] [CrossRef] [PubMed]
- Ishikawa, M.; Yamasato, K. The genus Marinilactibacillus. In Lactic Acid Bacteria: Biodiversity and Taxonomy; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2014; pp. 125–134. [Google Scholar]
- Huang, J.; Yang, J.; Jiang, H.; Wu, G.; Liu, W.; Wang, B.; Xiao, H.; Han, J. Microbial Responses to Simulated Salinization and Desalinization in the Sediments of the Qinghai-Tibetan Lakes. Front. Microbiol. 2020, 11, 1772. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yang, J.; Jiang, H. Research progress on microbes involved in lacustrine sulfur cycling. Acta Microbiol. Sin. 2020, 60, 1177–1191. [Google Scholar] [CrossRef]
- Menendez-Serra, M.; Triado-Margarit, X.; Casamayor, E.O. Ecological and Metabolic Thresholds in the Bacterial, Protist, and Fungal Microbiome of Ephemeral Saline Lakes (Monegros Desert, Spain). Microb. Ecol. 2021, 82, 885–896. [Google Scholar] [CrossRef]
- Sorokin, D.Y.; Berben, T.; Melton, E.D.; Overmars, L.; Vavourakis, C.D.; Muyzer, G. Microbial diversity and biogeochemical cycling in soda lakes. Extremophiles 2014, 18, 791–809. [Google Scholar] [CrossRef]
- Bougouffa, S.; Yang, J.K.; Lee, O.O.; Wang, Y.; Batang, Z.; Al-Suwailem, A.; Qiana, P.Y. Distinctive Microbial Community Structure in Highly Stratified Deep-Sea Brine Water Columns. Appl. Environ. Microbiol. 2013, 79, 3425–3437. [Google Scholar] [CrossRef]
- Oren, A. Halophilic microbial communities and their environments. Curr. Opin. Biotechnol. 2015, 33, 119–124. [Google Scholar] [CrossRef]
- Morrissey, E.M.; Franklin, R.B. Evolutionary history influences the salinity preference of bacterial taxa in wetland soils. Front. Microbiol. 2015, 6, 1013. [Google Scholar] [CrossRef]
- Jackson, C.R.; Vallaire, S.C. Effects of salinity and nutrients on microbial assemblages in louisiana wetland sediments. Wetlands 2009, 29, 277–287. [Google Scholar] [CrossRef]
- Pavloudi, C.; Kristoffersen, J.B.; Oulas, A.; De Troch, M.; Arvanitidis, C. Sediment microbial taxonomic and functional diversity in a natural salinity gradient challenge Remane’s “species minimum” concept. PeerJ 2017, 5, e3687. [Google Scholar] [CrossRef] [PubMed]
- Edmonds, J.W.; Weston, N.B.; Joye, S.B.; Mou, X.Z.; Moran, M.A. Microbial Community Response to Seawater Amendment in Low-Salinity Tidal Sediments. Microb. Ecol. 2009, 58, 558–568. [Google Scholar] [CrossRef] [PubMed]
- Roychoudhury, A.N.; Cowan, D.; Porter, D.; Valverde, A. Dissimilatory sulphate reduction in hypersaline coastal pans: An integrated microbiological and geochemical study. Geobiology 2013, 11, 224–233. [Google Scholar] [CrossRef] [PubMed]
- Sorokin, D.Y.; Zacharova, E.E.; Pimenov, N.V.; Tourova, T.P.; Panteleeva, A.N.; Muyzer, G. Sulfidogenesis in hypersaline chloride-sulfate lakes of Kulunda Steppe (Altai, Russia). Fems Microbiol. Ecol. 2012, 79, 445–453. [Google Scholar] [CrossRef]
- Jin, Z.; Ci, M.; Yang, W.; Shen, D.; Hu, L.; Fang, C.; Long, Y. Sulfate reduction behavior in the leachate saturated zone of landfill sites. Sci. Total Environ. 2020, 730, 138946. [Google Scholar] [CrossRef]
- Fortunato, C.S.; Crump, B.C. Microbial Gene Abundance and Expression Patterns across a River to Ocean Salinity Gradient. PLoS ONE 2015, 10, e0140578. [Google Scholar] [CrossRef]
- Baker, B.J.; De Anda, V.; Seitz, K.W.; Dombrowski, N.; Santoro, A.E.; Lloyd, K.G. Diversity, ecology and evolution of Archaea. Nat. Microbiol. 2020, 5, 887–900. [Google Scholar] [CrossRef]
- Megyes, M.; Moga, J.; Strat, D.; Borsodi, A.K. Bacterial and Archaeal Taxonomic Diversity of Mud Volcanoes (Beciu, Romania) via Metagenomic Approach. Geomicrobiol. J. 2021, 38, 532–539. [Google Scholar] [CrossRef]
- Maignien, L.; Parkes, R.J.; Cragg, B.; Niemann, H.; Knittel, K.; Coulon, S.; Akhmetzhanov, A.; Boon, N. Anaerobic oxidation of methane in hypersaline cold seep sediments. Fems Microbiol. Ecol. 2013, 83, 214–231. [Google Scholar] [CrossRef]
- Liu, Y.; Priscu, J.C.; Xiong, J.; Conrad, R.; Vick-Majors, T.; Chu, H.; Hou, J. Salinity drives archaeal distribution patterns in high altitude lake sediments on the Tibetan Plateau. Fems Microbiol. Ecol. 2016, 92, fiw033. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Deng, S.; Huang, Q.; Dong, H.; Yu, B. Response of Aerobic Anoxygenic Phototrophic Bacterial Diversity to Environment Conditions in Saline Lakes and Daotang River on the Tibetan Plateau, NW China. Geomicrobiol. J. 2010, 27, 400–408. [Google Scholar] [CrossRef]
- Tang, X.; Xie, G.; Shao, K.; Sai, B.; Chen, Y.; Gao, G. Influence of Salinity on the Bacterial Community Composition in Lake Bosten, a Large Oligosaline Lake in Arid Northwestern China. Appl. Environ. Microbiol. 2012, 78, 4748–4751. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Yao, T.; Pearce, D.A.; Jiao, N.; Zeng, Y.; Guo, B.; Liu, Y. Bacteria in the lakes of the Tibetan Plateau and polar regions. Sci. Total Environ. 2021, 754, 142248. [Google Scholar] [CrossRef]
- Lindström, E.S.; Eiler, A.; Langenheder, S.; Bertilsson, S.; Drakare, S.; Ragnarsson, H.; Tranvik, L.J. Does ecosystem size determine aquatic bacterial richness? Comment. Ecology 2007, 88, 242–255. [Google Scholar] [CrossRef]
- Zhong, Z.; Liu, Y.; Miao, L.; Wang, F.; Chu, L.; Wang, J.; Liu, Z. Prokaryotic Community Structure Driven by Salinity and Ionic Concentrations in Plateau Lakes of the Tibetan Plateau. Appl. Environ. Microbiol. 2016, 82, 1846–1858. [Google Scholar] [CrossRef]
- Martiny, J.B.; Bohannan, B.J.; Brown, J.H.; Colwell, R.K.; Fuhrman, J.A.; Green, J.L.; Horner-Devine, M.C.; Kane, M.; Krumins, J.A.; Kuske, C.R.; et al. Microbial biogeography: Putting microorganisms on the map. Nat. Rev. Microbiol. 2006, 4, 102–112. [Google Scholar] [CrossRef]
- Logares, R.; Lindstrom, E.S.; Langenheder, S.; Logue, J.B.; Paterson, H.; Laybourn-Parry, J.; Rengefors, K.; Tranvik, L.; Bertilsson, S. Biogeography of bacterial communities exposed to progressive long-term environmental change. ISME J. 2013, 7, 937–948. [Google Scholar] [CrossRef]
- Jia, Q.; Liu, X.; Wang, H.; Liu, S.; Luo, Y.; Chen, L.; Zheng, M. Bio-ecological resources of saline lakes in Tibet and their economic prospect. Sci. Technol. Rev. 2017, 35, 19–26. [Google Scholar]
- Cai, P.; Chen, C.; Liu, Z.; Yang, T. Effect of Lighting on Vertical Distribution of Cyanobacteria in Hongfeng Reservoir in Summer. Guizhou Agric. Sci. 2013, 41, 176–179. [Google Scholar]
- He, H. Study on Planktonic and Sedimentary Microbial Community Structure and Molecular Ecological Network in the Weihe River Mainstem and Its Typical Tributaries from the Northern Foot of the Qinling Mountains. Master’s Thesis, Xi’an University of Technology, Xi’an, China, 2020. [Google Scholar]
- Sorokin, D.Y.; Tourova, T.P.; Sukhacheva, M.V.; Mardanov, A.V.; Ravin, N.V. Bacterial chitin utilisation at extremely haloalkaline conditions. Extremophiles 2012, 16, 883–894. [Google Scholar] [CrossRef] [PubMed]
- Blumel, M.; Suling, J.; Imhoff, J.F. Depth-specific distribution of Bacteroidetes in the oligotrophic eastern Mediterranean Sea. Aquat. Microb. Ecol. 2007, 46, 209–224. [Google Scholar] [CrossRef]
- Demergasso, C.; Casamayor, E.O.; Chong, G.; Galleguillos, P.; Escudero, L.; Pedros-Alio, C. Distribution of prokaryotic genetic diversity in athalassohaline lakes of the Atacama Desert, Northern Chile. FEMS Microbiol. Ecol. 2004, 48, 57–69. [Google Scholar] [CrossRef] [PubMed]
- O’Malley, M.A. The nineteenth century roots of ‘everything is everywhere’. Nat. Rev. Microbiol. 2007, 5, 647–651. [Google Scholar] [CrossRef]
- de Wit, R.; Bouvier, T. ‘Everything is everywhere, but the environment selects’; what did Baas Becking and Beijerinck really say? Environ. Microbiol. 2006, 8, 755–758. [Google Scholar] [CrossRef]
- Huang, S.; Liu, Y.; Hu, A.; Liu, X.; Chen, F.; Yao, T.; Jiao, N. Genetic Diversity of Picocyanobacteria in Tibetan Lakes: Assessing the Endemic and Universal Distributions. Appl. Environ. Microbiol. 2014, 80, 7640–7650. [Google Scholar] [CrossRef]








| Lake | Longitude (E) | Latitude (N) | Altitude (m) | Distance to Glaciers (km) | S(PSU) | pH | DO (mg/L) | T (°C) | W (%) | OM (%) | Chl-a (μg/g) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| BZC | 88°38′42.50″ | 32°04′57.88″ | 4536 | 81.00 | 0.64 | 8.9 | 6.81 | 20.6 | 15.83 | 3.41 | 0.06 |
| CXBC | 87°52′21.06″ | 31°55′45.66″ | 4499 | 44.00 | 2.89 | 8.96 | 6.42 | 19.18 | 23.57 | 0.33 | 0.16 |
| GGC | 89°10′30.44″ | 32°21′23.87″ | 4675 | 120.00 | 4.67 | 8.98 | 7.26 | 12.74 | 13.59 | 1.60 | 0.67 |
| PSEC | 89°29′58.92″ | 32°22′29.93″ | 4609 | 91.00 | 4.7 | 9.00 | 6.62 | 13.53 | 15.95 | 0.49 | 1.66 |
| GJLC | 88°31′18.15″ | 32°01′40.35″ | 4532 | 83.00 | 10.2 | 8.94 | 5.38 | 16.14 | 15.18 | 0.39 | 0.05 |
| DZC | 87°23′02.71″ | 31°49′40.52″ | 4472 | 28.00 | 11.82 | 8.19 | 6.62 | 18.86 | 21.29 | 1.51 | 0.39 |
| YNPC | 89°47′49″ | 32°19′46.64″ | 4633 | 124.00 | 32.31 | 9.03 | 3.58 | 18.55 | 14.58 | 0.87 | 1.60 |
| ADEC | 89°31′36″ | 32°40′00″ | 4854 | 143.00 | 33.88 | 8.87 | 5.33 | 13.28 | 12.93 | 0.37 | 0.04 |
| BGC | 89°25′49.61″ | 31°44′21.86″ | 4531 | 100.00 | 44.25 | 9.24 | 7.2 | 14.4 | 36.40 | 1.88 | 1.53 |
| GTC | 86°39′51.08″ | 33°10′17.90″ | 4872 | 7.00 | 45.52 | 8.85 | 3.73 | 13.78 | 16.99 | 1.03 | 0.11 |
| YBCK | 86°40′06.09″ | 32°59′31.10″ | 4559 | 14.00 | 49.85 | 8.76 | 5.36 | 16.32 | 58.33 | 12.54 | 1.62 |
| DXC | 86°47′30.24″ | 31°35′26.16″ | 4469 | 6.00 | 106.9 | 8.80 | 12.82 | 15.78 | 10.63 | 0.61 | 0.02 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, Y.; He, L.; Wang, Z.; Liang, T.; Sun, S.; Liu, X. Salinity Shapes the Microbial Communities in Surface Sediments of Salt Lakes on the Tibetan Plateau, China. Water 2022, 14, 4043. https://doi.org/10.3390/w14244043
He Y, He L, Wang Z, Liang T, Sun S, Liu X. Salinity Shapes the Microbial Communities in Surface Sediments of Salt Lakes on the Tibetan Plateau, China. Water. 2022; 14(24):4043. https://doi.org/10.3390/w14244043
Chicago/Turabian StyleHe, Yuanyuan, Lele He, Zhe Wang, Ting Liang, Shichun Sun, and Xiaoshou Liu. 2022. "Salinity Shapes the Microbial Communities in Surface Sediments of Salt Lakes on the Tibetan Plateau, China" Water 14, no. 24: 4043. https://doi.org/10.3390/w14244043
APA StyleHe, Y., He, L., Wang, Z., Liang, T., Sun, S., & Liu, X. (2022). Salinity Shapes the Microbial Communities in Surface Sediments of Salt Lakes on the Tibetan Plateau, China. Water, 14(24), 4043. https://doi.org/10.3390/w14244043







