Application of the Human Viral Surrogate Pepper Mild Mottle Virus for Wastewater Fecal Pollution Management
Abstract
:1. Introduction
2. Viral Contamination of Treated Wastewater
3. Human Health Risk of Virus-Associated Water Pollution
| Virus | Size of Viral Particle | Zoonotic Transmission | Aquatic Environment | References |
|---|---|---|---|---|
| Mastadenovirus A–F | 70–90 nm | No | Wastewater | [79,80] |
| Torque teno virus | 30 nm | Yes | River | [79,80,81] |
| Astrovirus | 28–30 nm | Potentially | Sewage water | [79,82,83] |
| Norovirus GI, GII | 35–40 nm | No | River | [79,84] |
| Sapovirus GI, GII | No | Wastewater and river | [75,79] | |
| Human-associated circovirus | 15–25 nm | No | Sewage | [85,86] |
| Hepatitis E virus type 1–4 | 27–34 nm | Yes | Tap and bottled water | [87,88] |
| Assorted papillomaviruses | 55 nm | No | Wastewater | [89,90] |
| Human bocavirus type 1–4 | 22 nm | No | Recycled water and sewage | [79,91] |
| Aichivirus A–B | 30–32 nm | No | Sewage and surface water | [92] |
| Cosavirus A | No | River and waste water | [85,93] | |
| Coxsackievirus B | No | Sewage water | [93,94] | |
| Enterovirus A–D | No | Groundwater | [93,95] | |
| Poliovirus type 1–3 | No | Wastewater | [93,96] | |
| Hepatitis A virus | 40–45 nm | No | Wastewater | [93,97] |
| BK polyomavirus | No | River and sewage water | [98] | |
| JC polyomavirus | No | Wastewater | [99] | |
| Rotavirus A | 60–80 nm | Potentially | Drinking water | [79,100] |
4. Management Strategies for Wastewater Pollution
4.1. Traditional Fecal Bacterial Indicators
4.2. Viral Indicators
| Organism | Habitat | Temperature | Duration (Days) | Log Reduction | Reference |
|---|---|---|---|---|---|
| Adenovirus | Groundwater | 4 | 132 | 1.00 | [119] |
| 20 | 36 | 1.00 | |||
| Adenovirus 40 | Seawater | 15 | 28 | 1.40 | [120] |
| 15 | 85 | 2.00 | |||
| Drinking water | 4 | 60 | 0.49 | ||
| 4 | 92 | 2.00 | |||
| Adenovirus 41 | Seawater | 15 | 28 | 1.60 | [120] |
| 15 | 77 | 2.00 | |||
| Drinking water | 4 | 60 | 1.00 | ||
| 4 | 304 | 2.00 | |||
| Rotavirus | Fresh water | 20 | 10 | 2.00 | [121] |
| 4 | 32 | 2.00 | |||
| Seawater | 37 | 7 | 5.00 | [122] | |
| Soil | 37 | 7 | 1.70 | [123] | |
| Drinking water | 20 | 64 | 2.00 | [124] | |
| Norovirus | Groundwater | 25 | 1266 | 1.79 | [125] |
| Mineral water | 25 | 80 | 1.30 | [126] | |
| 4 | 80 | 0.89 | |||
| Tap water | 25 | 80 | 0.80 | ||
| 4 | 80 | 3.00 | |||
| Hepatitis A virus | Seawater | 20 | 28 | 4.00 | [127] |
| Artificial seawater | 25 | 11 | 1.00 | [128] | |
| 24 | 19 | 1.00 | |||
| Drinking water | 4 | 60 | 1.60 | [120] | |
| 4 | 56 | 2.00 | |||
| Bottled water | 21 | 21 | 1.99 | [129] | |
| Astrovirus | Tap water | 20 | 30 | 2.00 | [130] |
| 4 | 60 | 2.00 |
4.3. Human Viral Surrogates
4.3.1. Coliphages
4.3.2. CrAssphage
4.3.3. Pepper Mild Mottle Virus
5. PMMoV Application as Fecal Indicator
5.1. Characteristics of Ideal Viral Indicator
5.1.1. Ease of Detection and Quantification
5.1.2. Human Waste Association
5.1.3. Presence in Wastewater at High Concentrations
5.1.4. Resistance to Wastewater Treatment
5.1.5. Persistence in the Aquatic Environment
PMMoV Occurrence in Freshwater Environments
PMMoV Occurrence in Marine Environment
5.1.6. Global Distribution and Temporal Stability
5.2. Suitability of PMMoV as a Viral Indicator of Human Fecal Pollution
5.2.1. Advantages of PMMoV as a Viral Indicator of Fecal Pollution
5.2.2. Limitations of Utilizing PMMoV as a Viral Indicator of Fecal Pollution
5.3. Application in Risk Assessment
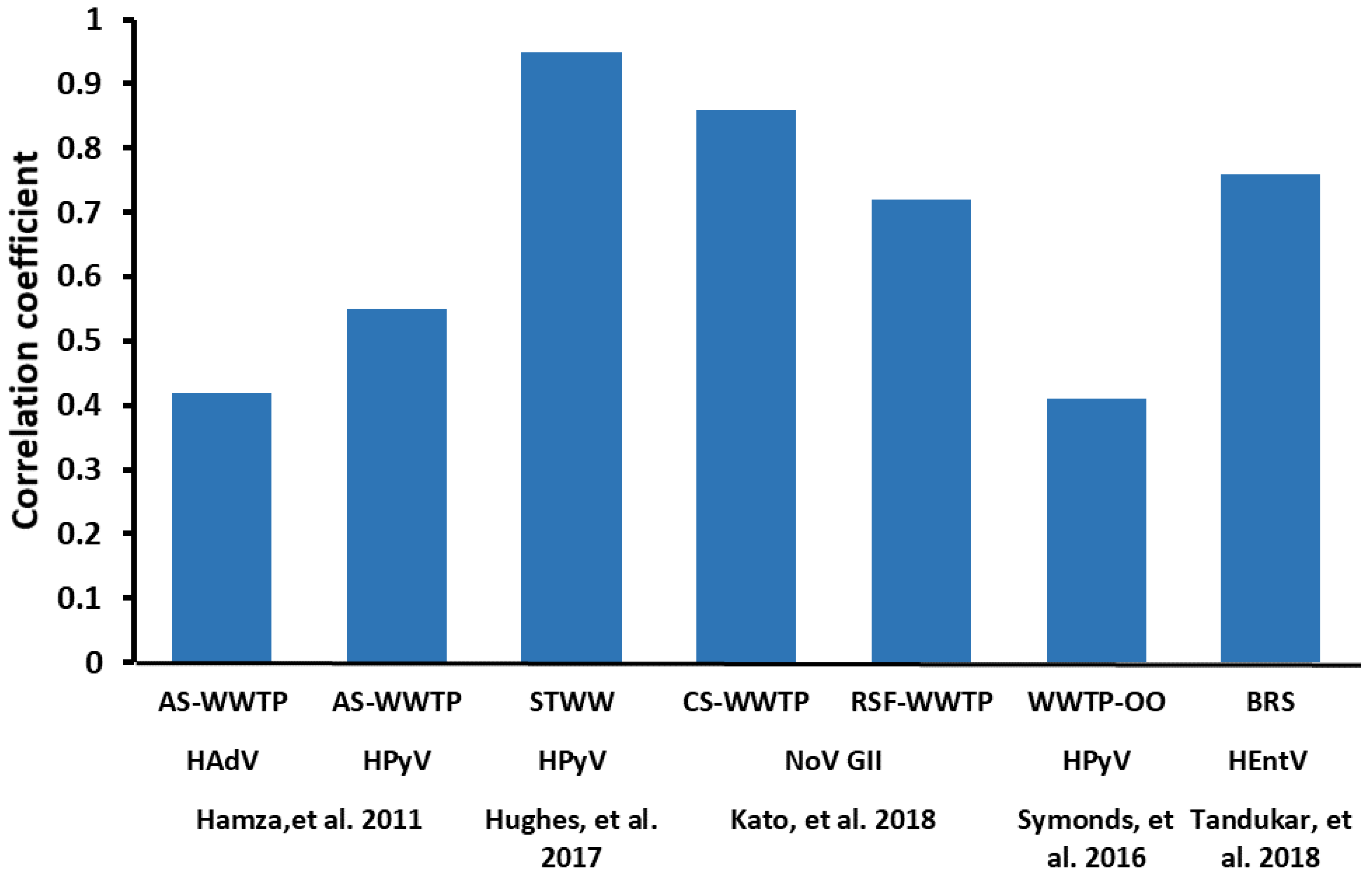
5.4. Correlation with Other Fecal Indicators
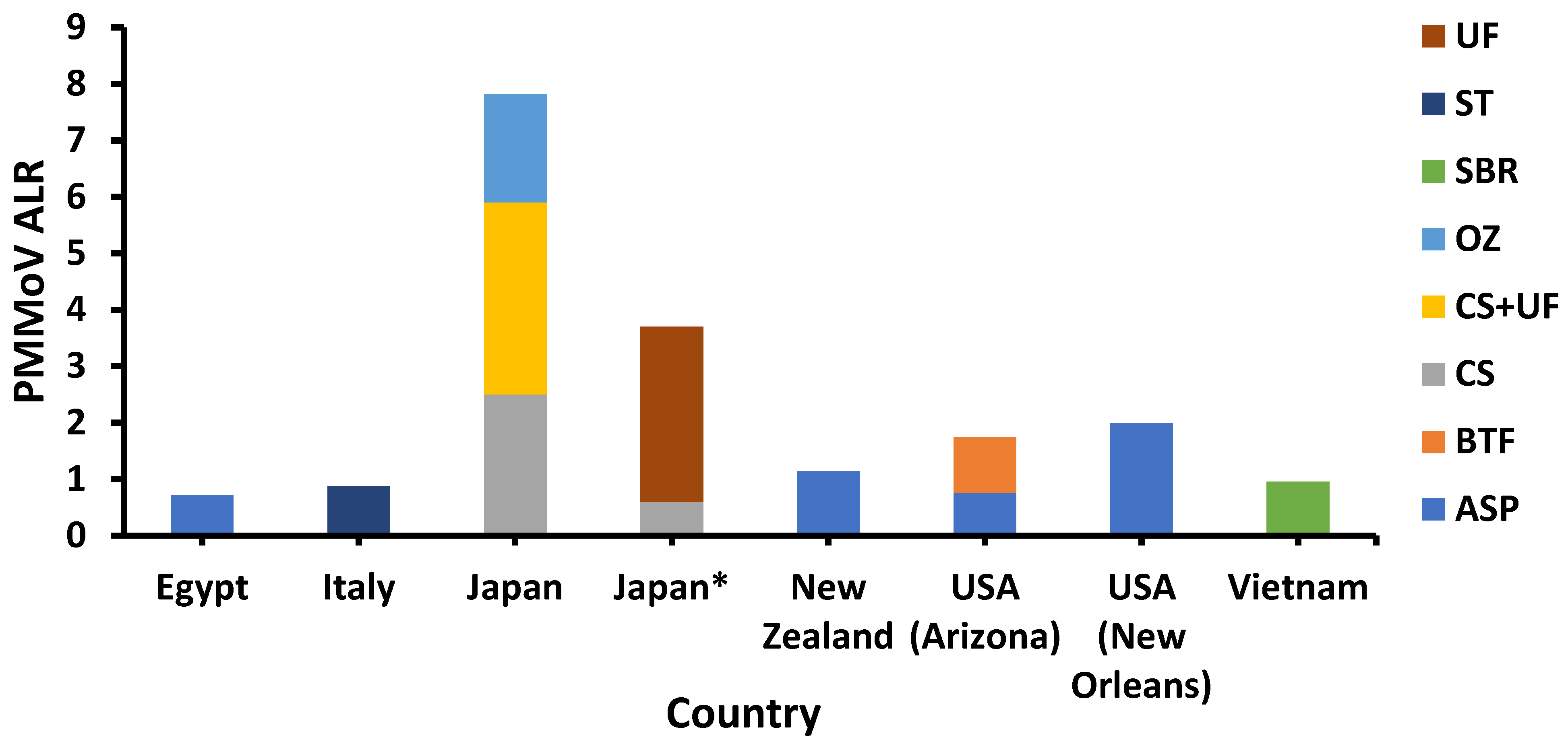
5.5. Removal in Wastewater and Wastewater Treatment Plant
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Karst, S.M. Viral Safeguard: The Enteric Virome Protects against Gut Inflammation. Immunity 2016, 44, 715–718. [Google Scholar] [CrossRef] [Green Version]
- Majumder, A.; Gupta, A.K.; Ghosal, P.S.; Varma, M. A Review on Hospital Wastewater Treatment: A Special Emphasis on Occurrence and Removal of Pharmaceutically Active Compounds, Resistant Microorganisms, and SARS-CoV-2. J. Environ. Chem. Eng. 2021, 9, 104812. [Google Scholar] [CrossRef] [PubMed]
- Virgin, H.W. The Virome in Mammalian Physiology and Disease. Cell 2014, 157, 142–150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Minot, S.; Grunberg, S.; Wu, G.D.; Lewis, J.D.; Bushman, F.D. Hypervariable Loci in the Human Gut Virome. Proc. Natl. Acad. Sci. USA 2012, 109, 3962–3966. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Virgin, H.W.; Wherry, E.J.; Ahmed, R. Redefining Chronic Viral Infection. Cell 2009, 138, 30–50. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Minot, S.; Bryson, A.; Chehoud, C.; Wu, G.D.; Lewis, J.D.; Bushman, F.D. Rapid Evolution of the Human Gut Virome. Proc. Natl. Acad. Sci. USA 2013, 110, 12450–12455. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Neil, J.A.; Cadwell, K. The Intestinal Virome and Immunity. J. Immunol. 2018, 201, 1615–1624. [Google Scholar] [CrossRef] [Green Version]
- Cadwell, K. The Virome in Host Health and Disease. Immunity 2015, 42, 805–813. [Google Scholar] [CrossRef] [Green Version]
- Espul, C.; Martínez, N.; Noel, J.S.; Cuello, H.; Abrile, C.; Grucci, S.; Glass, R.; Berke, T.; Matson, D.O. Prevalence and Characterization of Astroviruses in Argentinean Children with Acute Gastroenteritis. J. Med. Virol. 2004, 72, 75–82. [Google Scholar] [CrossRef]
- Lartey, B.L.; Quaye, O.; Damanka, S.A.; Agbemabiese, C.A.; Armachie, J.; Dennis, F.E.; Enweronu-Laryea, C.; Armah, G.E. Understanding Pediatric Norovirus Epidemiology: A Decade of Study among Ghanaian Children. Viruses 2020, 12, 1321. [Google Scholar] [CrossRef]
- Phan, T.; Ide, T.; Komoto, S.; Khamrin, P.; Pham, N.T.K.; Okitsu, S.; Taniguchi, K.; Nishimura, S.; Maneekarn, N.; Hayakawa, S. Genomic Analysis of Group A Rotavirus G12P [8] Including a New Japanese Strain Revealed Evidence for Intergenotypic Recombination in VP7 and VP4 Genes. Infect. Genet. Evol. 2021, 87, 104656. [Google Scholar] [CrossRef] [PubMed]
- Gorgich, M.; Mata, T.M.; Martins, A.; Caetano, N.S.; Formigo, N. Application of Domestic Greywater for Irrigating Agricultural Products: A Brief Study. Energy Rep. 2020, 6, 811–817. [Google Scholar] [CrossRef]
- Angelakis, A.N.; Asano, T.; Bahri, A.; Jimenez, B.E.; Tchobanoglous, G. Water Reuse: From Ancient to Modern Times and the Future. Front. Environ. Sci. 2018, 6, 26. [Google Scholar] [CrossRef] [Green Version]
- Libutti, A.; Gatta, G.; Gagliardi, A.; Vergine, P.; Pollice, A.; Beneduce, L.; Disciglio, G.; Tarantino, E. Agro-Industrial Wastewater Reuse for Irrigation of a Vegetable Crop Succession under Mediterranean Conditions. Agric. Water Manag. 2018, 196, 1–14. [Google Scholar] [CrossRef]
- Fountoulakis, M.S.; Markakis, N.; Petousi, I.; Manios, T. Single House On-Site Grey Water Treatment Using a Submerged Membrane Bioreactor for Toilet Flushing. Sci. Total Environ. 2016, 551, 706–711. [Google Scholar] [CrossRef] [PubMed]
- Becerra-Castro, C.; Lopes, A.R.; Vaz-Moreira, I.; Silva, E.F.; Manaia, C.M.; Nunes, O.C. Wastewater Reuse in Irrigation: A Microbiological Perspective on Implications in Soil Fertility and Human and Environmental Health. Environ. Int. 2015, 75, 117–135. [Google Scholar] [CrossRef] [Green Version]
- Jennings, W.C.; Chern, E.C.; O’Donohue, D.; Kellogg, M.G.; Boehm, A.B. Frequent Detection of a Human Fecal Indicator in the Urban Ocean: Environmental Drivers and Covariation with Enterococci. Environ. Sci. Process. Impacts 2018, 20, 480–492. [Google Scholar] [CrossRef]
- McGinnis, S.; Spencer, S.; Firnstahl, A.; Stokdyk, J.; Borchardt, M.; McCarthy, D.T.; Murphy, H.M. Human Bacteroides and Total Coliforms as Indicators of Recent Combined Sewer Overflows and Rain Events in Urban Creeks. Sci. Total Environ. 2018, 630, 967–976. [Google Scholar] [CrossRef]
- McMinn, B.R.; Ashbolt, N.J.; Korajkic, A. Bacteriophages as Indicators of Faecal Pollution and Enteric Virus Removal. Lett. Appl. Microbiol. 2017, 65, 11–26. [Google Scholar] [CrossRef] [Green Version]
- Figueras, M.J.; Borrego, J.J. New Perspectives in Monitoring Drinking Water Microbial Quality. Int. J. Environ. Res. Public Health 2010, 7, 4179–4202. [Google Scholar] [CrossRef]
- Boehm, A.B.; Van De Werfhorst, L.C.; Griffith, J.F.; Holden, P.A.; Jay, J.A.; Shanks, O.C.; Wang, D.; Weisberg, S.B. Performance of Forty-One Microbial Source Tracking Methods: A Twenty-Seven Lab Evaluation Study. Water Res. 2013, 47, 6812–6828. [Google Scholar] [CrossRef] [PubMed]
- Tillett, B.J.; Sharley, D.; Almeida, M.I.G.; Valenzuela, I.; Hoffmann, A.A.; Pettigrove, V. A Short Work-Flow to Effectively Source Faecal Pollution in Recreational Waters–a Case Study. Sci. Total Environ. 2018, 644, 1503–1510. [Google Scholar] [CrossRef] [PubMed]
- Kirs, M.; Kisand, V.; Wong, M.; Caffaro-Filho, R.A.; Moravcik, P.; Harwood, V.J.; Yoneyama, B.; Fujioka, R.S. Multiple Lines of Evidence to Identify Sewage as the Cause of Water Quality Impairment in an Urbanized Tropical Watershed. Water Res. 2017, 116, 23–33. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, W.; Staley, C.; Sadowsky, M.J.; Gyawali, P.; Sidhu, J.P.; Palmer, A.; Beale, D.J.; Toze, S. Toolbox Approaches Using Molecular Markers and 16S RRNA Gene Amplicon Data Sets for Identification of Fecal Pollution in Surface Water. Appl. Environ. Microbiol. 2015, 81, 7067–7077. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Symonds, E.M.; Sinigalliano, C.; Gidley, M.; Ahmed, W.; McQuaig-Ulrich, S.M.; Breitbart, M. Faecal Pollution along the Southeastern Coast of Florida and Insight into the Use of Pepper Mild Mottle Virus as an Indicator. J. Appl. Microbiol. 2016, 121, 1469–1481. [Google Scholar] [CrossRef]
- Ikner, L.A.; Gerba, C.P.; Bright, K.R. Concentration and Recovery of Viruses from Water: A Comprehensive Review. Food Environ. Virol. 2012, 4, 41–67. [Google Scholar] [CrossRef]
- Falman, J.C.; Fagnant-Sperati, C.S.; Kossik, A.L.; Boyle, D.S.; Meschke, J.S. Evaluation of Secondary Concentration Methods for Poliovirus Detection in Wastewater. Food Environ. Virol. 2019, 11, 20–31. [Google Scholar] [CrossRef] [Green Version]
- Elsäßer, D.; Ho, J.; Niessner, R.; Tiehm, A.; Seidel, M. Heterogeneous Asymmetric Recombinase Polymerase Amplification (HaRPA) for Rapid Hygiene Control of Large-Volume Water Samples. Anal. Biochem. 2018, 546, 58–64. [Google Scholar] [CrossRef]
- Gonzales-Gustavson, E.; Cárdenas-Youngs, Y.; Calvo, M.; da Silva, M.F.M.; Hundesa, A.; Amorós, I.; Moreno, Y.; Moreno-Mesonero, L.; Rosell, R.; Ganges, L. Characterization of the Efficiency and Uncertainty of Skimmed Milk Flocculation for the Simultaneous Concentration and Quantification of Water-Borne Viruses, Bacteria and Protozoa. J. Microbiol. Methods 2017, 134, 46–53. [Google Scholar] [CrossRef]
- Rusinol, M.; Fernandez-Cassi, X.; Timoneda, N.; Carratala, A.; Abril, J.F.; Silvera, C.; Figueras, M.J.; Gelati, E.; Rodó, X.; Kay, D. Evidence of Viral Dissemination and Seasonality in a Mediterranean River Catchment: Implications for Water Pollution Management. J. Environ. Manag. 2015, 159, 58–67. [Google Scholar] [CrossRef]
- Dhakar, V.; Geetanjali, A.S. Role of Pepper Mild Mottle Virus as a Tracking Tool for Fecal Pollution in Aquatic Environments. Arch. Microbiol. 2022, 204, 513. [Google Scholar] [CrossRef] [PubMed]
- Koonin, E.V.; Dolja, V.V.; Krupovic, M.; Kuhn, J.H. Viruses Defined by the Position of the Virosphere within the Replicator Space. Microbiol. Mol. Biol. Rev. 2021, 85, e00120–e00193. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.; Gupta, R.; Singh, R.L. Microbes and Environment. In Principles and Applications of Environmental Biotechnology for a Sustainable Future; Springer: Berlin/Heidelberg, Germany, 2017; pp. 43–84. [Google Scholar]
- Melkikh, A.V. Viruses, Immunity and Evolution. Biosystems 2022, 220, 104761. [Google Scholar] [CrossRef] [PubMed]
- Hernroth, B.E.; Conden-Hansson, A.-C.; Rehnstam-Holm, A.-S.; Girones, R.; Allard, A.K. Environmental Factors Influencing Human Viral Pathogens and Their Potential Indicator Organisms in the Blue Mussel, Mytilus Edulis: The First Scandinavian Report. Appl. Environ. Microbiol. 2002, 68, 4523–4533. [Google Scholar] [CrossRef] [Green Version]
- Kotwal, G.; Cannon, J.L. Environmental Persistence and Transfer of Enteric Viruses. Curr. Opin. Virol. 2014, 4, 37–43. [Google Scholar] [CrossRef]
- Tiwari, R.C.; Dikshit, M.; Mittal, B.; Sharma, V.B. Effects of Water Pollution-A Review Article. J. Ayurveda Integr. Med. Sci. 2022, 7, 63–68. [Google Scholar]
- Roos, Y.H. Water and Pathogenic Viruses Inactivation—Food Engineering Perspectives. Food Eng. Rev. 2020, 12, 251–267. [Google Scholar] [CrossRef]
- Lanrewaju, A.A.; Enitan-Folami, A.M.; Sabiu, S.; Edokpayi, J.N.; Swalaha, F.M. Global Public Health Implications of Human Exposure to Viral Contaminated Water. Front. Microbiol. 2022, 13, 981896. [Google Scholar] [CrossRef]
- Mihelcic, J.R.; Naughton, C.C.; Verbyla, M.E.; Zhang, Q.; Schweitzer, R.W.; Oakley, S.M.; Wells, E.C.; Whiteford, L.M. The Grandest Challenge of All: The Role of Environmental Engineering to Achieve Sustainability in the World’s Developing Regions. Environ. Eng. Sci. 2017, 34, 16–41. [Google Scholar] [CrossRef]
- Trussell, R.R.; Salveson, A.; Snyder, S.A.; Trussell, R.S.; Gerrity, D.; Pecson, B.M. Potable Reuse: State of the Science Report and Equivalency Criteria for Treatment Trains. WRRF 11-02; WateReuse Research Foundation: Alexandria, VA, USA, 2013; p. 21. Available online: http://www.twdb.texas.gov/publications/reports/contracted_reports/doc/1248321508_Vol2.pdf (accessed on 1 November 2022).
- Carvajal, G.; Roser, D.J.; Sisson, S.A.; Keegan, A.; Khan, S.J. Modelling Pathogen Log10 Reduction Values Achieved by Activated Sludge Treatment Using Naïve and Semi Naïve Bayes Network Models. Water Res. 2015, 85, 304–315. [Google Scholar] [CrossRef]
- Kobayashi, Y.; Ashbolt, N.J.; Davies, E.G.; Liu, Y. Life Cycle Assessment of Decentralized Greywater Treatment Systems with Reuse at Different Scales in Cold Regions. Environ. Int. 2020, 134, 105215. [Google Scholar] [CrossRef] [PubMed]
- Morrison, C.M.; Betancourt, W.Q.; Quintanar, D.R.; Lopez, G.U.; Pepper, I.L.; Gerba, C.P. Potential Indicators of Virus Transport and Removal during Soil Aquifer Treatment of Treated Wastewater Effluent. Water Res. 2020, 177, 115812. [Google Scholar] [CrossRef]
- Haramoto, E.; Kitajima, M.; Hata, A.; Torrey, J.R.; Masago, Y.; Sano, D.; Katayama, H. A Review on Recent Progress in the Detection Methods and Prevalence of Human Enteric Viruses in Water. Water Res. 2018, 135, 168–186. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.C.; Bischel, H.N.; Goel, R.; Rosso, D.; Sherchan, S.P.; Whiteson, K.L.; Yan, T.; Solo-Gabriele, H.M. Integrating Virus Monitoring Strategies for Safe Non-Potable Water Reuse. Water 2022, 14, 1187. [Google Scholar] [CrossRef]
- Kitajima, M.; Sassi, H.P.; Torrey, J.R. Pepper Mild Mottle Virus as a Water Quality Indicator. NPJ Clean Water 2018, 1, 19. [Google Scholar] [CrossRef] [Green Version]
- Kristanti, R.A.; Hadibarata, T.; Syafrudin, M.; Yılmaz, M.; Abdullah, S. Microbiological Contaminants in Drinking Water: Current Status and Challenges. Water. Air. Soil Pollut. 2022, 233, 299. [Google Scholar] [CrossRef]
- Okoh, A.I.; Sibanda, T.; Gusha, S.S. Inadequately Treated Wastewater as a Source of Human Enteric Viruses in the Environment. Int. J. Environ. Res. Public Health 2010, 7, 2620–2637. [Google Scholar] [CrossRef] [Green Version]
- Bouseettine, R.; Hassou, N.; Bessi, H.; Ennaji, M.M. Waterborne Transmission of Enteric Viruses and Their Impact on Public Health. In Emerging and Reemerging Viral Pathogens; Elsevier: Amsterdam, The Netherlands, 2020; pp. 907–932. [Google Scholar]
- Liang, G.; Bushman, F.D. The Human Virome: Assembly, Composition and Host Interactions. Nat. Rev. Microbiol. 2021, 19, 514–527. [Google Scholar] [CrossRef]
- Cooper, D.M.; McDonald, J.E.; Malham, S.K.; de Rougemont, A.; Jones, D.L. Seasonal and Spatial Dynamics of Enteric Viruses in Wastewater and in Riverine and Estuarine Receiving Waters. Sci. Total Environ. 2018, 634, 1174–1183. [Google Scholar]
- Kauppinen, A.; Pitkänen, T.; Miettinen, I.T. Persistent Norovirus Contamination of Groundwater Supplies in Two Waterborne Outbreaks. Food Environ. Virol. 2018, 10, 39–50. [Google Scholar] [CrossRef]
- Walker, D.I.; Adriaenssens, E.M.; McDonald, J.E.; Hillary, L.S.; Malham, S.K.; Jones, D.L. Viral Indicators for Tracking Domestic Wastewater Contamination in the Aquatic Environment. Water Res. 2020, 181, 115926. [Google Scholar]
- Chatziprodromidou, I.P.; Bellou, M.; Vantarakis, G.; Vantarakis, A. Viral outbreaks linked to fresh produce consumption: A systematic review. J. Appl. Microbiol. 2018, 124, 932–942. [Google Scholar] [CrossRef] [PubMed]
- Al-Ansari, N.; Abbas, N.; Laue, J.; Knutsson, S. Water Scarcity: Problems and Possible Solutions. J. Earth Sci. Geotech. Eng. 2021, 11, 243–312. [Google Scholar]
- Corpuz, M.V.A.; Buonerba, A.; Vigliotta, G.; Zarra, T.; Ballesteros, F., Jr.; Campiglia, P.; Belgiorno, V.; Korshin, G.; Naddeo, V. Viruses in Wastewater: Occurrence, Abundance and Detection Methods. Sci. Total Environ. 2020, 745, 140910. [Google Scholar] [CrossRef]
- Errani, F.; Ciulli, S.; Mandrioli, L.; Serratore, P.; Volpe, E. Detection of Human and Fish Viruses in Marine Gastropods. Animals 2022, 12, 2122. [Google Scholar] [CrossRef]
- Bányai, K.; Estes, M.K.; Martella, V.; Parashar, U.D. Viral Gastroenteritis. Lancet 2018, 392, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Harper, A.; Vijayakumar, V.; Ouwehand, A.C.; Ter Haar, J.; Obis, D.; Espadaler, J.; Binda, S.; Desiraju, S.; Day, R. Viral Infections, the Microbiome and Probiotics. Front. Cell Infect. Microbiol. 2021, 10, 596166. [Google Scholar] [CrossRef]
- Lopman, B.A.; Steele, D.; Kirkwood, C.D.; Parashar, U.D. The Vast and Varied Global Burden of Norovirus: Prospects for Prevention and Control. PLoS Med. 2016, 13, e1001999. [Google Scholar] [CrossRef]
- Montasser, K.A.; Youssef, M.I.; Ghandour, A.A.; Kamal, M. Infection with Adenovirus, Rotavirus, and Coinfection among Hospitalized Children with Gastroenteritis in an Egyptian University Hospital. J. Med. Virol. 2022, 94, 4950–4958. [Google Scholar] [CrossRef]
- Nour, I.; Hanif, A.; Ryan, M.; Eifan, S. Insights into Gastrointestinal Virome: Etiology and Public Exposure. Water 2021, 13, 2794. [Google Scholar] [CrossRef]
- Torre, P.; Aglitti, A.; Masarone, M.; Persico, M. Viral Hepatitis: Milestones, Unresolved Issues, and Future Goals. World J. Gastroenterol. 2021, 27, 4603. [Google Scholar] [CrossRef] [PubMed]
- Upfold, N.S.; Luke, G.A.; Knox, C. Occurrence of Human Enteric Viruses in Water Sources and Shellfish: A Focus on Africa. Food Environ. Virol. 2021, 13, 1–31. [Google Scholar] [CrossRef] [PubMed]
- Joshi, M.S.; Lole, K.S.; Barve, U.S.; Salve, D.S.; Ganorkar, N.N.; Chavan, N.A.; Shinde, M.S.; Gopalkrishna, V. Investigation of a Large Waterborne Acute Gastroenteritis Outbreak Caused by Group B Rotavirus in Maharashtra State, India. J. Med. Virol. 2019, 91, 1877–1881. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kauppinen, A.; Pitkänen, T.; Al-Hello, H.; Maunula, L.; Hokajärvi, A.-M.; Rimhanen-Finne, R.; Miettinen, I.T. Two Drinking Water Outbreaks Caused by Wastewater Intrusion Including Sapovirus in Finland. Int. J. Environ. Res. Public Health 2019, 16, 4376. [Google Scholar] [CrossRef]
- Naik, S.R.; Aggarwal, R.; Salunke, P.N.; Mehrotra, N.N. A Large Waterborne Viral Hepatitis E Epidemic in Kanpur, India. Bull. World Health Organ. 1992, 70, 597. [Google Scholar]
- Rachmadi, A.T.; Torrey, J.R.; Kitajima, M. Human Polyomavirus: Advantages and Limitations as a Human-Specific Viral Marker in Aquatic Environments. Water Res. 2016, 105, 456–469. [Google Scholar] [CrossRef]
- Di Bonito, P.; Iaconelli, M.; Gheit, T.; Tommasino, M.; Della Libera, S.; Bonadonna, L.; La Rosa, G. Detection of Oncogenic Viruses in Water Environments by a Luminex-Based Multiplex Platform for High Throughput Screening of Infectious Agents. Water Res. 2017, 123, 549–555. [Google Scholar] [CrossRef]
- Hamza, H.; Hamza, I.A. Oncogenic Papillomavirus and Polyomavirus in Urban Sewage in Egypt. Sci. Total Environ. 2018, 610, 1413–1420. [Google Scholar] [CrossRef]
- Fratini, M.; Di Bonito, P.; La Rosa, G. Oncogenic Papillomavirus and Polyomavirus in Water Environments: Is There a Potential for Waterborne Transmission? Food Environ. Virol. 2014, 6, 1–12. [Google Scholar] [CrossRef]
- Allander, T.; Tammi, M.T.; Eriksson, M.; Bjerkner, A.; Tiveljung-Lindell, A.; Andersson, B. Cloning of a Human Parvovirus by Molecular Screening of Respiratory Tract Samples. Proc. Natl. Acad. Sci. USA 2005, 102, 12891–12896. [Google Scholar] [CrossRef] [Green Version]
- Myrmel, M.; Lange, H.; Rimstad, E. A 1-Year Quantitative Survey of Noro-, Adeno-, Human Boca-, and Hepatitis E Viruses in Raw and Secondarily Treated Sewage from Two Plants in Norway. Food Environ. Virol. 2015, 7, 213–223. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Katayama, H.; Phanuwan, C.; Ohgaki, S. Quantitative Detection of Sapoviruses in Wastewater and River Water in Japan. Lett. Appl. Microbiol. 2008, 46, 408–413. [Google Scholar] [CrossRef] [PubMed]
- Adriaenssens, E.M.; McDonald, J.E.; Jones, D.L.; Allison, H.E.; McCarthy, A.J. Tracing the Fate of Wastewater Viruses Reveals Catchment-Scale Virome Diversity and Connectivity. Water Res. 2021, 203, 117568. [Google Scholar] [CrossRef] [PubMed]
- Bibby, K.; Peccia, J. Identification of Viral Pathogen Diversity in Sewage Sludge by Metagenome Analysis. Environ. Sci. Technol. 2013, 47, 1945–1951. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lebarbenchon, C.; Yang, M.; Keeler, S.P.; Ramakrishnan, M.A.; Brown, J.D.; Stallknecht, D.E.; Sreevatsan, S. Viral Replication, Persistence in Water and Genetic Characterization of Two Influenza A Viruses Isolated from Surface Lake Water. PLoS ONE 2011, 6, e26566. [Google Scholar] [CrossRef]
- King, A.M.; Adams, M.J.; Carstens, E.B.; Lefkowitz, E.J. Virus Taxonomy: Classification and Nomenclature of Viruses. In Virus Taxonomy: Classification and Nomenclature of Viruses; Springer: Berlin/Heidelberg, Germany, 2012; p. 1327. [Google Scholar]
- Lun, J.H.; Crosbie, N.D.; White, P.A. Genetic Diversity and Quantification of Human Mastadenoviruses in Wastewater from Sydney and Melbourne, Australia. Sci. Total Environ. 2019, 675, 305–312. [Google Scholar] [CrossRef]
- Verani, M.; Casini, B.; Battistini, R.; Pizzi, F.; Rovini, E.; Carducci, A. One-Year Monthly Monitoring of Torque Teno Virus (TTV) in River Water in Italy. Water Sci. Technol. 2006, 54, 191–195. [Google Scholar] [CrossRef]
- De Benedictis, P.; Schultz-Cherry, S.; Burnham, A.; Cattoli, G. Astrovirus Infections in Humans and Animals–Molecular Biology, Genetic Diversity, and Interspecies Transmissions. Infect. Genet. Evol. 2011, 11, 1529–1544. [Google Scholar] [CrossRef]
- Hamad, A.Y.; Mohammed, N.S.; Hasan, A.S. Molecular Detection and Phylogenetic Analysis of Astrovirus and Noroviruses from Sewage Water in Diyala-Iraq. J. Pharm. Negat. Results 2022, 13, 339–347. [Google Scholar]
- Pérez-Sautu, U.; Sano, D.; Guix, S.; Kasimir, G.; Pintó, R.M.; Bosch, A. Human Norovirus Occurrence and Diversity in the Llobregat River Catchment, Spain. Environ. Microbiol. 2012, 14, 494–502. [Google Scholar] [CrossRef]
- Blinkova, O.; Rosario, K.; Li, L.; Kapoor, A.; Slikas, B.; Bernardin, F.; Breitbart, M.; Delwart, E. Frequent Detection of Highly Diverse Variants of Cardiovirus, Cosavirus, Bocavirus, and Circovirus in Sewage Samples Collected in the United States. J. Clin. Microbiol. 2009, 47, 3507–3513. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Breitbart, M.; Delwart, E.; Rosario, K.; Segalés, J.; Varsani, A.; Consortium, I.R. ICTV Virus Taxonomy Profile: Circoviridae. J. Gen. Virol. 2017, 98, 1997. [Google Scholar] [CrossRef] [PubMed]
- Purdy, M.A.; Harrison, T.J.; Jameel, S.; Meng, X.-J.; Okamoto, H.; Van der Poel, W.H.M.; Smith, D.B.; Consortium, I.R. ICTV Virus Taxonomy Profile: Hepeviridae. J. Gen. Virol. 2017, 98, 2645. [Google Scholar] [CrossRef] [PubMed]
- Hennechart-Collette, C.; Dehan, O.; Laurentie, M.; Fraisse, A.; Martin-Latil, S.; Perelle, S. Method for Detecting Norovirus, Hepatitis A and Hepatitis E Viruses in Tap and Bottled Drinking Water. Int. J. Food Microbiol. 2022, 377, 109757. [Google Scholar] [CrossRef]
- Ahmed, N.I.; Elmahdy, E.M.; Allayh, A.K.; Mohamed, E.-C.B.; Loutfy, S.A.; Barakat, A.; Ali, M.A. Prevalence of Human Polyomavirus and Papillomavirus in Wastewater and in Stool of Egyptian Patients. Egypt. J. Aquat. Biol. Fish. 2019, 23, 29–41. [Google Scholar] [CrossRef] [Green Version]
- Van Doorslaer, K.; Chen, Z.; Bernard, H.-U.; Chan, P.K.; DeSalle, R.; Dillner, J.; Forslund, O.; Haga, T.; McBride, A.A.; Villa, L.L. ICTV Virus Taxonomy Profile: Papillomaviridae. J. Gen. Virol. 2018, 99, 989–990. [Google Scholar] [CrossRef]
- Booranathawornsom, T.; Pombubpa, K.; Tipayamongkholgul, M.; Kittigul, L. Molecular Characterization of Human Bocavirus in Recycled Water and Sewage Sludge in Thailand. Infect. Genet. Evol. 2022, 100, 105276. [Google Scholar] [CrossRef]
- Lodder, W.J.; Rutjes, S.A.; Takumi, K.; de Roda Husman, A.M. Aichi Virus in Sewage and Surface Water, the Netherlands. Emerg. Infect. Dis. 2013, 19, 1222. [Google Scholar] [CrossRef]
- Prevost, B.; Lucas, F.S.; Goncalves, A.; Richard, F.; Moulin, L.; Wurtzer, S. Large Scale Survey of Enteric Viruses in River and Waste Water Underlines the Health Status of the Local Population. Environ. Int. 2015, 79, 42–50. [Google Scholar] [CrossRef]
- El-Senousy, W.M.; Abdel-Moneim, A.; Abdel-Latif, M.; El-Hefnawy, M.H.; Khalil, R.G. Coxsackievirus B4 as a Causative Agent of Diabetes Mellitus Type 1: Is There a Role of Inefficiently Treated Drinking Water and Sewage in Virus Spreading? Food Environ. Virol. 2018, 10, 89–98. [Google Scholar] [CrossRef]
- Hsu, B.-M.; Chen, C.-H.; Wan, M.-T. Genetic Diversity of Epidemic Enterovirus 71 Strains Recovered from Clinical and Environmental Samples in Taiwan. Virus Res. 2007, 126, 69–75. [Google Scholar] [CrossRef]
- Bredykhina, M.; Shtepa, O.; Rezvykh, V.; Paliychuk, O.; Yurchenko, O.; Kovalenko, S.; Hernets, I. Wastewater as an Indicator of Virus Circulation among Population of Dnipropetrovsk Oblast, Ukraine. Online J. Public Health Inform. 2019, 11, e420. [Google Scholar] [CrossRef]
- Matos, A.; Mesquito, J.R.; Goncalves, D.; Abreu-Silva, J.; Luxo, C.; Nascimento, M.S. First Detection and Molecular Characterization of Hepatitis E Virus in Water from Wastewater Treatment Plants in Portugal. Ann. Agric. Environ. Med. 2018, 25, 364–367. [Google Scholar] [CrossRef] [Green Version]
- Moens, U.; Calvignac-Spencer, S.; Lauber, C.; Ramqvist, T.; Feltkamp, M.C.; Daugherty, M.D.; Verschoor, E.J.; Ehlers, B. ICTV Virus Taxonomy Profile: Polyomaviridae. J. Gen. Virol. 2017, 98, 1159–1160. [Google Scholar] [CrossRef] [PubMed]
- Levican, J.; Levican, A.; Ampuero, M.; Gaggero, A. JC Polyomavirus Circulation in One-Year Surveillance in Wastewater in Santiago, Chile. Infect. Genet. Evol. 2019, 71, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Miura, T.; Kadoya, S.; Takino, H.; Sano, D.; Akiba, M. Temporal Variations of Human and Animal Rotavirus A Genotypes in Surface Water Used for Drinking Water Production. Front. Microbiol. 2022, 13, 912147. [Google Scholar] [CrossRef] [PubMed]
- Pachepsky, Y.A.; Allende, A.; Boithias, L.; Cho, K.; Jamieson, R.; Hofstra, N.; Molina, M. Microbial Water Quality: Monitoring and Modeling. J. Environ. Qual. 2018, 47, 931–938. [Google Scholar] [CrossRef] [Green Version]
- Charles, K.J.; Nowicki, S.; Bartram, J.K. A Framework for Monitoring the Safety of Water Services: From Measurements to Security. NPJ Clean Water 2020, 3, 36. [Google Scholar] [CrossRef]
- Ishii, S.; Sadowsky, M.J. Escherichia Coli in the Environment: Implications for Water Quality and Human Health. Microbes Environ. 2008, 23, 101–108. [Google Scholar] [CrossRef] [Green Version]
- Slade, J.S.; Ford, B.J. Discharge to the Environment of Viruses in Wastewater, Sludges, and Aerosols. In Viral Pollution of the Environment; CRC Press: Boca Raton, FL, USA, 2018; pp. 3–16. [Google Scholar]
- Rajapaksha, P.; Elbourne, A.; Gangadoo, S.; Brown, R.; Cozzolino, D.; Chapman, J. A Review of Methods for the Detection of Pathogenic Microorganisms. Analyst 2019, 144, 396–411. [Google Scholar] [CrossRef]
- Edge, T.A.; Boehm, A.B. Classical and Molecular Methods to Measure Fecal Bacteria. Fecal Bact. 2010, 2, 241–273. [Google Scholar] [CrossRef]
- Sadeghi, S.; Nikaeen, M.; Mohammadi, F.; Nafez, A.H.; Gholipour, S.; Shamsizadeh, Z.; Hadi, M. Microbial Characteristics of Municipal Solid Waste Compost: Occupational and Public Health Risks from Surface Applied Compost. Waste Manag. 2022, 144, 98–105. [Google Scholar] [CrossRef] [PubMed]
- Rock, C.M.; Brassill, N.; Dery, J.L.; Carr, D.; McLain, J.E.; Bright, K.R.; Gerba, C.P. Review of Water Quality Criteria for Water Reuse and Risk-Based Implications for Irrigated Produce under the FDA Food Safety Modernization Act, Produce Safety Rule. Environ. Res. 2019, 172, 616–629. [Google Scholar] [CrossRef]
- Ho, J.Y.; Lavinya, A.A.; Kay, D.S.W.; Lee, C.I.S.; Razmi, A.H.; Walsh, C.L.; Goodson, M.L.; Eswaran, J. Towards an Integrated Approach to Improve the Understanding of the Relationships Between Water-Borne Infections and Health Outcomes: Using Malaysia as a Detailed Case Study. Front. Water 2022, 4, 779860. [Google Scholar] [CrossRef]
- Bisseux, M.; Colombet, J.; Mirand, A.; Roque-Afonso, A.-M.; Abravanel, F.; Izopet, J.; Archimbaud, C.; Peigue-Lafeuille, H.; Debroas, D.; Bailly, J.-L. Monitoring Human Enteric Viruses in Wastewater and Relevance to Infections Encountered in the Clinical Setting: A One-Year Experiment in Central France, 2014 to 2015. Eurosurveillance 2018, 23, 17-00183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sinclair, R.G.; Rose, J.B.; Hashsham, S.A.; Gerba, C.P.; Haas, C.N. Criteria for Selection of Surrogates Used to Study the Fate and Control of Pathogens in the Environment. Appl. Environ. Microbiol. 2012, 78, 1969–1977. [Google Scholar] [CrossRef] [PubMed]
- Jankowski, P.; Gan, J.; Le, T.; McKennitt, M.; Garcia, A.; Yanaç, K.; Yuan, Q.; Uyaguari-Diaz, M. Metagenomic Community Composition and Resistome Analysis in a Full-Scale Cold Climate Wastewater Treatment Plant. Environ. Microb. 2022, 17, 3. [Google Scholar] [CrossRef]
- Savichtcheva, O.; Okabe, S. Alternative Indicators of Fecal Pollution: Relations with Pathogens and Conventional Indicators, Current Methodologies for Direct Pathogen Monitoring and Future Application Perspectives. Water Res. 2006, 40, 2463–2476. [Google Scholar] [CrossRef]
- McCall, C.; Xagoraraki, I. Metagenomic Approaches for Detecting Viral Diversity in Water Environments. J. Environ. Eng. 2019, 148, 04019039. [Google Scholar] [CrossRef]
- Morsy El-Senousy, W. Suitability of Some Viruses as Indices of Viral Pollution of Water. Egypt. J. Aquat. Biol. Fish. 2021, 25, 1049–1084. [Google Scholar] [CrossRef]
- Franco, E.; Meleleo, C.; Serino, L.; Sorbara, D.; Zaratti, L. Hepatitis A: Epidemiology and Prevention in Developing Countries. World J. Hepatol. 2012, 4, 68. [Google Scholar] [CrossRef]
- Ahmed, W.; Bivins, A.; Payyappat, S.; Cassidy, M.; Harrison, N.; Besley, C. Distribution of Human Fecal Marker Genes and Their Association with Pathogenic Viruses in Untreated Wastewater Determined Using Quantitative PCR. Water Res. 2022, 226, 119093. [Google Scholar] [CrossRef]
- Canh, V.D.; Lien, N.T.; Nga, T.T.V. Evaluation of the Suitability of Pepper Mild Mottle Virus (PMMoV) as an Indicator Virus for Water Safety and Quality. J. Sci. Technol. Civ. Eng. 2022, 16, 76–88. [Google Scholar] [CrossRef]
- Ogorzaly, L.; Bertrand, I.; Paris, M.; Maul, A.; Gantzer, C. Occurrence, Survival, and Persistence of Human Adenoviruses and F-Specific RNA Phages in Raw Groundwater. Appl. Environ. Microbiol. 2010, 76, 8019. [Google Scholar] [CrossRef] [Green Version]
- Enriquez, C.E.; Hurst, C.J.; Gerba, C.P. Survival of the Enteric Adenoviruses 40 and 41 in Tap, Sea, and Waste Water. Water Res. 1995, 29, 2548–2553. [Google Scholar] [CrossRef]
- Rzeżutka, A.; Cook, N. Survival of Human Enteric Viruses in the Environment and Food. FEMS Microbiol. Rev. 2004, 28, 441–453. [Google Scholar] [CrossRef] [PubMed]
- Pinon, A.; Vialette, M. Survival of Viruses in Water. Intervirology 2018, 61, 214–222. [Google Scholar] [CrossRef] [PubMed]
- Davidson, P.C.; Kuhlenschmidt, T.B.; Bhattarai, R.; Kalita, P.K.; Kuhlenschmidt, M.S. Investigation of Rotavirus Survival in Different Soil Fractions and Temperature Conditions. J. Environ. Prot. 2013, 4, 34107. [Google Scholar] [CrossRef] [Green Version]
- Raphael, R.A.; Sattar, S.A.; Springthorpe, V.S. Long-Term Survival of Human Rotavirus in Raw and Treated River Water. Can. J. Microbiol. 1985, 31, 124–128. [Google Scholar] [CrossRef]
- Seitz, S.R.; Leon, J.S.; Schwab, K.J.; Lyon, G.M.; Dowd, M.; McDaniels, M.; Abdulhafid, G.; Fernandez, M.L.; Lindesmith, L.C.; Baric, R.S. Norovirus Infectivity in Humans and Persistence in Water. Appl. Environ. Microbiol. 2011, 77, 6884. [Google Scholar] [CrossRef] [Green Version]
- Ngazoa, E.S.; Fliss, I.; Jean, J. Quantitative Study of Persistence of Human Norovirus Genome in Water Using TaqMan Real-Time RT-PCR. J. Appl. Microbiol. 2008, 104, 707–715. [Google Scholar] [CrossRef]
- Callahan, K.M.; Taylor, D.J.; Sobsey, M.D. Comparative Survival of Hepatitis A Virus, Poliovirus and Indicator Viruses in Geographically Diverse Seawaters. Water Sci. Technol. 1995, 31, 189–193. [Google Scholar] [CrossRef]
- Crance, J.M.; Gantzer, C.; Schwartzbrod, L.; Deloince, R. Effect of Temperature on the Survival of Hepatitis A Virus and Its Capsidal Antigen in Synthetic Seawater. Environ. Toxicol. Water Qual. Int. J. 1998, 13, 89–92. [Google Scholar] [CrossRef]
- Leblanc, D.; Gagné, M.-J.; Poitras, É.; Brassard, J. Persistence of Murine Norovirus, Bovine Rotavirus, and Hepatitis A Virus on Stainless Steel Surfaces, in Spring Water, and on Blueberries. Food Microbiol. 2019, 84, 103257. [Google Scholar] [CrossRef] [PubMed]
- Abad, F.X.; Pintó, R.M.; Villena, C.; Gajardo, R.; Bosch, A. Astrovirus Survival in Drinking Water. Appl. Environ. Microbiol. 1997, 63, 3119–3122. [Google Scholar] [CrossRef] [Green Version]
- Sharma, S.; Chatterjee, S.; Datta, S.; Prasad, R.; Dubey, D.; Prasad, R.K.; Vairale, M.G. Bacteriophages and Its Applications: An Overview. Folia Microbiol. 2017, 62, 17–55. [Google Scholar] [CrossRef]
- Elbahnasawy, M.A.; ElSayed, E.E.; Azzam, M.I. Newly Isolated Coliphages for Bio-Controlling Multidrug-Resistant Escherichia Coli Strains. Environ. Nanotechnol. Monit. Manag. 2021, 16, 100542. [Google Scholar] [CrossRef]
- Li, E.; Saleem, F.; Edge, T.A.; Schellhorn, H.E. Biological Indicators for Fecal Pollution Detection and Source Tracking: A Review. Processes 2021, 9, 2058. [Google Scholar] [CrossRef]
- Fitzmorris-Brisolara, K.; Maal-Bared, R.; Worley-Morse, T.; Danley-Thomson, A.; Sobsey, M. Monitoring Coliphages to Reduce Waterborne Infectious Disease Transmission in the One Water Framework. Int. J. Hyg. Environ. Health 2022, 240, 113921. [Google Scholar] [CrossRef]
- Smither, S.J.; Lear-Rooney, C.; Biggins, J.; Pettitt, J.; Lever, M.S.; Olinger, G.G., Jr. Comparison of the Plaque Assay and 50% Tissue Culture Infectious Dose Assay as Methods for Measuring Filovirus Infectivity. J. Virol. Methods 2013, 193, 565–571. [Google Scholar] [CrossRef]
- Anderson-Coughlin, B.L.; Craighead, S.; Kelly, A.; Gartley, S.; Vanore, A.; Johnson, G.; Jiang, C.; Haymaker, J.; White, C.; Foust, D. Enteric Viruses and Pepper Mild Mottle Virus Show Significant Correlation in Select Mid-Atlantic Agricultural Waters. Appl. Environ. Microbiol. 2021, 87, e00211–e00221. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, W.; Kitajima, M.; Tandukar, S.; Haramoto, E. Recycled Water Safety: Current Status of Traditional and Emerging Viral Indicators. Curr. Opin. Environ. Sci. Health 2020, 16, 62–72. [Google Scholar] [CrossRef]
- Jones, T.H.; Brassard, J.; Topp, E.; Wilkes, G.; Lapen, D.R. Waterborne Viruses and F-Specific Coliphages in Mixed-Use Watersheds: Microbial Associations, Host Specificities, and Affinities with Environmental/Land Use Factors. Appl. Environ. Microbiol. 2017, 83, e02763–e02816. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- European Union. Directive (EU) 2020/2184 of the European Parliament and of the Council of 16 December 2020 on the Quality of Water Intended for Human Consumption. Off. J. Eur. Union 2020, 435, 1–62. [Google Scholar]
- Jofre, J.; Blanch, A.R.; Lucena, F.; Muniesa, M. Bacteriophages Infecting Bacteroides as a Marker for Microbial Source Tracking. Water Res. 2014, 55, 1–11. [Google Scholar] [CrossRef]
- Koonin, E.V.; Yutin, N. The CrAss-like Phage Group: How Metagenomics Reshaped the Human Virome. Trends Microbiol. 2020, 28, 349–359. [Google Scholar] [CrossRef]
- Malla, B.; Ghaju Shrestha, R.; Tandukar, S.; Sherchand, J.B.; Haramoto, E. Performance Evaluation of Human-Specific Viral Markers and Application of Pepper Mild Mottle Virus and CrAssphage to Environmental Water Samples as Fecal Pollution Markers in the Kathmandu Valley, Nepal. Food Environ. Virol. 2019, 11, 274–287. [Google Scholar] [CrossRef]
- Stachler, E.; Kelty, C.; Sivaganesan, M.; Li, X.; Bibby, K.; Shanks, O.C. Quantitative CrAssphage PCR Assays for Human Fecal Pollution Measurement. Environ. Sci. Technol. 2017, 51, 9146–9154. [Google Scholar] [CrossRef]
- Rosario, K.; Symonds, E.M.; Sinigalliano, C.; Stewart, J.; Breitbart, M. Pepper Mild Mottle Virus as an Indicator of Fecal Pollution. Appl. Environ. Microbiol. 2009, 75, 7261–7267. [Google Scholar] [CrossRef] [Green Version]
- Hamza, H.; Rizk, N.M.; Gad, M.A.; Hamza, I.A. Pepper Mild Mottle Virus in Wastewater in Egypt: A Potential Indicator of Wastewater Pollution and the Efficiency of the Treatment Process. Arch. Virol. 2019, 164, 2707–2713. [Google Scholar] [CrossRef]
- Hasiów-Jaroszewska, B.; Budzyńska, D.; Zarzyńska-Nowak, A. First Report of Pepper Mild Mottle Virus in Peppers in Poland. Plant Dis. 2019, 103, 1441. [Google Scholar] [CrossRef]
- Zhang, T.; Breitbart, M.; Lee, W.H.; Run, J.-Q.; Wei, C.L.; Soh, S.W.L.; Hibberd, M.L.; Liu, E.T.; Rohwer, F.; Ruan, Y. RNA Viral Community in Human Feces: Prevalence of Plant Pathogenic Viruses. PLoS Biol. 2006, 4, e3. [Google Scholar] [CrossRef] [Green Version]
- Symonds, E.M.; Nguyen, K.H.; Harwood, V.J.; Breitbart, M. Pepper Mild Mottle Virus: A Plant Pathogen with a Greater Purpose in (Waste) Water Treatment Development and Public Health Management. Water Res. 2018, 144, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Betancourt, W.Q.; Kitajima, M.; Wing, A.D.; Regnery, J.; Drewes, J.E.; Pepper, I.L.; Gerba, C.P. Assessment of Virus Removal by Managed Aquifer Recharge at Three Full-Scale Operations. J. Environ. Sci. Health Part A 2014, 49, 1685–1692. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, K.; Nakada, N.; Hanamoto, S.; Inaba, M.; Katayama, H.; Do, A.T.; Nga, T.T.V.; Oguma, K.; Hayashi, T.; Takizawa, S. Pepper Mild Mottle Virus as an Indicator and a Tracer of Fecal Pollution in Water Environments: Comparative Evaluation with Wastewater-Tracer Pharmaceuticals in Hanoi, Vietnam. Sci. Total Environ. 2015, 506, 287–298. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Colson, P.; Richet, H.; Desnues, C.; Balique, F.; Moal, V.; Grob, J.-J.; Berbis, P.; Lecoq, H.; Harlé, J.-R.; Berland, Y. Pepper Mild Mottle Virus, a Plant Virus Associated with Specific Immune Responses, Fever, Abdominal Pains, and Pruritus in Humans. PLoS ONE 2010, 5, e10041. [Google Scholar] [CrossRef]
- Hamza, I.A.; Jurzik, L.; Überla, K.; Wilhelm, M. Evaluation of Pepper Mild Mottle Virus, Human Picobirnavirus and Torque Teno Virus as Indicators of Fecal Contamination in River Water. Water Res. 2011, 45, 1358–1368. [Google Scholar] [CrossRef]
- Haramoto, E.; Kitajima, M.; Kishida, N.; Konno, Y.; Katayama, H.; Asami, M.; Akiba, M. Occurrence of Pepper Mild Mottle Virus in Drinking Water Sources in Japan. Appl. Environ. Microbiol. 2013, 79, 7413–7418. [Google Scholar] [CrossRef] [Green Version]
- Kitajima, M.; Iker, B.C.; Pepper, I.L.; Gerba, C.P. Relative Abundance and Treatment Reduction of Viruses during Wastewater Treatment Processes—Identification of Potential Viral Indicators. Sci. Total Environ. 2014, 488, 290–296. [Google Scholar] [CrossRef]
- Symonds, E.M.; Rosario, K.; Breitbart, M. Pepper Mild Mottle Virus: Agricultural Menace Turned Effective Tool for Microbial Water Quality Monitoring and Assessing (Waste) Water Treatment Technologies. PLoS Pathog. 2019, 15, e1007639. [Google Scholar] [CrossRef]
- Harwood, V.J.; Boehm, A.B.; Sassoubre, L.M.; Vijayavel, K.; Stewart, J.R.; Fong, T.-T.; Caprais, M.-P.; Converse, R.R.; Diston, D.; Ebdon, J. Performance of Viruses and Bacteriophages for Fecal Source Determination in a Multi-Laboratory, Comparative Study. Water Res. 2013, 47, 6929–6943. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gyawali, P.; Croucher, D.; Ahmed, W.; Devane, M.; Hewitt, J. Evaluation of Pepper Mild Mottle Virus as an Indicator of Human Faecal Pollution in Shellfish and Growing Waters. Water Res. 2019, 154, 370–376. [Google Scholar] [CrossRef] [PubMed]
- Leland, D.S.; Ginocchio, C.C. Role of Cell Culture for Virus Detection in the Age of Technology. Clin. Microbiol. Rev. 2007, 20, 49–78. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mostafa-Hedeab, G.; Allayeh, A.K.; Elhady, H.A.; Eledrdery, A.Y.; Mraheil, M.A.; Mostafa, A. Viral Eco-Genomic Tools: Development and Implementation for Aquatic Biomonitoring. Int. J. Environ. Res. Public Health 2022, 19, 7707. [Google Scholar] [CrossRef] [PubMed]
- Ashbolt, N.J. Microbial Contamination of Drinking Water and Human Health from Community Water Systems. Curr. Environ. Health Rep. 2015, 2, 95–106. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tandukar, S.; Sherchan, S.P.; Haramoto, E. Applicability of CrAssphage, Pepper Mild Mottle Virus, and Tobacco Mosaic Virus as Indicators of Reduction of Enteric Viruses during Wastewater Treatment. Sci. Rep. 2020, 10, 3616. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, J.; Ganesh, A. Water Quality Indicators: Bacteria, Coliphages, Enteric Viruses. Int. J. Environ. Health Res. 2013, 23, 484–506. [Google Scholar] [CrossRef] [PubMed]
- Herzog, A.B.; Bhaduri, P.; Stedtfeld, R.D.; Seyrig, G.; Ahmad, F.; Dave, P.K.; Hashsham, S.A. Detection and Occurrence of Indicator Organisms and Pathogens. Water Environ. Res. 2010, 82, 883–907. [Google Scholar] [CrossRef]
- Cashdollar, J.L.; Wymer, L. Methods for Primary Concentration of Viruses from Water Samples: A Review and Meta-Analysis of Recent Studies. J. Appl. Microbiol. 2013, 115, 1–11. [Google Scholar] [CrossRef]
- Ibrahim, Y.; Ouda, M.; Kadadou, D.; Banat, F.; Naddeo, V.; Alsafar, H.; Yousef, A.F.; Barceló, D.; Hasan, S.W. Detection and Removal of Waterborne Enteric Viruses from Wastewater: A Comprehensive Review. J. Environ. Chem. Eng. 2021, 9, 105613. [Google Scholar] [CrossRef]
- Torii, S.; Oishi, W.; Zhu, Y.; Thakali, O.; Malla, B.; Yu, Z.; Zhao, B.; Arakawa, C.; Kitajima, M.; Hata, A. Comparison of Five Polyethylene Glycol Precipitation Procedures for the RT-QPCR Based Recovery of Murine Hepatitis Virus, Bacteriophage Phi6, and Pepper Mild Mottle Virus as a Surrogate for SARS-CoV-2 from Wastewater. Sci. Total Environ. 2022, 807, 150722. [Google Scholar] [CrossRef] [PubMed]
- Sylvestre, É.; Prévost, M.; Burnet, J.-B.; Pang, X.; Qiu, Y.; Smeets, P.; Medema, G.; Hachad, M.; Dorner, S. Demonstrating the Reduction of Enteric Viruses by Drinking Water Treatment during Snowmelt Episodes in Urban Areas. Water Res. X 2021, 11, 100091. [Google Scholar] [CrossRef] [PubMed]
- Xiao, M.; Tian, F.; Liu, X.; Zhou, Q.; Pan, J.; Luo, Z.; Yang, M.; Yi, C. Virus Detection: From State-of-the-Art Laboratories to Smartphone-Based Point-of-Care Testing. Adv. Sci. 2022, 9, 2105904. [Google Scholar] [CrossRef] [PubMed]
- Sedji, M.I.; Varbanov, M.; Meo, M.; Colin, M.; Mathieu, L.; Bertrand, I. Quantification of Human Adenovirus and Norovirus in River Water in the North-East of France. Environ. Sci. Pollut. Res. 2018, 25, 30497–30507. [Google Scholar] [CrossRef] [PubMed]
- Mannion, F.; Hillary, L.S.; Malham, S.K.; Walker, D.I. Emerging Technologies for the Rapid Detection of Enteric Viruses in the Aquatic Environment. Curr. Opin. Environ. Sci. Health 2020, 16, 1–6. [Google Scholar]
- Fuhrmeister, E.R.; Ercumen, A.; Pickering, A.J.; Jeanis, K.M.; Ahmed, M.; Brown, S.; Arnold, B.F.; Hubbard, A.E.; Alam, M.; Sen, D. Predictors of Enteric Pathogens in the Domestic Environment from Human and Animal Sources in Rural Bangladesh. Environ. Sci. Technol. 2019, 53, 10023–10033. [Google Scholar] [CrossRef] [Green Version]
- Staggemeier, R.; Bortoluzzi, M.; Heck, T.M.D.S.; Spilki, F.R.; Almeida, S.E.D.M. Quantitative vs. Conventional PCR for Detection of Human Adenoviruses in Water and Sediment Samples. Rev. Inst. Med. Trop. São Paulo 2015, 57, 299–303. [Google Scholar] [CrossRef] [Green Version]
- Paruch, L.; Paruch, A.M. An Overview of Microbial Source Tracking Using Host-Specific Genetic Markers to Identify Origins of Fecal Contamination in Different Water Environments. Water 2022, 14, 1809. [Google Scholar] [CrossRef]
- Symonds, E.M.; Young, S.; Verbyla, M.E.; McQuaig-Ulrich, S.M.; Ross, E.; Jiménez, J.A.; Harwood, V.J.; Breitbart, M. Microbial Source Tracking in Shellfish Harvesting Waters in the Gulf of Nicoya, Costa Rica. Water Res. 2017, 111, 177–184. [Google Scholar]
- Symonds, E.M.; Verbyla, M.E.; Lukasik, J.O.; Kafle, R.C.; Breitbart, M.; Mihelcic, J.R. A Case Study of Enteric Virus Removal and Insights into the Associated Risk of Water Reuse for Two Wastewater Treatment Pond Systems in Bolivia. Water Res. 2014, 65, 257–270. [Google Scholar]
- Hughes, B.; Beale, D.J.; Dennis, P.G.; Cook, S.; Ahmed, W. Cross-Comparison of Human Wastewater-Associated Molecular Markers in Relation to Fecal Indicator Bacteria and Enteric Viruses in Recreational Beach Waters. Appl. Environ. Microbiol. 2017, 83, e00028–e00117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, S.; Hata, A.; Yamashita, N.; Tanaka, H. Evaluation of Virus Reduction by Ultrafiltration with Coagulation–Sedimentation in Water Reclamation. Food Environ. Virol. 2017, 9, 453–463. [Google Scholar] [CrossRef]
- Sangsanont, J.; The Dan, D.; Thi Viet Nga, T.; Katayama, H.; Furumai, H. Detection of Pepper Mild Mottle Virus as an Indicator for Drinking Water Quality in Hanoi, Vietnam, in Large Volume of Water after Household Treatment. J. Environ. Sci. Health Part A 2016, 51, 1100–1106. [Google Scholar] [CrossRef] [PubMed]
- Asami, T.; Katayama, H.; Torrey, J.R.; Visvanathan, C.; Furumai, H. Evaluation of Virus Removal Efficiency of Coagulation-Sedimentation and Rapid Sand Filtration Processes in a Drinking Water Treatment Plant in Bangkok, Thailand. Water Res. 2016, 101, 84–94. [Google Scholar] [CrossRef] [PubMed]
- Kato, R.; Asami, T.; Utagawa, E.; Furumai, H.; Katayama, H. Pepper Mild Mottle Virus as a Process Indicator at Drinking Water Treatment Plants Employing Coagulation-Sedimentation, Rapid Sand Filtration, Ozonation, and Biological Activated Carbon Treatments in Japan. Water Res. 2018, 132, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Garcia, A.; Le, T.; Jankowski, P.; Yanaç, K.; Yuan, Q.; Uyaguari-Diaz, M.I. Quantification of Human Enteric Viruses as Alternative Indicators of Fecal Pollution to Evaluate Wastewater Treatment Processes. PeerJ 2022, 10, e12957. [Google Scholar] [CrossRef]
- Pariente, M.I.; Segura, Y.; Álvarez-Torrellas, S.; Casas, J.A.; de Pedro, Z.M.; Diaz, E.; García, J.; López-Muñoz, M.J.; Marugán, J.; Mohedano, A.F. Critical Review of Technologies for the On-Site Treatment of Hospital Wastewater: From Conventional to Combined Advanced Processes. J. Environ. Manag. 2022, 320, 115769. [Google Scholar] [CrossRef]
- Shirasaki, N.; Matsushita, T.; Matsui, Y.; Yamashita, R. Evaluation of the Suitability of a Plant Virus, Pepper Mild Mottle Virus, as a Surrogate of Human Enteric Viruses for Assessment of the Efficacy of Coagulation–Rapid Sand Filtration to Remove Those Viruses. Water Res. 2018, 129, 460–469. [Google Scholar] [CrossRef] [Green Version]
- González-Fernández, A.; Symonds, E.M.; Gallard-Gongora, J.F.; Mull, B.; Lukasik, J.O.; Navarro, P.R.; Aguilar, A.B.; Peraud, J.; Brown, M.L.; Alvarado, D.M. Relationships among Microbial Indicators of Fecal Pollution, Microbial Source Tracking Markers, and Pathogens in Costa Rican Coastal Waters. Water Res. 2021, 188, 116507. [Google Scholar] [CrossRef]
- Shrestha, S.; Shrestha, S.; Shindo, J.; Sherchand, J.B.; Haramoto, E. Virological Quality of Irrigation Water Sources and Pepper Mild Mottle Virus and Tobacco Mosaic Virus as Index of Pathogenic Virus Contamination Level. Food Environ. Virol. 2018, 10, 107–120. [Google Scholar] [CrossRef]
- Tandukar, S.; Sherchand, J.B.; Bhandari, D.; Sherchan, S.P.; Malla, B.; Ghaju Shrestha, R. Presence of Human Enteric Viruses, Protozoa, and Indicators of Pathogens in the Bagmati River, Nepal. Pathogens 2018, 7, 38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bonanno Ferraro, G.; Suffredini, E.; Mancini, P.; Veneri, C.; Iaconelli, M.; Bonadonna, L.; Montagna, M.T.; De Giglio, O.; La Rosa, G. Pepper Mild Mottle Virus as Indicator of Pollution: Assessment of Prevalence and Concentration in Different Water Environments in Italy. Food Environ. Virol. 2021, 13, 117–125. [Google Scholar] [CrossRef] [PubMed]
- Rashed, M.K.; El-Senousy, W.M.; Sayed, E.T.A.E.; AlKhazindar, M. Infectious Pepper Mild Mottle Virus and Human Adenoviruses as Viral Indices in Sewage and Water Samples. Food Environ. Virol. 2022, 14, 246–257. [Google Scholar] [CrossRef] [PubMed]
- Hata, A.; Shirasaka, Y.; Ihara, M.; Yamashita, N.; Tanaka, H. Spatial and Temporal Distributions of Enteric Viruses and Indicators in a Lake Receiving Municipal Wastewater Treatment Plant Discharge. Sci. Total Environ. 2021, 780, 146607. [Google Scholar] [CrossRef]
- Han, T.-H.; Kim, S.-C.; Kim, S.-T.; Chung, C.-H.; Chung, J.-Y. Detection of Norovirus Genogroup IV, Klassevirus, and Pepper Mild Mottle Virus in Sewage Samples in South Korea. Arch. Virol. 2014, 159, 457–463. [Google Scholar] [CrossRef]
- Gyawali, P.; Devane, M.; Scholes, P.; Hewitt, J. Application of CrAssphage, F-RNA Phage and Pepper Mild Mottle Virus as Indicators of Human Faecal and Norovirus Contamination in Shellfish. Sci. Total Environ. 2021, 783, 146848. [Google Scholar] [CrossRef]
- Rosiles-González, G.; Ávila-Torres, G.; Moreno-Valenzuela, O.A.; Acosta-González, G.; Leal-Bautista, R.M.; Grimaldo-Hernández, C.D.; Brown, J.K.; Chaidez-Quiroz, C.; Betancourt, W.Q.; Gerba, C.P. Occurrence of Pepper Mild Mottle Virus (PMMoV) in Groundwater from a Karst Aquifer System in the Yucatan Peninsula, Mexico. Food Environ. Virol. 2017, 9, 487–497. [Google Scholar] [CrossRef]
- Ahmed, W.; Bivins, A.; Bertsch, P.M.; Bibby, K.; Gyawali, P.; Sherchan, S.P.; Simpson, S.L.; Thomas, K.V.; Verhagen, R.; Kitajima, M. Intraday Variability of Indicator and Pathogenic Viruses in 1-h and 24-h Composite Wastewater Samples: Implications for Wastewater-Based Epidemiology. Environ. Res. 2021, 193, 110531. [Google Scholar] [CrossRef]
- Holm, R.H.; Nagarkar, M.; Yeager, R.A.; Talley, D.; Chaney, A.C.; Rai, J.P.; Mukherjee, A.; Rai, S.N.; Bhatnagar, A.; Smith, T. Surveillance of RNase P, PMMoV, and CrAssphage in wastewater as indicators of human fecal concentration across urban sewer neighborhoods, Kentucky. FEMS Microbes. 2022, 3, 1–12. [Google Scholar] [CrossRef]
- Adriaenssens, E.M.; Walker, D.I.; McDonald, J.E.; Malham, S.K.; Jones, D.L. Critical Evaluation of CrAssphage as a Molecular Marker for Human-Derived Wastewater Contamination in the Aquatic Environment. Food Environ. Virol. 2019, 11, 113–119. [Google Scholar]
- Elmahdy, E.M.; Shaheen, M.N.; Rizk, N.M.; Saad-Hussein, A. Quantitative Detection of Human Adenovirus and Human Rotavirus Group A in Wastewater and El-Rahawy Drainage Canal Influencing River Nile in the North of Giza, Egypt. Food Environ. Virol. 2020, 12, 218–225. [Google Scholar] [CrossRef] [PubMed]
- Hamza, I.A.; Jurzik, L.; Stang, A.; Sure, K.; Überla, K.; Wilhelm, M. Detection of Human Viruses in Rivers of a Densly-Populated Area in Germany Using a Virus Adsorption Elution Method Optimized for PCR Analyses. Water Res. 2009, 43, 2657–2668. [Google Scholar] [CrossRef] [PubMed]
- Kishida, N.; Morita, H.; Haramoto, E.; Asami, M.; Akiba, M. One-Year Weekly Survey of Noroviruses and Enteric Adenoviruses in the Tone River Water in Tokyo Metropolitan Area, Japan. Water Res. 2012, 46, 2905–2910. [Google Scholar] [CrossRef] [PubMed]
- Canh, V.D.; Torii, S.; Furumai, H.; Katayama, H. Application of Capsid Integrity (RT-) QPCR to Assessing Occurrence of Intact Viruses in Surface Water and Tap Water in Japan. Water Res. 2021, 189, 116674. [Google Scholar] [CrossRef] [PubMed]
- Tran, N.H.; Gin, K.Y.-H.; Ngo, H.H. Fecal Pollution Source Tracking Toolbox for Identification, Evaluation and Characterization of Fecal Contamination in Receiving Urban Surface Waters and Groundwater. Sci. Total Environ. 2015, 538, 38–57. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Ram, B.; Honda, R.; Poopipattana, C.; Canh, V.D.; Chaminda, T.; Furumai, H. Concurrence of Antibiotic Resistant Bacteria (ARB), Viruses, Pharmaceuticals and Personal Care Products (PPCPs) in Ambient Waters of Guwahati, India: Urban Vulnerability and Resilience Perspective. Sci. Total Environ. 2019, 693, 133640. [Google Scholar] [CrossRef] [PubMed]
- Armah, E.K.; Chetty, M.; Adedeji, J.A.; Kukwa, D.T.; Mutsvene, B.; Shabangu, K.P.; Bakare, B.F. Emerging Trends in Wastewater Treatment Technologies: The Current Perspective. In Promising Techniques for Wastewater Treatment and Water Quality Assessment; Intechopen: London, UK, 2021; Volume 1, p. 71. [Google Scholar]
- Greaves, J.; Stone, D.; Wu, Z.; Bibby, K. Persistence of Emerging Viral Fecal Indicators in Large-Scale Freshwater Mesocosms. Water Res. X 2020, 9, 100067. [Google Scholar] [CrossRef]
- Bibby, K.; Crank, K.; Greaves, J.; Li, X.; Wu, Z.; Hamza, I.A.; Stachler, E. Metagenomics and the Development of Viral Water Quality Tools. NPJ Clean Water 2019, 2, 9. [Google Scholar] [CrossRef] [Green Version]
- Calderon, J.S.; Verbyla, M.E.; Gil, M.; Pinongcos, F.; Kinoshita, A.M.; Mladenov, N. Persistence of Fecal Indicators and Microbial Source Tracking Markers in Water Flushed from Riverbank Soils. Water. Air. Soil Pollut. 2022, 233, 83. [Google Scholar] [CrossRef]
- Fong, T.-T.; Lipp, E.K. Enteric Viruses of Humans and Animals in Aquatic Environments: Health Risks, Detection, and Potential Water Quality Assessment Tools. Microbiol. Mol. Biol. Rev. 2005, 69, 357–371. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Q.; Gallard, J.; Wu, B.; Harwood, V.J.; Sadowsky, M.J.; Hamilton, K.A.; Ahmed, W. Synergy between Quantitative Microbial Source Tracking (QMST) and Quantitative Microbial Risk Assessment (QMRA): A Review and Prospectus. Environ. Int. 2019, 130, 104703. [Google Scholar] [CrossRef] [PubMed]
- Carducci, A.; Donzelli, G.; Cioni, L.; Federigi, I.; Lombardi, R.; Verani, M. Quantitative Microbial Risk Assessment for Workers Exposed to Bioaerosol in Wastewater Treatment Plants Aimed at the Choice and Setup of Safety Measures. Int. J. Environ. Res. Public. Health 2018, 15, 1490. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schijven, J.F.; Maria de Roda Husman, A. Applications of Quantitative Microbial Source Tracking (QMST) and Quantitative Microbial Risk Assessment (QMRA). In Microbial Source Tracking: Methods, Applications, and Case Studies; Springer: Berlin/Heidelberg, Germany, 2011; pp. 559–583. [Google Scholar]
- Bivins, A.; Crank, K.; Greaves, J.; North, D.; Wu, Z.; Bibby, K. Cross-Assembly Phage and Pepper Mild Mottle Virus as Viral Water Quality Monitoring Tools—Potential, Research Gaps, and Way Forward. Curr. Opin. Environ. Sci. Health 2020, 16, 54–61. [Google Scholar] [CrossRef]
- Crank, K.; Petersen, S.; Bibby, K. Quantitative Microbial Risk Assessment of Swimming in Sewage Impacted Waters Using CrAssphage and Pepper Mild Mottle Virus in a Customizable Model. Environ. Sci. Technol. Lett. 2019, 6, 571–577. [Google Scholar] [CrossRef]
- Panasiuk, O.; Hedström, A.; Marsalek, J.; Ashley, R.M.; Viklander, M. Contamination of Stormwater by Wastewater: A Review of Detection Methods. J. Environ. Manag. 2015, 152, 241–250. [Google Scholar] [CrossRef]
- World Health Organization. Strengthening Drinking-Water Surveillance Using Risk-Based Approaches; WHO: Geneve, Switzerland, 2019. [Google Scholar]
- Ahmed, W.; Payyappat, S.; Cassidy, M.; Besley, C. Enhanced Insights from Human and Animal Host-Associated Molecular Marker Genes in a Freshwater Lake Receiving Wet Weather Overflows. Sci. Rep. 2019, 9, 12503. [Google Scholar] [CrossRef]

| Selection Criteria | Applicability of PMMoV | Reason | Reference |
|---|---|---|---|
| Could be detected in all water types | Applicable | High prevalence in all types of water | [144] |
| Simple methodology of testing | Applicable | Can be examined alongside the other viral pathogens | [150] |
| Comparatively more durable than the most enteric pathogens | Applicable | Enduring compared to human enteric viruses | [199] |
| The incidence of indicator is associated with enteric viruses | Applicable | Greater frequency than the majority of enteric pathogens | [69] |
| There is a relation between the indicator prevalence and the level of fecal contamination. | Applicable | However, it can be too persistent to detect new contamination. | [152] |
| No aquatic growth | Applicable | Without its host plant, there is no replication | [201] |
| Member of the microflora of warm-blooded animals | Applicable | Highly abundant in human feces | [151] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Maniah, K.; Nour, I.; Hanif, A.; Yassin, M.T.; Alkathiri, A.; Alharbi, Y.; Alotaibi, R.; Al-Anazi, A.E.; Eifan, S. Application of the Human Viral Surrogate Pepper Mild Mottle Virus for Wastewater Fecal Pollution Management. Water 2022, 14, 4033. https://doi.org/10.3390/w14244033
Maniah K, Nour I, Hanif A, Yassin MT, Alkathiri A, Alharbi Y, Alotaibi R, Al-Anazi AE, Eifan S. Application of the Human Viral Surrogate Pepper Mild Mottle Virus for Wastewater Fecal Pollution Management. Water. 2022; 14(24):4033. https://doi.org/10.3390/w14244033
Chicago/Turabian StyleManiah, Khalid, Islam Nour, Atif Hanif, Mohamed Taha Yassin, Abdulrahman Alkathiri, Yazeed Alharbi, Riyadh Alotaibi, Abdullah E. Al-Anazi, and Saleh Eifan. 2022. "Application of the Human Viral Surrogate Pepper Mild Mottle Virus for Wastewater Fecal Pollution Management" Water 14, no. 24: 4033. https://doi.org/10.3390/w14244033
APA StyleManiah, K., Nour, I., Hanif, A., Yassin, M. T., Alkathiri, A., Alharbi, Y., Alotaibi, R., Al-Anazi, A. E., & Eifan, S. (2022). Application of the Human Viral Surrogate Pepper Mild Mottle Virus for Wastewater Fecal Pollution Management. Water, 14(24), 4033. https://doi.org/10.3390/w14244033






