Biodegradation and Decolorization of Textile Azo Dyes by Paramecium caudatum Isolated from Industrial Wastewater
Abstract
:1. Introduction
2. Materials and Methods
2.1. Dyes Used
2.2. Isolation, Purification and Maintenance of Paramecium sp.
2.3. Molecular Characterization of Paramecium sp.
2.4. Decolorization at Different Temperatures and pH
2.5. Decolorization Potential of Paramecium sp.
2.6. Growth of Paramecium sp. at Different Temperatures and pH values in the Lag Phase
2.7. Analysis of Dye Degraded Products
2.8. Estimation of Glutathione Contents
2.9. Statistical Analysis
3. Results
3.1. Paramecium Caudatum (UR03) as Efficient Dye Decolorizer
3.2. Decolorization of RR2, BB5 and LY
3.3. Optimum Temperature and pH for Maximum Growth
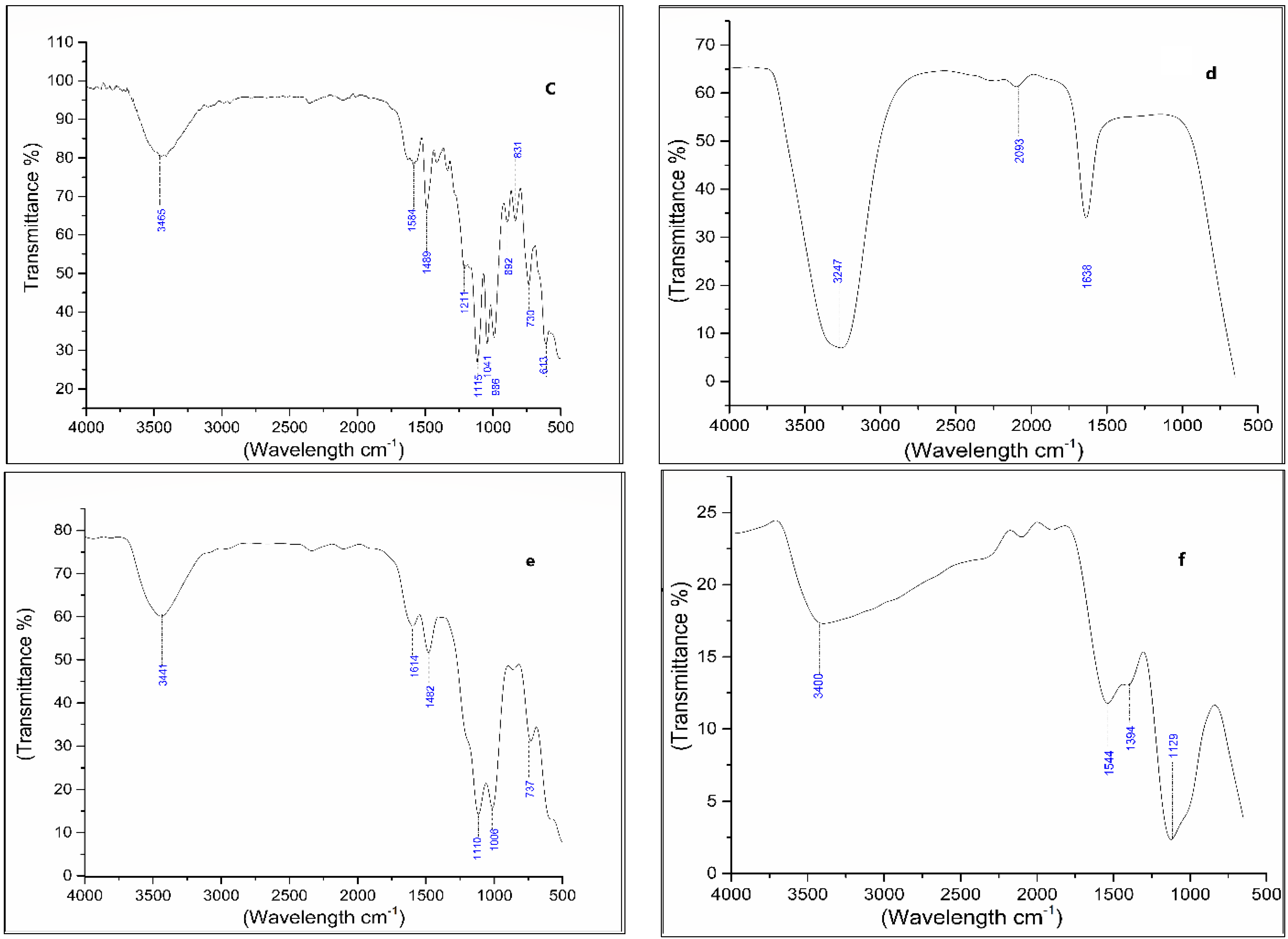
3.4. Degradation Products of Azo Dyes
3.5. Total Protein Contents
3.6. Glutathione Contents
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ramzan, U.; Shakoori, F.R.; Shakoori, A.R.; Abbas, S.Z.; Wabaidur, S.M.; Eldesoky, G.E.; Islam, M.A.; Rafatullah, M. Biodegradation and decolorization of Reactive Red 2 azo dye by Paramecium jenningsi and Paramecium multimicronucleatum in industrial wastewater. Biomass Convers. Biorefinery 2022, 1–9. [Google Scholar] [CrossRef]
- Mustafa, G.; Zahid, M.T.; Ali, S.; Abbas, S.Z.; Rafatullah, M. Biodegradation and discoloration of disperse blue-284 textile dye by Klebsiella pneumoniae GM-04 bacterial isolate. J. King Saud Univ. 2021, 33, 101442. [Google Scholar] [CrossRef]
- Prasad, A.S.A.; Satyanarayana, V.S.V.; Rao, K.V.B. Biotransformation of Direct Blue 1 by a moderately halophilic bacterium Marinobacter sp. strain HBRA and toxicity assessment of degraded metabolites. J. Hazard. Mater. 2013, 262, 674–684. [Google Scholar] [CrossRef] [PubMed]
- Adane, T.; Adugna, A.T.; Alemayehu, E. Textile Industry Effluent Treatment Techniques. J. Chem. 2021, 2021, 5314404. [Google Scholar] [CrossRef]
- Rafiq, A.; Ikram, M.; Ali, S.; Niaz, F.; Khan, M.; Khan, Q.; Maqbool, M. Photocatalytic degradation of dyes using semiconductor photocatalysts to clean industrial water pollution. J. Ind. Eng. Chem. 2021, 97, 111–128. [Google Scholar] [CrossRef]
- Mani, A.; Hameed, S.A.S. Improved bacterial-fungal consortium as an alternative approach for enhanced decolourisation and degradation of azo dyes: A review. Nat. Environ. Pollut. Technol. 2019, 18, 49–64. [Google Scholar]
- Anjaneya, O.; Shrishailnath, S.S.; Guruprasad, K.; Nayak, A.S.; Mashetty, S.B.; Karegoudar, T.B. Decolourization of Amaranth dye by bacterial biofilm in batch and continuous packed bed bioreactor. Int. Biodeterior. Biodegrad. 2013, 79, 64–72. [Google Scholar] [CrossRef]
- Mnif, I.; Maktouf, S.; Fendri, R.; Kriaa, M.; Ellouze, S.; Ghribi, D. Improvement of methyl orange dye biotreatment by a novel isolated strain, Aeromonas veronii GRI, by SPB1 biosurfactant addition. Environ. Sci. Pollut. Res. 2016, 23, 1742–1754. [Google Scholar] [CrossRef]
- Balapure, K.; Bhatt, N.; Madamwar, D. Mineralization of reactive azo dyes present in simulated textile waste water using down flow microaerophilic fixed film bioreactor. Bioresour. Technol. 2015, 175, 1–7. [Google Scholar] [CrossRef]
- Chen, B.-Y.; Lin, K.-W.; Wang, Y.-M.; Yen, C.-Y. Revealing interactive toxicity of aromatic amines to azo dye decolorizer Aeromonas hydrophila. J. Hazard. Mater. 2009, 166, 187–194. [Google Scholar] [CrossRef]
- Manatunga, D.C.; de Silva, R.M.; Nalin de Silva, K.M.; de Silva, N.; Premalal, E.V.A. Metal and polymer-mediated synthesis of porous crystalline hydroxyapatite nanocomposites for environmental remediation. R. Soc. Open Sci. 2018, 5, 171557. [Google Scholar] [CrossRef]
- Soni, P.; Sharma, S.; Sharma, S.; Kumar, S.; Sharma, K.P. A comparative study on the toxic effects of textile dye wastewaters(untreated and treated) on mortality and RBC of a freshwater fish Gambusia affinis(Baird and Gerard). J. Environ. Biol. 2006, 27, 623–628. [Google Scholar]
- Zaharia, C.; Suteu, D.; Muresan, A.; Muresan, R.; Popescu, A. Textile wastewater treatment by homogenous oxidation with hydrogen peroxide. Environ. Eng. Manag. J. 2009, 8, 1359–1369. [Google Scholar] [CrossRef]
- Deniz, F. Color removal from aqueous solutions of metal-containing dye using pine cone. Desalin. Water Treat. 2013, 51, 4573–4581. [Google Scholar] [CrossRef]
- Thanavel, M.; Kadam, S.K.; Biradar, S.P.; Govindwar, S.P.; Jeon, B.H.; Sadasivam, S.K. Combined biological and advanced oxidation process for decolorization of textile dyes. SN Appl. Sci. 2019, 1, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Bafana, A.; Devi, S.S.; Chakrabarti, T. Azo dyes: Past, present and the future. Environ. Rev. 2011, 19, 350–371. [Google Scholar] [CrossRef]
- Shah, K. Biodegradation of azo dye compounds. Int. Res. J. Biochem. Biotechnol. 2014, 1, 5–13. [Google Scholar]
- Cao, J.; Sanganyado, E.; Liu, W.; Zhang, W.; Liu, Y. Decolorization and detoxification of Direct Blue 2B by indigenous bacterial consortium. J. Environ. Manag. 2019, 242, 229–237. [Google Scholar] [CrossRef]
- Ebrahimi, R.; Maleki, A.; Zandsalimi, Y.; Ghanbari, R.; Shahmoradi, B.; Rezaee, R.; Safari, M.; Joo, S.W.; Daraei, H.; Puttaiah, S.H. Photocatalytic degradation of organic dyes using WO3-doped ZnO nanoparticles fixed on a glass surface in aqueous solution. J. Ind. Eng. Chem. 2019, 73, 297–305. [Google Scholar] [CrossRef]
- Ajaz, M.; Shakeel, S.; Rehman, A. Microbial use for azo dye degradation—A strategy for dye bioremediation. Int. Microbiol. 2020, 23, 149–159. [Google Scholar] [CrossRef]
- Dong, H.; Guo, T.; Zhang, W.; Ying, H.; Wang, P.; Wang, Y.; Chen, Y. Biochemical characterization of a novel azoreductase from Streptomyces sp.: Application in eco-friendly decolorization of azo dye wastewater. Int. J. Biol. Macromol. 2019, 140, 1037–1046. [Google Scholar] [CrossRef]
- Ikram, M.; Naeem, M.; Zahoor, M.; Rahim, A.; Hanafiah, M.M.; Oyekanmi, A.A.; Shah, A.B.; Mahnashi, M.H.; Al Ali, A.; Jalal, N.A. Biodegradation of Azo Dye Methyl Red by Pseudomonas aeruginosa: Optimization of Process Conditions. Int. J. Environ. Res. Public Health 2022, 19, 9962. [Google Scholar] [CrossRef]
- Selvaraj, V.; Karthika, T.S.; Mansiya, C.; Alagar, M. An over review on recently developed techniques, mechanisms and intermediate involved in the advanced azo dye degradation for industrial applications. J. Mol. Struct. 2021, 1224, 129195. [Google Scholar] [CrossRef]
- Pandey, A.; Singh, P.; Iyengar, L. Bacterial decolorization and degradation of azo dyes. Int. Biodeterior. Biodegrad. 2007, 59, 73–84. [Google Scholar] [CrossRef]
- Chang, J.-S.; Chen, B.-Y.; Lin, Y.S. Stimulation of bacterial decolorization of an azo dye by extracellular metabolites from Escherichia coli strain NO3. Bioresour. Technol. 2004, 91, 243–248. [Google Scholar] [CrossRef]
- Naresh, B.; Jaydip, J.; Prabhat, B.; Rajkumar, P. Recent biological technologies for textile effluent treatment. Int. Res. J. Biol. Sci 2013, 2, 77–82. [Google Scholar]
- Davies, L.C.; Pedro, I.S.; Novais, J.M.; Martins-Dias, S. Aerobic degradation of acid orange 7 in a vertical-flow constructed wetland. Water Res. 2006, 40, 2055–2063. [Google Scholar] [CrossRef]
- Kamika, I.; Momba, M.N.B. Comparing the tolerance limits of selected bacterial and protozoan species to nickel in wastewater systems. Sci. Total Environ. 2011, 410, 172–181. [Google Scholar] [CrossRef]
- Singh, D.; Goswami, R.K.; Agrawal, K.; Chaturvedi, V.; Verma, P. Bio-inspired remediation of wastewater: A contemporary approach for environmental clean-up. Curr. Res. Green Sustain. Chem. 2022, 5, 100261. [Google Scholar] [CrossRef]
- Gomiero, A.; Dagnino, A.; Nasci, C.; Viarengo, A. The use of protozoa in ecotoxicology: Application of multiple endpoint tests of the ciliate E. crassus for the evaluation of sediment quality in coastal marine ecosystems. Sci. Total Environ. 2013, 442, 534–544. [Google Scholar] [CrossRef]
- Shakoori, A.R.; Rehman, A. Multiple metal resistance in the ciliate protozoan, Vorticella microstoma, isolated from industrial effluents and its potential in bioremediation of toxic wastes. Bull. Environ. Contam. Toxicol. 2004, 72, 1046–1051. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: New York, NY, USA, 1989; ISBN 0879693096. [Google Scholar]
- Regensbogenova, M.; Kisidayova, S.; Michalowski, T.; Javorsky, P.; Moon-Van Der Staay, S.Y.; Moon-Van Der Staay, G.W.M.; Hackstein, J.H.P.; McEwan, N.R.; Jouany, J.P.; Newbold, J.C.; et al. Rapid identification of rumen protozoa by restriction analysis of amplified 18S rRNA gene. Acta Protozool. 2004, 43, 219–224. [Google Scholar]
- Thanavel, M.; Bankole, P.O.; Kadam, S.; Govindwar, S.P.; Sadasivam, S.K. Desulfonation of the textile azo dye Acid Fast Yellow MR by newly isolated Aeromonas hydrophila SK16. Water Resour. Ind. 2019, 22, 100116. [Google Scholar] [CrossRef]
- Kalyani, D.C.; Telke, A.A.; Dhanve, R.S.; Jadhav, J.P. Ecofriendly biodegradation and detoxification of Reactive Red 2 textile dye by newly isolated Pseudomonas sp. SUK1. J. Hazard. Mater. 2009, 163, 735–742. [Google Scholar] [CrossRef] [PubMed]
- Phugare, S.S.; Kalyani, D.C.; Surwase, S.N.; Jadhav, J.P. Ecofriendly degradation, decolorization and detoxification of textile effluent by a developed bacterial consortium. Ecotoxicol. Environ. Saf. 2011, 74, 1288–1296. [Google Scholar] [CrossRef]
- Bradford, M.M. A Rapid and Sensitive Method for the Quantitation Microgram Quantities of Protein Utilizing the Principle of Protein-Dye Binding. Anal Biochem. 1976, 254, 248–254. [Google Scholar] [CrossRef]
- Israr, M.; Sahi, S.V.; Jain, J. Cadmium accumulation and antioxidative responses in the Sesbania drummondii callus. Arch. Environ. Contam. Toxicol. 2006, 50, 121–127. [Google Scholar] [CrossRef]
- Yaseen, D.A.; Scholz, M. Textile dye wastewater characteristics and constituents of synthetic effluents: A critical review. Int. J. Environ. Sci. Technol. 2019, 16, 1193–1226. [Google Scholar] [CrossRef] [Green Version]
- Mani, S.; Chowdhary, P.; Bharagava, R.N. Textile wastewater dyes: Toxicity profile and treatment approaches. In Emerging and Eco-Friendly Approaches for Waste Management; Springer: Berlin/Heidelberg, Germany, 2019; pp. 219–244. [Google Scholar]
- Jamee, R.; Siddique, R. Biodegradation of synthetic dyes of textile effluent by microorganisms: An environmentally and economically sustainable approach. Eur. J. Microbiol. Immunol. 2019, 9, 114–118. [Google Scholar] [CrossRef]
- Djekoun, M.; Berrebbah, H.; Djebar, M.R. In Vivo Cytotoxicity Assessment of Thiram: Physiological and Biochemical Changes in Paramecium sp. Int. J. Sci. Res. Sci. Eng. Technol. 2015, 1, 320–326. [Google Scholar]
- Rehman, A.; Shakoori, F.R.; Shakoori, A.R. Multiple heavy metal tolerant ciliates, Oxytricha fallax and Paramecium caudatum, isolated from industrial effluents and their potential use in wastewater treatment. Pak. J. Zool. 2010, 42, 301–309. [Google Scholar]
- Ponraj, M.; Gokila, K.; Zambare, V. Bacterial decolorization of textile dye-Orange 3R. Int. J. Adv. Biotechnol. Res. 2011, 2, 168–177. [Google Scholar]
- Maas, R.; Chaudhari, S. Adsorption and biological decolourization of azo dye Reactive Red 2 in semicontinuous anaerobic reactors. Process Biochem. 2005, 40, 699–705. [Google Scholar] [CrossRef] [Green Version]
- Zahid, M.T.; Shakoori, F.R.; Zulfiqar, S.; Al-Ghanim, K.A.; Shakoori, A.R. Growth characteristics, metal uptake and expression analysis of copper metallothionein in a newly reported ciliate, Tetrahymena farahensis. Pak. J. Zool 2018, 50, 1171–1181. [Google Scholar] [CrossRef]
- Dopheide, A.; Lear, G.; Stott, R.; Lewis, G. Preferential feeding by the ciliates Chilodonella and Tetrahymena spp. and effects of these protozoa on bacterial biofilm structure and composition. Appl. Environ. Microbiol. 2011, 77, 4564–4572. [Google Scholar] [CrossRef] [Green Version]
- Khan, Z.; Jain, K.; Soni, A.; Madamwar, D. Microaerophilic degradation of sulphonated azo dye - Reactive Red 195 by bacterial consortium AR1 through co-metabolism. Int. Biodeterior. Biodegrad. 2014, 94, 167–175. [Google Scholar] [CrossRef]
- Amamra, R.; Djebar, M.R.; Grara, N.; Moumeni, O.; Otmani, H.; Alayat, A.; Berrebbah, H. Cypermethrin-induces oxidative stress to the freshwater Ciliate model: Paramecium tetraurelia. Annu. Res. Rev. Biol. 2015, 385–399. [Google Scholar] [CrossRef]
- Anzenbacher, P.; Anzenbacherova, E. Cytochromes P450 and metabolism of xenobiotics. Cell. Mol. Life Sci. C. 2001, 58, 737–747. [Google Scholar] [CrossRef]
- Benlaifa, M.; Djebar, M.R.; Berredjem, H.; Benamara, M.; Ouali, K.; Djebar, H. Stress induced by cadmium: Its effects on growth respiratory metabolism, antioxidant enzymes and reactive oxygen species (ROS) of Paramecium sp. Int. J. Pharm. Sci. Rev. Res. 2016, 38, 276–281. [Google Scholar]
- Benbouzid, H.; Berrebbah, H.; Djebar, M.R. Toxicity of the chlorfenapyr: Growth inhibition and induction of oxidative stress on a freshwater protozoan: Paramecium sp. Adv. Environ. Biol. 2015, 9, 281–285. [Google Scholar]
- Giustarini, D.; Colombo, G.; Garavaglia, M.L.; Astori, E.; Portinaro, N.M.; Reggiani, F.; Badalamenti, S.; Aloisi, A.M.; Santucci, A.; Rossi, R. Assessment of glutathione/glutathione disulphide ratio and S-glutathionylated proteins in human blood, solid tissues, and cultured cells. Free Radic. Biol. Med. 2017, 112, 360–375. [Google Scholar] [CrossRef] [PubMed]
- Musaogullari, A.; Chai, Y.-C. Redox regulation by protein S-glutathionylation: From molecular mechanisms to implications in health and disease. Int. J. Mol. Sci. 2020, 21, 8113. [Google Scholar] [CrossRef] [PubMed]
- Bjørklund, G.; Tinkov, A.A.; Hosnedlová, B.; Kizek, R.; Ajsuvakova, O.P.; Chirumbolo, S.; Skalnaya, M.G.; Peana, M.; Dadar, M.; El-Ansary, A. The role of glutathione redox imbalance in autism spectrum disorder: A review. Free Radic. Biol. Med. 2020, 160, 149–162. [Google Scholar] [CrossRef] [PubMed]








| Isolate | RR2 (%) | RB5 (%) | LY (%) |
|---|---|---|---|
| UR01 | 70.88 | 70.00 | 63.55 |
| UR02 | 80.00 | 75.02 | 70.00 |
| UR03 | 90.86 | 83.06 | 85.43 |
| UR04 | 80.00 | 60.08 | 74.00 |
| UR05 | 40.22 | 51.00 | 50.00 |
| Variable | Total Protein Content (ug/mL) |
|---|---|
| Control | 25.363 ± 0.27 a |
| RR2 | 45.790 ± 0.15 |
| RB5 | 60.837 ± 0.56 |
| LY | 36.073 ± 0.21 |
| Groups | GSH (mM g−1) | GSSG (mM g−1) | GSH + GSSG (mM g−1) | GSH/GSSG Ratio |
|---|---|---|---|---|
| Control | 2.898 ± 0.04 a | 0.764 ± 0.04 a | 3.662 ± 0.01 a | 3.818 ± 0.25 a |
| RR2 | 4.463 ± 0.04 | 2.737 ± 0.02 | 7.200 ± 0.02 | 1.631 ± 0.03 |
| RB5 | 4.155 ± 0.02 | 1.421 ± 0.03 | 5.576 ± 0.01 | 2.926 ± 0.08 |
| LY | 3.575 ± 0.02 | 0.562 ± 0.02 | 4.136 ± 0.01 | 6.377 ± 0.21 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramzan, U.; Shakoori, F.R.; Zahid, M.T.; Majeed, W.; Zahra, I.; Abbas, S.Z.; Hedfi, A.; Hassan, S.; Shakoori, A.R.; Mutery, A.A. Biodegradation and Decolorization of Textile Azo Dyes by Paramecium caudatum Isolated from Industrial Wastewater. Water 2022, 14, 3553. https://doi.org/10.3390/w14213553
Ramzan U, Shakoori FR, Zahid MT, Majeed W, Zahra I, Abbas SZ, Hedfi A, Hassan S, Shakoori AR, Mutery AA. Biodegradation and Decolorization of Textile Azo Dyes by Paramecium caudatum Isolated from Industrial Wastewater. Water. 2022; 14(21):3553. https://doi.org/10.3390/w14213553
Chicago/Turabian StyleRamzan, Uzma, Farah Rauf Shakoori, Muhammad Tariq Zahid, Waqar Majeed, Itrat Zahra, Syed Zaghum Abbas, Amor Hedfi, Sabry Hassan, Abdul Rauf Shakoori, and Abdullah Al Mutery. 2022. "Biodegradation and Decolorization of Textile Azo Dyes by Paramecium caudatum Isolated from Industrial Wastewater" Water 14, no. 21: 3553. https://doi.org/10.3390/w14213553
APA StyleRamzan, U., Shakoori, F. R., Zahid, M. T., Majeed, W., Zahra, I., Abbas, S. Z., Hedfi, A., Hassan, S., Shakoori, A. R., & Mutery, A. A. (2022). Biodegradation and Decolorization of Textile Azo Dyes by Paramecium caudatum Isolated from Industrial Wastewater. Water, 14(21), 3553. https://doi.org/10.3390/w14213553










