Transcriptome Analysis of the Toxic Effects of Amisulbrom and Isoflucypram on Zebrafish (Danio rerio) Larvae
Abstract
:1. Introduction
2. Materials and Methods
2.1. Zebrafish Husbandry
2.2. Exposure Experiments
2.3. RNA Isolation and Sequencing
2.4. Bioinformatic Analyses
2.5. qRT-PCR Verification
2.6. Statistical Analyses
3. Results
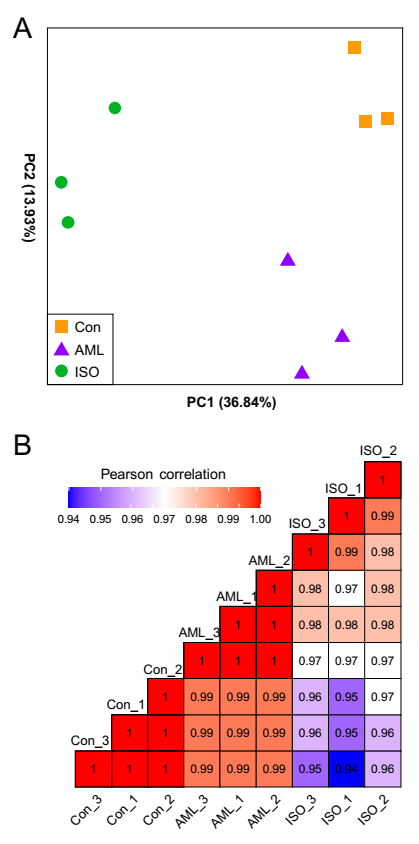
3.1. Sequencing and Read Mapping
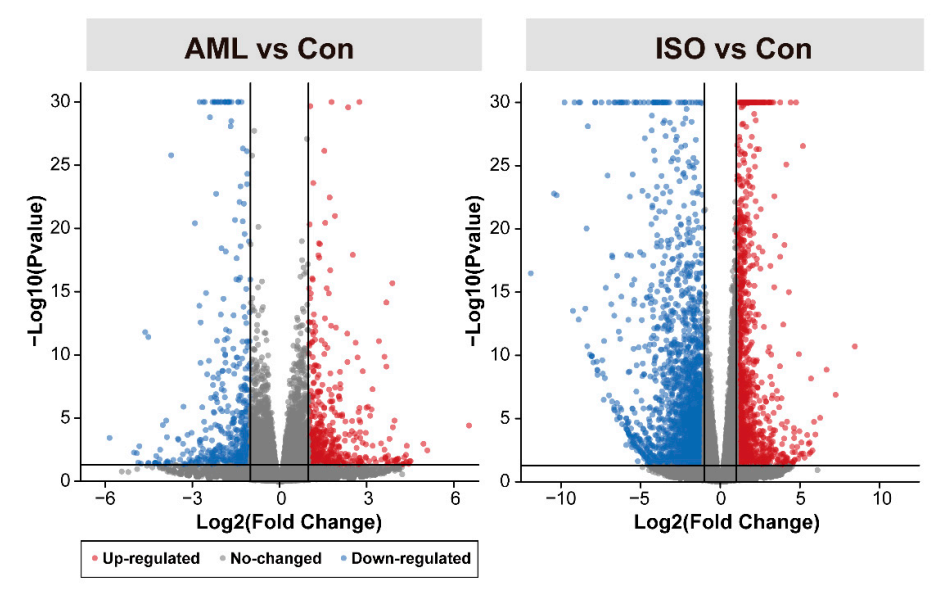
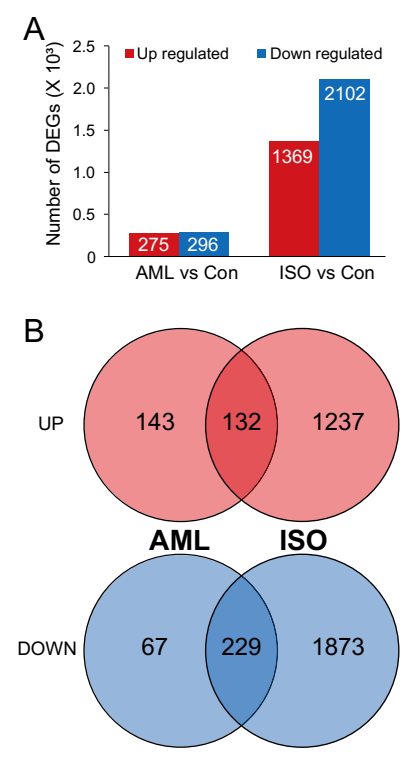
3.2. Identification of Differentially Expressed Genes (DEGs)
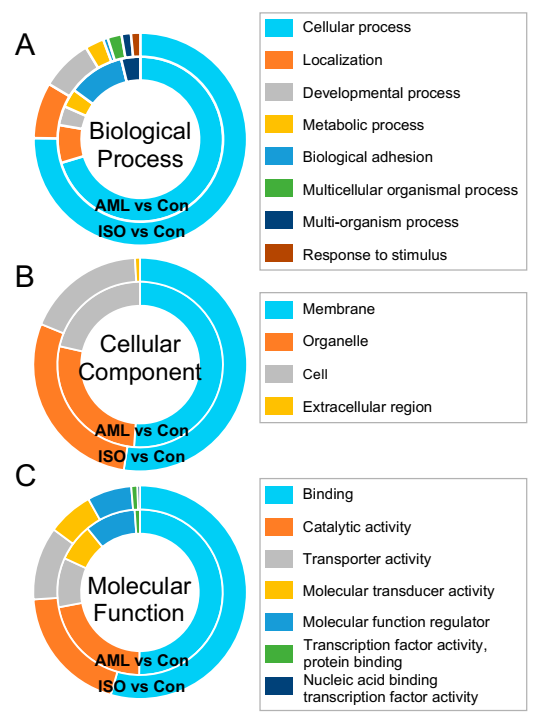
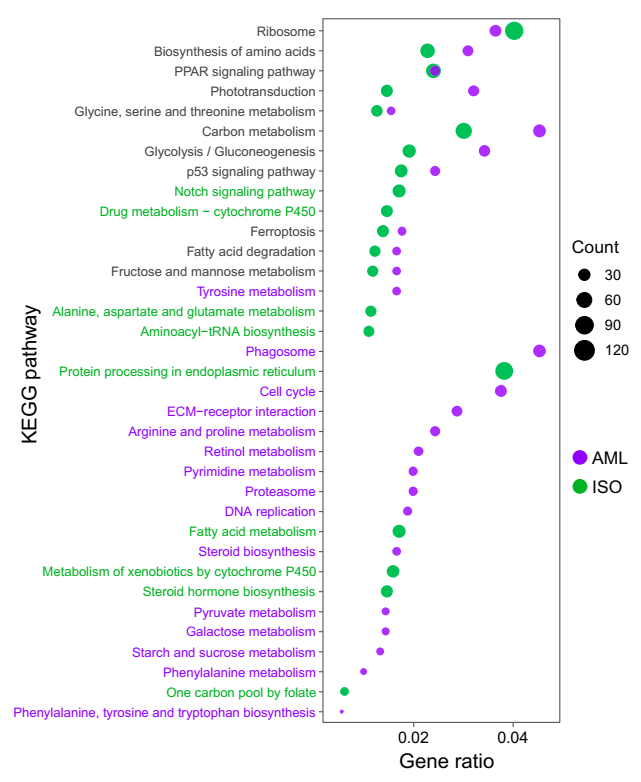
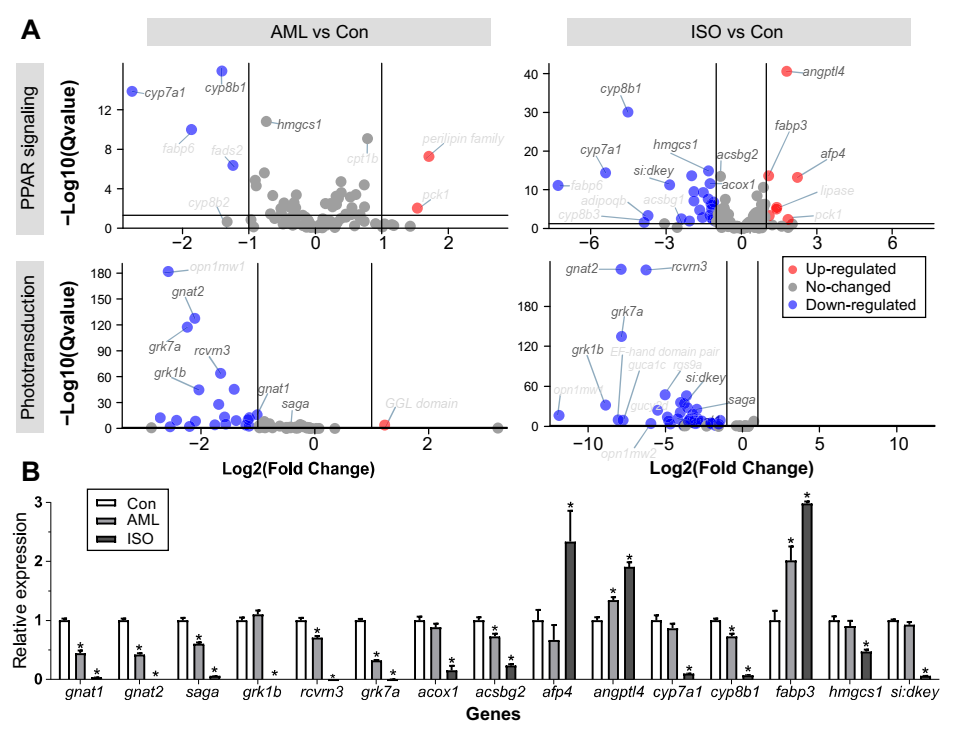
3.3. Enrichment Analysis
3.4. PCR Verification
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- USEPA. U.S. Pesticide Fact Sheet Number 016330: Amisulbrom; Office of Chemical Safety and Pollution Prevention, United States Environmental Protection Agency: Boston, MA, USA, 2011. [Google Scholar]
- Pedersen, M.; Wegner, C.; Phansak, P.; Sarath, G.; Gaussoin, R.; Schlegel, V. Monitoring wheat mitochondrial compositional and respiratory changes using Fourier transform mid-infrared spectroscopy in response to agrochemical treatments. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2017, 173, 727–732. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Desbordes, P.; Essigmann, B.; Gary, S.; Gutbrod, O.; Maue, M.; Schwarz, H.G. Isoflucypram, the first representative of a new succinate dehydrogenase inhibitor fungicide subclass: Its chemical discovery and unusual binding mode. Pest Manag. Sci. 2020, 76, 3340–3347. [Google Scholar] [CrossRef] [PubMed]
- Fontaine, S.; Remuson, F.; Caddoux, L.; Barres, B. Investigation of the sensitivity of Plasmopara viticola to amisulbrom and ametoctradin in French vineyards using bioassays and molecular tools. Pest. Manag. Sci. 2019, 75, 2115–2123. [Google Scholar] [PubMed]
- Lewis, K.A.; Tzilivakis, J.; Warner, D.; Green, A. An international database for pesticide risk assessments and management. Hum. Ecol. Risk Assess. 2016, 22, 1050–1064. [Google Scholar] [CrossRef] [Green Version]
- EFSA. Conclusion on the peer review of the pesticide risk assessment of the active substance amisulbrom. EFSA J. 2014, 12, 3237. [Google Scholar]
- Chen, X.; Li, W. Isoflucypram cardiovascular toxicity in zebrafish (Danio rerio). Sci. Total Environ. 2021, 787, 147529. [Google Scholar] [CrossRef]
- Ma, X.; Li, W. Amisulbrom causes cardiovascular toxicity in zebrafish (Danio rerio). Chemosphere 2021, 283, 131236. [Google Scholar] [CrossRef]
- Hrdlickova, R.; Toloue, M.; Tian, B. RNA-Seq methods for transcriptome analysis. Wiley Interdiscip. Rev. RNA 2017, 8, e1364. [Google Scholar] [CrossRef] [Green Version]
- Hsu, L.S.; Chiou, B.H.; Hsu, T.W.; Wang, C.C.; Chen, S.C. The regulation of transcriptome responses in zebrafish embryo exposure to triadimefon. Environ. Toxicol. 2017, 32, 217–226. [Google Scholar] [CrossRef]
- Li, W.; Yuan, M.; Wu, Y.; Liu, X. Bixafen exposure induces developmental toxicity in zebrafish (Danio rerio) embryos. Environ. Res. 2020, 189, 109923. [Google Scholar] [CrossRef]
- Jiang, J.; Wu, S.; Lv, L.; Liu, X.; Chen, L.; Zhao, X.; Wang, Q. Mitochondrial dysfunction, apoptosis and transcriptomic alterations induced by four strobilurins in zebrafish (Danio rerio) early life stages. Environ. Pollut. 2019, 253, 722–730. [Google Scholar] [CrossRef]
- Valadas, J.; Mocelin, R.; Sachett, A.; Marcon, M.; Zanette, R.A.; Dallegrave, E.; Herrmann, A.P.; Piato, A. Propiconazole induces abnormal behavior and oxidative stress in zebrafish. Environ. Sci. Pollut. Res. Int. 2019, 26, 27808–27815. [Google Scholar] [CrossRef]
- Bevilaqua, F.; Sachett, A.; Chitolina, R.; Garbinato, C.; Gasparetto, H.; Marcon, M.; Mocelin, R.; Dallegrave, E.; Conterato, G.; Piato, A.; et al. A mixture of fipronil and fungicides induces alterations on behavioral and oxidative stress parameters in zebrafish. Ecotoxicology 2020, 29, 140–147. [Google Scholar] [CrossRef]
- Kumar, N.; Willis, A.; Satbhai, K.; Ramalingam, L.; Schmitt, C.; Moustaid-Moussa, N.; Crago, J. Developmental toxicity in embryo-larval zebrafish (Danio rerio) exposed to strobilurin fungicides (azoxystrobin and pyraclostrobin). Chemosphere 2020, 241, 124980. [Google Scholar] [CrossRef]
- Westerfield, M. The Zebrafish Book: A Guide for the Laboratory Use of Zebrafish (Danio rerio), 5th ed.; University of Oregon Press: Eugene, OR, USA, 2007. [Google Scholar]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef] [Green Version]
- Kovaka, S.; Zimin, A.V.; Pertea, G.M.; Razaghi, R.; Salzberg, S.L.; Pertea, M. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 2019, 20, 278. [Google Scholar] [CrossRef] [Green Version]
- Varet, H.; Brillet-Guéguen, L.; Coppée, J.Y.; Dillies, M.A. SARTools: A DESeq2- and EdgeR-Based R pipeline for comprehensive differential analysis of RNA-Seq data. PLoS ONE 2016, 11, e0157022. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [Green Version]
- Jung, J.H.; Moon, Y.S.; Kim, B.M.; Lee, Y.M.; Kim, M.; Rhee, J.S. Comparative analysis of distinctive transcriptome profiles with biochemical evidence in bisphenol S- and benzo[a]pyrene-exposed liver tissues of the olive flounder Paralichthys olivaceus. PLoS ONE 2018, 13, e0196425. [Google Scholar] [CrossRef]
- Sonnack, L.; Klawonn, T.; Kriehuber, R.; Hollert, H.; Schäfers, C.; Fenske, M. Comparative analysis of the transcriptome responses of zebrafish embryos after exposure to low concentrations of cadmium, cobalt and copper. Comp. Biochem. Physiol. Part D Genom. Proteom. 2018, 25, 99–108. [Google Scholar] [CrossRef]
- Christou, M.; Fraser, T.W.K.; Berg, V.; Ropstad, E.; Kamstra, J.H. Calcium signaling as a possible mechanism behind increased locomotor response in zebrafish larvae exposed to a human relevant persistent organic pollutant mixture or PFOS. Environ. Res. 2020, 187, 109702. [Google Scholar] [CrossRef]
- Kvandová, M.; Majzúnová, M.; Dovinová, I. The role of PPARgamma in cardiovascular diseases. Physiol. Res. 2016, 65, S343–S363. [Google Scholar] [CrossRef]
- Li, J.; Tripathi, R.C.; Tripathi, B.J. Drug-induced ocular disorders. Drug Saf. 2008, 31, 127–141. [Google Scholar] [CrossRef]
- Wang, H.; Meng, Z.; Liu, F.; Zhou, L.; Su, M.; Meng, Y.; Zhang, S.; Liao, X.; Cao, Z.; Lu, H. Characterization of boscalid-induced oxidative stress and neurodevelopmental toxicity in zebrafish embryos. Chemosphere 2020, 238, 124753. [Google Scholar] [CrossRef]
- Qian, L.; Qi, S.; Wang, Z.; Magnuson, J.T.; Volz, D.C.; Schlenk, D.; Jiang, J.; Wang, C. Environmentally relevant concentrations of boscalid exposure affects the neurobehavioral response of zebrafish by disrupting visual and nervous systems. J. Hazard. Mater. 2021, 404, 124083. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xiao, P.; Li, W.; Lu, J.; Zhang, H. Transcriptome Analysis of the Toxic Effects of Amisulbrom and Isoflucypram on Zebrafish (Danio rerio) Larvae. Water 2022, 14, 272. https://doi.org/10.3390/w14020272
Xiao P, Li W, Lu J, Zhang H. Transcriptome Analysis of the Toxic Effects of Amisulbrom and Isoflucypram on Zebrafish (Danio rerio) Larvae. Water. 2022; 14(2):272. https://doi.org/10.3390/w14020272
Chicago/Turabian StyleXiao, Peng, Wenhua Li, Jinfang Lu, and He Zhang. 2022. "Transcriptome Analysis of the Toxic Effects of Amisulbrom and Isoflucypram on Zebrafish (Danio rerio) Larvae" Water 14, no. 2: 272. https://doi.org/10.3390/w14020272






