Pathogenesis of the Candida parapsilosis Complex in the Model Host Caenorhabditis elegans
Abstract
1. Introduction
2. Materials and Methods
2.1. Strains and Media
2.2. Caenorhabditis elegans Liquid Medium Killing Assays
2.3. Antifungal Drug Treatments
2.4. Microscopic Studies
2.5. Quantitative RT-PCR Analyses of Candida parapsilosis Infected Nematodes
2.6. Statistics
3. Results
3.1. Killing Caenorhabditis elegans by Candida parapsilosis Species Complex
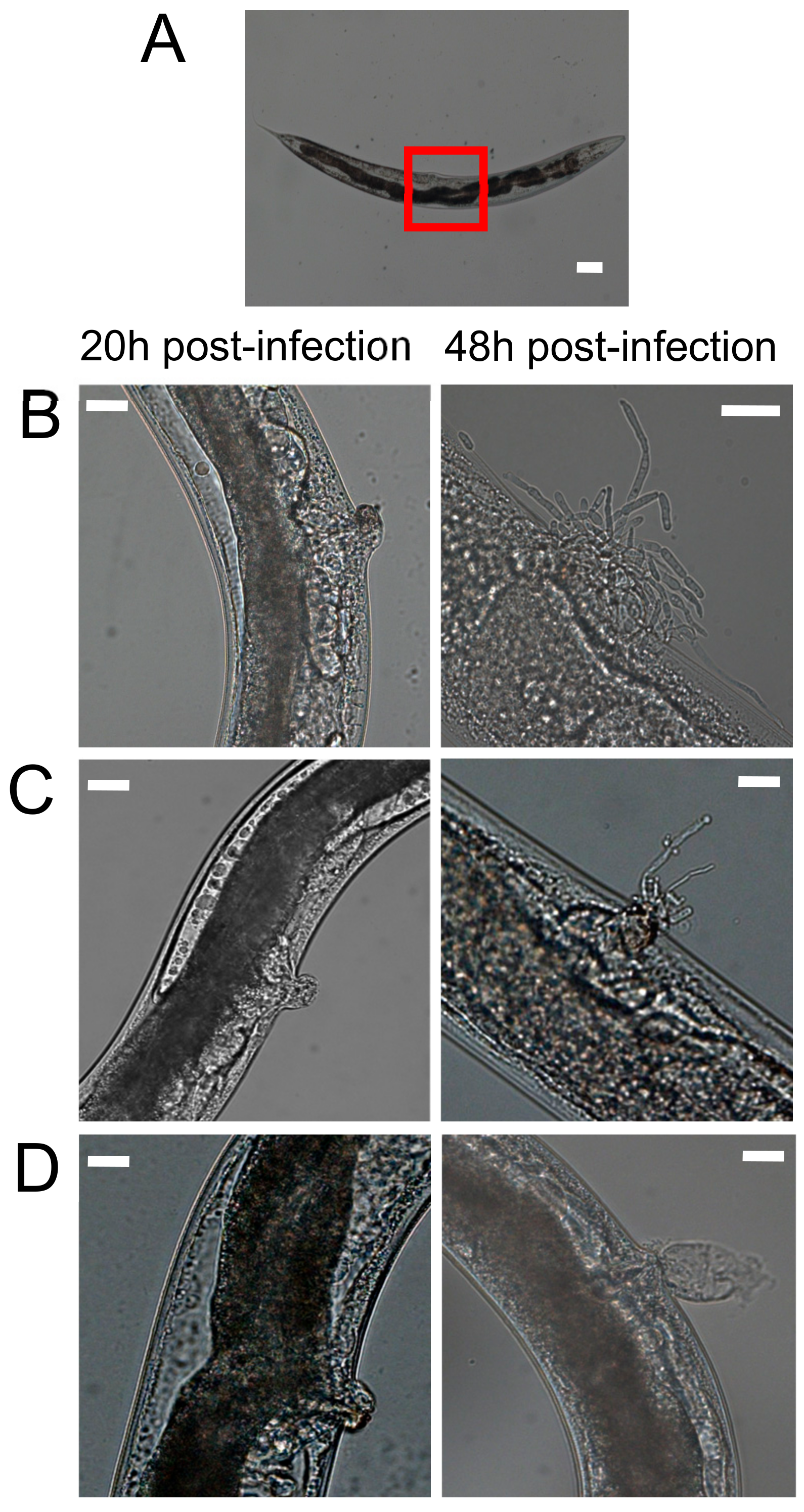
3.2. Hyphal Formation of Candida parapsilosis Species Complex within Caenorhabditis elegans
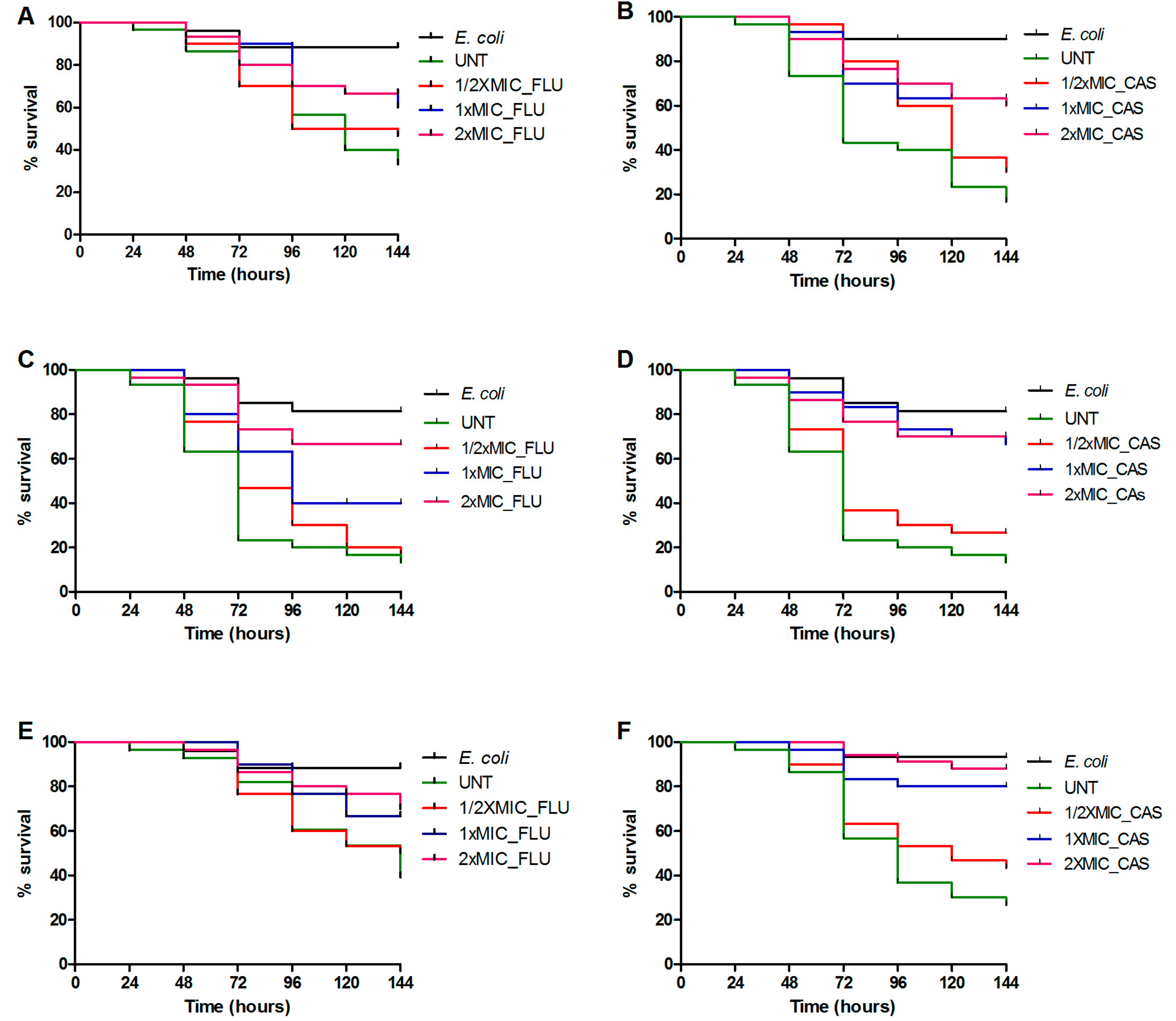
3.3. Treatment with Antifungal Drugs
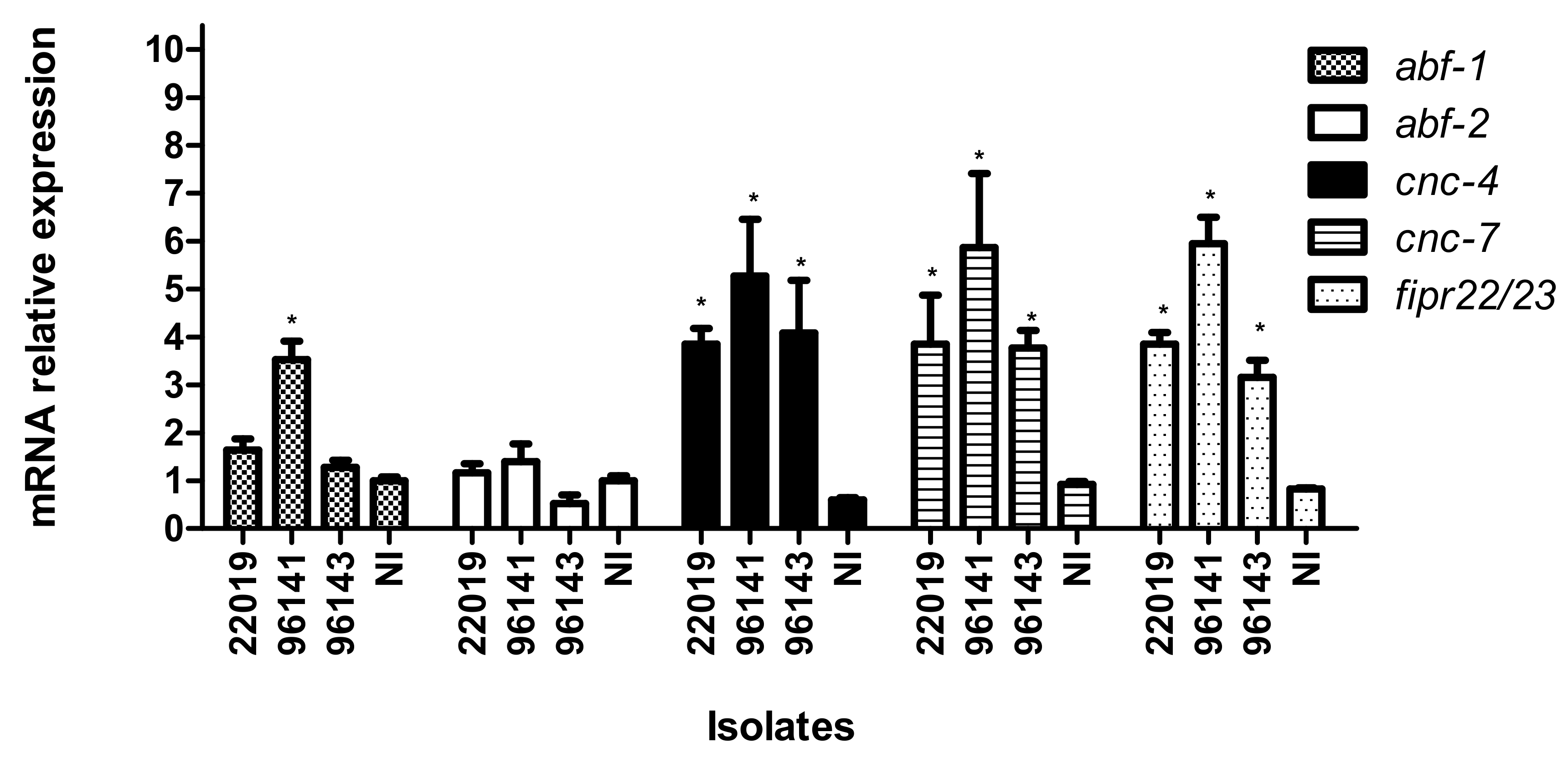
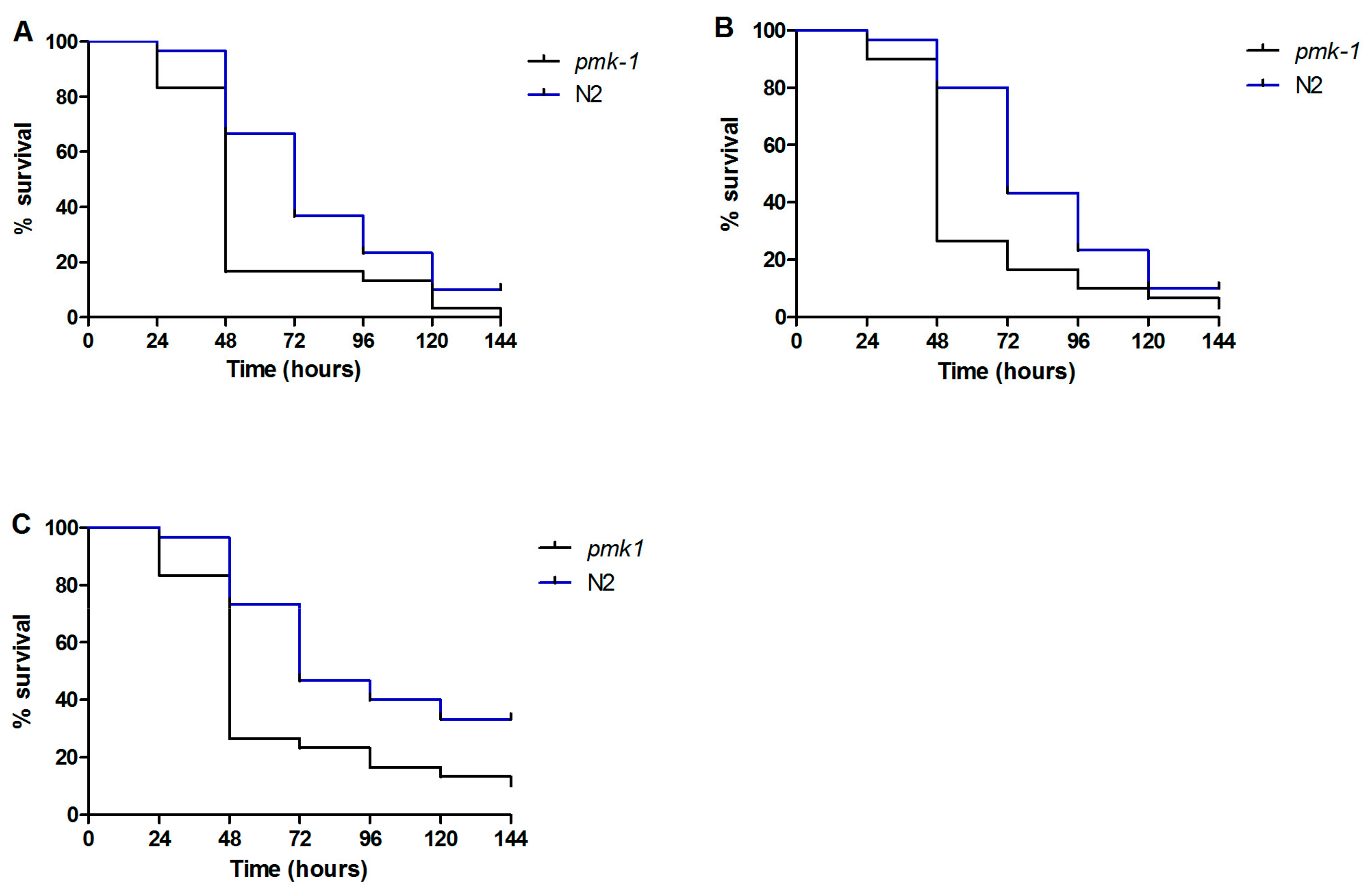
3.4. Caenorhabditis elegans Immune Response to Candida parapsilosis Complex Infection
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Pammi, M.; Holland, L.; Butler, G.; Gacser, A.; Bliss, J.M. Candida parapsilosis is a significant neonatal pathogen: A systematic review and meta-analysis. Pediatr. Infect. Dis. J. 2013, 32, e206–e216. [Google Scholar] [CrossRef] [PubMed]
- Trofa, D.; Gacser, A.; Nosanchuk, J.D. Candida parapsilosis, an emerging fungal pathogen. Clin. Microbiol. Rev. 2008, 21, 606–625. [Google Scholar] [CrossRef] [PubMed]
- Quindos, G. Epidemiology of candidaemia and invasive candidiasis. A changing face. Rev. Iberoam. Micol. 2014, 31, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Tavanti, A.; Davidson, A.D.; Gow, N.A.; Maiden, M.C.; Odds, F.C. Candida orthopsilosis and Candida metapsilosis spp. Nov. to replace Candida parapsilosis groups II and III. J. Clin. Microbiol. 2005, 43, 284–292. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Lopez, A.; Alastruey-Izquierdo, A.; Rodriguez, D.; Almirante, B.; Pahissa, A.; Rodriguez-Tudela, J.L.; Cuenca-Estrella, M.; Barcelona Candidemia Project Study Group. Prevalence and susceptibility profile of Candida metapsilosis and Candida orthopsilosis: Results from population-based surveillance of candidemia in Spain. Antimicrob. Agents Chemother. 2008, 52, 1506–1509. [Google Scholar] [CrossRef] [PubMed]
- Lockhart, S.R.; Messer, S.A.; Pfaller, M.A.; Diekema, D.J. Geographic distribution and antifungal susceptibility of the newly described species Candida orthopsilosis and Candida metapsilosis in comparison to the closely related species Candida parapsilosis. J. Clin. Microbiol. 2008, 46, 2659–2664. [Google Scholar] [CrossRef] [PubMed]
- Goncalves, S.S.; Amorim, C.S.; Nucci, M.; Padovan, A.C.; Briones, M.R.; Melo, A.S.; Colombo, A.L. Prevalence rates and antifungal susceptibility profiles of the Candida parapsilosis species complex: Results from a nationwide surveillance of candidaemia in Brazil. Clin. Microbiol. Infect. 2010, 16, 885–887. [Google Scholar] [CrossRef] [PubMed]
- Nucci, M.; Queiroz-Telles, F.; Alvarado-Matute, T.; Tiraboschi, I.N.; Cortes, J.; Zurita, J.; Guzman-Blanco, M.; Santolaya, M.E.; Thompson, L.; Sifuentes-Osornio, J.; et al. Epidemiology of candidemia in Latin America: A laboratory-based survey. PLoS ONE 2013, 8, e59373. [Google Scholar] [CrossRef] [PubMed]
- Marcos-Zambrano, L.J.; Escribano, P.; Sanchez, C.; Munoz, P.; Bouza, E.; Guinea, J. Antifungal resistance to fluconazole and echinocandins is not emerging in yeast isolates causing fungemia in a Spanish tertiary care center. Antimicrob. Agents Chemother. 2014, 58, 4565–4572. [Google Scholar] [CrossRef] [PubMed]
- Colombo, A.L.; Guimaraes, T.; Sukienik, T.; Pasqualotto, A.C.; Andreotti, R.; Queiroz-Telles, F.; Nouer, S.A.; Nucci, M. Prognostic factors and historical trends in the epidemiology of candidemia in critically ill patients: An analysis of five multicenter studies sequentially conducted over a 9-year period. Intens. Care Med. 2014, 40, 1489–1498. [Google Scholar] [CrossRef] [PubMed]
- Pfaller, M.A.; Jones, R.N.; Castanheira, M. Regional data analysis of Candida non-albicans strains collected in United States medical sites over a 6-year period, 2006–2011. Mycoses 2014, 57, 602–611. [Google Scholar] [CrossRef] [PubMed]
- Arvanitis, M.; Glavis-Bloom, J.; Mylonakis, E. Invertebrate models of fungal infection. Biochim. Biophys. Acta 2013, 1832, 1378–1383. [Google Scholar] [CrossRef] [PubMed]
- Desalermos, A.; Fuchs, B.B.; Mylonakis, E. Selecting an invertebrate model host for the study of fungal pathogenesis. PLoS Pathog. 2012, 8, e1002451. [Google Scholar] [CrossRef] [PubMed]
- Muhammed, M.; Coleman, J.J.; Mylonakis, E. Caenorhabditis elegans: A nematode infection model for pathogenic fungi. Methods Mol. Biol. 2012, 845, 447–454. [Google Scholar] [PubMed]
- Junqueira, J.C. Galleria mellonella as a model host for human pathogens: Recent studies and new perspectives. Virulence 2012, 3, 474–476. [Google Scholar] [CrossRef] [PubMed]
- Fallon, J.P.; Reeves, E.P.; Kavanagh, K. The Aspergillus fumigatus toxin fumagillin suppresses the immune response of Galleria mellonella larvae by inhibiting the action of haemocytes. Microbiology 2011, 157, 1481–1488. [Google Scholar] [CrossRef] [PubMed]
- Gago, S.; Garcia-Rodas, R.; Cuesta, I.; Mellado, E.; Alastruey-Izquierdo, A. Candida parapsilosis, Candida orthopsilosis, and Candida metapsilosis virulence in the non-conventional host Galleria mellonella. Virulence 2014, 5, 278–285. [Google Scholar] [CrossRef] [PubMed]
- Breger, J.; Fuchs, B.B.; Aperis, G.; Moy, T.I.; Ausubel, F.M.; Mylonakis, E. Antifungal chemical compounds identified using a C. elegans pathogenicity assay. PLoS Pathog. 2007, 3, e18. [Google Scholar] [CrossRef] [PubMed]
- Pukkila-Worley, R.; Ausubel, F.M.; Mylonakis, E. Candida albicans infection of Caenorhabditis elegans induces antifungal immune defenses. PLoS Pathog. 2011, 7, e1002074. [Google Scholar] [CrossRef] [PubMed]
- Mylonakis, E.; Moreno, R.; El Khoury, J.B.; Idnurm, A.; Heitman, J.; Calderwood, S.B.; Ausubel, F.M.; Diener, A. Galleria mellonella as a model system to study Cryptococcus neoformans pathogenesis. Infect. Immun. 2005, 73, 3842–3850. [Google Scholar] [CrossRef] [PubMed]
- Pukkila-Worley, R.; Peleg, A.Y.; Tampakakis, E.; Mylonakis, E. Candida albicans hyphal formation and virulence assessed using a Caenorhabditis elegans infection model. Eukaryot. Cell 2009, 8, 1750–1758. [Google Scholar] [CrossRef] [PubMed]
- Ortega-Riveros, M.; De-la-Pinta, I.; Marcos-Arias, C.; Ezpeleta, G.; Quindos, G.; Eraso, E. Usefulness of the non-conventional Caenorhabditis elegans model to assess Candida virulence. Mycopathologia 2017, 182, 785–795. [Google Scholar] [CrossRef] [PubMed]
- Brenner, S. The genetics of Caenorhabditis elegans. Genetics 1974, 77, 71–94. [Google Scholar] [PubMed]
- Kim, D.H.; Feinbaum, R.; Alloing, G.; Emerson, F.E.; Garsin, D.A.; Inoue, H.; Tanaka-Hino, M.; Hisamoto, N.; Matsumoto, K.; Tan, M.W.; et al. A conserved p38 map kinase pathway in Caenorhabditis elegans innate immunity. Science 2002, 297, 623–626. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Li, D.; Xi, L.; Mylonakis, E. Caenorhabditis elegans: A simple nematode infection model for Penicillium marneffei. PLoS ONE 2014, 9, e108764. [Google Scholar] [CrossRef] [PubMed]
- Rohlfing, A.K.; Miteva, Y.; Hannenhalli, S.; Lamitina, T. Genetic and physiological activation of osmosensitive gene expression mimics transcriptional signatures of pathogen infection in C. elegans. PLoS ONE 2010, 5, e9010. [Google Scholar] [CrossRef] [PubMed]
- Hoogewijs, D.; Houthoofd, K.; Matthijssens, F.; Vandesompele, J.; Vanfleteren, J.R. Selection and validation of a set of reliable reference genes for quantitative sod gene expression analysis in C. elegans. BMC Mol. Biol. 2008, 9, 9. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Ebata, A.; Dong, Y.; Rizki, G.; Iwata, T.; Lee, S.S. Caenorhabditis elegans HCF-1 functions in longevity maintenance as a DAF-16 regulator. PLoS Biol. 2008, 6, e233. [Google Scholar] [CrossRef] [PubMed]
- Pujol, N.; Cypowyj, S.; Ziegler, K.; Millet, A.; Astrain, A.; Goncharov, A.; Jin, Y.; Chisholm, A.D.; Ewbank, J.J. Distinct innate immune responses to infection and wounding in the C. elegans epidermis. Curr. Biol. 2008, 18, 481–489. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, D.; Smith, M.A.; Zhang, B.; Pan, X. Selection of reliable reference genes in Caenorhabditis elegans for analysis of nanotoxicity. PLoS ONE 2012, 7, e31849. [Google Scholar] [CrossRef] [PubMed]
- Kato, Y.; Aizawa, T.; Hoshino, H.; Kawano, K.; Nitta, K.; Zhang, H. abf-1 and abf-2, ASABF-type antimicrobial peptide genes in Caenorhabditis elegans. Biochem. J. 2002, 361, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Treviño-Rangel Rde, J.; Rodriguez-Sanchez, I.P.; Elizondo-Zertuche, M.; Martinez-Fierro, M.L.; Garza-Veloz, I.; Romero-Diaz, V.J.; Gonzalez, J.G.; Gonzalez, G.M. Evaluation of in vivo pathogenicity of Candida parapsilosis, Candida orthopsilosis, and Candida metapsilosis with different enzymatic profiles in a murine model of disseminated candidiasis. Med. Mycol. 2014, 52, 240–245. [Google Scholar] [CrossRef] [PubMed]
- Blanco-Blanco, M.T.; Gomez-Garcia, A.C.; Hurtado, C.; Galan-Ladero, M.A.; Lozano Mdel, C.; Garcia-Tapias, A.; Blanco, M.T. Candida orthopsilosis fungemias in a Spanish tertiary care hospital: Incidence, epidemiology and antifungal susceptibility. Rev. Iberoam. Micol. 2014, 31, 145–148. [Google Scholar] [CrossRef] [PubMed]
- Gacser, A.; Trofa, D.; Schafer, W.; Nosanchuk, J.D. Targeted gene deletion in Candida parapsilosis demonstrates the role of secreted lipase in virulence. J. Clin. Investig. 2007, 117, 3049–3058. [Google Scholar] [CrossRef] [PubMed]
- Nemeth, T.; Toth, A.; Szenzenstein, J.; Horvath, P.; Nosanchuk, J.D.; Grozer, Z.; Toth, R.; Papp, C.; Hamari, Z.; Vagvolgyi, C.; et al. Characterization of virulence properties in the C. parapsilosis sensu lato species. PLoS ONE 2013, 8, e68704. [Google Scholar] [CrossRef] [PubMed]
- Bertini, A.; De Bernardis, F.; Hensgens, L.A.; Sandini, S.; Senesi, S.; Tavanti, A. Comparison of Candida parapsilosis, Candida orthopsilosis, and Candida metapsilosis adhesive properties and pathogenicity. Int. J. Med. Microbiol. 2013, 303, 98–103. [Google Scholar] [CrossRef] [PubMed]
- Liu, H. Transcriptional control of dimorphism in Candida albicans. Curr. Opin. Microbiol. 2001, 4, 728–735. [Google Scholar] [CrossRef]
- Navarro-Garcia, F.; Sanchez, M.; Nombela, C.; Pla, J. Virulence genes in the pathogenic yeast Candida albicans. FEMS Microbiol. Rev. 2001, 25, 245–268. [Google Scholar] [CrossRef] [PubMed]
- Engelmann, I.; Pujol, N. Innate immunity in C. elegans. Adv. Exp. Med. Biol. 2010, 708, 105–121. [Google Scholar] [PubMed]
- Ermolaeva, M.A.; Schumacher, B. Insights from the worm: The C. elegans model for innate immunity. Semin. Immunol. 2014, 26, 303–309. [Google Scholar] [CrossRef] [PubMed]
- Engelmann, I.; Griffon, A.; Tichit, L.; Montanana-Sanchis, F.; Wang, G.; Reinke, V.; Waterston, R.H.; Hillier, L.W.; Ewbank, J.J. A comprehensive analysis of gene expression changes provoked by bacterial and fungal infection in C. elegans. PLoS ONE 2011, 6, e19055. [Google Scholar] [CrossRef] [PubMed]
- Zugasti, O.; Bose, N.; Squiban, B.; Belougne, J.; Kurz, C.L.; Schroeder, F.C.; Pujol, N.; Ewbank, J.J. Activation of a G protein-coupled receptor by its endogenous ligand triggers the innate immune response of Caenorhabditis elegans. Nat. Immunol. 2014, 15, 833–838. [Google Scholar] [CrossRef] [PubMed]
- Kato, Y.; Komatsu, S. ASABF, a novel cysteine-rich antibacterial peptide isolated from the nematode Ascaris suum. Purification, primary structure, and molecular cloning of cDNA. J. Biol. Chem. 1996, 271, 30493–30498. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Kato, Y. Common structural properties specifically found in the CSαβ-type antimicrobial peptides in nematodes and mollusks: Evidence for the same evolutionary origin? Dev. Comp. Immunol. 2003, 27, 499–503. [Google Scholar] [CrossRef]
- Means, T.K.; Mylonakis, E.; Tampakakis, E.; Colvin, R.A.; Seung, E.; Puckett, L.; Tai, M.F.; Stewart, C.R.; Pukkila-Worley, R.; Hickman, S.E.; et al. Evolutionarily conserved recognition and innate immunity to fungal pathogens by the scavenger receptors SCARF1 and CD36. J. Exp. Med. 2009, 206, 637–653. [Google Scholar] [CrossRef] [PubMed]
- Couillault, C.; Pujol, N.; Reboul, J.; Sabatier, L.; Guichou, J.F.; Kohara, Y.; Ewbank, J.J. TlR-independent control of innate immunity in Caenorhabditis elegans by the TIR domain adaptor protein TIR-1, an ortholog of human SARM. Nat. Immunol. 2004, 5, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Pujol, N.; Zugasti, O.; Wong, D.; Couillault, C.; Kurz, C.L.; Schulenburg, H.; Ewbank, J.J. Anti-fungal innate immunity in C. elegans is enhanced by evolutionary diversification of antimicrobial peptides. PLoS Pathog. 2008, 4, e1000105. [Google Scholar] [CrossRef] [PubMed]
- Zugasti, O.; Ewbank, J.J. Neuroimmune regulation of antimicrobial peptide expression by a noncanonical TGF-β signaling pathway in Caenorhabditis elegans epidermis. Nat. Immunol. 2009, 10, 249–256. [Google Scholar] [CrossRef] [PubMed]
| Strain | Description | Purpose | Reference |
|---|---|---|---|
| Candida species | |||
| C. parapsilosis (sensu stricto) ATCC 22019 | WT a | All experiments | ATCC |
| Candida orthopsilosis ATCC 96141 | WT | All experiments | ATCC |
| Candida metapsilosis ATCC 93143 | WT | All experiments | ATCC |
| C. elegans | |||
| N2 | WT | Immunity response | [23] |
| glp-4; sek-1 | glp-4(bn2) I; sek-1(km4) | Killing assay, treatment with antifungal drugs, microscopic studies | [24] |
| pmk-1 | pmk-1(km25) | Immunity response | [24] |
| MIC (µg/mL) | ||
|---|---|---|
| Strain | Fluconazole | Caspofungin |
| ATCC 22019 | 1.0 | 0.5 |
| ATCC 96141 | 1.0 | 0.5 |
| ATCC 96143 | 1.0 | 0.5 |
| Oligonucleotide a | Sequence 5′ to 3′ | Reference |
|---|---|---|
| ABF-1/Fw | CTGCCTTCTCCTTGTTCTCCTACT | [19] |
| ABF-1/Rv | CCTCTGCATTACCGGAACATC | [19] |
| ABF-2/Fw | TTTCCTTGCACTTCTCCTGG | This study b |
| ABF-2/Rv | CGGTTCCACAGTTTTGCATAC | This study |
| CNC-4/Fw | ACAATGGGGCTACGGTCCATAT | This study |
| CNC-4/Rv | ACTTTCCAATGAGCATTCCGAGGA | This study |
| CNC-7/Fw | CAGGTTCAATGCAGTATGGCTATGG | This study |
| CNC-7/Rv | GGACGGTACATTCCCATACC | This study |
| FIPR-22/23 Fw | GCTGAAGCTCCACACATCC | [19] |
| FIPR-22/23 Rv | TATCCCATTCCTCCGTATCC | [19] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Souza, A.C.R.; Burgwyn Fuchs, B.; De Souza Alves, V.; Jayamani, E.; Colombo, A.L.; Mylonakis, E. Pathogenesis of the Candida parapsilosis Complex in the Model Host Caenorhabditis elegans. Genes 2018, 9, 401. https://doi.org/10.3390/genes9080401
Souza ACR, Burgwyn Fuchs B, De Souza Alves V, Jayamani E, Colombo AL, Mylonakis E. Pathogenesis of the Candida parapsilosis Complex in the Model Host Caenorhabditis elegans. Genes. 2018; 9(8):401. https://doi.org/10.3390/genes9080401
Chicago/Turabian StyleSouza, Ana Carolina Remondi, Beth Burgwyn Fuchs, Viviane De Souza Alves, Elamparithi Jayamani, Arnaldo Lopes Colombo, and Eleftherios Mylonakis. 2018. "Pathogenesis of the Candida parapsilosis Complex in the Model Host Caenorhabditis elegans" Genes 9, no. 8: 401. https://doi.org/10.3390/genes9080401
APA StyleSouza, A. C. R., Burgwyn Fuchs, B., De Souza Alves, V., Jayamani, E., Colombo, A. L., & Mylonakis, E. (2018). Pathogenesis of the Candida parapsilosis Complex in the Model Host Caenorhabditis elegans. Genes, 9(8), 401. https://doi.org/10.3390/genes9080401








