An ‛Aukward’ Tale: A Genetic Approach to Discover the Whereabouts of the Last Great Auks
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Information
2.2. DNA Extraction
2.3. Data Generation
2.4. Read Processing
3. Results
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Higuchi, R.; Bowman, B.; Freiberger, M.; Ryder, O.A.; Wilson, A.C. DNA sequences from the quagga, an extinct member of the horse family. Nature 1984, 312, 282–284. [Google Scholar] [CrossRef]
- Orlando, L.; Ginolhac, A.; Zhang, G.; Froese, D.; Albrechtsen, A.; Stiller, M.; Schubert, M.; Cappellini, E.; Petersen, B.; Moltke, I.; et al. Recalibrating equus evolution using the genome sequence of an early middle pleistocene horse. Nature 2013, 499, 74–78. [Google Scholar] [CrossRef]
- Fuller, E. The Great Auk; Errol Fuller: Kent, UK, 1999; 448p, ISBN 0-9533553-0-6. [Google Scholar]
- Grieve, S. The Great Auk, or Garefowl. Its History, Archaeology and Remains, Digitally Printed Version 2015 ed.; Cambridge University Press: Cambridge, UK, 1885; ISBN 978-1-108-08147-4. [Google Scholar]
- Bengtson, S.-A. Breeding ecology and extinction of the great auk (Pinguinus impennis): Anecdotal evidence and conjectures. Auk 1984, 101, 1–12. [Google Scholar]
- Gaskell, J. Who Killed the Great Auk? Oxford University Press: New York, NY, USA, 2000; 240p, ISBN 978-0-19856478-2. [Google Scholar]
- Newton, A. XlII.—Abstract of Mr. J. Wolley’s researches in Iceland respecting the gare-fowl or great auk (Alca impennis, linn.). Ibis 1861, 3, 374–399. [Google Scholar] [CrossRef]
- Meldgaard, M. The great auk, Pinguinus impennis (L.) in greenland. Hist. Biol. 1988, 1, 145–178. [Google Scholar] [CrossRef]
- Serjeantson, D. The great auk and the gannet: A prehistoric perspective on the extinction of the great auk. Int. J. Osteoarchaeol. 2001, 11, 43–55. [Google Scholar] [CrossRef]
- Montevecchi, W.A.; David, A. Kirk. Great Auk (Pinguinus impennis) Birds of North America Online; Rodewald, P.G., Ed.; Cornell Lab of Ornithology: Ithaca, NY, USA, 1996. [Google Scholar]
- International Union for Conservation of Nature, Birdlife International and Handbook of the Birds of the World (2016). Pinguinus impennis, The IUCN Red List of Threatened Species. Version 2016–3; 2016. Available online: http://maps.iucnredlist.org/map.html?id=22694856 (accessed on 16 May 2017).
- ArcGIS. Arcgis Desktop, Arcmap; 10.5.0.6491; ESRI: Redlands, CA, USA, 2016. [Google Scholar]
- Knapp, M.; Clarke, A.C.; Horsburgh, K.A.; Matisoo-Smith, E.A. Setting the stage—Building and working in an ancient DNA laboratory. Ann. Anat. 2012, 194, 3–6. [Google Scholar] [CrossRef] [PubMed]
- Dabney, J.; Knapp, M.; Glocke, I.; Gansauge, M.-T.; Weihmann, A.; Nickel, B.; Valdiosera, C.; García, N.; Pääbo, S.; Arsuaga, J.-L.; et al. Complete mitochondrial genome sequence of a middle pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc. Natl. Acad. Sci. USA 2013, 110, 15758–15763. [Google Scholar] [CrossRef]
- Gilbert, M.T.P.; Tomsho, L.P.; Rendulic, S.; Packard, M.; Drautz, D.I.; Sher, A.; Tikhonov, A.; Dalén, L.; Kuznetsova, T.; Kosintsev, P.; et al. Whole-genome shotgun sequencing of mitochondria from ancient hair shafts. Science 2007, 317, 1927–1930. [Google Scholar] [CrossRef] [PubMed]
- Campos, P.F.; Gilbert, T.M.P. DNA extraction from keratin and chitin. In Ancient DNA: Methods and Protocols; Shapiro, B., Hofreiter, M., Eds.; Humana Press: Totowa, NJ, USA, 2012; pp. 43–49. ISBN 978-1-61779-516-9. [Google Scholar]
- Gansauge, M.-T.; Meyer, M. Single-stranded DNA library preparation for the sequencing of ancient or damaged DNA. Nat. Protoc. 2013, 8, 737–748. [Google Scholar] [CrossRef]
- Bennett, E.A.; Massilani, D.; Lizzo, G.; Daligault, J.; Geigl, E.M.; Grange, T. Library construction for ancient genomics: Single strand or double strand? BioTechniques 2014, 56, 289–290, 292–286, 298, passim. [Google Scholar] [CrossRef]
- Meyer, M.; Kircher, M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb. Protoc. 2010, 2010. [Google Scholar] [CrossRef]
- MYcroarry. Mybaits Manual—Sequence Enrichment for Targeted Sequencing v2.3.1; MYcroarry, 2014; Available online: http://www.mycroarray.com/mybaits/manuals.html (accessed on 13 June 2017).
- Schubert, M.; Ermini, L.; Sarkissian, C.D.; Jónsson, H.; Ginolhac, A.; Schaefer, R.; Martin, M.D.; Fernández, R.; Kircher, M.; McCue, M.; et al. Characterization of ancient and modern genomes by snp detection and phylogenomic and metagenomic analysis using paleomix. Nat. Protoc. 2014, 9, 1056–1082. [Google Scholar] [CrossRef]
- Lindgreen, S. Adapterremoval: Easy cleaning of next-generation sequencing reads. BMC Res. Notes 2012, 5, 337. [Google Scholar] [CrossRef]
- Schubert, M.; Lindgreen, S.; Orlando, L. Adapterremoval v2: Rapid adapter trimming, identification, and read merging. BMC Res. Notes 2016, 9, 88. [Google Scholar] [CrossRef]
- Anmarkrud, J.A.; Lifjeld, J.T. Complete mitochondrial genomes of eleven extinct or possibly extinct bird species. Mol. Ecol. Resour. 2017, 17, 334–341. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with burrows–wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- BroadInstitute. Picard v1.82. Available online: http://broadinstitute.github.io/picard/ (accessed on 13 June 2017).
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; 1000 Genome Project Data Processing Subgroup. The sequence alignment/map format and samtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The genome analysis toolkit: A mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Jónsson, H.; Ginolhac, A.; Schubert, M.; Johnson, P.L.F.; Orlando, L. Mapdamage2.0: Fast approximate bayesian estimates of ancient DNA damage parameters. Bioinformatics 2013, 29, 1682–1684. [Google Scholar] [CrossRef]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and vcftools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Gouy, M.; Guindon, S.; Gascuel, O. Seaview version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 2010, 27, 221–224. [Google Scholar] [CrossRef]
- Hall, T.A. Bioedit: A user-friendly biological sequence alignment editor and analysis program for windows 95/98/nt. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Milne, I.; Stephen, G.; Bayer, M.; Cock, P.J.A.; Pritchard, L.; Cardle, L.; Shaw, P.D.; Marshall, D. Using tablet for visual exploration of second-generation sequencing data. Brief. Bioinform. 2013, 14, 193–202. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. Mega7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. Jmodeltest 2: More models, new heuristics and high-performance computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef]
- Guindon, S.; Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef]
- Hasegawa, M.; Kishino, H.; Yano, T. Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J. Mol. Evol. 1985, 22, 160–174. [Google Scholar] [CrossRef]
- Thomas, J.E.; Haile, J.; Martin, M.D.; Samaniego Castuita, J.A; Niemann, J.; Sinding, .M.-H.S; Sandoval-Velasco, M..; Soares, A.E.R.; Rawlence, N.J.; Fuller, E.; et al. The evolution and extinction of the Great Auk. (manuscript in preparation).
- Boessenkool, S.; Star, B.; Scofield, R.P.; Seddon, P.J.; Waters, J.M. Lost in translation or deliberate falsification? Genetic analyses reveal erroneous museum data for historic penguin specimens. Proc. R. Soc. B Biol. Sci. 2010, 277, 1057–1064. [Google Scholar] [CrossRef]
- Rawlence, N.J.; Kennedy, M.; Waters, J.M.; Scofield, R.P. Morphological and ancient DNA analyses reveal inaccurate labels on two of buller’s bird specimens. J. R. Soc. N. Z. 2014, 44, 163–169. [Google Scholar] [CrossRef]
- Shepherd, L.D.; Tennyson, A.J.D.; Lambert, D.M. Using ancient DNA to enhance museum collections: A case study of rare kiwi (Apteryx spp.) specimens. J. R. Soc. N. Z. 2013, 43, 119–127. [Google Scholar] [CrossRef]
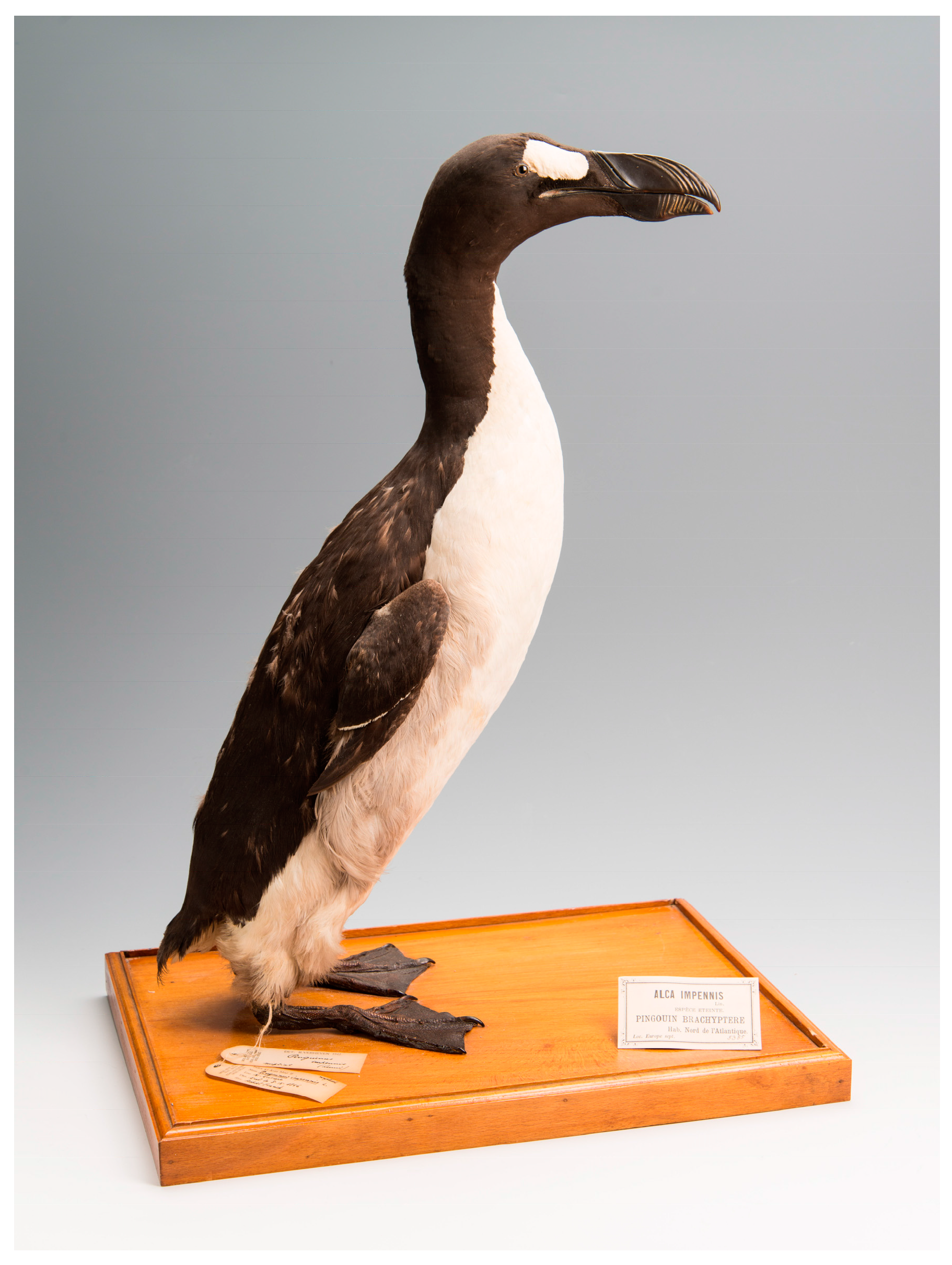

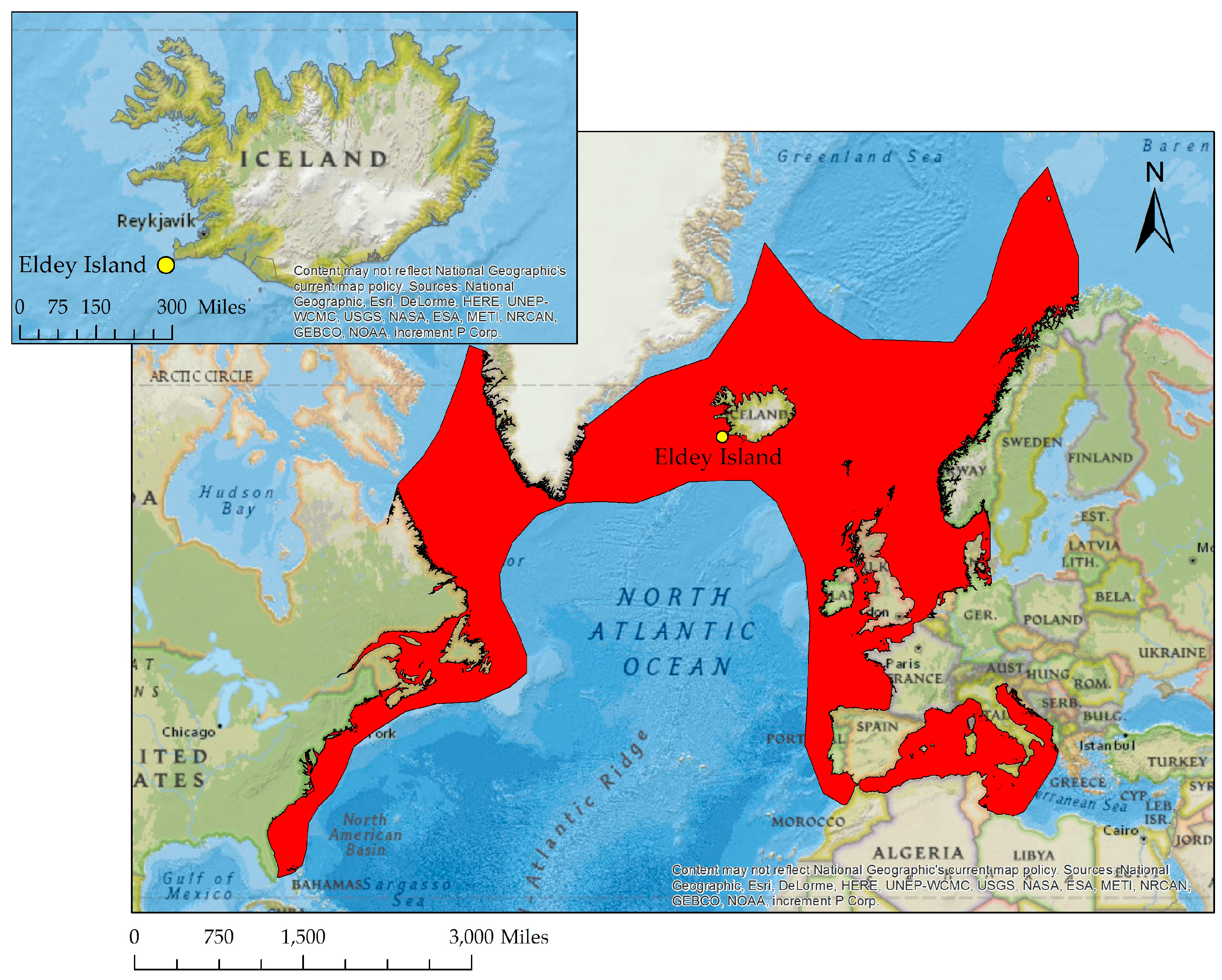

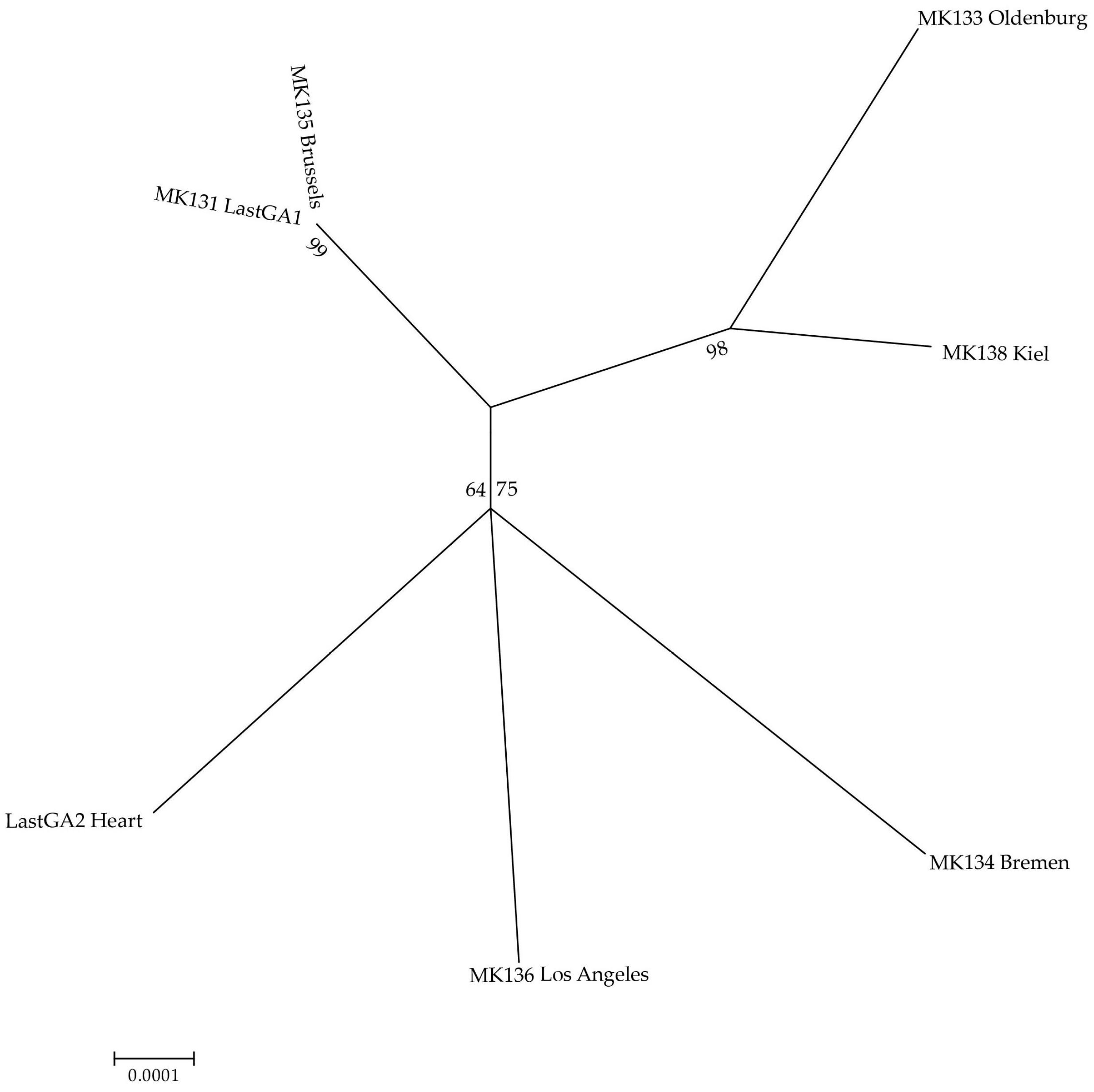
| Lab ID | Bird Name, Number & Description | Origin & Date | Institution | Curator/Collector | Institution Number | Sample Type/Sampling Method |
|---|---|---|---|---|---|---|
| MK131 | Last Great Auk 1 Oesophagus (male) | Eldey Island, Iceland. Date: June 1844 | Natural History Museum of Denmark. Copenhagen, Denmark | J. Fjeldså/ J. Thomas | NHMD 153069 | Oesophagus. Tissue cut from end of oesophagus. |
| MK132 | Last Great Auk 2 Oesophagus (female) | Eldey Island, Iceland. Date: June 1844 | Natural History Museum of Denmark. Copenhagen, Denmark | J. Fjeldså/ J. Thomas | NHMD 153070 | Oesophagus. Tissue cut from end of oesophagus. |
| MK133 | The Oldenburg Auk Fuller: Bird no. 47, Grieve: no. 57, Hahn: no. 77 Adult in summer plumage | Iceland. Probably Eldey. Date: Unknown | Landesmuseum Natur und Mensch Oldenburg. Germany | C. Barilaro | AVE 8086 | Body tissue. Tissue cut from body of bird under wing. |
| MK134 | The Bremen Auk Fuller: Bird no. 36, Grieve: no. 10, Hahn: no. 71 Adult in summer plumage | Unknown. Probably Eldey. Date: Unknown | Übersee-Museum Bremen. Germany | M. Stiller | RKNr. 2392 | Toepad tissue. Tissue cut from feet. |
| MK135 | The Brussels Auk Fuller: Bird no. 3, Grieve: no. 15, Hahn: no. 6 Adult in summer plumage | Probably Eldey Date: Unknown perhaps June, 1844 | Institut Royal des Sciences Naturelles de Belgique. Brussels, Belgium | G. Lenglet | RBINS 5355 | Toepad tissue. Tissue cut from feet |
| MK136 | Dawson Rowley’s Los Angeles Auk Fuller: Bird no. 73, Grieve no. 13, Hahn: no. 5 Adult in summer plumage, said to be female | Iceland. Probably Eldey. Date: Unknown perhaps June, 1844 | Natural History Museum of Los Angeles County. USA | K. Garett | LACM 76476 | Feather. Feathers plucked from body of bird. |
| MK138 | The Schleswig-Holstein Auk Fuller: Bird no. 42, Grieve: no. 31, Hahn: no. 74 Adult in summer plumage | Unknown Date: Unknown | Zoologisches Museum der Christian-Albrechts Universität zu Kiel. Germany | D. Brandis/ L. Rosotta | cat. No. A0585 | Toepad tissue. Tissue cut from feet. |
| LastGA2_Heart | Last Great Auk 2 Heart (female) | Eldey Island, Iceland. Date: June 1844 | Natural History Museum of Denmark. Copenhagen, Denmark | J. Fjeldså/ J. Haile | NHMD 153070 | Heart. Tissue cut from aorta. |
| Sample | GenBank Accession Number | Number of Reads | Number of Unique Reads Mapping to Reference Mitogenome | Estimated Coverage from Unique Hits | Relaxed Settings Sequence Length (bp 1) | Strict Settings Sequence Length (bp) |
|---|---|---|---|---|---|---|
| MK131 | MF188883 | 300754 (read pairs) | 30,297 | 74.40 | 16,001 | 15,067 |
| MK132 | NA | 550631 (read pairs) | 2366 | 6.23 | 13,267 | 3312 |
| MK133 | MF188884 | 429392 (read pairs) | 8750 | 23.04 | 16,251 | 14,240 |
| MK134 | MF188885 | 343766 (read pairs) | 86,325 | 288.62 | 16,607 | 16,526 |
| MK135 | MF188886 | 579992 (read pairs) | 27,767 | 88.90 | 16,554 | 16,356 |
| MK136 | MF188887 | 563635 (read pairs) | 24,401 | 67.83 | 16,330 | 15,833 |
| MK138 | MF188888 | 10796460 (SE 2 reads) | 2799 | 9.76 | 16509 | 7866 |
| LastGA2_Heart | MF188889 | 957970612 (SE reads) | 121,886 | 430.09 | 16,698 | 16,649 |
| MK131 | MK133 | MK134 | MK135 | MK136 | MK138 | LastGA2_Heart | |
|---|---|---|---|---|---|---|---|
| MK131_LastGA1 | |||||||
| MK133_Oldenburg | 17 | ||||||
| MK134_Bremen | 18 | 23 | |||||
| MK135_Brussels | 0 | 17 | 18 | ||||
| MK136_LA | 16 | 23 | 20 | 16 | |||
| MK138_Kiel | 14 | 11 | 20 | 14 | 20 | ||
| LastGA2_Heart | 16 | 23 | 20 | 16 | 18 | 20 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thomas, J.E.; Carvalho, G.R.; Haile, J.; Martin, M.D.; Castruita, J.A.S.; Niemann, J.; Sinding, M.-H.S.; Sandoval-Velasco, M.; Rawlence, N.J.; Fuller, E.; et al. An ‛Aukward’ Tale: A Genetic Approach to Discover the Whereabouts of the Last Great Auks. Genes 2017, 8, 164. https://doi.org/10.3390/genes8060164
Thomas JE, Carvalho GR, Haile J, Martin MD, Castruita JAS, Niemann J, Sinding M-HS, Sandoval-Velasco M, Rawlence NJ, Fuller E, et al. An ‛Aukward’ Tale: A Genetic Approach to Discover the Whereabouts of the Last Great Auks. Genes. 2017; 8(6):164. https://doi.org/10.3390/genes8060164
Chicago/Turabian StyleThomas, Jessica E., Gary R. Carvalho, James Haile, Michael D. Martin, Jose A. Samaniego Castruita, Jonas Niemann, Mikkel-Holger S. Sinding, Marcela Sandoval-Velasco, Nicolas J. Rawlence, Errol Fuller, and et al. 2017. "An ‛Aukward’ Tale: A Genetic Approach to Discover the Whereabouts of the Last Great Auks" Genes 8, no. 6: 164. https://doi.org/10.3390/genes8060164
APA StyleThomas, J. E., Carvalho, G. R., Haile, J., Martin, M. D., Castruita, J. A. S., Niemann, J., Sinding, M.-H. S., Sandoval-Velasco, M., Rawlence, N. J., Fuller, E., Fjeldså, J., Hofreiter, M., Stewart, J. R., Gilbert, M. T. P., & Knapp, M. (2017). An ‛Aukward’ Tale: A Genetic Approach to Discover the Whereabouts of the Last Great Auks. Genes, 8(6), 164. https://doi.org/10.3390/genes8060164








