Eukaryotic Replicative Helicase Subunit Interaction with DNA and Its Role in DNA Replication
Abstract
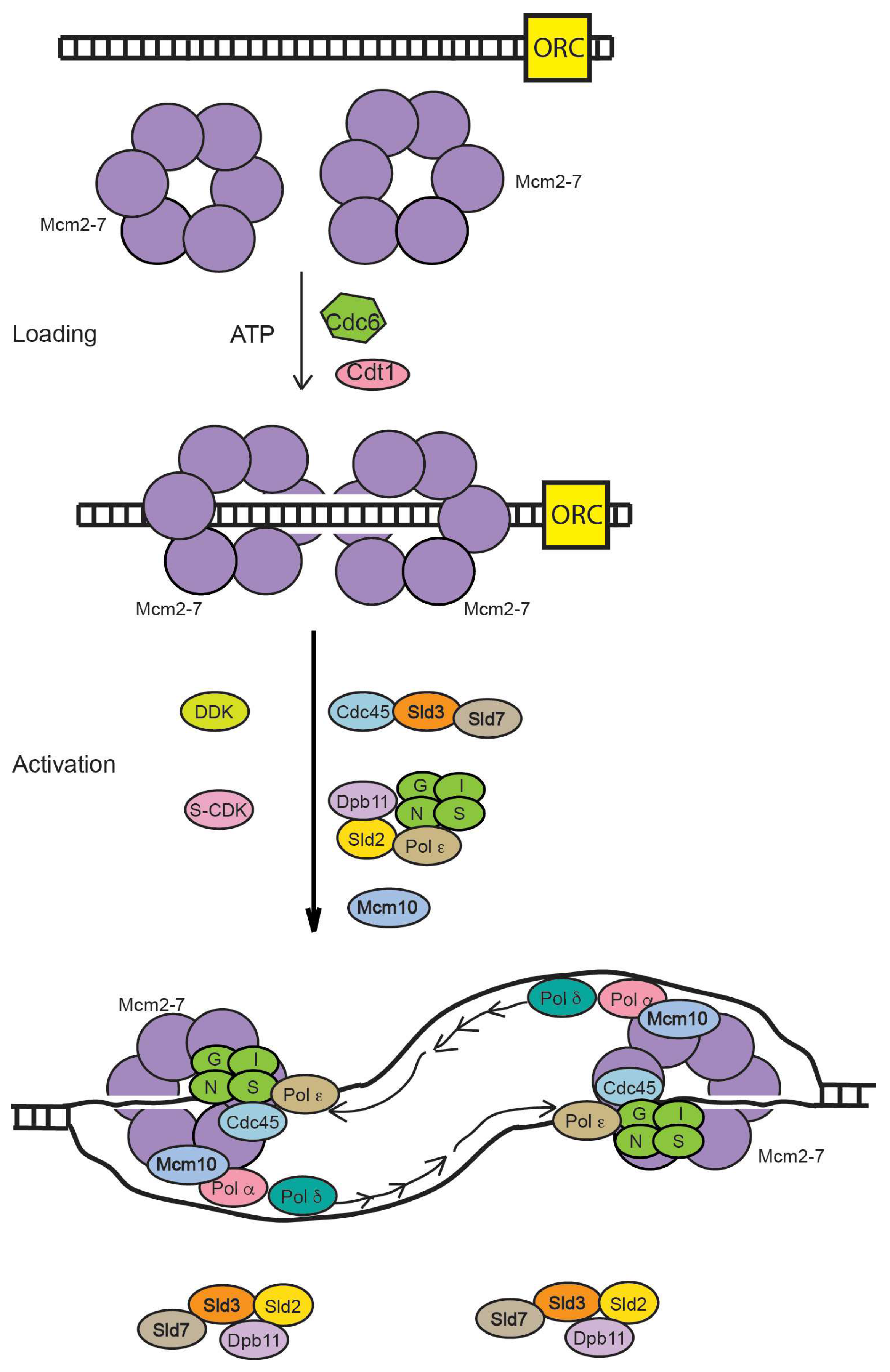
:1. Introduction to DNA Replication and Mcm2-7
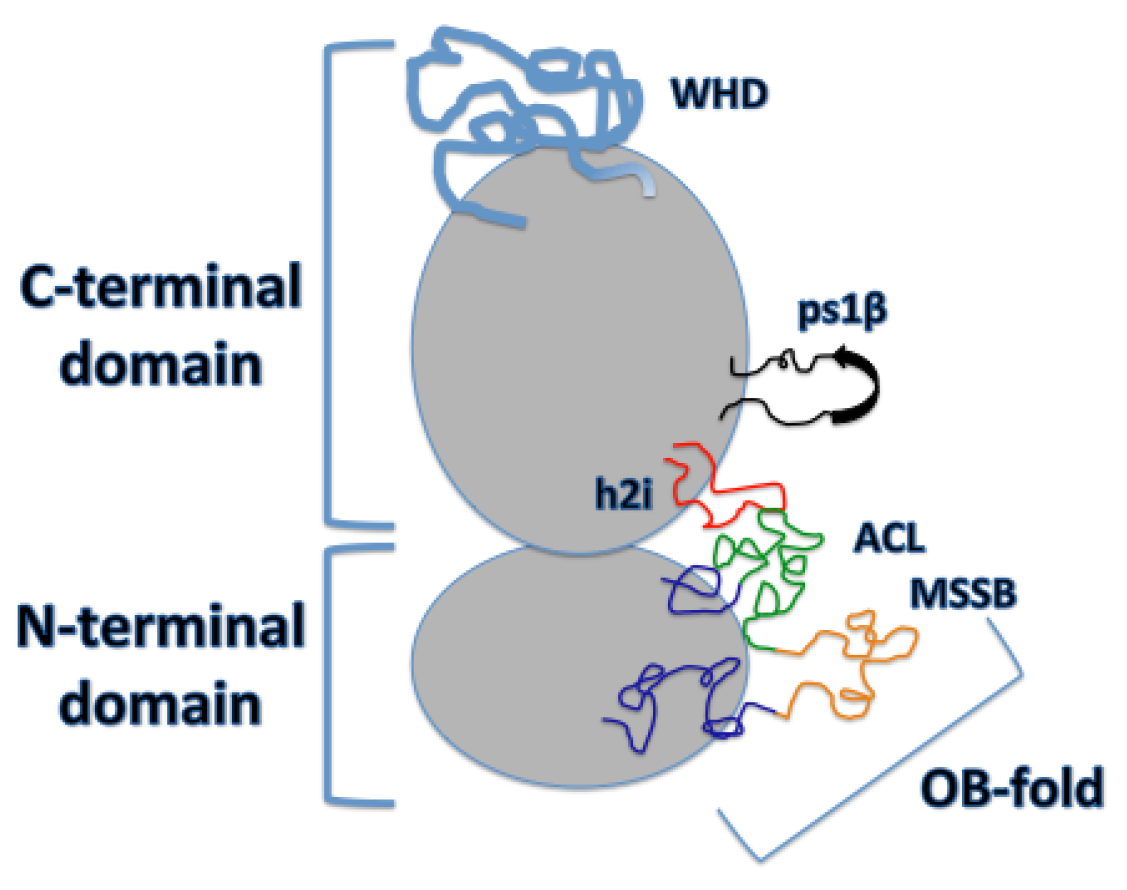
2. DNA Binding Structures of Mcm2-7
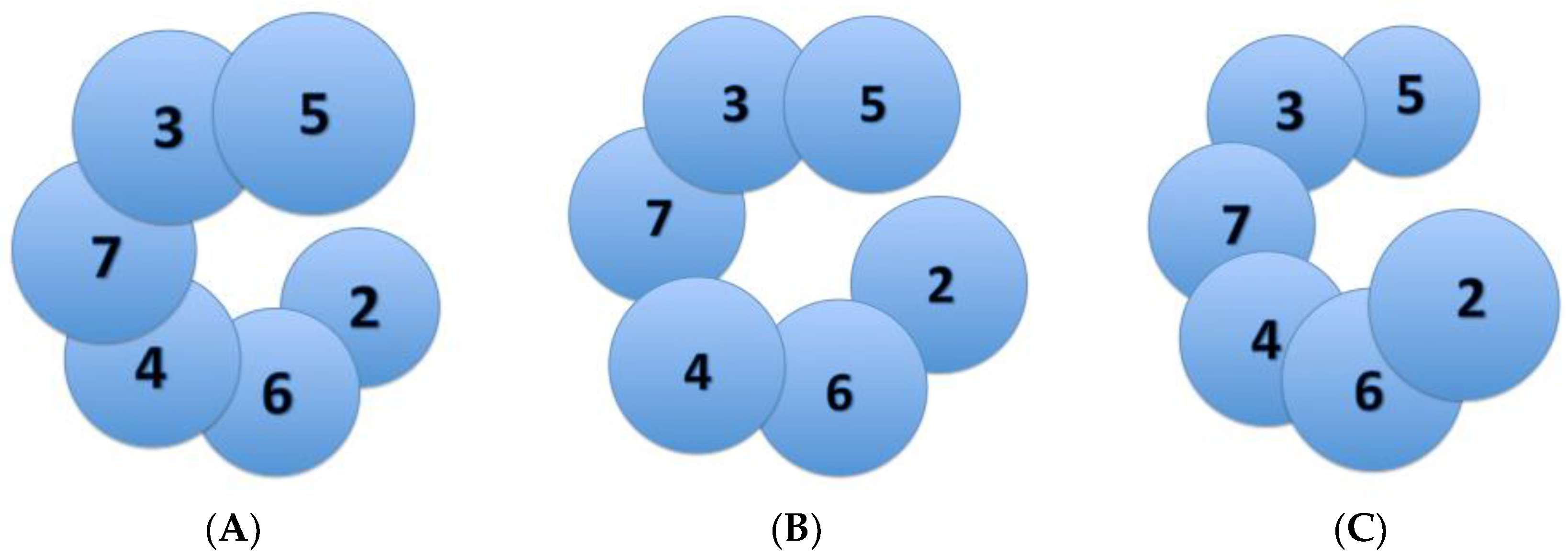
3. Varying Mcm2-7 Conformations Leading to CMG Activation
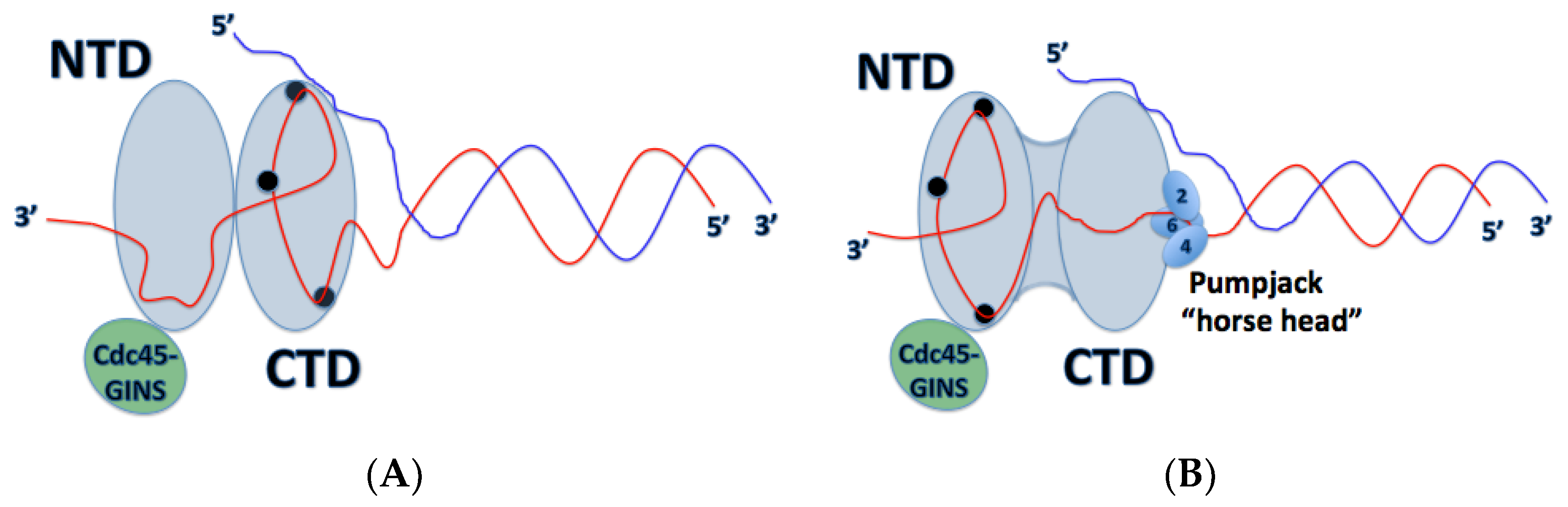
4. Mechanism of DNA Unwinding and Translocation by the Mcm2-7 Helicase
5. Cdc45 Architecture
6. Cdc45 Interaction with DNA
7. GINS Architecture
8. GINS Interaction with DNA
9. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bell, S.; Labib, K. Chromosome Duplication in Saccharomyces cerevisiae. Genetics 2016, 203, 1027067. [Google Scholar] [CrossRef] [PubMed]
- Parker, M.; Botchan, M.; Berger, J. Mechanisms and regulation of DNA replication initiation in eukaryotes. Crit. Rev. Biochem. Mol. Biol. 2017, 52, 1–41. [Google Scholar] [CrossRef] [PubMed]
- Bochman, M.; Schwacha, A. The Mcm2-7 complex has in vitro helicase activity. Mol. Cell 2008, 31, 287–293. [Google Scholar] [CrossRef] [PubMed]
- Ilves, I.; Petojevic, T.; Pesavento, J.; Botchan, M. Activation of the MCM2-7 Helicase by Association with Cdc45 and GINS Proteins. Mol. Cell 2010, 37, 247–258. [Google Scholar] [CrossRef] [PubMed]
- Yeeles, J.; Deegan, T.; Janska, A.; Early, A.; Diffley, J. Regulated eukaryotic DNA replication origin firing with purified proteins. Nature 2015, 519, 431–435. [Google Scholar] [CrossRef] [PubMed]
- Lyubimov, A.; Strycharska, M.; Berger, J. The nuts and bolts of ring-translocase structure and mechanism. Curr. Opin. Struct. Biol. 2011, 21, 240–248. [Google Scholar] [CrossRef] [PubMed]
- Evrin, C.; Clarke, P.; Zech, J.; Lurz, R.; Sun, J.; Uhle, S.; Li, H.; Stillman, B.; Speck, C. A double-hexameric MCM2-7 complex is loaded onto origin DNA during licensing of eukaryotic DNA replication. Proc. Natl. Acad. Sci. USA 2009, 106, 20240–20245. [Google Scholar] [CrossRef] [PubMed]
- Remus, D.; Beuron, F.; Tolun, G.; Griffith, J.; Morris, E.; Diffley, J. Concerted loading of Mcm2-7 double hexamers around DNA during DNA replication origin licensing. Cell 2009, 139, 719–730. [Google Scholar] [CrossRef] [PubMed]
- Martinez, M.; Jones, J.; Irina, B.; Kaplan, D. Origin DNA Melting—An Essential Process with Divergent Mechanisms. Genes 2017, 8, 26. [Google Scholar] [CrossRef] [PubMed]
- Barry, E.R.; McGeoch, A.T.; Kelman, Z.; Bell, S.D. Archaeal MCM has separable processivity, substrate choice and helicase domains. Nucleic Acids Res. 2007, 35, 988–998. [Google Scholar] [CrossRef] [PubMed]
- Pucci, B.; De Felice, M.; Rocco, M.; Esposito, F.; De Falco, M.; Esposito, L.; Rossi, M.; Pisani, F.M. Modular organization of the Sulfolobus solfataricus mini-chromosome maintenance protein. J. Biol. Chem. 2007, 282, 12574–12582. [Google Scholar] [CrossRef] [PubMed]
- Atanassova, N.; Grainge, I. Biochemical characterization of the minichromosome maintenance (MCM) protein of the crenarchaeote Aeropyrum pernix and its interactions with the origin recognition complex (ORC) proteins. Biochemistry 2008, 47, 13362–13370. [Google Scholar] [CrossRef] [PubMed]
- Froelich, C.; Kang, S.; Epling, L.; Bell, S.; Enemark, E. A conserved MCM single-stranded DNA binding element is essential for replication initiation. eLife 2014, 3, e019993. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.; Arachea, B.; Leslie, B.; Enemark, E. Analysis of the crystal structure of an active MCM hexamer. eLife 2014, 3, e03433. [Google Scholar] [CrossRef] [PubMed]
- Barry, E.; Lovett, J.; Costa, A.; Lea, S.; Bell, S. Intersubunit allosteric communication mediated by a conserved loop in the MCM helicase. Proc. Natl. Acad. Sci. USA 2009, 106, 1051–1056. [Google Scholar] [CrossRef] [PubMed]
- Sakakibara, N.; Kasiviswanathan, R.; Melamud, E.; Han, M.; Schwarz, F.; Kelman, Z. Coupling of DNA binding and helicase activity is mediated by a conserved loop in the MCM protein. Nucleic Acids Res. 2008, 36, 1309–1320. [Google Scholar] [CrossRef] [PubMed]
- McGeoch, A.T.; Trakselis, M.A.; Laskey, R.A.; Bell, S.D. Organization of the archaeal MCM complex on DNA and implications for the helicase mechanism. Nat. Struct. Mol. Biol. 2005, 12, 756–762. [Google Scholar] [CrossRef] [PubMed]
- Jenkinson, E.R.; Chong, J.P. Minichromosome maintenance helicase activity is controlled by N- and C-terminal motifs and requires the ATPase domain helix-2 insert. Proc. Natl. Acad. Sci. USA 2006, 103, 7613–7618. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Z.; Bai, L.; Sun, J.; Georgescu, R.; Liu, J.; O′Donnell, M.E.; Li, H. Structure of the eukaryotic replicative CMG helicase suggests a pumpjack motion for translocation. Nat. Struct. Mol. Biol. 2016, 23, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Costa, A.; llves, I.; Tamberg, N.; Petojevic, T.; Nogales, E.; Botchan, M.R.; Berger, J.M. The structural basis for MCM2-7 helicase activation by GINS and Cdc45. Nat. Struct. Mol. Biol. 2011, 18, 471–477. [Google Scholar] [CrossRef] [PubMed]
- Boskovic, J.; Bragado-Nilsson, E.; Prabhakar, B.; Yefimenko, I.; Martinez-Gago, J.; Muñoz, S.; Méndez, J.; Montoya, G. Molecular architecture of the recombinant human MCM2-7 helicase in complex with nucleotides and DNA. Cell Cycle 2016, 15, 2431–2440. [Google Scholar] [CrossRef] [PubMed]
- Costa, A.; Renault, L.; Swuec, P.; Petojevic, T.; Pesavento, J.; Ilves, I.; MacLellan-Gibson, K.; Fleck, R.A.; Botchan, M.R.; Berger, J.M. DNA binding polarity, dimerization and ATPase ring remodeling in the CMG helicase of the eukaryotic replisome. eLife 2014, 3, e03273. [Google Scholar] [CrossRef] [PubMed]
- Ali, F.; Renault, L.; Gannon, J.; Gahlon, H.; Kotecha, A.; Zhou, J.; Rueda, D.; Costa, A. Cryo-EM structures of the eukaryotic replicative helicase bound to a translocation substrate. Nat. Commun. 2016, 7, 10708. [Google Scholar]
- Bruck, I.; Kaplan, D.L. The Dbf4-Cdc7 kinase promotes Mcm2-7 ring opening to allow for single-stranded DNA extrusion and helicase assembly. J. Biol. Chem. 2015, 290, 1210–1221. [Google Scholar] [CrossRef] [PubMed]
- Dhingra, N.; Bruck, I.; Smith, S.; Ning, B.; Kaplan, D. Dpb11 helps control assembly of the Cdc45-Mcm2-7-GINS replication fork helicase. J. Biol. Chem. 2015, 290, 7586–7601. [Google Scholar] [CrossRef] [PubMed]
- Bruck, I.; Kaplan, D. Origin Single-stranded DNA Releases Sld3 Protein from the Mcm2-7 Complex, Allowing the GINS Tetramer to Bind the Mcm2-7 Complex. J. Biol. Chem. 2011, 286, 18602–18613. [Google Scholar] [CrossRef] [PubMed]
- Bruck, I.; Kaplan, D. The replication initiation protein Sld2 regulates helicase assembly. J. Biol. Chem. 2014, 289, 1948–1959. [Google Scholar] [CrossRef] [PubMed]
- Perez-Arnaiz, P.; Kaplan, D. An Mcm10 mutant defective in ssDNA binding shows defects in DNA replication initiation. J. Mol. Biol. 2016, 428, 4608–4625. [Google Scholar] [CrossRef] [PubMed]
- Georgescu, R.; Yuan, Z.; Bai, L.; Santos, R.; Sun, J.; Zhang, D.; Yurieva, O.; Li, H.; O′Donnell, M.E. Structure of the eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation. Proc. Natl. Acad. Sci. USA 2017, 114, 697–706. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Syed, S.; Enemark, E.; Schuck, S.; Stenlund, A.; Ha, T.; Joshua-Tor, L. Dynamic look at DNA unwinding by a replicative helicase. Proc. Natl. Acad. Sci. USA 2014, 111, E827–E835. [Google Scholar] [CrossRef] [PubMed]
- Enemark, E.; Joshua-Tor, L. Mechanism of DNA translocation in a replicative hexameric helicase. Nature 2006, 442, 270–275. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Shi, Y.; Georgescu, R.; Yuan, Z.; Chait, B.; Li, H.; O′Donnell, M.E. The architecture of a eukaryotic replisome. Nat. Struct. Mol. Biol. 2015, 22, 976–982. [Google Scholar] [CrossRef] [PubMed]
- Lõoke, M.; Maloney, M.; Bell, S. Mcm10 regulates DNA replication elongation by stimulating the CMG replicative helicase. Genes Dev. 2017, 31, 291–305. [Google Scholar] [CrossRef] [PubMed]
- Van Deursen, F.; Sengupta, S.; De Piccoli, G.; Sanchez-Diaz, A.; Labib, K. Mcm10 associates with the loaded DNA helicase at replication origins and defines a novel step in its activation. EMBO J. 2012, 31, 2195–2206. [Google Scholar] [CrossRef] [PubMed]
- Kanke, M.; Kodama, Y.; Takahashi, T.; Nakagawa, T.; Masukata, H. Mcm10 plays an essential role in origin DNA unwinding after loading of the CMG components. EMBO J. 2012, 31, 2182–2194. [Google Scholar] [CrossRef] [PubMed]
- Maric, M.; Maculins, T.; De Piccoli, G.; Labib, K. Cdc48 and a ubiquitin ligase drive disassembly of the CMG helicase at the end of DNA replication. Science 2014, 346, 1253596. [Google Scholar] [CrossRef] [PubMed]
- Moreno, S.; Bailey, R.; Campion, N.; Herron, S.; Gambus, A. Polyubiquitylation drives replisome disassembly at the termination of DNA replication. Science 2014, 346, 477–481. [Google Scholar] [CrossRef] [PubMed]
- Franz, A.; Orth, M.; Pirson, P.; Sonneville, R.; Blow, J.; Gartner, A.; Stemmann, O.; Hoppe, T. CDC-48/p97 coordinates CDT-1 degradation with GINS chromatin dissociation to ensure faithful DNA replication. Mol. Cell 2011, 44, 85–96. [Google Scholar] [CrossRef] [PubMed]
- Buchberger, A.; Schindelin, H.; Hanzelmann, P. Control of p97 function by cofactor binding. FEBS Lett. 2015, 589, 2578–2589. [Google Scholar] [CrossRef] [PubMed]
- Aparicio, O.M.; Weinstein, D.M.; Bell, S.P. Components and dynamics of DNA replication complexes in S. cerevisiae: Redistribution of MCM proteins and Cdc45p during S phase. Cell 1997, 91, 59–69. [Google Scholar] [CrossRef]
- Sanchez-Pulido, L.; Ponting, C. Cdc45: The missing RecJ ortholog in eukaryotes? Bioinformatics 2011, 27, 1885–1888. [Google Scholar] [CrossRef] [PubMed]
- Krastanova, I.; Sannino, V.; Amenitsch, H.; Gileadi, O.; Pisani, F.; Onesti, S. Structural and functional insights into the DNA replication factor Cdc45 reveal an evolutionary relationship to the DHH family of phosphoesterases. J. Biol. Chem. 2012, 287, 4121–4128. [Google Scholar] [CrossRef] [PubMed]
- Simon, A.C.; Sannino, V.; Costanzo, V.; Pellegrini, L. Structure of human Cdc45 and implications for CMG helicase function. Nat. Commun. 2016, 7, 11638. [Google Scholar] [CrossRef] [PubMed]
- Bruck, I.; Kaplan, D. Cdc45 protein-single-stranded DNA interaction is important for stalling the helicase during replication stress. J. Biol. Chem. 2013, 288, 7550–7563. [Google Scholar] [CrossRef] [PubMed]
- Szambowska, A.; Tessmer, I.; Kursula, P.; Usskilat, C.; Prus, P.; Pospiech, H.; Grosse, F. DNA binding properties of human Cdc45 suggest a function as molecular wedge for DNA unwinding. Nucleic Acids Res. 2014, 42, 2308–2319. [Google Scholar] [CrossRef] [PubMed]
- Szambowska, A.; Tessmer, I.; Prus, P.; Schlott, B.; Pospiech, H.; Grosse, F. Cdc45-induced loading of human RPA onto single-stranded DNA. Nucleic Acids Res. 2017. [Google Scholar] [CrossRef] [PubMed]
- Petojevic, T.; Pesavento, J.; Costa, A.; Liang, J.; Wang, Z.; Berger, J.; Botchan, M.R. Cdc45 (cell division cycle protein 45) guards the gate of the Eukaryote Replisome helicase stabilizing leading strand engagement. Proc. Natl. Acad. Sci. USA 2015, 112, E249–E258. [Google Scholar] [CrossRef] [PubMed]
- Bochman, M.; Schwacha, A. The Saccharomyces cerevisiae Mcm6/2 and Mcm5/3 ATPase active sites contribute to the function of the putative Mcm2-7 ‘gate’. Nucleic Acids Res. 2010, 38, 6078–6088. [Google Scholar] [CrossRef] [PubMed]
- Takayama, Y.; Kamimura, Y.; Okawa, M.; Muramatsu, S.; Sugino, A.; Araki, H. GINS, a novel multiprotein complex required for chromosomal DNA replication in budding yeast. Genes Dev. 2003, 17, 1153–1165. [Google Scholar] [CrossRef] [PubMed]
- Boskovic, J.; Coloma, J.; Aparicio, T.; Zhou, M.; Robinson, C.; Méndez, J.; Montoya, G. Molecular architecture of the human GINS complex. EMBO Rep. 2007, 8, 678–684. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.P.; Wang, G.; Bermudez, V.; Hurwitz, J.; Chen, X. Crystal structure of the GINS complex and functional insights into its role in DNA replication. Proc. Natl. Acad. Sci. USA 2007, 104, 12685–12690. [Google Scholar] [CrossRef] [PubMed]
- Seki, T.; Akita, M.; Kamimura, Y.; Muramatsu, S.; Araki, H.; Sugino, A. GINS is a DNA polymerase epsilon accessory factor during chromosomal DNA replication in budding yeast. J. Biol. Chem. 2006, 281, 21422–21432. [Google Scholar] [CrossRef] [PubMed]
- Pacek, M.; Tutter, A.; Kubota, Y.; Takisawa, H.; Walter, J. Localization of MCM2-7, Cdc45, and GINS to the site of DNA unwinding during eukaryotic DNA replication. Mol. Cell 2006, 21, 581–587. [Google Scholar] [CrossRef] [PubMed]
- Gambus, A.; Jones, R.C.; Sanchez-Diaz, A.; Kanemaki, M.; van Deursen, F.; Edmondson, R.D.; Labib, K. GINS maintains association of Cdc45 with MCM in replisome progression complexes at eukaryotic DNA replication forks. Nat. Cell Biol. 2006, 8, 358–366. [Google Scholar] [CrossRef] [PubMed]
- Carroni, M.; De March, M.; Medagli, B.; Krastanova, I.; Taylor, I.; Amenitsch, H.; Araki, H.; Pisani, F.M.; Patwardhan, A.; Onesti, S. New insights into the GINS complex explain the controversy between existing structural models. Sci. Rep. 2017, 7, 40188. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, Y.; Puddu, F.; Costanzo, V. RAD51- and MRE11-dependent reassembly of uncoupled CMG helicase complex at collapsed replication forks. Nat. Struct. Mol. Biol. 2011, 19, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Schwacha, A.; Bell, S.P. Interactions between two catalytically distinct MCM subgroups are essential for coordinated ATP hydrolysis and DNA replication. Mol. Cell 2001, 8, 1093–1104. [Google Scholar] [CrossRef]
- Fu, Y.; Yardimci, H.; Long, D.; Ho, T.; Guainazzi, A.; Bermudez, V.; Hurwitz, J.; van Oijen, A.; Schärer, O.D.; Walter, J.C. Selective bypass of a lagging strand roadblock by the eukaryotic replicative DNA helicase. Cell 2011, 146, 931–941. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martinez, M.P.; Wacker, A.L.; Bruck, I.; Kaplan, D.L. Eukaryotic Replicative Helicase Subunit Interaction with DNA and Its Role in DNA Replication. Genes 2017, 8, 117. https://doi.org/10.3390/genes8040117
Martinez MP, Wacker AL, Bruck I, Kaplan DL. Eukaryotic Replicative Helicase Subunit Interaction with DNA and Its Role in DNA Replication. Genes. 2017; 8(4):117. https://doi.org/10.3390/genes8040117
Chicago/Turabian StyleMartinez, Matthew P., Amanda L. Wacker, Irina Bruck, and Daniel L. Kaplan. 2017. "Eukaryotic Replicative Helicase Subunit Interaction with DNA and Its Role in DNA Replication" Genes 8, no. 4: 117. https://doi.org/10.3390/genes8040117
APA StyleMartinez, M. P., Wacker, A. L., Bruck, I., & Kaplan, D. L. (2017). Eukaryotic Replicative Helicase Subunit Interaction with DNA and Its Role in DNA Replication. Genes, 8(4), 117. https://doi.org/10.3390/genes8040117





