Non‐Canonical Replication Initiation: You’re Fired!
Abstract
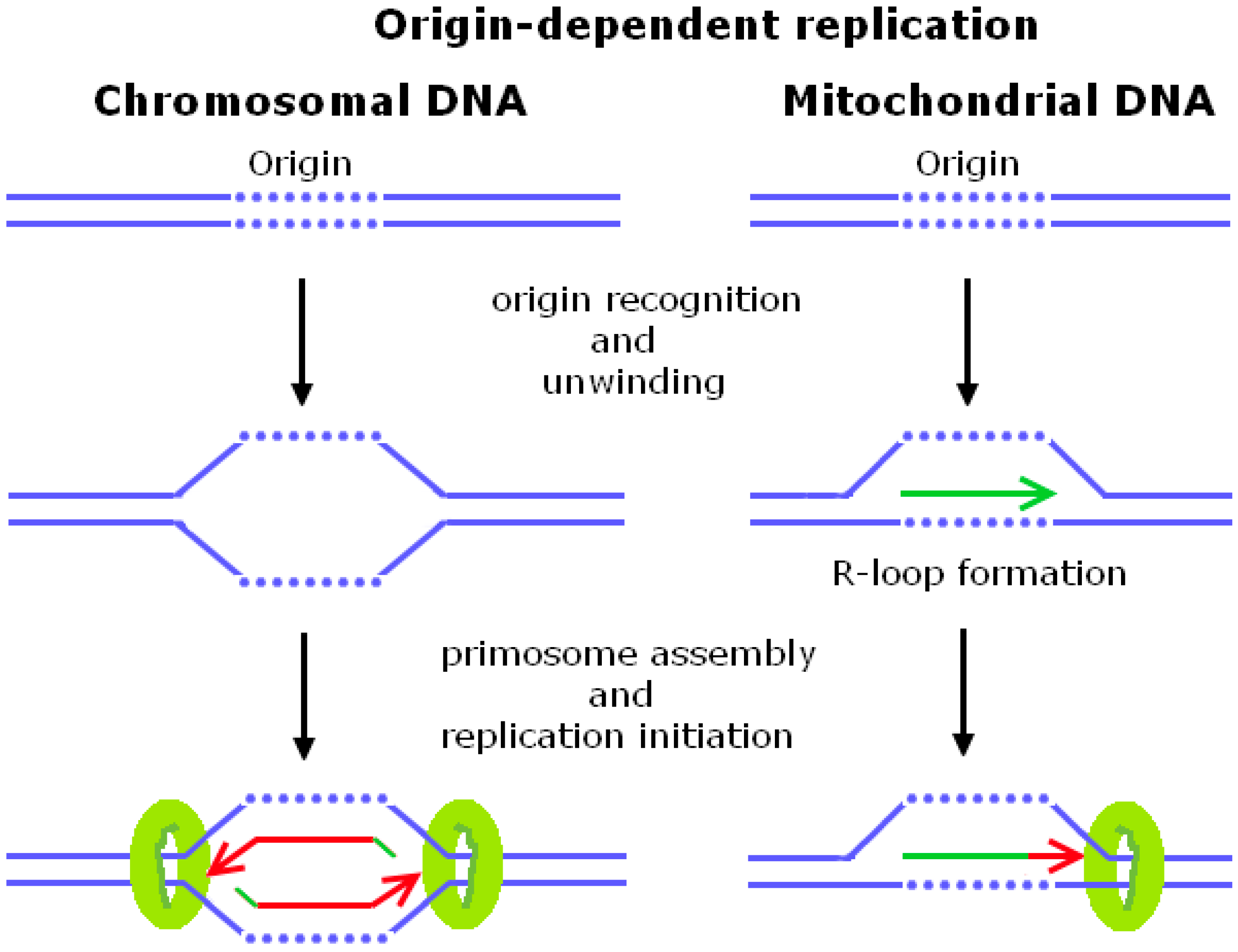
:1. Origin-Dependent Replication
1.1. Chromosomal DNA Replication Initiation in Escherichia coli and Saccharomyces cerevisiae
1.2. Mitochondrial DNA Replication Initiation
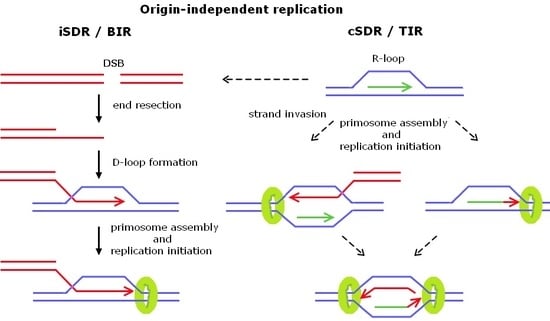
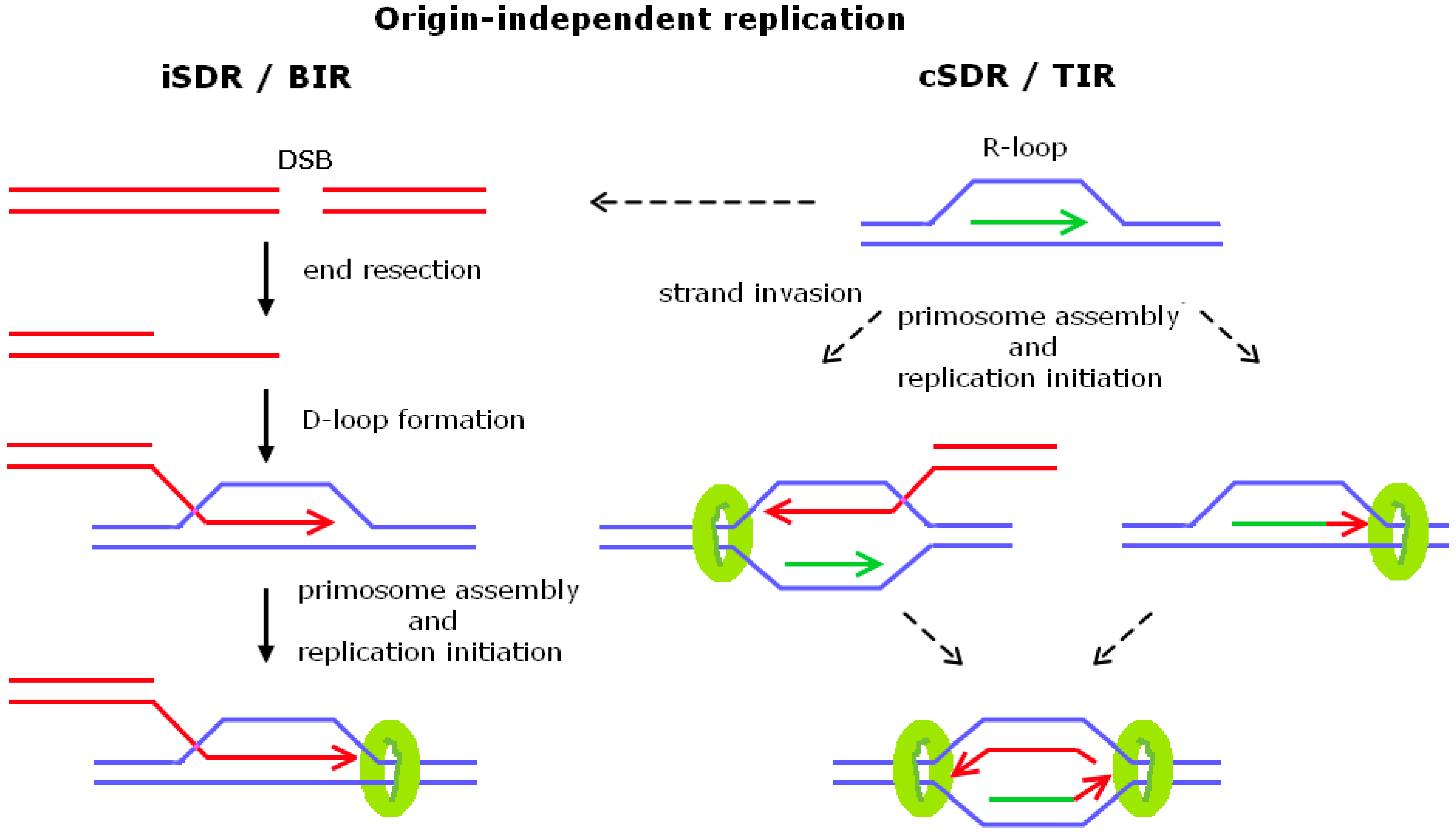
2. Origin-Independent Replication
2.1. Break-Induced Replication
2.2. Transcription-Initiated Replication
3. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Fuller, R.S.; Funnell, B.E.; Kornberg, A. The DnaA protein complex with the E. coli chromosomal replication origin (oriC) and other DNA sites. Cell 1984, 38, 889–900. [Google Scholar] [CrossRef]
- Hwang, D.S.; Kornberg, A. Opening of the replication origin of Escherichia coli by DnaA protein with protein HU or IHF. J. Biol. Chem. 1992, 267, 23083–23086. [Google Scholar] [PubMed]
- Boye, E.; Lobner-Olesen, A.; Skarstad, K. Limiting DNA replication to once and only once. EMBO Rep. 2000, 1, 479–483. [Google Scholar] [CrossRef] [PubMed]
- Mott, M.L.; Berger, J.M. DNA replication initiation: Mechanisms and regulation in bacteria. Nat. Rev. Microbiol. 2007, 5, 343–354. [Google Scholar] [CrossRef] [PubMed]
- Wolanski, M.; Donczew, R.; Zawilak-Pawlik, A.; Zakrzewska-Czerwinska, J. OriC-encoded instructions for the initiation of bacterial chromosome replication. Front. Microbiol. 2014, 5, 735. [Google Scholar] [PubMed]
- Kohara, Y.; Akiyama, K.; Isono, K. The physical map of the whole E. coli chromosome: Application of a new strategy for rapid analysis and sorting of a large genomic library. Cell 1987, 50, 495–508. [Google Scholar] [CrossRef]
- Speck, C.; Messer, W. Mechanism of origin unwinding: Sequential binding of DnaA to double- and single-stranded DNA. EMBO J. 2001, 20, 1469–1476. [Google Scholar] [CrossRef] [PubMed]
- Wold, S.; Skarstad, K.; Steen, H.B.; Stokke, T.; Boye, E. The initiation mass for DNA replication in Escherichia coli K-12 is dependent on growth rate. EMBO J. 1994, 13, 2097–2102. [Google Scholar] [PubMed]
- Yamaki, H.; Ohtsubo, E.; Nagai, K.; Maeda, Y. The oriC unwinding by dam methylation in Escherichia coli. Nucleic Acids Res. 1988, 16, 5067–5073. [Google Scholar] [CrossRef] [PubMed]
- Campbell, J.L.; Kleckner, N. E. coli oriC and the dnaA gene promoter are sequestered from dam methyltransferase following the passage of the chromosomal replication fork. Cell 1990, 62, 967–979. [Google Scholar] [CrossRef]
- Boye, E.; Stokke, T.; Kleckner, N.; Skarstad, K. Coordinating DNA replication initiation with cell growth: Differential roles for DnaA and SeqA proteins. Proc. Natl. Acad. Sci. USA 1996, 93, 12206–12211. [Google Scholar] [CrossRef] [PubMed]
- Torheim, N.K.; Boye, E.; Løbner-Olesen, A.; Stokke, T.; Skarstad, K. The Escherichia coli SeqA protein destabilizes mutant DnaA204 protein. Mol. Microbiol. 2000, 37, 629–638. [Google Scholar] [CrossRef] [PubMed]
- Fujimitsu, K.; Senriuchi, T.; Katayama, T. Specific genomic sequences of E. coli promote replicational initiation by directly reactivating ADP-DnaA. Genes Dev. 2009, 23, 1221–1233. [Google Scholar] [CrossRef] [PubMed]
- Bramhill, D.; Kornberg, A. Duplex opening by DnaA protein at novel sequences in initiation of replication at the origin of the E. coli chromosome. Cell 1988, 52, 743–755. [Google Scholar] [CrossRef]
- Chang, P.; Marians, K.J. Identification of a region of Escherichia coli DnaB required for functional interaction with DnaG at the replication fork. J. Biol. Chem. 2000, 275, 26187–26195. [Google Scholar] [CrossRef] [PubMed]
- Chodavarapu, S.; Kaguni, J.M. Replication initiation in bacteria. In The Enzymes; Academic Press: New York, NY, USA, 2016; Volume 39, Chapter 1; pp. 1–30. [Google Scholar]
- O’Donnell, M.E.; Kornberg, A. Complete replication of templates by Escherichia coli DNA polymerase III holoenzyme. J. Biol. Chem. 1985, 260, 12884–12889. [Google Scholar] [PubMed]
- Lewis, J.S.; Jergic, S.; Dixon, N.E. The E. coli DNA replication fork. In The Enzymes; Academic Press: New York, NY, USA, 2016; Volume 39, Chapter 2; pp. 31–88. [Google Scholar]
- Gilbert, D.M. Replication origins in yeast versus metazoa: Separation of the haves and the have nots. Curr. Opin. Genet. Dev. 1998, 8, 194–199. [Google Scholar] [CrossRef]
- Yamashita, M.; Hori, Y.; Shinomiya, T.; Obuse, C.; Tsurimoto, T.; Yoshikawa, H.; Shirahige, K. The efficiency and timing of initiation of replication of multiple replicons of Saccharomyces cerevisiae chromosome VI. Genes Cells 1997, 2, 655–665. [Google Scholar] [CrossRef] [PubMed]
- Das, S.P.; Borrman, T.; Liu, V.W.; Yang, S.C.; Bechhoefer, J.; Rhind, N. Replication timing is regulated by the number of MCMs loaded at origins. Genome Res. 2015, 25, 1886–1892. [Google Scholar] [CrossRef] [PubMed]
- Peace, J.M.; Ter-Zakarian, A.; Aparicio, O.M. Rif1 regulates initiation timing of late replication origins throughout the S. cerevisiae genome. PLoS ONE 2014, 9, e98501. [Google Scholar] [CrossRef] [PubMed]
- Goren, A.; Cedar, H. Replicating by the clock. Nat. Rev. Mol. Cell Biol. 2003, 4, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Stinchcomb, D.T.; Struhl, K.; Davis, R.W. Isolation and characterisation of a yeast chromosomal replicator. Nature 1979, 282, 39–43. [Google Scholar] [CrossRef] [PubMed]
- Bell, S.P.; Stillman, B. ATP-dependent recognition of eukaryotic origins of DNA replication by a multiprotein complex. Nature 1992, 357, 128–134. [Google Scholar] [CrossRef] [PubMed]
- Marahrens, Y.; Stillman, B. A yeast chromosomal origin of DNA replication defined by multiple functional elements. Science 1992, 255, 817–823. [Google Scholar] [CrossRef] [PubMed]
- Nieduszynski, C.A.; Knox, Y.; Donaldson, A.D. Genome-wide identification of replication origins in yeast by comparative genomics. Genes Dev. 2006, 20, 1874–1879. [Google Scholar] [CrossRef] [PubMed]
- Rao, H.; Stillman, B. The origin recognition complex interacts with a bipartite DNA binding site within yeast replicators. Proc. Natl. Acad. Sci. USA 1995, 92, 2224–2228. [Google Scholar] [CrossRef] [PubMed]
- Speck, C.; Chen, Z.; Li, H.; Stillman, B. ATPase-dependent cooperative binding of ORC and Cdc6 to origin DNA. Nat. Struct. Mol. Biol. 2005, 12, 965–971. [Google Scholar] [CrossRef] [PubMed]
- Aparicio, O.M.; Weinstein, D.M.; Bell, S.P. Components and dynamics of DNA replication complexes in S. cerevisiae: Redistribution of MCM proteins and Cdc45p during S phase. Cell 1997, 91, 59–69. [Google Scholar] [CrossRef]
- Kawasaki, Y.; Kim, H.D.; Kojima, A.; Seki, T.; Sugino, A. Reconstitution of Saccharomyces cerevisiae prereplicative complex assembly in vitro. Genes Cells 2006, 11, 745–756. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, S.; Diffley, J.F. Interdependent nuclear accumulation of budding yeast Cdt1 and Mcm2–7 during G1 phase. Nat. Cell Biol. 2002, 4, 198–207. [Google Scholar] [CrossRef] [PubMed]
- Chesnokov, I.N. Multiple functions of the origin recognition complex. Int. Rev. Cytol. 2007, 256, 69–109. [Google Scholar] [PubMed]
- Tanaka, S.; Umemori, T.; Hirai, K.; Muramatsu, S.; Kamimura, Y.; Araki, H. CDK-dependent phosphorylation of Sld2 and Sld3 initiates DNA replication in budding yeast. Nature 2007, 445, 328–332. [Google Scholar] [CrossRef] [PubMed]
- Sheu, Y.J.; Stillman, B. Cdc7-Dbf4 phosphorylates MCM proteins via a docking site-mediated mechanism to promote S phase progression. Mol. Cell 2006, 24, 101–113. [Google Scholar] [CrossRef] [PubMed]
- Zou, L.; Stillman, B. Formation of a preinitiation complex by S-phase cyclin CDK-dependent loading of Cdc45p onto chromatin. Science 1998, 280, 593–596. [Google Scholar] [CrossRef] [PubMed]
- Muramatsu, S.; Hirai, K.; Tak, Y.S.; Kamimura, Y.; Araki, H. CDK-dependent complex formation between replication proteins Dpb11, Sld2, Polε, and GINS in budding yeast. Genes Dev. 2010, 24, 602–612. [Google Scholar] [CrossRef] [PubMed]
- Homesley, L.; Lei, M.; Kawasaki, Y.; Sawyer, S.; Christensen, T.; Tye, B.K. Mcm10 and the Mcm2–7 complex interact to initiate DNA synthesis and to release replication factors from origins. Genes Dev. 2000, 14, 913–926. [Google Scholar] [PubMed]
- Gambus, A.; Jones, R.C.; Sanchez-Diaz, A.; Kanemaki, M.; van Deursen, F.; Edmondson, R.D.; Labib, K. GINS maintains association of Cdc45 with MCM in replisome progression complexes at eukaryotic DNA replication forks. Nat. Cell Biol. 2006, 8, 358–366. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, S.; Araki, H. Helicase activation and establishment of replication forks at chromosomal origins of replication. Cold Spring Harb. Perspect. Biol. 2013, 5, a010371. [Google Scholar] [CrossRef] [PubMed]
- Moyer, S.E.; Lewis, P.W.; Botchan, M.R. Isolation of the Cdc45/Mcm2–7/GINS (CMG) complex, a candidate for the eukaryotic DNA replication fork helicase. Proc. Natl. Acad. Sci. USA 2006, 103, 10236–10241. [Google Scholar] [CrossRef] [PubMed]
- Foiani, M.; Marini, F.; Gamba, D.; Lucchini, G.; Plevani, P. The B subunit of the DNA polymerase alpha-primase complex in Saccharomyces cerevisiae executes an essential function at the initial stage of DNA replication. Mol. Cell. Biol. 1994, 14, 923–933. [Google Scholar] [CrossRef] [PubMed]
- Garg, P.; Burgers, P.M. DNA polymerases that propagate the eukaryotic DNA replication fork. Crit. Rev. Biochem. Mol. Biol. 2005, 40, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Weinreich, M.; Liang, C.; Chen, H.H.; Stillman, B. Binding of cyclin-dependent kinases to ORC and Cdc6p regulates the chromosome replication cycle. Proc. Natl. Acad. Sci. USA 2001, 98, 11211–11217. [Google Scholar] [CrossRef] [PubMed]
- Labib, K.; Diffley, J.F.; Kearsey, S.E. G1-phase and B-type cyclins exclude the DNA-replication factor Mcm4 from the nucleus. Nat. Cell Biol. 1999, 1, 415–422. [Google Scholar] [PubMed]
- Nguyen, V.Q.; Co, C.; Li, J.J. Cyclin-dependent kinases prevent DNA re-replication through multiple mechanisms. Nature 2001, 411, 1068–1073. [Google Scholar] [CrossRef] [PubMed]
- Moll, T.; Tebb, G.; Surana, U.; Robitsch, H.; Nasmyth, K. The role of phosphorylation and the CDC28 protein kinase in cell cycle-regulated nuclear import of the S. cerevisiae transcription factor SWI5. Cell 1991, 66, 743–758. [Google Scholar] [CrossRef]
- Drury, L.S.; Perkins, G.; Diffley, J.F. The Cdc4/34/53 pathway targets Cdc6p for proteolysis in budding yeast. EMBO J. 1997, 16, 5966–5976. [Google Scholar] [CrossRef] [PubMed]
- Drury, L.S.; Perkins, G.; Diffley, J.F. The cyclin-dependent kinase Cdc28p regulates distinct modes of Cdc6p proteolysis during the budding yeast cell cycle. Curr. Biol. 2000, 10, 231–240. [Google Scholar] [CrossRef]
- Honey, S.; Futcher, B. Roles of the CDK Phosphorylation Sites of Yeast Cdc6 in Chromatin Binding and Rereplication. Mol. Biol. Cell 2007, 18, 1324–1336. [Google Scholar] [CrossRef] [PubMed]
- Archambault, V.; Ikui, A.E.; Drapkin, B.J.; Cross, F.R. Disruption of mechanisms that prevent rereplication triggers a DNA damage response. Mol. Cell. Biol. 2005, 25, 6707–6721. [Google Scholar] [CrossRef] [PubMed]
- Green, B.M.; Morreale, R.J.; Ozaydin, B.; Derisi, J.L.; Li, J.J. Genome-wide mapping of DNA synthesis in Saccharomyces cerevisiae reveals that mechanisms preventing reinitiation of DNA replication are not redundant. Mol. Biol. Cell 2006, 17, 2401–2414. [Google Scholar] [CrossRef] [PubMed]
- Blow, J.J.; Dutta, A. Preventing re-replication of chromosomal DNA. Nat. Rev. Mol. Cell Biol. 2005, 6, 476–486. [Google Scholar] [CrossRef] [PubMed]
- Vassilev, A.; Lee, C.Y.; Vassilev, B.; Zhu, W.; Ormanoglu, P.; Martin, S.E.; DePamphilis, M.L. Identification of genes that are essential to restrict genome duplication to once per cell division. Oncotarget 2016, 7, 34956–34976. [Google Scholar] [CrossRef] [PubMed]
- Kasamatsu, H.; Robberson, D.L.; Vinograd, J. A novel closed-circular mitochondrial DNA with properties of a replicating intermediate. Proc. Natl. Acad. Sci. USA 1971, 68, 2252–2257. [Google Scholar] [CrossRef] [PubMed]
- Aguilera, A.; Garcia-Muse, T. R loops: From transcription byproducts to threats to genome stability. Mol. Cell 2012, 46, 115–124. [Google Scholar] [CrossRef] [PubMed]
- Lujan, S.A.; Williams, J.S.; Kunkel, T.A. DNA polymerases divide the labor of genome replication. Trends Cell Biol. 2016, 26, 640–654. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Clayton, D.A. RNA-DNA hybrid formation at the human mitochondrial heavy-strand origin ceases at replication start sites: An implication for RNA-DNA hybrids serving as primers. EMBO J. 1996, 15, 3135–3143. [Google Scholar] [PubMed]
- Nicholls, T.J.; Minczuk, M. In D-loop: 40 years of mitochondrial 7S DNA. Exp. Gerontol. 2014, 56, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Baldacci, G.; Cherif-Zahar, B.; Bernardi, G. The initiation of DNA replication in the mitochondrial genome of yeast. EMBO J. 1984, 3, 2115–2120. [Google Scholar] [PubMed]
- Shadel, G.S. Yeast as a model for human mtDNA replication. Am. J. Hum. Genet. 1999, 65, 1230–1237. [Google Scholar] [CrossRef] [PubMed]
- Williamson, D. The curious history of yeast mitochondrial DNA. Nat. Rev. Genet. 2002, 3, 475–481. [Google Scholar] [PubMed]
- Wanrooij, P.H.; Uhler, J.P.; Shi, Y.; Westerlund, F.; Falkenberg, M.; Gustafsson, C.M. A hybrid G-quadruplex structure formed between RNA and DNA explains the extraordinary stability of the mitochondrial R-loop. Nucleic Acids Res. 2012, 40, 10334–10344. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Sandoval, E.; Diaz-Quezada, C.; Velazquez, G.; Arroyo-Navarro, L.F.; Almanza-Martinez, N.; Trasvina-Arenas, C.H.; Brieba, L.G. Yeast mitochondrial RNA polymerase primes mitochondrial DNA polymerase at origins of replication and promoter sequences. Mitochondrion 2015, 24, 22–31. [Google Scholar] [CrossRef] [PubMed]
- Sedman, T.; Gaidutsik, I.; Villemson, K.; Hou, Y.; Sedman, J. Double-stranded DNA-dependent ATPase Irc3p is directly involved in mitochondrial genome maintenance. Nucleic Acids Res. 2014, 42, 13214–13227. [Google Scholar] [CrossRef] [PubMed]
- Van Dyck, E.; Foury, F.; Stillman, B.; Brill, S.J. A single-stranded DNA binding protein required for mitochondrial DNA replication in S. cerevisiae is homologous to E. coli SSB. EMBO J. 1992, 11, 3421–3430. [Google Scholar] [PubMed]
- Holmes, J.B.; Akman, G.; Wood, S.R.; Sakhuja, K.; Cerritelli, S.M.; Moss, C.; Bowmaker, M.R.; Jacobs, H.T.; Crouch, R.J.; Holt, I.J. Primer retention owing to the absence of RNase H1 is catastrophic for mitochondrial DNA replication. Proc. Natl. Acad. Sci. USA 2015, 112, 9334–9339. [Google Scholar] [CrossRef] [PubMed]
- Cerritelli, S.M.; Frolova, E.G.; Feng, C.; Grinberg, A.; Love, P.E.; Crouch, R.J. Failure to produce mitochondrial DNA results in embryonic lethality in Rnaseh1 null mice. Mol. Cell 2003, 11, 807–815. [Google Scholar] [CrossRef]
- Holt, I.J. Mitochondrial DNA replication and repair: All a flap. Trends Biochem. Sci. 2009, 34, 358–365. [Google Scholar] [CrossRef] [PubMed]
- Ciesielski, G.L.; Oliveira, M.T.; Kaguni, L.S. Animal mitochondrial DNA replication. In The Enzymes; Academic Press: New York, NY, USA, 2016; Volume 39, Chapter 8; pp. 255–292. [Google Scholar]
- Gerhold, J.M.; Aun, A.; Sedman, T.; Joers, P.; Sedman, J. Strand invasion structures in the inverted repeat of Candida albicans mitochondrial DNA reveal a role for homologous recombination in replication. Mol. Cell 2010, 39, 851–861. [Google Scholar] [CrossRef] [PubMed]
- Gustafsson, C.M.; Falkenberg, M.; Larsson, N.G. Maintenance and expression of mammalian mitochondrial DNA. Annu. Rev. Biochem. 2016, 85, 133–160. [Google Scholar] [CrossRef] [PubMed]
- Clayton, D.A. Replication of animal mitochondrial DNA. Cell 1982, 28, 693–705. [Google Scholar] [CrossRef]
- Holt, I.J.; Lorimer, H.E.; Jacobs, H.T. Coupled leading- and lagging-strand synthesis of mammalian mitochondrial DNA. Cell 2000, 100, 515–524. [Google Scholar] [CrossRef]
- Yasukawa, T.; Reyes, A.; Cluett, T.J.; Yang, M.Y.; Bowmaker, M.; Jacobs, H.T.; Holt, I.J. Replication of vertebrate mitochondrial DNA entails transient ribonucleotide incorporation throughout the lagging strand. EMBO J. 2006, 25, 5358–5371. [Google Scholar] [CrossRef] [PubMed]
- Fangman, W.L.; Henly, J.W.; Churchill, G.; Brewer, B.J. Stable maintenance of a 35-base-pair yeast mitochondrial genome. Mol. Cell. Biol. 1989, 9, 1917–1921. [Google Scholar] [CrossRef] [PubMed]
- Fangman, W.L.; Henly, J.W.; Brewer, B.J. RPO41-independent maintenance of [rho-] mitochondrial DNA in Saccharomyces cerevisiae. Mol. Cell. Biol. 1990, 10, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Lorimer, H.E.; Brewer, B.J.; Fangman, W.L. A test of the transcription model for biased inheritance of yeast mitochondrial DNA. Mol. Cell. Biol. 1995, 15, 4803–4809. [Google Scholar] [CrossRef] [PubMed]
- Ling, F.; Hori, A.; Shibata, T. DNA recombination-initiation plays a role in the extremely biased inheritance of yeast [rho-] mitochondrial DNA that contains the replication origin ori5. Mol. Cell. Biol. 2007, 27, 1133–1145. [Google Scholar] [CrossRef] [PubMed]
- Ling, F.; Hori, A.; Yoshitani, A.; Niu, R.; Yoshida, M.; Shibata, T. Din7 and Mhr1 expression levels regulate double-strand-break-induced replication and recombination of mtDNA at ori5 in yeast. Nucleic Acids Res. 2013, 41, 5799–5816. [Google Scholar] [CrossRef] [PubMed]
- Engel, M.L.; Hines, J.C.; Ray, D.S. The Crithidia fasciculata RNH1 gene encodes both nuclear and mitochondrial isoforms of RNase H. Nucleic Acids Res. 2001, 29, 725–731. [Google Scholar] [CrossRef] [PubMed]
- Crider, D.G.; Garcia-Rodriguez, L.J.; Srivastava, P.; Peraza-Reyes, L.; Upadhyaya, K.; Boldogh, I.R.; Pon, L.A. Rad53 is essential for a mitochondrial DNA inheritance checkpoint regulating G1 to S progression. J. Cell Biol. 2012, 198, 793–798. [Google Scholar] [CrossRef] [PubMed]
- Futami, K.; Shimamoto, A.; Furuichi, Y. Mitochondrial and nuclear localization of human Pif1 helicase. Biol. Pharm. Bull. 2007, 30, 1685–1692. [Google Scholar] [CrossRef] [PubMed]
- Sanders, C.M. Human Pif1 helicase is a G-quadruplex DNA-binding protein with G-quadruplex DNA-unwinding activity. Biochem. J. 2010, 430, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Bochman, M.L.; Sabouri, N.; Zakian, V.A. Unwinding the functions of the Pif1 family helicases. DNA Repair (Amst) 2010, 9, 237–249. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.; Liu, Y. Borrowing nuclear DNA helicases to protect mitochondrial DNA. Int. J. Mol. Sci. 2015, 16, 10870–10887. [Google Scholar] [CrossRef] [PubMed]
- Anand, R.P.; Lovett, S.T.; Haber, J.E. Break-induced DNA replication. Cold Spring Harb. Perspect. Biol. 2013, 5, a010397. [Google Scholar] [CrossRef] [PubMed]
- Asai, T.; Sommer, S.; Bailone, A.; Kogoma, T. Homologous recombination-dependent initiation of DNA replication from DNA damage-inducible origins in Escherichia coli. EMBO J. 1993, 12, 3287–3295. [Google Scholar] [PubMed]
- Asai, T.; Bates, D.B.; Kogoma, T. DNA replication triggered by double-stranded breaks in E. coli: Dependence on homologous recombination functions. Cell 1994, 78, 1051–1061. [Google Scholar] [CrossRef]
- Kogoma, T. Stable DNA replication: Interplay between DNA replication, homologous recombination, and transcription. Microbiol. Mol. Biol. Rev. 1997, 61, 212–238. [Google Scholar] [PubMed]
- Kuzminov, A.; Stahl, F.W. Double-strand end repair via the RecBC pathway in Escherichia coli primes DNA replication. Genes Dev. 1999, 13, 345–356. [Google Scholar] [CrossRef] [PubMed]
- Magee, T.R.; Kogoma, T. Requirement of RecBC enzyme and an elevated level of activated RecA for induced stable DNA replication in Escherichia coli. J. Bacteriol. 1990, 172, 1834–1839. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Xu, L.; Sandler, S.J.; Marians, K.J. Replication fork assembly at recombination intermediates is required for bacterial growth. Proc. Natl. Acad. Sci. USA 1999, 96, 3552–3555. [Google Scholar] [CrossRef] [PubMed]
- Gabbai, C.B.; Marians, K.J. Recruitment to stalled replication forks of the PriA DNA helicase and replisome-loading activities is essential for survival. DNA Repair (Amst) 2010, 9, 202–209. [Google Scholar] [CrossRef] [PubMed]
- Asai, T.; Imai, M.; Kogoma, T. DNA damage-inducible replication of the Escherichia coli chromosome is initiated at separable sites within the minimal oriC. J. Mol. Biol. 1994, 235, 1459–1469. [Google Scholar] [CrossRef] [PubMed]
- Voelkel-Meiman, K.; Roeder, G.S. Gene conversion tracts stimulated by HOT1-promoted transcription are long and continuous. Genetics 1990, 126, 851–867. [Google Scholar] [PubMed]
- Morrow, D.M.; Connelly, C.; Hieter, P. “Break copy” duplication: A model for chromosome fragment formation in Saccharomyces cerevisiae. Genetics 1997, 147, 371–382. [Google Scholar] [PubMed]
- Davis, A.P.; Symington, L.S. RAD51-dependent break-induced replication in yeast. Mol. Cell. Biol. 2004, 24, 2344–2351. [Google Scholar] [CrossRef] [PubMed]
- Lydeard, J.R.; Jain, S.; Yamaguchi, M.; Haber, J.E. Break-induced replication and telomerase-independent telomere maintenance require Pol32. Nature 2007, 448, 820–823. [Google Scholar] [CrossRef] [PubMed]
- Kraus, E.; Leung, W.Y.; Haber, J.E. Break-induced replication: A review and an example in budding yeast. Proc. Natl. Acad. Sci. USA 2001, 98, 8255–8262. [Google Scholar] [CrossRef] [PubMed]
- Malkova, A.; Ira, G. Break-induced replication: Functions and molecular mechanism. Curr. Opin. Genet. Dev. 2013, 23, 271–279. [Google Scholar] [CrossRef] [PubMed]
- Malkova, A.; Naylor, M.L.; Yamaguchi, M.; Ira, G.; Haber, J.E. RAD51-dependent break-induced replication differs in kinetics and checkpoint responses from RAD51-mediated gene conversion. Mol. Cell. Biol. 2005, 25, 933–944. [Google Scholar] [CrossRef] [PubMed]
- Malkova, A.; Signon, L.; Schaefer, C.B.; Naylor, M.L.; Theis, J.F.; Newlon, C.S.; Haber, J.E. RAD51-independent break-induced replication to repair a broken chromosome depends on a distant enhancer site. Genes Dev. 2001, 15, 1055–1060. [Google Scholar] [CrossRef] [PubMed]
- Godin, S.K.; Sullivan, M.R.; Bernstein, K.A. Novel insights into RAD51 activity and regulation during homologous recombination and DNA replication. Biochem. Cell Biol. 2016, 94, 407–418. [Google Scholar] [CrossRef] [PubMed]
- Ira, G.; Haber, J.E. Characterization of RAD51-independent break-induced replication that acts preferentially with short homologous sequences. Mol. Cell. Biol. 2002, 22, 6384–6392. [Google Scholar] [CrossRef] [PubMed]
- Verma, P.; Greenberg, R.A. Noncanonical views of homology-directed DNA repair. Genes Dev. 2016, 30, 1138–1154. [Google Scholar] [CrossRef] [PubMed]
- Donnianni, R.A.; Symington, L.S. Break-induced replication occurs by conservative DNA synthesis. Proc. Natl. Acad. Sci. USA 2013, 110, 13475–13480. [Google Scholar] [CrossRef] [PubMed]
- Jain, S.; Sugawara, N.; Lydeard, J.; Vaze, M.; Tanguy Le Gac, N.; Haber, J.E. A recombination execution checkpoint regulates the choice of homologous recombination pathway during DNA double-strand break repair. Genes Dev. 2009, 23, 291–303. [Google Scholar] [CrossRef] [PubMed]
- McEachern, M.J.; Haber, J.E. Break-induced replication and recombinational telomere elongation in yeast. Annu. Rev. Biochem. 2006, 75, 111–135. [Google Scholar] [CrossRef] [PubMed]
- Lydeard, J.R.; Lipkin-Moore, Z.; Sheu, Y.J.; Stillman, B.; Burgers, P.M.; Haber, J.E. Break-induced replication requires all essential DNA replication factors except those specific for pre-RC assembly. Genes Dev. 2010, 24, 1133–1144. [Google Scholar] [CrossRef] [PubMed]
- Wilson, M.A.; Kwon, Y.; Xu, Y.; Chung, W.H.; Chi, P.; Niu, H.; Mayle, R.; Chen, X.; Malkova, A.; Sung, P.; et al. Pif1 helicase and Polδ promote recombination-coupled DNA synthesis via bubble migration. Nature 2013, 502, 393–396. [Google Scholar] [CrossRef] [PubMed]
- Saini, N.; Ramakrishnan, S.; Elango, R.; Ayyar, S.; Zhang, Y.; Deem, A.; Ira, G.; Haber, J.E.; Lobachev, K.S.; Malkova, A. Migrating bubble during break-induced replication drives conservative DNA synthesis. Nature 2013, 502, 389–392. [Google Scholar] [CrossRef] [PubMed]
- Fangman, W.L.; Brewer, B.J. Activation of replication origins within yeast chromosomes. Annu. Rev. Cell Biol. 1991, 7, 375–402. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Sterling, J.; Storici, F.; Resnick, M.A.; Gordenin, D.A. Hypermutability of damaged single-strand DNA formed at double-strand breaks and uncapped telomeres in yeast Saccharomyces cerevisiae. PLoS Genet. 2008, 4, e1000264. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.E.; Llorente, B.; Symington, L.S. Template switching during break-induced replication. Nature 2007, 447, 102–105. [Google Scholar] [CrossRef] [PubMed]
- Llorente, B.; Smith, C.E.; Symington, L.S. Break-induced replication: What is it and what is it for? Cell Cycle 2008, 7, 859–864. [Google Scholar] [CrossRef] [PubMed]
- Mayle, R.; Campbell, I.M.; Beck, C.R.; Yu, Y.; Wilson, M.; Shaw, C.A.; Bjergbaek, L.; Lupski, J.R.; Ira, G. DNA REPAIR. Mus81 and converging forks limit the mutagenicity of replication fork breakage. Science 2015, 349, 742–747. [Google Scholar] [CrossRef] [PubMed]
- Pardo, B.; Aguilera, A. Complex chromosomal rearrangements mediated by break-induced replication involve structure-selective endonucleases. PLoS Genet. 2012, 8, e1002979. [Google Scholar] [CrossRef] [PubMed]
- Lovett, S.T. Connecting replication and recombination. Mol. Cell 2003, 11, 554–556. [Google Scholar] [CrossRef]
- Heller, R.C.; Marians, K.J. Replisome assembly and the direct restart of stalled replication forks. Nat. Rev. Mol. Cell Biol. 2006, 7, 932–943. [Google Scholar] [CrossRef] [PubMed]
- Sandler, S.J.; Marians, K.J. Role of PriA in replication fork reactivation in Escherichia coli. J. Bacteriol. 2000, 182, 9–13. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Zhao, X. A new MCM modification cycle regulates DNA replication initiation. Nat. Struct. Mol. Biol. 2016, 23, 209–216. [Google Scholar] [CrossRef] [PubMed]
- Maric, M.; Maculins, T.; De Piccoli, G.; Labib, K. Cdc48 and a ubiquitin ligase drive disassembly of the CMG helicase at the end of DNA replication. Science 2014, 346, 1253596. [Google Scholar] [CrossRef] [PubMed]
- Moreno, S.P.; Bailey, R.; Campion, N.; Herron, S.; Gambus, A. Polyubiquitylation drives replisome disassembly at the termination of DNA replication. Science 2014, 346, 477–481. [Google Scholar] [CrossRef] [PubMed]
- Makovets, S.; Blackburn, E.H. DNA damage signalling prevents deleterious telomere addition at DNA breaks. Nat. Cell Biol. 2009, 11, 1383–1386. [Google Scholar] [CrossRef] [PubMed]
- Vasianovich, Y.; Harrington, L.A.; Makovets, S. Break-induced replication requires DNA damage-induced phosphorylation of Pif1 and leads to telomere lengthening. PLoS Genet. 2014, 10, e1004679. [Google Scholar] [CrossRef] [PubMed]
- Aguilera, A.; Garcia-Muse, T. Causes of genome instability. Annu. Rev. Genet. 2013, 47, 1–32. [Google Scholar] [CrossRef] [PubMed]
- Costantino, L.; Koshland, D. The Yin and Yang of R-loop biology. Curr. Opin. Cell Biol. 2015, 34, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Hamperl, S.; Cimprich, K.A. The contribution of co-transcriptional RNA:DNA hybrid structures to DNA damage and genome instability. DNA Repair (Amst) 2014, 19, 84–94. [Google Scholar] [CrossRef] [PubMed]
- Groh, M.; Gromak, N. Out of balance: R-loops in human disease. PLoS Genet. 2014, 10, e1004630. [Google Scholar] [CrossRef] [PubMed]
- Santos-Pereira, J.M.; Aguilera, A. R loops: New modulators of genome dynamics and function. Nat. Rev. Genet. 2015, 16, 583–597. [Google Scholar] [CrossRef] [PubMed]
- Skourti-Stathaki, K.; Kamieniarz-Gdula, K.; Proudfoot, N.J. R-loops induce repressive chromatin marks over mammalian gene terminators. Nature 2014, 516, 436–439. [Google Scholar] [CrossRef] [PubMed]
- Gaillard, H.; Aguilera, A. Transcription as a threat to genome integrity. Annu. Rev. Biochem. 2016, 85, 291–317. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, S.; Masukata, H.; Tomizawa, J. Multiple mechanisms for initiation of ColE1 DNA replication: DNA synthesis in the presence and absence of ribonuclease H. Cell 1987, 51, 1113–1122. [Google Scholar] [CrossRef]
- Masukata, H.; Dasgupta, S.; Tomizawa, J. Transcriptional activation of ColE1 DNA synthesis by displacement of the nontranscribed strand. Cell 1987, 51, 1123–1130. [Google Scholar] [CrossRef]
- Marians, K.J. Prokaryotic DNA replication. Annu. Rev. Biochem. 1992, 61, 673–719. [Google Scholar] [CrossRef] [PubMed]
- Mosig, G. The essential role of recombination in phage T4 growth. Annu. Rev. Genet. 1987, 21, 347–371. [Google Scholar] [CrossRef] [PubMed]
- de Massy, B.; Fayet, O.; Kogoma, T. Multiple origin usage for DNA replication in sdrA(rnh) mutants of Escherichia coli K-12. Initiation in the absence of oriC. J. Mol. Biol. 1984, 178, 227–236. [Google Scholar] [CrossRef]
- Kogoma, T. Absence of RNase H allows replication of pBR322 in Escherichia coli mutants lacking DNA polymerase I. Proc. Natl. Acad. Sci. USA 1984, 81, 7845–7849. [Google Scholar] [CrossRef] [PubMed]
- Von Meyenburg, K.; Boye, E.; Skarstad, K.; Koppes, L.; Kogoma, T. Mode of initiation of constitutive stable DNA replication in RNase H-defective mutants of Escherichia coli K-12. J. Bacteriol. 1987, 169, 2650–2658. [Google Scholar] [CrossRef] [PubMed]
- Horiuchi, T.; Maki, H.; Sekiguchi, M. RNase H-defective mutants of Escherichia coli: A possible discriminatory role of RNase H in initiation of DNA replication. Mol. Gen. Genet. 1984, 195, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, T.; Pickett, G.G.; Kogoma, T.; Kornberg, A. RNase H confers specificity in the DnaA-dependent initiation of replication at the unique origin of the Escherichia coli chromosome in vivo and in vitro. Proc. Natl. Acad. Sci. USA 1984, 81, 1040–1044. [Google Scholar] [CrossRef] [PubMed]
- Maduike, N.Z.; Tehranchi, A.K.; Wang, J.D.; Kreuzer, K.N. Replication of the Escherichia coli chromosome in RNase HI-deficient cells: Multiple initiation regions and fork dynamics. Mol. Microbiol. 2014, 91, 39–56. [Google Scholar] [CrossRef] [PubMed]
- Dimude, J.U.; Stockum, A.; Midgley-Smith, S.L.; Upton, A.L.; Foster, H.A.; Khan, A.; Saunders, N.J.; Retkute, R.; Rudolph, C.J. The consequences of replicating in the wrong orientation: Bacterial chromosome duplication without an active replication origin. MBio 2015, 6, e01294-15. [Google Scholar] [CrossRef] [PubMed]
- Hong, X.; Cadwell, G.W.; Kogoma, T. Escherichia coli RecG and RecA proteins in R-loop formation. EMBO J. 1995, 14, 2385–2392. [Google Scholar] [PubMed]
- Hong, X.; Cadwell, G.W.; Kogoma, T. Activation of stable DNA replication in rapidly growing Escherichia coli at the time of entry to stationary phase. Mol. Microbiol. 1996, 21, 953–961. [Google Scholar] [CrossRef] [PubMed]
- Drolet, M.; Phoenix, P.; Menzel, R.; Masse, E.; Liu, L.F.; Crouch, R.J. Overexpression of RNase H partially complements the growth defect of an Escherichia coli delta topA mutant: R-loop formation is a major problem in the absence of DNA topoisomerase I. Proc. Natl. Acad. Sci. USA 1995, 92, 3526–3530. [Google Scholar] [CrossRef] [PubMed]
- Martel, M.; Balleydier, A.; Sauriol, A.; Drolet, M. Constitutive stable DNA replication in Escherichia coli cells lacking type 1A topoisomerase activity. DNA Repair (Amst) 2015, 35, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Sandler, S.J. Requirements for replication restart proteins during constitutive stable DNA replication in Escherichia coli K-12. Genetics 2005, 169, 1799–1806. [Google Scholar] [CrossRef] [PubMed]
- Usongo, V.; Drolet, M. Roles of type 1A topoisomerases in genome maintenance in Escherichia coli. PLoS Genet. 2014, 10, e1004543. [Google Scholar] [CrossRef] [PubMed]
- Kasahara, M.; Clikeman, J.A.; Bates, D.B.; Kogoma, T. RecA protein-dependent R-loop formation in vitro. Genes Dev. 2000, 14, 360–365. [Google Scholar] [PubMed]
- Zaitsev, E.N.; Kowalczykowski, S.C. A novel pairing process promoted by Escherichia coli RecA protein: Inverse DNA and RNA strand exchange. Genes Dev. 2000, 14, 740–749. [Google Scholar] [PubMed]
- Wimberly, H.; Shee, C.; Thornton, P.C.; Sivaramakrishnan, P.; Rosenberg, S.M.; Hastings, P.J. R-loops and nicks initiate DNA breakage and genome instability in non-growing Escherichia coli. Nat. Commun. 2013, 4, 2115. [Google Scholar] [CrossRef] [PubMed]
- Usongo, V.; Martel, M.; Balleydier, A.; Drolet, M. Mutations reducing replication from R-loops suppress the defects of growth, chromosome segregation and DNA supercoiling in cells lacking topoisomerase I and RNase HI activity. DNA Repair (Amst) 2016, 40, 1–17. [Google Scholar] [CrossRef] [PubMed]
- El Hage, A.; Webb, S.; Kerr, A.; Tollervey, D. Genome-wide distribution of RNA-DNA hybrids identifies RNase H targets in tRNA genes, retrotransposons and mitochondria. PLoS Genet. 2014, 10, e1004716. [Google Scholar] [CrossRef] [PubMed]
- Chan, Y.A.; Aristizabal, M.J.; Lu, P.Y.; Luo, Z.; Hamza, A.; Kobor, M.S.; Stirling, P.C.; Hieter, P. Genome-wide profiling of yeast DNA:RNA hybrid prone sites with DRIP-chip. PLoS Genet. 2014, 10, e1004288. [Google Scholar] [CrossRef] [PubMed]
- Mischo, H.E.; Gomez-Gonzalez, B.; Grzechnik, P.; Rondon, A.G.; Wei, W.; Steinmetz, L.; Aguilera, A.; Proudfoot, N.J. Yeast Sen1 helicase protects the genome from transcription-associated instability. Mol. Cell 2011, 41, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Alzu, A.; Bermejo, R.; Begnis, M.; Lucca, C.; Piccini, D.; Carotenuto, W.; Saponaro, M.; Brambati, A.; Cocito, A.; Foiani, M.; et al. Senataxin associates with replication forks to protect fork integrity across RNA-polymerase-II-transcribed genes. Cell 2012, 151, 835–846. [Google Scholar] [CrossRef] [PubMed]
- Salvi, J.S.; Chan, J.N.; Szafranski, K.; Liu, T.T.; Wu, J.D.; Olsen, J.B.; Khanam, N.; Poon, B.P.; Emili, A.; Mekhail, K. Roles for Pbp1 and caloric restriction in genome and lifespan maintenance via suppression of RNA-DNA hybrids. Dev. Cell 2014, 30, 177–191. [Google Scholar] [CrossRef] [PubMed]
- El Hage, A.; French, S.L.; Beyer, A.L.; Tollervey, D. Loss of Topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev. 2010, 24, 1546–1558. [Google Scholar] [CrossRef] [PubMed]
- Stuckey, R.; Garcia-Rodriguez, N.; Aguilera, A.; Wellinger, R.E. Role for RNA:DNA hybrids in origin-independent replication priming in a eukaryotic system. Proc. Natl. Acad. Sci. USA 2015, 112, 5779–5784. [Google Scholar] [CrossRef] [PubMed]
- Pomerantz, R.T.; O’Donnell, M. The replisome uses mRNA as a primer after colliding with RNA polymerase. Nature 2008, 456, 762–766. [Google Scholar] [CrossRef] [PubMed]
- Pomerantz, R.T.; O’Donnell, M. Direct restart of a replication fork stalled by a head-on RNA polymerase. Science 2010, 327, 590–592. [Google Scholar] [CrossRef] [PubMed]
- Longhese, M.P.; Plevani, P.; Lucchini, G. Replication factor A is required in vivo for DNA replication, repair, and recombination. Mol. Cell. Biol. 1994, 14, 7884–7890. [Google Scholar] [CrossRef] [PubMed]
- Bartrand, A.J.; Iyasu, D.; Brush, G.S. DNA stimulates Mec1-mediated phosphorylation of replication protein A. J. Biol. Chem. 2004, 279, 26762–26767. [Google Scholar] [CrossRef] [PubMed]
- Foiani, M.; Liberi, G.; Lucchini, G.; Plevani, P. Cell cycle-dependent phosphorylation and dephosphorylation of the yeast DNA polymerase alpha-primase B subunit. Mol. Cell. Biol. 1995, 15, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Rippe, K.; Luke, B. TERRA and the state of the telomere. Nat. Struct. Mol. Biol. 2015, 22, 853–858. [Google Scholar] [CrossRef] [PubMed]
- Balk, B.; Maicher, A.; Dees, M.; Klermund, J.; Luke-Glaser, S.; Bender, K.; Luke, B. Telomeric RNA-DNA hybrids affect telomere-length dynamics and senescence. Nat. Struct. Mol. Biol. 2013, 20, 1199–1205. [Google Scholar] [CrossRef] [PubMed]
- Arora, R.; Lee, Y.; Wischnewski, H.; Brun, C.M.; Schwarz, T.; Azzalin, C.M. RNase H1 regulates TERRA-telomeric DNA hybrids and telomere maintenance in ALT tumour cells. Nat. Commun. 2014, 5, 5220. [Google Scholar] [CrossRef] [PubMed]
- Flynn, R.L.; Cox, K.E.; Jeitany, M.; Wakimoto, H.; Bryll, A.R.; Ganem, N.J.; Bersani, F.; Pineda, J.R.; Suva, M.L.; Benes, C.H.; et al. Alternative lengthening of telomeres renders cancer cells hypersensitive to ATR inhibitors. Science 2015, 347, 273–277. [Google Scholar] [CrossRef] [PubMed]
- Law, M.J.; Lower, K.M.; Voon, H.P.; Hughes, J.R.; Garrick, D.; Viprakasit, V.; Mitson, M.; De Gobbi, M.; Marra, M.; Morris, A.; et al. ATR-X syndrome protein targets tandem repeats and influences allele-specific expression in a size-dependent manner. Cell 2010, 143, 367–378. [Google Scholar] [CrossRef] [PubMed]
- Kogoma, T.; Lark, K.G. Characterization of the replication of Escherichia coli DNA in the absence of protein synthesis: Stable DNA replication. J. Mol. Biol. 1975, 94, 243–256. [Google Scholar] [CrossRef]
- Lombrana, R.; Almeida, R.; Alvarez, A.; Gomez, M. R-loops and initiation of DNA replication in human cells: A missing link? Front. Genet. 2015, 6, 158. [Google Scholar] [CrossRef] [PubMed]
- Leonard, A.C.; Mechali, M. DNA replication origins. Cold Spring Harb. Perspect. Biol. 2013, 5, a010116. [Google Scholar] [CrossRef] [PubMed]
- Norseen, J.; Thomae, A.; Sridharan, V.; Aiyar, A.; Schepers, A.; Lieberman, P.M. RNA-dependent recruitment of the origin recognition complex. EMBO J. 2008, 27, 3024–3035. [Google Scholar] [CrossRef] [PubMed]
- Gomez, M.; Antequera, F. Overreplication of short DNA regions during S phase in human cells. Genes Dev. 2008, 22, 375–385. [Google Scholar] [CrossRef] [PubMed]
- Lombrana, R.; Alvarez, A.; Fernandez-Justel, J.M.; Almeida, R.; Poza-Carrion, C.; Gomes, F.; Calzada, A.; Requena, J.M.; Gomez, M. Transcriptionally driven DNA replication program of the human parasite Leishmania major. Cell Rep. 2016, 16, 1774–1786. [Google Scholar] [CrossRef] [PubMed]
| Origin-Dependent Replication | E. coli | S. cerevisiae | |
|---|---|---|---|
| Chromosomal DNA Replication | Chromosomal DNA Replication | Mitochondrial DNA Replication | |
| Origin | OriC | ARS | OriH, OriL |
| DNA unwinding | DnaA, DnaB, DnaC, SSB | Cdc45, GINS, Mcm2–7, Mcm10, RPA | Rpo41, Irc3, Rim1 |
| Replication priming/elongation | DnaG, DNA Pol III | DNA Pol-α-primase, DNA Pol-ε and Pol-δ | Rpo41, DNA Pol-γ |
| E. coli | Function | iSDR | cSDR |
| End processing | RecBCD | RecBCD | |
| Strand invasion | RecA | RecA | |
| DNA unwinding | DnaBC, PriAB | DnaBC, PriAB | |
| RecG | ? | ||
| DnaT | ? | ||
| Replication priming/elongation | DnaG, DNA Pol III | DnaG, DNA Pol I/Pol III | |
| Resolution | RuvABC | ? | |
| S. cerevisiae | Function | BIR | TIR |
| End processing | MRX (Mre11-Rad50-Xrs2) | ? | |
| Strand invasion | Rad51*, Rad52, Rad54, Rad55, Rad57 | ? | |
| DNA unwinding | Cdc45-MCM-GINS, DDK, Mcm10, Ctf4, RPA, Pif1 | RNA:DNA hybrid | |
| Replication priming/elongation | Pol-α-primase, Pol-δ, Pol32* | ? | |
| Resolution | Mus81-MMS4, Slx1–Slx4, Yen1 | ? |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ravoitytė, B.; Wellinger, R.E. Non‐Canonical Replication Initiation: You’re Fired! Genes 2017, 8, 54. https://doi.org/10.3390/genes8020054
Ravoitytė B, Wellinger RE. Non‐Canonical Replication Initiation: You’re Fired! Genes. 2017; 8(2):54. https://doi.org/10.3390/genes8020054
Chicago/Turabian StyleRavoitytė, Bazilė, and Ralf Erik Wellinger. 2017. "Non‐Canonical Replication Initiation: You’re Fired!" Genes 8, no. 2: 54. https://doi.org/10.3390/genes8020054
APA StyleRavoitytė, B., & Wellinger, R. E. (2017). Non‐Canonical Replication Initiation: You’re Fired! Genes, 8(2), 54. https://doi.org/10.3390/genes8020054