NAD1 Controls Defense-Like Responses in Medicago truncatula Symbiotic Nitrogen Fixing Nodules Following Rhizobial Colonization in a BacA-Independent Manner
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material, Growth Condition and Bacterial Strains
2.2. DNA Isolation and Genetic Mapping
2.3. Microscopy Analyses
2.4. Transmission and Scanning Electron Microscopy Analysis of the nad1-3 Mutant
2.5. Gene Expression Analysis
2.6. Generating Constructs and Complementation Experiments Accomplished by Agrobacterium rhizogenes Transformation
2.7. Transient Gene Expression in Nicotiana benthamiana Epidermal Cells
3. Results and Discussion
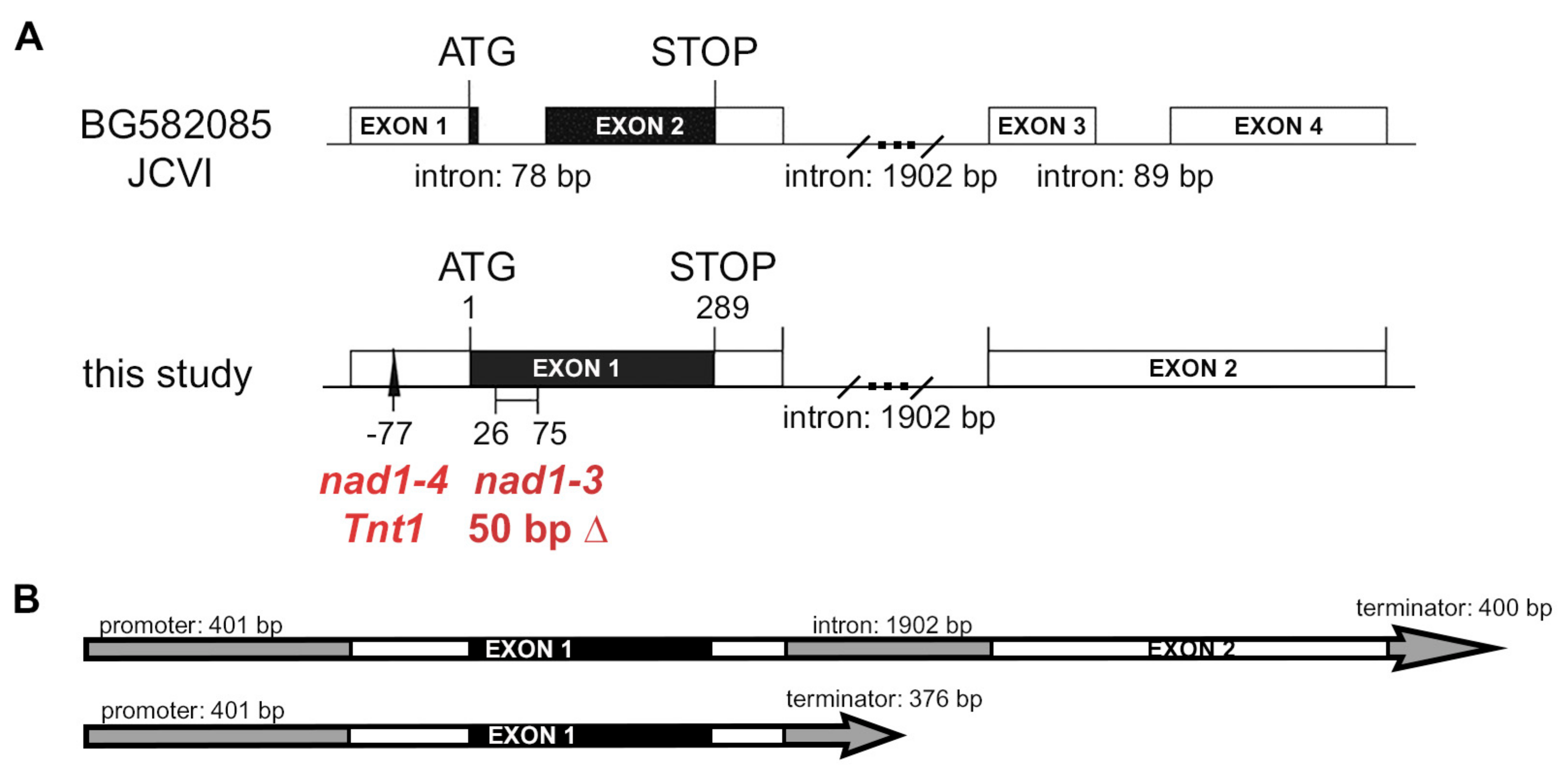
3.1. Mutants Defective in Symbiotic Nitrogen Fixation and Showing Induced Defense Responses Are Deficient in the NAD1 Gene
3.2. Both Exons of NAD1 Are Required for Complete Rescue of the Symbiotic Phenotype of nad1-3 and nad1-4 Mutants
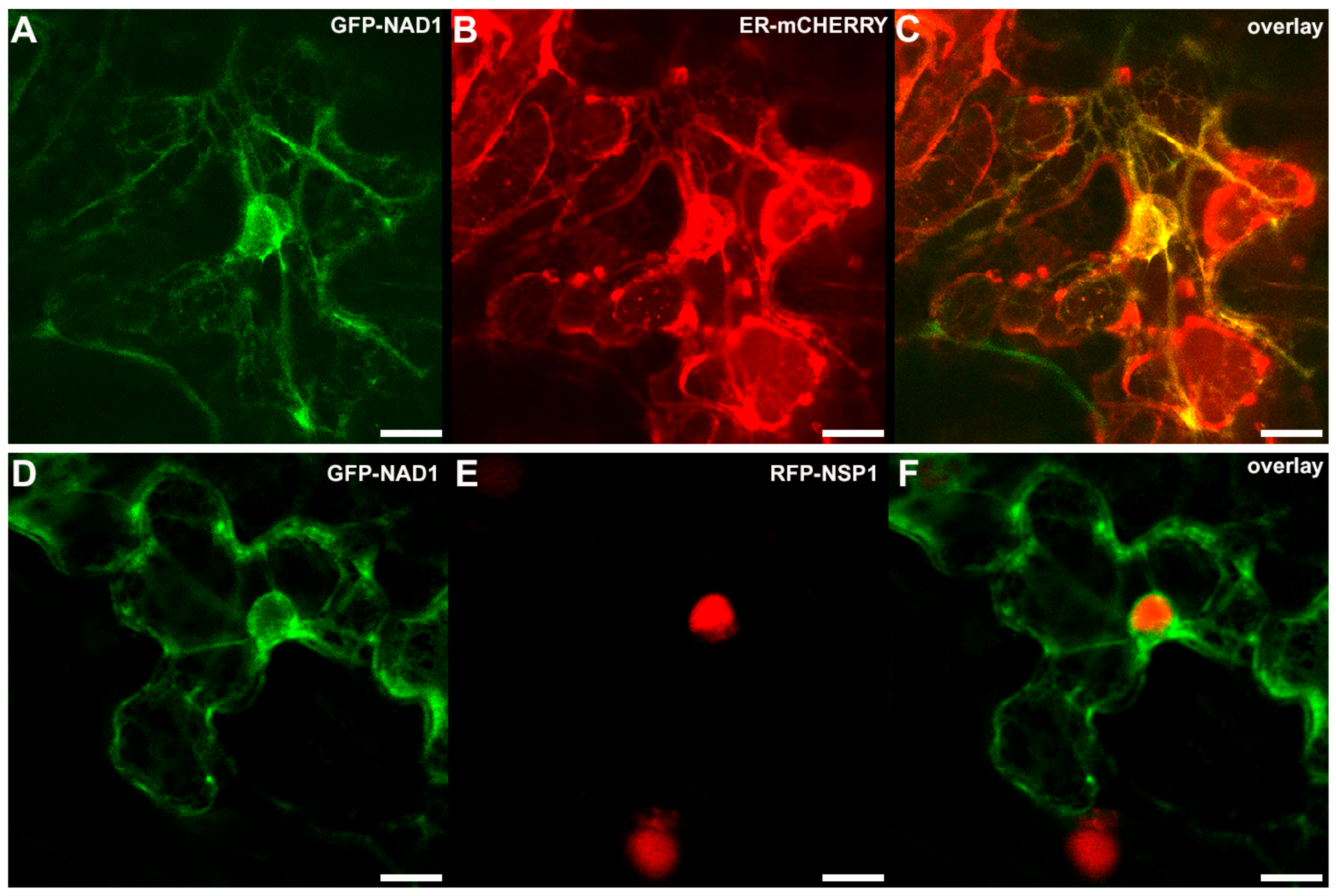
3.3. NAD1 Is Expressed in Infected Cells and the Gene Product Localizes to the Endoplasmic Reticulum
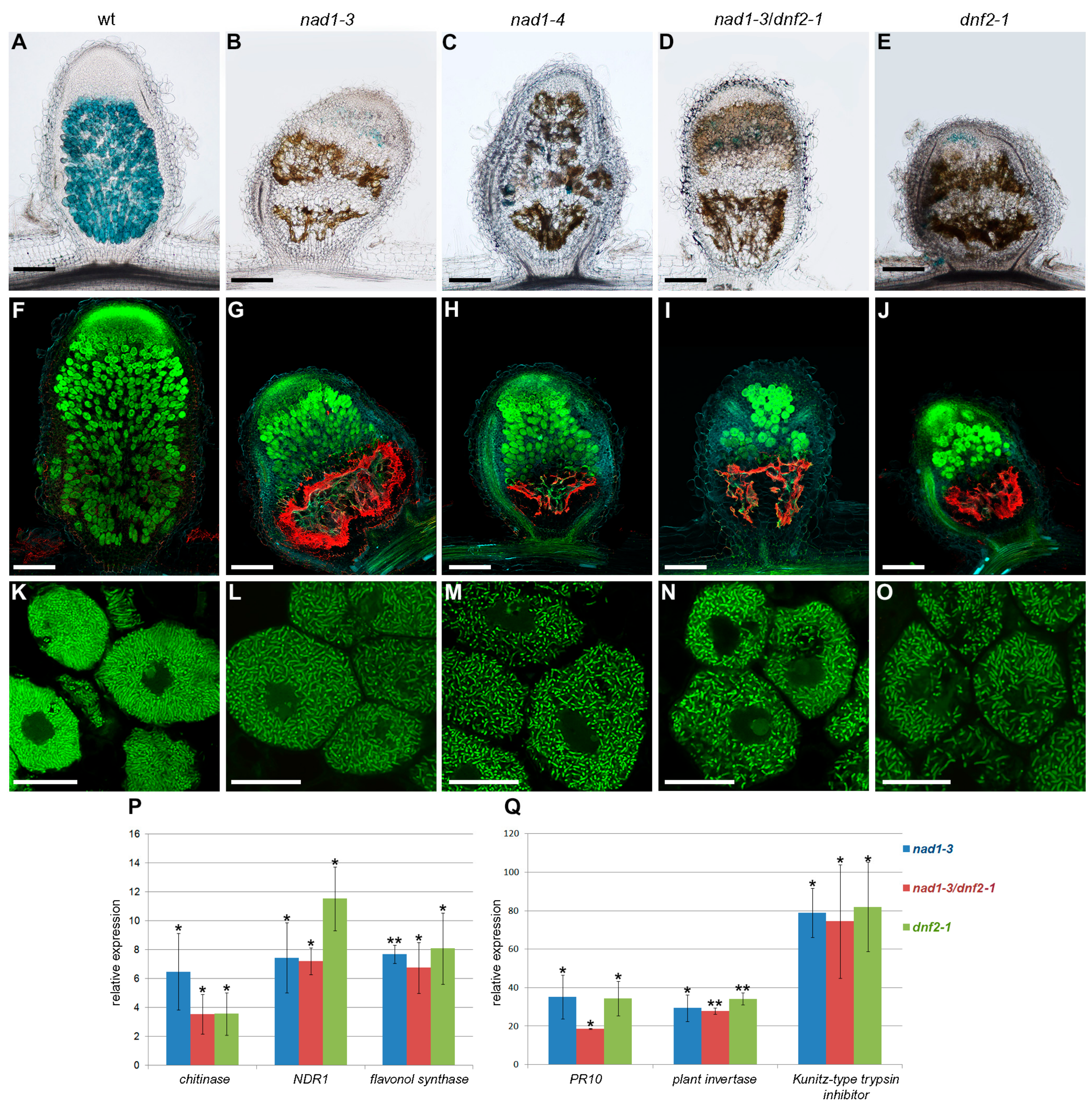
3.4. The Defense Responses Are Induced Simultaneously in nad1 and dnf2 Mutants
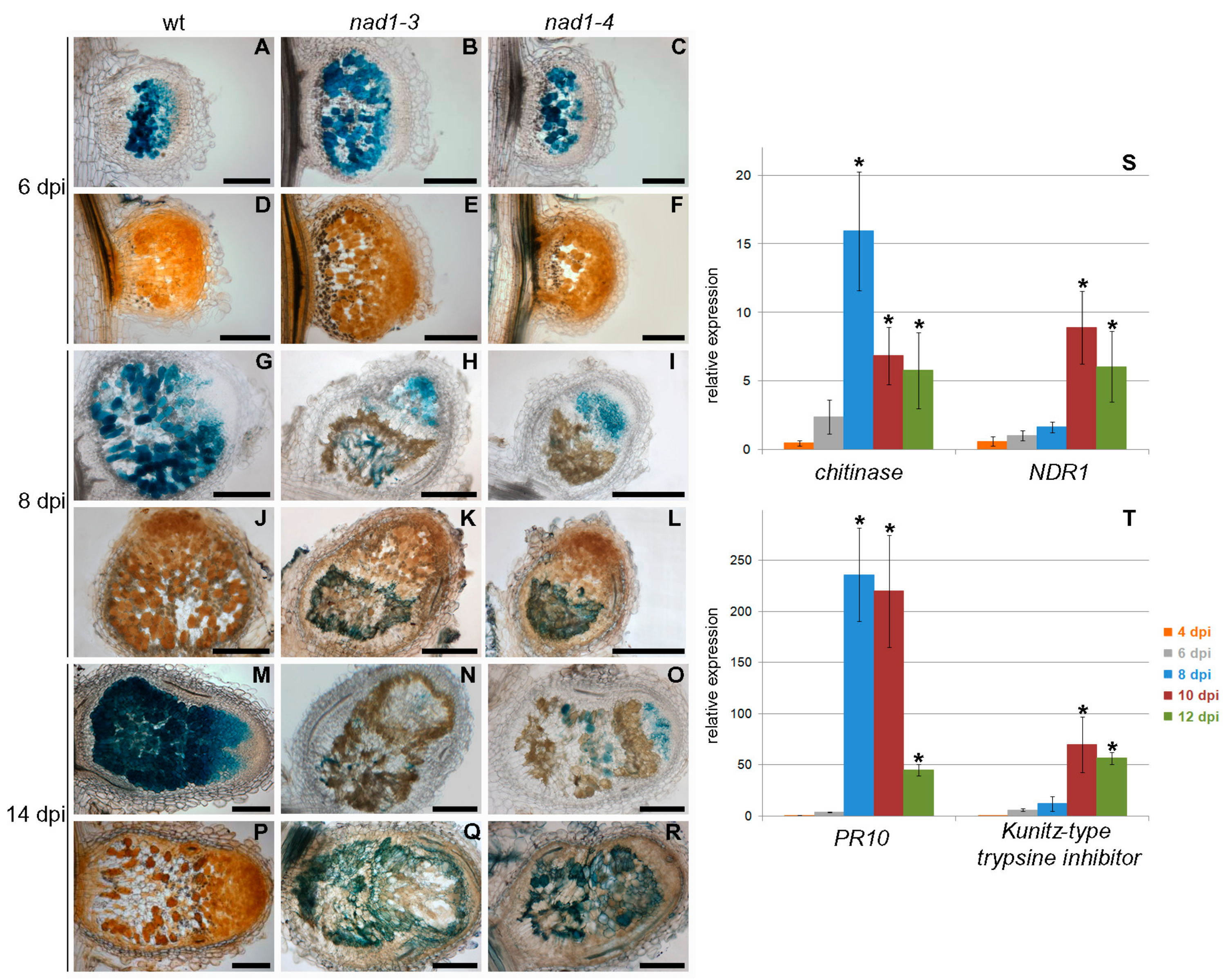
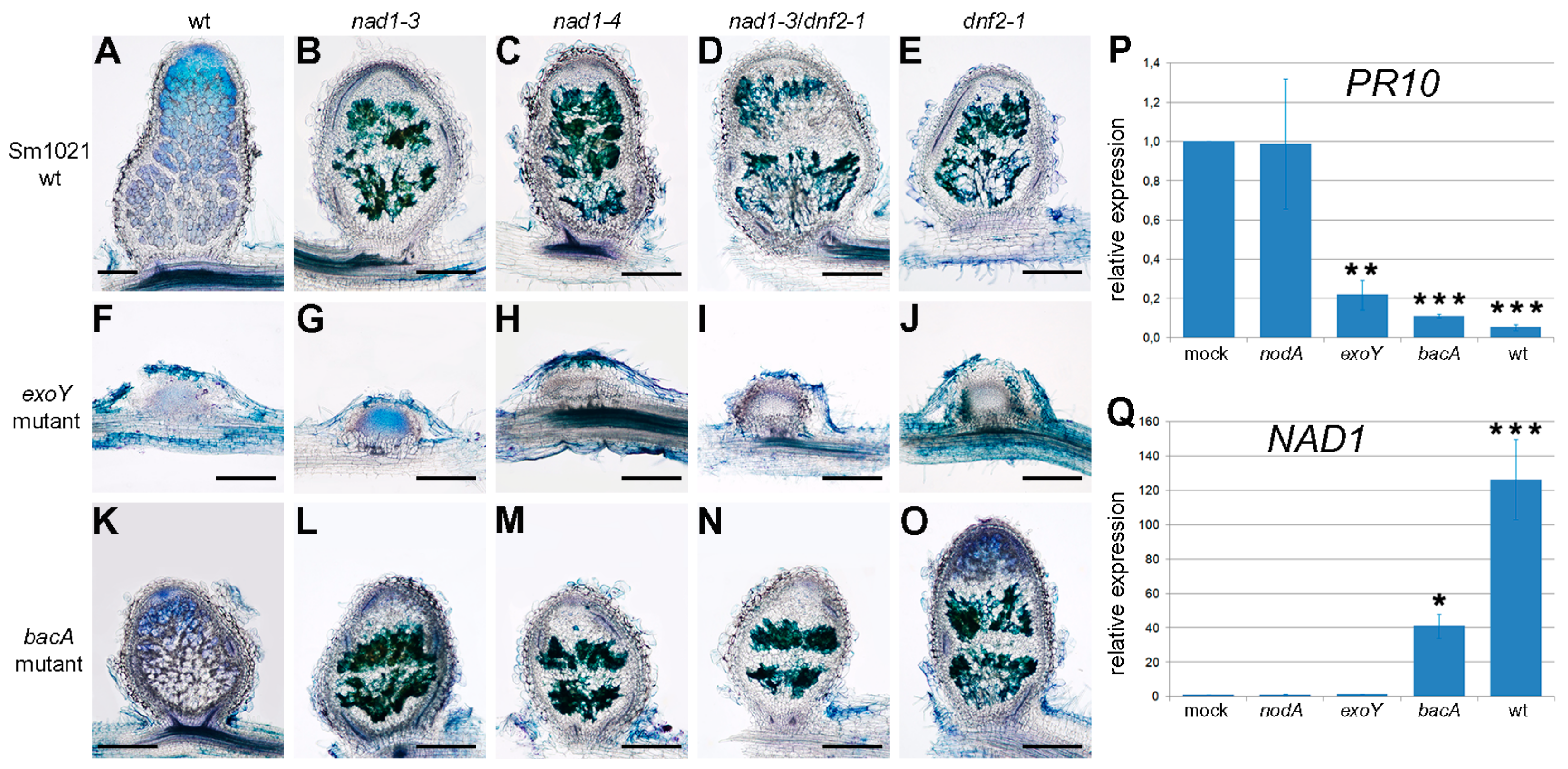
3.5. Defense-Like Responses Are Induced upon the Internalization of the Symbiotic Nodule Cells by Rhizobia and Rapidly Induce the Degradation of Host and Bacterial Cells
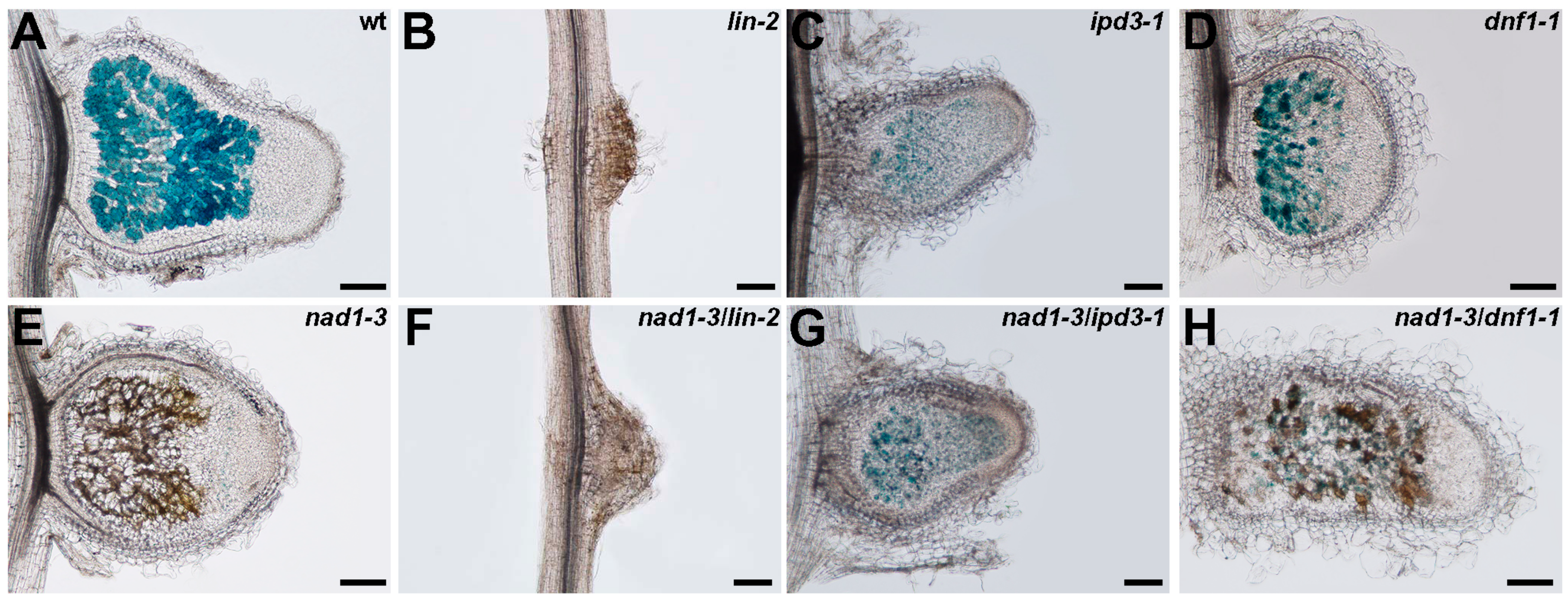
3.6. The Function of NAD1 Precedes DNF1 and NAD1 Acts Independently and Prior to Rhizobial BacA in the Symbiotic Process
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Oldroyd, G.E.D.; Downie, J.A. Coordinating nodule morphogenesis with rhizobial infection in legumes. Annu. Rev. Plant Biol. 2008, 59, 519–546. [Google Scholar] [CrossRef] [PubMed]
- Oldroyd, G.E.D.; Murray, J.D.; Poole, P.S.; Downie, J.A. The rules of engagement in the legume-rhizobial symbiosis. Annu. Rev. Genet. 2011, 45, 119–144. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.M.; Kobayashi, H.; Davies, B.W.; Taga, M.E.; Walker, G.C. How rhizobial symbionts invade plants: The Sinorhizobium-Medicago model. Nat. Rev. Microbiol. 2007, 5, 619–633. [Google Scholar] [CrossRef] [PubMed]
- Vasse, J.; de Billy, F.; Camut, S.; Truchet, G. Correlation between ultrastructural differentiation of bacteroids and nitrogen fixation in alfalfa nodules. J. Bacteriol. 1990, 172, 4295–4306. [Google Scholar] [CrossRef] [PubMed]
- Mergaert, P.; Uchiumi, T.; Alunni, B.; Evanno, G.; Cheron, A.; Catrice, O.; Mausset, A.E.; Barloy-Hubler, F.; Galibert, F.; Kondorosi, A.; et al. Eukaryotic control on bacterial cell cycle and differentiation in the Rhizobium-legume symbiosis. Proc. Natl. Acad. Sci. USA 2006, 103, 5230–5235. [Google Scholar] [CrossRef] [PubMed]
- Van de Velde, W.; Zehirov, G.; Szatmari, A.; Debreczeny, M.; Ishihara, H.; Kevei, Z.; Farkas, A.; Mikulass, K.; Nagy, A.; Tiricz, H.; et al. Plant peptides govern terminal differentiation of bacteria in symbiosis. Science 2010, 327, 1122–1126. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Griffitts, J.; Starker, C.; Fedorova, E.; Limpens, E.; Ivanov, S.; Bisseling, T.; Long, S. A nodule-specific protein secretory pathway required for nitrogen-fixing symbiosis. Science 2010, 327, 1126–1129. [Google Scholar] [CrossRef] [PubMed]
- Haag, A.F.; Baloban, M.; Sani, M.; Kerscher, B.; Pierre, O.; Farkas, A.; Longhi, R.; Boncompagni, E.; Herouart, D.; Dall’Angelo, S.; et al. Protection of Sinorhizobium against host cysteine-rich antimicrobial peptides is critical for symbiosis. PLoS Biol. 2011, 9, e1001169. [Google Scholar] [CrossRef] [PubMed]
- Libault, M.; Farmer, A.; Brechenmacher, L.; Drnevich, J.; Langley, R.J.; Bilgin, D.D.; Radwan, O.; Neece, D.J.; Clough, S.J.; May, G.D.; et al. Complete transcriptome of the soybean root hair cell, a single-cell model, and its alteration in response to Bradyrhizobium japonicum infection. Plant Physiol. 2010, 152, 541–552. [Google Scholar] [CrossRef] [PubMed]
- Lohar, D.P.; Sharopova, N.; Endre, G.; Penuela, S.; Samac, D.; Town, C.; Silverstein, K.A.T.; VandenBosch, K.A. Transcript analysis of early nodulation events in Medicago truncatula. Plant Physiol. 2006, 140, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Kouchi, H.; Shimomura, K.; Hata, S.; Hirota, A.; Wu, G.J.; Kumagai, H.; Tajima, S.; Suganuma, N.; Suzuki, A.; Aoki, T.; et al. Large-scale analysis of gene expression profiles during early stages of root nodule formation in a model legume, Lotus japonicus. DNA Res. 2004, 11, 263–274. [Google Scholar] [CrossRef] [PubMed]
- Sewelam, N.; Kazan, K.; Schenk, P.M. Global plant stress signaling: Reactive oxygen species at the cross-road. Front. Plant Sci. 2016, 7, 187. [Google Scholar] [CrossRef] [PubMed]
- Vasse, J.; Debilly, F.; Truchet, G. Abortion of infection during the Rhizobium meliloti-alfalfa symbiotic interaction is accompanied by a hypersensitive reaction. Plant J. 1993, 4, 555–566. [Google Scholar] [CrossRef]
- Jamet, A.; Mandon, K.; Puppo, A.; Herouart, D. H2O2 is required for optimal establishment of the Medicago sativa/Sinorhizobium meliloti symbiosis. J. Bacteriol. 2007, 189, 8741–8745. [Google Scholar] [CrossRef] [PubMed]
- Montiel, J.; Arthikala, M.K.; Cardenas, L.; Quinto, C. Legume NADPH oxidases have crucial roles at different stages of nodulation. Int. J. Mol. Sci. 2016, 17, 680. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.M.; Sharopova, N.; Lohar, D.P.; Zhang, J.Q.; VandenBosch, K.A.; Walker, G.C. Differential response of the plant Medicago truncatula to its symbiont Sinorhizobium meliloti or an exopolysaccharide-deficient mutant. Proc. Natl. Acad. Sci. USA 2008, 105, 704–709. [Google Scholar] [CrossRef] [PubMed]
- Niehaus, K.; Kapp, D.; Puhler, A. Plant defense and delayed infection of alfalfa pseudonodules induced by an exopolysaccharide (EPS-I)-deficient Rhizobium meliloti mutant. Planta 1993, 190, 415–425. [Google Scholar] [CrossRef]
- Cao, Y.R.; Halane, M.K.; Gassmann, W.; Stacey, G. The role of plant innate immunity in the legume-rhizobium symbiosis. Annu. Rev. Plant Biol. 2017, 68, 535–561. [Google Scholar] [CrossRef] [PubMed]
- Bourcy, M.; Brocard, L.; Pislariu, C.I.; Cosson, V.; Mergaert, P.; Tadege, M.; Mysore, K.S.; Udvardi, M.K.; Gourion, B.; Ratet, P. Medicago truncatula DNF2 is a PI-PLC-XD-containing protein required for bacteroid persistence and prevention of nodule early senescence and defense-like reactions. New Phytol. 2013, 197, 1250–1261. [Google Scholar] [CrossRef] [PubMed]
- Berrabah, F.; Bourcy, M.; Eschstruth, A.; Cayrel, A.; Guefrachi, I.; Mergaert, P.; Wen, J.Q.; Jean, V.; Mysore, K.S.; Gourion, B.; et al. A nonRD receptor-like kinase prevents nodule early senescence and defense-like reactions during symbiosis. New Phytol. 2014, 203, 1305–1314. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Yu, H.X.; Luo, L.; Duan, L.J.; Cai, L.Y.; He, X.X.; Wen, J.Q.; Mysore, K.S.; Li, G.L.; Xiao, A.F.; et al. Nodules with Activated Defense 1 is required for maintenance of rhizobial endosymbiosis in Medicago truncatula. New Phytol. 2016, 212, 176–191. [Google Scholar] [CrossRef] [PubMed]
- Sinharoy, S.; Torres-Jerez, I.; Bandyopadhyay, K.; Kereszt, A.; Pislariu, C.I.; Nakashima, J.; Benedito, V.A.; Kondorosi, E.; Udvardi, M.K. The C2H2 transcription factor Regulator of Symbiosome Differentiation represses transcription of the secretory pathway gene VAMP721a and promotes symbiosome development in Medicago truncatula. Plant Cell 2013, 25, 3584–3601. [Google Scholar] [CrossRef] [PubMed]
- Berrabah, F.; Ratet, P.; Gourion, B. Multiple steps control immunity during the intracellular accommodation of rhizobia. J. Exp. Bot. 2015, 66, 1977–1985. [Google Scholar] [CrossRef] [PubMed]
- Chabaud, M.; Lichtenzveig, J.; Ellwood, S.; Pfaff, T.; Journet, E. Vernalization, crossings and testing for pollen viability. In The Medicago Truncatula Handbook; Mathesius, U., Journet, E.P., Sumner, L.W., Eds.; The Samuel Roberts Noble Foundation: Ardmore, OK, USA, 2007; pp. 1–13. [Google Scholar]
- Auriac, M.; Timmers, A. Nodulation studies in the model legume Medicago truncatula: Advantages of using the constitutive EF1a promoter and limitations in detecting fluorescent reporter proteins in nodule tissues. Mol. Plant Microbe Interact. 2007, 20, 1040–1047. [Google Scholar] [CrossRef] [PubMed]
- Glazebrook, J.; Walker, G.C. Genetic techniques in Rhizobium meliloti. Methods Enzymol. 1991, 204, 398–418. [Google Scholar] [PubMed]
- Debelle, F.; Rosenberg, C.; Vasse, J.; Maillet, F.; Martinez, E.; Denarie, J.; Truchet, G. Assignment of symbiotic developmental phenotypes to common and specific nodulation (nod) genetic loci of Rhizobium meliloti. J. Bacteriol. 1986, 168, 1075–1086. [Google Scholar] [CrossRef] [PubMed]
- Martin, M.O.; Long, S.R. Generalized transduction in Rhizobium meliloti. J. Bacteriol. 1984, 159, 125–129. [Google Scholar] [PubMed]
- Ferguson, G.P.; Roop, R.M.; Walker, G.C. Deficiency of a Sinorhizobium meliloti bacA mutant in alfalfa symbiosis correlates with alteration of the cell envelope. J. Bacteriol. 2002, 184, 5625–5632. [Google Scholar] [CrossRef] [PubMed]
- Journet, E.; de Carvalho-Niebel, F.; Andriankaja, A.; Huguet, T.; Barker, D. Rhizobial inoculation and nodulation of Medicago truncatula. In The Medicago Truncatula Handbook; Mathesius, U., Journet, E.P., Sumner, L.W., Eds.; The Samuel Roberts Noble Foundation: Ardmore, OK, USA, 2007; pp. 1–6. [Google Scholar]
- Chabaud, M.; Boisson-Dernier, A.; Zhang, J.; Taylor, C.; Yu, O.; Barker, D. Agrobacterium rhizogenes-mediated root transformation. In The Medicago Truncatula Handbook; Mathesius, U., Journet, E.P., Sumner, L.W., Eds.; The Samuel Roberts Noble Foundation: Ardmore, OK, USA, 2007; pp. 1–8. [Google Scholar]
- Choi, H.K.; Kim, D.; Uhm, T.; Limpens, E.; Lim, H.; Mun, J.H.; Kalo, P.; Penmetsa, R.V.; Seres, A.; Kulikova, O.; et al. A sequence-based genetic map of Medicago truncatula and comparison of marker colinearity with M. sativa. Genetics 2004, 166, 1463–1502. [Google Scholar] [CrossRef] [PubMed]
- Mun, J.H.; Kim, D.J.; Choi, H.K.; Gish, J.; Debelle, F.; Mudge, J.; Denny, R.; Endre, G.; Saurat, O.; Dudez, A.M.; et al. Distribution of microsatellites in the genome of Medicago truncatula: A resource of genetic markers that integrate genetic and physical maps. Genetics 2006, 172, 2541–2555. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Russell, D. Molecular Cloning. A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2001. [Google Scholar]
- Domonkos, A.; Horvath, B.; Marsh, J.F.; Halasz, G.; Ayaydin, F.; Oldroyd, G.E.D.; Kalo, P. The identification of novel loci required for appropriate nodule development in Medicago truncatula. BMC Plant Biol. 2013, 13, 157. [Google Scholar] [CrossRef] [PubMed]
- Montiel, J.; Downie, J.A.; Farkas, A.; Bihari, P.; Herczeg, R.; Balint, B.; Mergaert, P.; Kereszt, A.; Kondorosi, E. Morphotype of bacteroids in different legumes correlates with the number and type of symbiotic NCR peptides. Proc. Natl. Acad. Sci. USA 2017, 114, 5041–5046. [Google Scholar] [CrossRef] [PubMed]
- Kakar, K.; Wandrey, M.; Czechowski, T.; Gaertner, T.; Scheible, W.; Stitt, M.; Torres-Jerez, I.; Xiao, Y.; Redman, J.; Wu, H.; et al. A community resource for high-throughput quantitative RT-PCR analysis of transcription factor gene expression in Medicago truncatula. Plant Methods 2008, 4, 18. [Google Scholar] [CrossRef] [PubMed]
- Horvath, B.; Domonkos, A.; Kereszt, A.; Szucs, A.; Abraham, E.; Ayaydin, F.; Boka, K.; Chen, Y.H.; Chen, R.J.; Murray, J.D.; et al. Loss of the nodule-specific cysteine rich peptide, NCR169, abolishes symbiotic nitrogen fixation in the Medicago truncatula dnf7 mutant. Proc. Natl. Acad. Sci. USA 2015, 112, 15232–15237. [Google Scholar] [CrossRef] [PubMed]
- Boisson-Dernier, A.; Chabaud, M.; Garcia, F.; Becard, G.; Rosenberg, C.; Barker, D. Agrobacterium rhizogenes-transformed roots of Medicago truncatula for the study of nitrogen-fixing and endomycorrhizal symbiotic associations. Mol. Plant Microbe Interact. 2001, 14, 695–700. [Google Scholar] [CrossRef] [PubMed]
- Smit, P.; Raedts, J.; Portyanko, V.; Debelle, F.; Gough, C.; Bisseling, T.; Geurts, R. NSP1 of the GRAS protein family is essential for rhizobial nod factor-induced transcription. Science 2005, 308, 1789–1791. [Google Scholar] [CrossRef] [PubMed]
- Curtis, M.D.; Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 2003, 133, 462–469. [Google Scholar] [CrossRef] [PubMed]
- Maniatis, T.; Fritsch, E.; Sambrook, J. Molecular Cloning: A Laboratory Manual; Cold Spring Laboratory Press: Cold Spring Harbor, NY, USA, 1982. [Google Scholar]
- Iantcheva, A.; Chabaud, M.; Cosson, V.; Barascud, M.; Schutz, B.; Primard-Brisset, C.; Durand, P.; Barker, D.; Vlahova, M.; Ratet, P. Osmotic shock improves Tnt1 transposition frequency in Medicago truncatula cv jemalong during in vitro regeneration. Plant Cell Rep. 2009, 28, 1563–1572. [Google Scholar] [CrossRef] [PubMed]
- Tadege, M.; Wen, J.Q.; He, J.; Tu, H.D.; Kwak, Y.; Eschstruth, A.; Cayrel, A.; Endre, G.; Zhao, P.X.; Chabaud, M.; et al. Large-scale insertional mutagenesis using the Tnt1 retrotransposon in the model legume Medicago truncatula. Plant J. 2008, 54, 335–347. [Google Scholar] [CrossRef] [PubMed]
- Starker, C.G.; Parra-Colmenares, A.L.; Smith, L.; Mitra, R.M.; Long, S.R. Nitrogen fixation mutants of Medicago truncatula fail to support plant and bacterial symbiotic gene expression. Plant Physiol. 2006, 140, 671–680. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.B.; Krishnakumar, V.; Bidwell, S.; Rosen, B.; Chan, A.N.; Zhou, S.G.; Gentzbittel, L.; Childs, K.L.; Yandell, M.; Gundlach, H.; et al. An improved genome release (version Mt4.0) for the model legume Medicago truncatula. BMC Genom. 2014, 15, 312. [Google Scholar] [CrossRef] [PubMed]
- Roux, B.; Rodde, N.; Jardinaud, M.F.; Timmers, T.; Sauviac, L.; Cottret, L.; Carrere, S.; Sallet, E.; Courcelle, E.; Moreau, S.; et al. An integrated analysis of plant and bacterial gene expression in symbiotic root nodules using laser-capture microdissection coupled to RNA sequencing. Plant J. 2014, 77, 817–837. [Google Scholar] [CrossRef] [PubMed]
- Nelson, B.K.; Cai, X.; Nebenfuhr, A. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J. 2007, 51, 1126–1136. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.W.; Brattain, M.G. The maximal size of protein to diffuse through the nuclear pore is larger than 60 kDa. FEBS Lett. 2007, 581, 3164–3170. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, D.S.; Blower, M.D. The endoplasmic reticulum: Structure, function and response to cellular signaling. Cell. Mol. Life Sci. 2016, 73, 79–94. [Google Scholar] [CrossRef] [PubMed]
- Eichmann, R.; Schafer, P. The endoplasmic reticulum in plant immunity and cell death. Front. Plant Sci. 2012, 3, 200. [Google Scholar] [CrossRef] [PubMed]
- Haynes, J.; Czymmek, K.; Carlson, C.; Veereshlingam, H.; Dickstein, R.; Sherrier, D. Rapid analysis of legume root nodule development using confocal microscopy. New Phytol. 2004, 163, 661–668. [Google Scholar] [CrossRef]
- Nicholson, R.L.; Hammerschmidt, R. Phenolic compounds and their role in disease resistance. Annu. Rev. Phytopathol. 1992, 30, 369–389. [Google Scholar] [CrossRef]
- Terpolilli, J.; O’Hara, G.; Tiwari, R.; Dilworth, M.; Howieson, J. The model legume Medicago truncatula A17 is poorly matched for N2 fixation with the sequenced microsymbiont Sinorhizobium meliloti 1021. New Phytol. 2008, 179, 62–66. [Google Scholar] [CrossRef] [PubMed]
- Leigh, J.A.; Signer, E.R.; Walker, G.C. Exopolysaccharide-deficient mutants of Rhizobium meliloti that form ineffective nodules. Proc. Natl. Acad. Sci. USA 1985, 82, 6231–6235. [Google Scholar] [CrossRef] [PubMed]
- Karunakaran, R.; Haag, A.F.; East, A.K.; Ramachandran, V.K.; Prell, J.; James, E.K.; Scocchi, M.; Ferguson, G.P.; Poole, P.S. BacA is essential for bacteroid development in nodules of galegoid, but not phaseoloid, legumes. J. Bacteriol. 2010, 192, 2920–2928. [Google Scholar] [CrossRef] [PubMed]
- Glazebrook, J.; Ichige, A.; Walker, G.C. A Rhizobium meliloti homolog of the Escherichia coli peptide-antibiotic transport protein SbmA is essential for bacteroid development. Genes Dev. 1993, 7, 1485–1497. [Google Scholar] [CrossRef] [PubMed]
- Debelle, F.; Plazanet, C.; Roche, P.; Pujol, C.; Savagnac, A.; Rosenberg, C.; Prome, J.C.; Denarie, J. The NodA proteins of Rhizobium meliloti and Rhizobium tropici specify the N-acylation of Nod factors by different fatty acids. Mol. Microbiol. 1996, 22, 303–314. [Google Scholar] [CrossRef] [PubMed]
- Kiss, E.; Olah, B.; Kalo, P.; Morales, M.; Heckmann, A.; Borbola, A.; Lozsa, A.; Kontar, K.; Middleton, P.; Downie, J.; et al. LIN, a novel type of U-Box/WD40 protein, controls early infection by rhizobia in legumes. Plant Physiol. 2009, 151, 1239–1249. [Google Scholar] [CrossRef] [PubMed]
- Horvath, B.; Yeun, L.H.; Domonkos, A.; Halasz, G.; Gobbato, E.; Ayaydin, F.; Miro, K.; Hirsch, S.; Sun, J.H.; Tadege, M.; et al. Medicago truncatula IPD3 is a member of the common symbiotic signaling pathway required for rhizobial and mycorrhizal symbioses. Mol. Plant Microbe Interact. 2011, 24, 1345–1358. [Google Scholar] [CrossRef] [PubMed]
- Gourion, B.; Berrabah, F.; Ratet, P.; Stacey, G. Rhizobium–legume symbioses: The crucial role of plant immunity. Trends Plant Sci. 2015, 20, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Zipfel, C.; Oldroyd, G.E.D. Plant signalling in symbiosis and immunity. Nature 2017, 543, 328–336. [Google Scholar] [CrossRef] [PubMed]

© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Domonkos, Á.; Kovács, S.; Gombár, A.; Kiss, E.; Horváth, B.; Kováts, G.Z.; Farkas, A.; Tóth, M.T.; Ayaydin, F.; Bóka, K.; et al. NAD1 Controls Defense-Like Responses in Medicago truncatula Symbiotic Nitrogen Fixing Nodules Following Rhizobial Colonization in a BacA-Independent Manner. Genes 2017, 8, 387. https://doi.org/10.3390/genes8120387
Domonkos Á, Kovács S, Gombár A, Kiss E, Horváth B, Kováts GZ, Farkas A, Tóth MT, Ayaydin F, Bóka K, et al. NAD1 Controls Defense-Like Responses in Medicago truncatula Symbiotic Nitrogen Fixing Nodules Following Rhizobial Colonization in a BacA-Independent Manner. Genes. 2017; 8(12):387. https://doi.org/10.3390/genes8120387
Chicago/Turabian StyleDomonkos, Ágota, Szilárd Kovács, Anikó Gombár, Ernő Kiss, Beatrix Horváth, Gyöngyi Z. Kováts, Attila Farkas, Mónika T. Tóth, Ferhan Ayaydin, Károly Bóka, and et al. 2017. "NAD1 Controls Defense-Like Responses in Medicago truncatula Symbiotic Nitrogen Fixing Nodules Following Rhizobial Colonization in a BacA-Independent Manner" Genes 8, no. 12: 387. https://doi.org/10.3390/genes8120387
APA StyleDomonkos, Á., Kovács, S., Gombár, A., Kiss, E., Horváth, B., Kováts, G. Z., Farkas, A., Tóth, M. T., Ayaydin, F., Bóka, K., Fodor, L., Ratet, P., Kereszt, A., Endre, G., & Kaló, P. (2017). NAD1 Controls Defense-Like Responses in Medicago truncatula Symbiotic Nitrogen Fixing Nodules Following Rhizobial Colonization in a BacA-Independent Manner. Genes, 8(12), 387. https://doi.org/10.3390/genes8120387





