A Spatial Control for Correct Timing of Gene Expression during the Escherichia coli Cell Cycle
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Plasmids
2.2. Growth Media and Conditions
2.3. Fluorescence Microscopy
2.4. Total RNA Extraction
2.5. Global Transcription Microarray Analysis
2.6. Relative Quantitative Real-Time PCR
2.7. Immunoprecipitation by GFP-Trap
2.8. SDS-PAGE and Mass Spectrometry Analysis
2.9. Bacterial Two Hybrid Analysis
2.10. Site-Directed Mutagenesis
3. Results
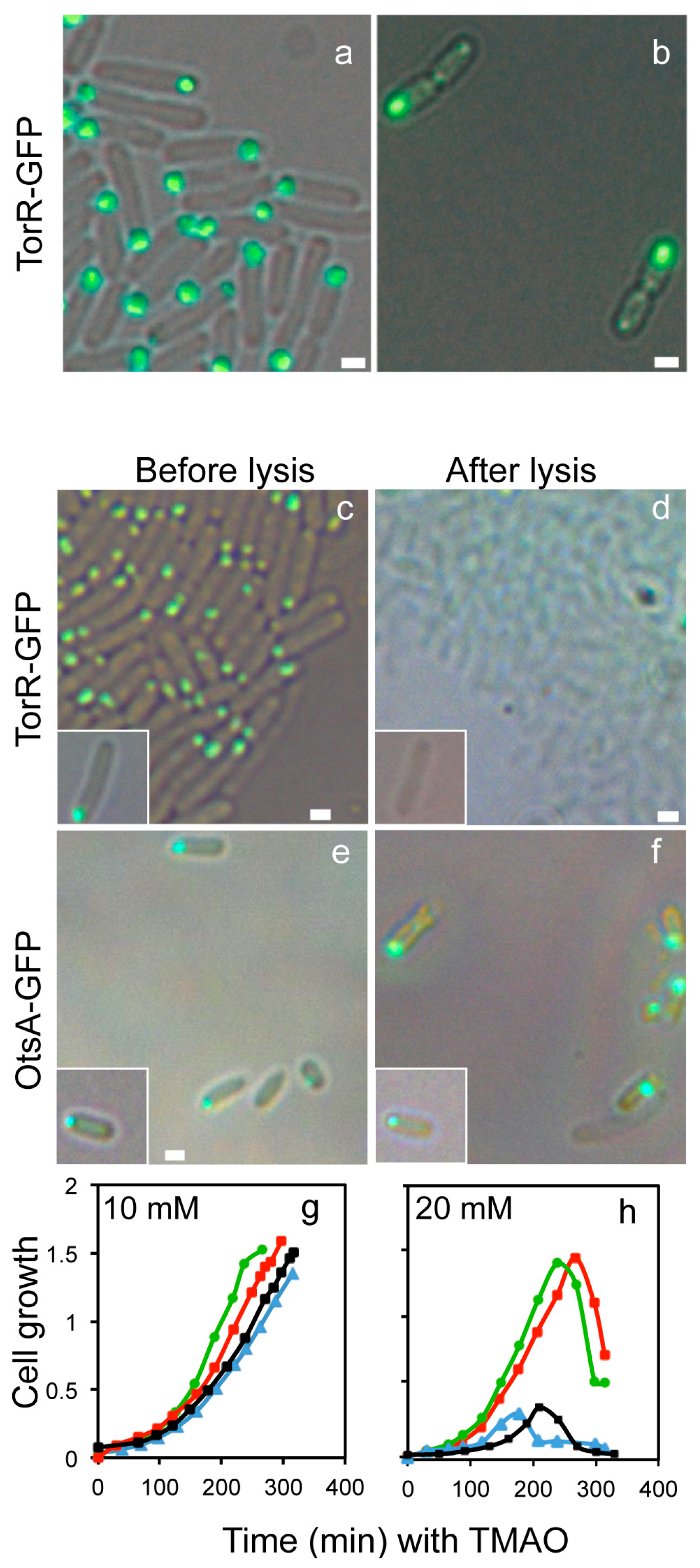
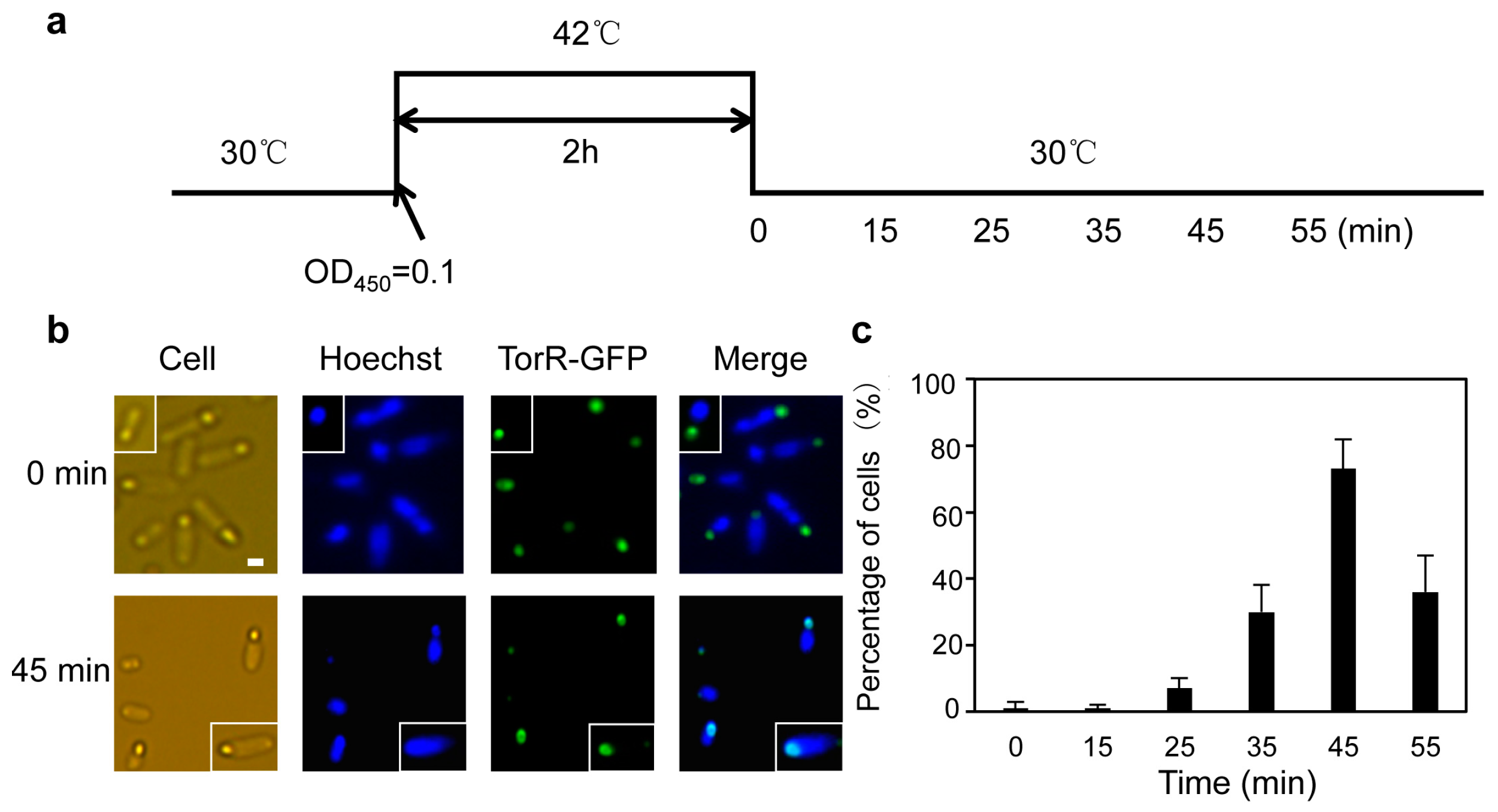
3.1. The TorR Protein Is Localized at the Old Cell Pole as a Functional Focus
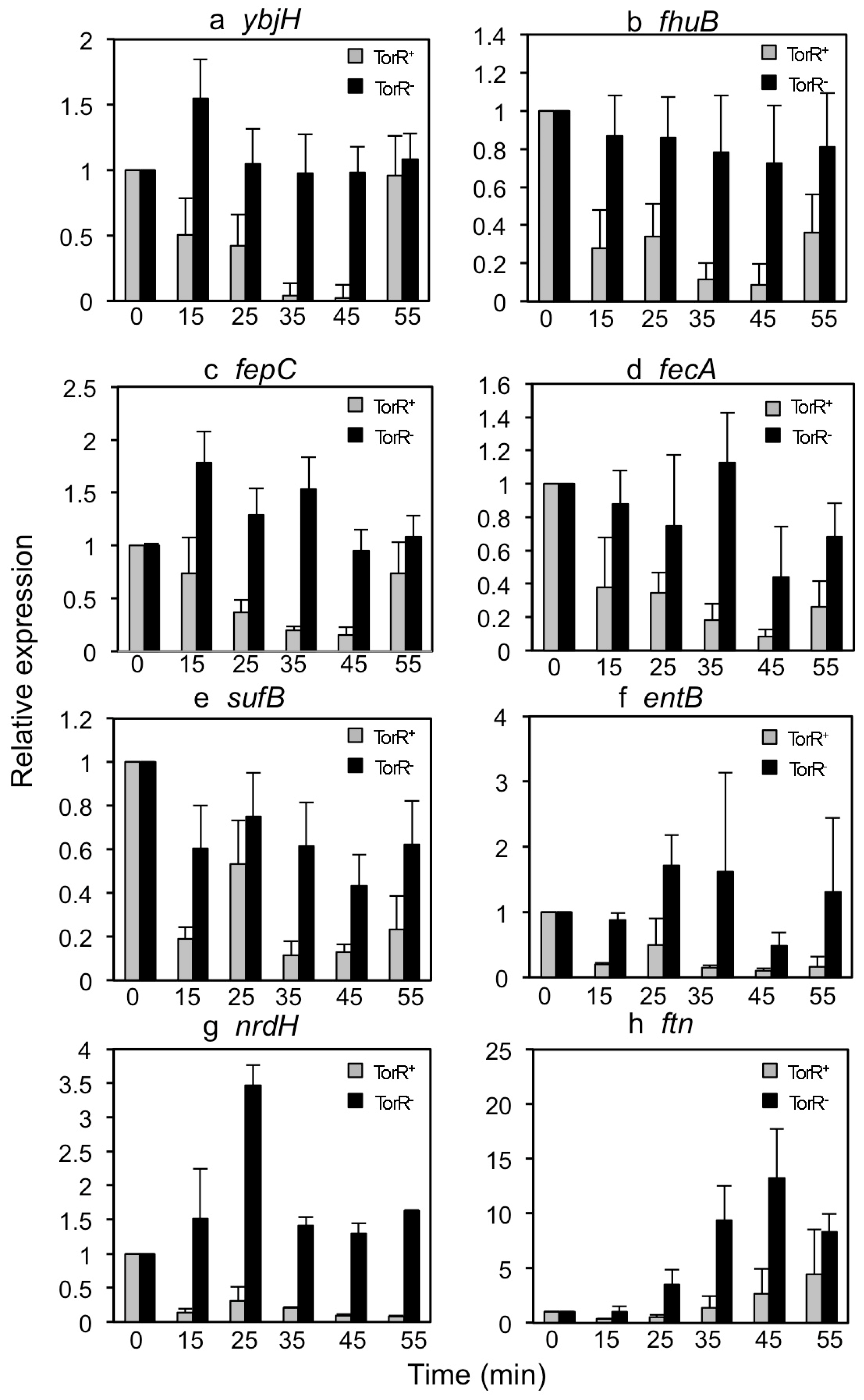
3.2. TorR Regulates Transcription of Several Genes in a Cell-Cycle-Dependent Manner
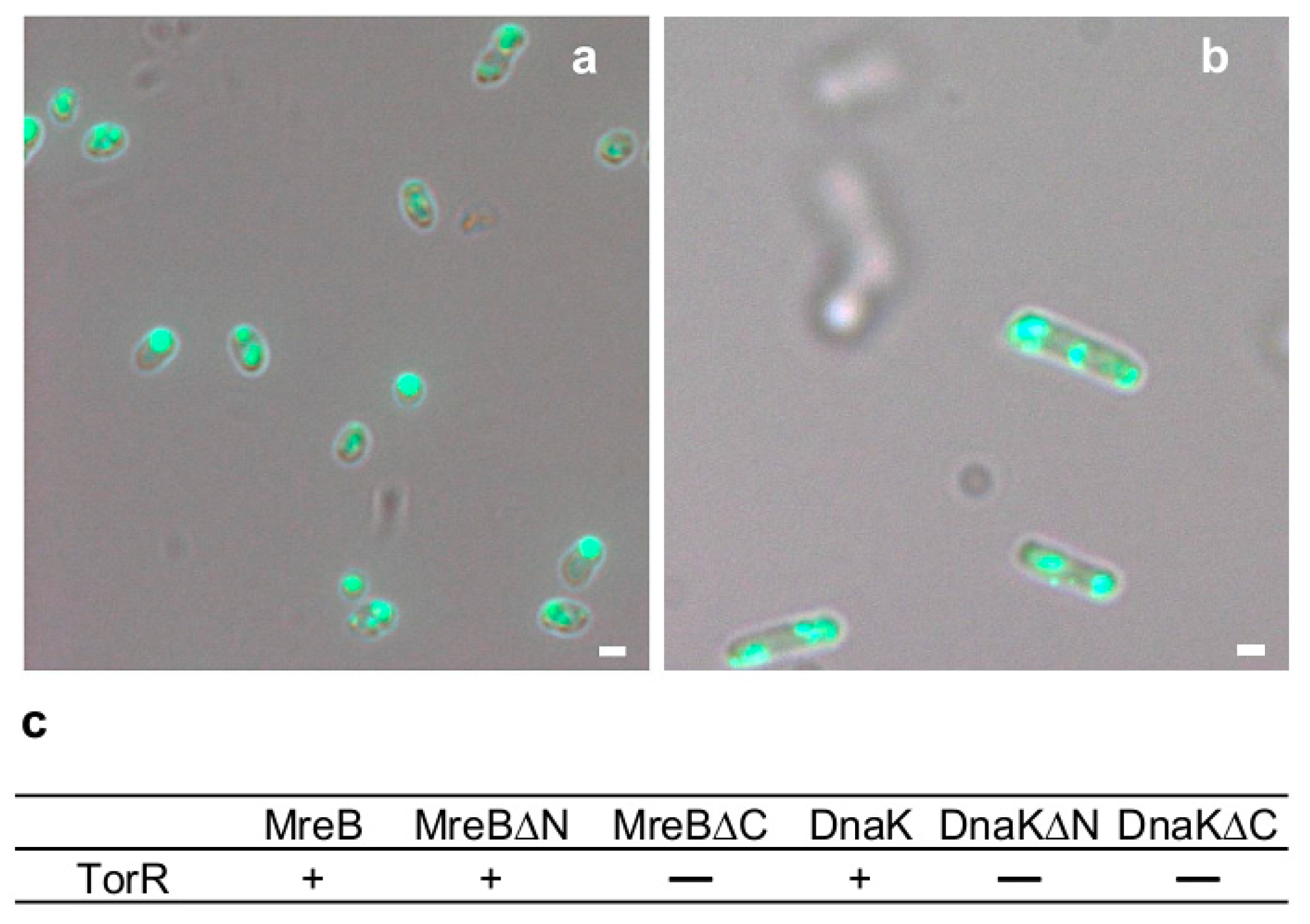
3.3. TorR Interacts with MreB and DnaK for Targeting the Old Cell Poles
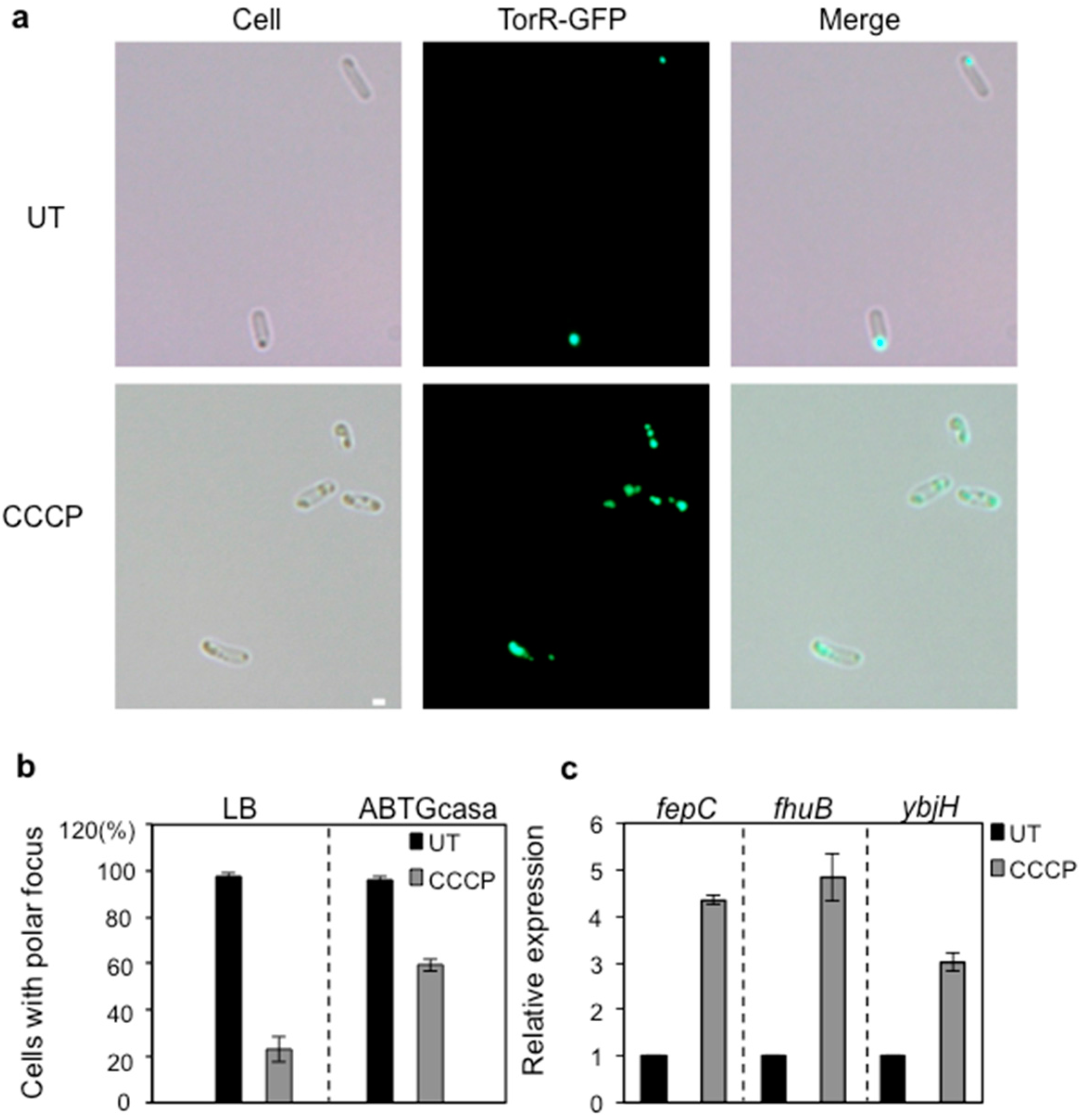
3.4. CCCP Treatment Affects the Polar Localization of TorR and Expression of the Target Genes
3.5. FtsZ Affects the TorR Foci Formation
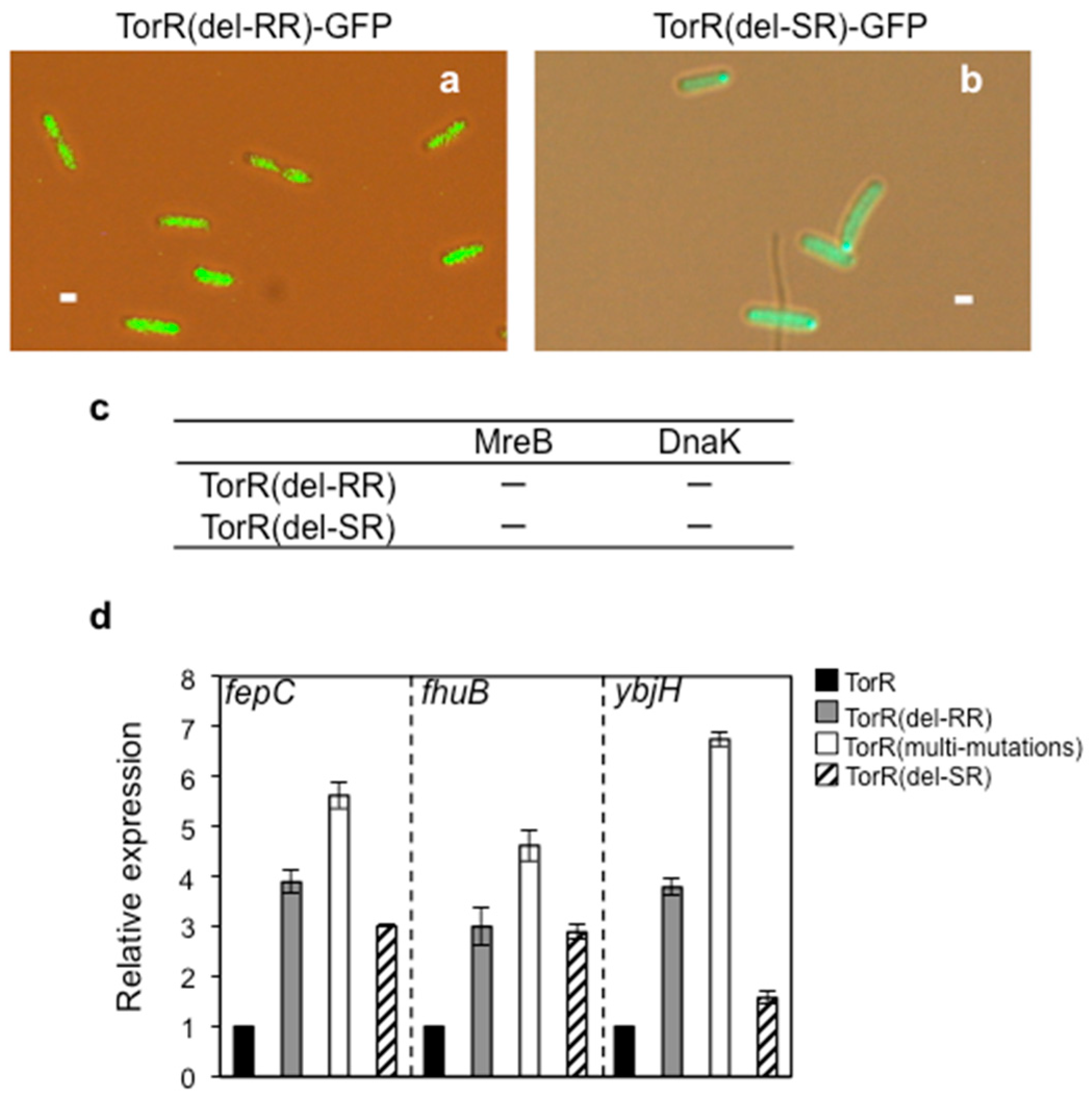
3.6. Interaction of TorR with MreB or DnaK is Responsible for Its Polar Localization and Subsequent Regulatory Effect on Gene Expression
4. Discussion
4.1. The TorR Protein Targets the Old Cell Poles by Interacting with MreB and DnaK Proteins
4.2. A Spatial Control for Correct Timing of Gene Expression during the Cell Cycle
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Cho, R.J.; Campbell, M.J.; Winzeler, E.A.; Steinmetz, L.; Conway, A.; Wodicka, L.; Wolfsberg, T.G.; Gabrielian, A.E.; Landsman, D.; Lockhart, D.J.; et al. A genome-wide transcriptional analysis of the mitotic cell cycle. Mol. Cell 1998, 2, 65–73. [Google Scholar] [CrossRef]
- Spellman, P.T.; Sherlock, G.; Zhang, M.Q.; Iyer, V.R.; Anders, K.; Eisen, M.B.; Brown, P.O.; Botstein, D.; Futcher, B. Comprehensive identification of cell cycle–regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol. Biol. Cell 1998, 9, 3273–3297. [Google Scholar] [CrossRef] [PubMed]
- Grunenfelder, B.; Rummel, G.; Vohradsky, J.; Roder, D.; Langen, H.; Jenal, U. Proteomic analysis of the bacterial cell cycle. Proc. Natl. Acad. Sci. USA 2001, 98, 4681–4686. [Google Scholar] [CrossRef] [PubMed]
- Laub, M.T.; McAdams, H.H.; Feldblyum, T.; Fraser, C.M.; Shapiro, L. Global analysis of the genetic network controlling a bacterial cell cycle. Science 2000, 290, 2144–2148. [Google Scholar] [CrossRef] [PubMed]
- Quon, K.C.; Marczynski, G.T.; Shapiro, L. Cell cycle control by an essential bacterial two-component signal transduction protein. Cell 1996, 84, 83–93. [Google Scholar] [CrossRef]
- Holtzendorff, J.; Hung, D.; Brende, P.; Reisenauer, A.; Viollier, P.H.; McAdams, H.H.; Shapiro, L. Oscillating global regulators control the genetic circuit driving a bacterial cell cycle. Science 2004, 304, 983–987. [Google Scholar] [CrossRef] [PubMed]
- Campbell, J.L.; Kleckner, N. E. coli oriC and the dnaA gene promoter are sequestered from dam methyltransferase following the passage of the chromosomal replication fork. Cell 1990, 62, 967–979. [Google Scholar] [CrossRef]
- Ogawa, T.; Okazaki, T. Cell cycle-dependent transcription from the gid and mioC promoters of Escherichia coli. J. Bacteriol. 1994, 176, 1609–1615. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Helmstetter, C.E. Relationship between ftsZ gene expression and chromosome replication in Escherichia coli. J. Bacteriol. 1994, 176, 6100–6106. [Google Scholar] [CrossRef] [PubMed]
- Garrido, T.; Sanchez, M.; Palacios, P.; Aldea, M.; Vicente, M. Transcription of ftsZ oscillates during the cell cycle of Escherichia coli. EMBO J. 1993, 12, 3957–3965. [Google Scholar] [PubMed]
- Speck, C.; Weigel, C.; Messer, W. ATP- and ADP-dnaA protein, a molecular switch in gene regulation. EMBO J. 1999, 18, 6169–6176. [Google Scholar] [CrossRef] [PubMed]
- Katayama, T.; Kubota, T.; Kurokawa, K.; Crooke, E.; Sekimizu, K. The initiator function of DnaA protein is negatively regulated by the sliding clamp of the E. coli chromosomal replicase. Cell 1998, 94, 61–71. [Google Scholar] [CrossRef]
- Shapiro, L.; McAdams, H.H.; Losick, R. Generating and exploiting polarity in bacteria. Science 2002, 298, 1942–1946. [Google Scholar] [CrossRef] [PubMed]
- Sourjik, V.; Berg, H.C. Localization of components of the chemotaxis machinery of Escherichia coli using fluorescent protein fusions. Mol. Microbiol. 2000, 37, 740–751. [Google Scholar] [CrossRef] [PubMed]
- De Boer, P.A.; Crossley, R.E.; Rothfield, L.I. A division inhibitor and a topological specificity factor coded for by the minicell locus determine proper placement of the division septum in E. coli. Cell 1989, 56, 641–649. [Google Scholar] [CrossRef]
- Hu, Z.; Lutkenhaus, J. Topological regulation of cell division in Escherichia coli involves rapid pole to pole oscillation of the division inhibitor MinC under the control of MinD and MinE. Mol. Microbiol. 1999, 34, 82–90. [Google Scholar] [CrossRef] [PubMed]
- Méjean, V.; Lobbi-Nivol, C.; Lepelletier, M.; Giordano, G.; Chippaux, M.; Pascal, M.C. TMAO anaerobic respiration in Escherichia coli: Involvement of the tor operon. Mol. Microbiol. 1994, 11, 1169–1179. [Google Scholar] [CrossRef] [PubMed]
- Ansaldi, M.; Simon, G.; Lepelletier, M.; Mejean, V. The TorR high-affinity binding site plays a key role in both torR autoregulation and torCAD operon expression in Escherichia coli. J. Bacteriol. 2000, 182, 961–966. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Jiaxin, Q.; Jing, L.; Hui, L.; Morigen, M. The effects of TorR protein on initiation of DNA replication in Escherichia coli. Yi Chuan Hered. 2015, 37, 302–308. [Google Scholar]
- Kitagawa, M.; Ara, T.; Arifuzzaman, M.; Ioka-Nakamichi, T.; Inamoto, E.; Toyonaga, H.; Mori, H. Complete set of ORF clones of Escherichia coli ASKA library (a complete set of E. coli K-12 ORF archive): Unique resources for biological research. DNA Res. 2006, 12, 291–299. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.V.; Ripp, S.; Replicon, J.; Ogunseitan, O.; Kokjohn, T.A. Virus-mediated gene transfer in freshwater environments. In Gene Transfers and Environment; Springer: New York, NY, USA, 1992; pp. 51–62. [Google Scholar]
- Karimova, G.; Dautin, N.; Ladant, D. Interaction network among Escherichia coli membrane proteins involved in cell division as revealed by bacterial two-hybrid analysis. J. Bacteriol. 2005, 187, 2233–2243. [Google Scholar] [CrossRef] [PubMed]
- Karimova, G.; Pidoux, J.; Ullmann, A.; Ladant, D. A bacterial two-hybrid system based on a reconstituted signal transduction pathway. Proc. Natl. Acad. Sci. USA 1998, 95, 5752–5756. [Google Scholar] [CrossRef] [PubMed]
- Bertani, G. Studies on lysogenesis I.: The mode of phage liberation by lysogenic Escherichia coli. J. Bacteriol. 1951, 62, 293–300. [Google Scholar] [PubMed]
- Morigen; Molina, F.; Skarstad, K. Deletion of the datA site does not affect once-per-cell-cycle timing but induces rifampin-resistant replication. J. Bacteriol. 2005, 187, 3913–3920. [Google Scholar] [CrossRef] [PubMed]
- Fleige, S.; Pfaffl, M.W. RNA integrity and the effect on the real-time qRT-PCR performance. Mol. Asp. Med. 2006, 27, 126–139. [Google Scholar] [CrossRef] [PubMed]
- Morigen; Flatten, I.; Skarstad, K. The Escherichia coli datA site promotes proper regulation of cell division. Microbiology 2014, 160 Pt 4, 703–710. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Fan, L.F.; Shi, Y.X.; Odsbu, I.; Morigen. NCBI Gene Expression Omnibus accession GSE72525. Available online: http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE72525 (accessed on 31 August 2016).
- Perkins, D.N.; Pappin, D.J.C.; Creasy, D.M.; Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 1999, 20, 3551–3567. [Google Scholar] [CrossRef]
- Morigen; Boye, E.; Skarstad, K.; Lobner-Olesen, A. Regulation of chromosomal replication by DnaA protein availability in Escherichia coli: Effects of the datA region. Biochim. Biophys. Acta 2001, 1521, 73–80. [Google Scholar] [CrossRef]
- Rousseau, A.; McEwen, A.G.; Poussin-Courmontagne, P.; Rognan, D.; Nomine, Y.; Rio, M.C.; Tomasetto, C.; Alpy, F. TRAF4 is a novel phosphoinositide-binding protein modulating tight junctions and favoring cell migration. PLoS Biol. 2013, 11, e1001726. [Google Scholar] [CrossRef] [PubMed]
- Sommer, E.; Koler, M.; Frank, V.; Sourjik, V.; Vaknin, A. The sensory histidine kinases TorS and EvgS tend to form clusters in Escherichia coli cells. PLoS ONE 2013, 8, e77708. [Google Scholar] [CrossRef] [PubMed]
- Lindner, A.B.; Madden, R.; Demarez, A.; Stewart, E.J.; Taddei, F. Asymmetric segregation of protein aggregates is associated with cellular aging and rejuvenation. Proc. Natl. Acad. Sci. USA 2008, 105, 3076–3081. [Google Scholar] [CrossRef] [PubMed]
- Winkler, J.; Seybert, A.; König, L.; Pruggnaller, S.; Haselmann, U.; Sourjik, V.; Weiss, M.; Frangakis, A.S.; Mogk, A.; Bukau, B. Quantitative and spatio-temporal features of protein aggregation in Escherichia coli and consequences on protein quality control and cellular ageing. EMBO J. 2010, 29, 910–923. [Google Scholar] [CrossRef] [PubMed]
- Fahnert, B.; Lilie, H.; Neubauer, P. Inclusion bodies: Formation and utilisation. Adv. Biochem. Eng. Biotechnol. 2004, 89, 93–142. [Google Scholar] [PubMed]
- Michaelis, A.M.; Gitai, Z. Dynamic polar sequestration of excess MurG may regulate enzymatic function. J. Bacteriol. 2010, 192, 4597–4605. [Google Scholar] [CrossRef] [PubMed]
- Chuang, S.E.; Burland, V.; Plunkett, G., 3rd; Daniels, D.L.; Blattner, F.R. Sequence analysis of four new heat-shock genes constituting the hslTS/ibpAB and hslVU operons in Escherichia coli. Gene 1993, 134, 1–6. [Google Scholar] [CrossRef]
- Li, G.; Young, K.D. Isolation and identification of new inner membrane-associated proteins that localize to cell poles in Escherichia coli. Mol. Microbiol. 2012, 84, 276–295. [Google Scholar] [CrossRef] [PubMed]
- Barrett, E.L.; Kwan, H.S. Bacterial reduction of trimethylamine oxide. Annu. Rev. Microbiol. 1985, 39, 131–149. [Google Scholar] [CrossRef] [PubMed]
- Ansaldi, M.; Théraulaz, L.; Baraquet, C.; Panis, G.; Méjean, V. Aerobic TMAO respiration in Escherichia coli. Mol. Microbiol. 2007, 66, 484–494. [Google Scholar] [CrossRef] [PubMed]
- Simon, G.; Méjean, V.; Jourlin, C.; Chippaux, M.; Pascal, M.-C. The torR gene of Escherichia coli encodes a response regulator protein involved in the expression of the trimethylamine N-oxide reductase genes. J. Bacteriol. 1994, 176, 5601–5606. [Google Scholar] [CrossRef] [PubMed]
- Pascal, M.C.; Lepelletier, M.; Giordano, G.; Chippaux, M. A regulatory mutant of the trimethylamine N-oxide reductase of Escherichia coli K12. FEMS Microbiol. Lett. 1991, 62, 297–300. [Google Scholar] [CrossRef] [PubMed]
- Withers, H.L.; Bernander, R. Characterization of dnaC2 and dnaC28 mutants by flow cytometry. J. Bacteriol. 1998, 180, 1624–1631. [Google Scholar] [PubMed]
- Carl, P.L. Escherichia coli mutants with temperature-sensitive synthesis of DNA. Mol. Gen. Genet. 1970, 109, 107–122. [Google Scholar] [CrossRef] [PubMed]
- Helmstetter, C.E.; Pierucci, O. Cell division during inhibition of deoxyribonucleic acid synthesis in Escherichia coli. J. Bacteriol. 1968, 95, 1627–1633. [Google Scholar] [PubMed]
- Oshima, T.; Aiba, H.; Masuda, Y.; Kanaya, S.; Sugiura, M.; Wanner, B.L.; Mori, H.; Mizuno, T. Transcriptome analysis of all two-component regulatory system mutants of Escherichia coli K-12. Mol. Microbiol. 2002, 46, 281–291. [Google Scholar] [CrossRef] [PubMed]
- Trinkle-Mulcahy, L.; Boulon, S.; Lam, Y.W.; Urcia, R.; Boisvert, F.M.; Vandermoere, F.; Morrice, N.A.; Swift, S.; Rothbauer, U.; Leonhardt, H. Identifying specific protein interaction partners using quantitative mass spectrometry and bead proteomes. J. Cell Biol. 2008, 183, 223–239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Teter, S.A.; Houry, W.A.; Ang, D.; Tradler, T.; Rockabrand, D.; Fischer, G.; Blum, P.; Georgopoulos, C.; Hart, F.U. Polypeptide flux through bacterial Hsp70: DnaK cooperates with trigger factor in chaperoning nascent chains. Cell 1999, 97, 755–765. [Google Scholar] [CrossRef]
- Wild, J.; Rossmeissl, P.; Walter, W.A.; Gross, C.A. Involvement of the DnaK-DnaJ-GrpE chaperone team in protein secretion in Escherichia coli. J. Bacteriol. 1996, 178, 3608–3613. [Google Scholar] [CrossRef] [PubMed]
- Wachi, M.; Doi, M.; Tamaki, S.; Park, W.; Nakajima-Iijima, S.; Matsuhashi, M. Mutant isolation and molecular cloning of mre genes, which determine cell shape, sensitivity to mecillinam, and amount of penicillin-binding proteins in Escherichia coli. J. Bacteriol. 1987, 169, 4935–4940. [Google Scholar] [CrossRef] [PubMed]
- Gitai, Z.; Dye, N.A.; Reisenauer, A.; Wachi, M.; Shapiro, L. MreB actin-mediated segregation of a specific region of a bacterial chromosome. Cell 2005, 120, 329–341. [Google Scholar] [CrossRef] [PubMed]
- Kruse, T.; Møller-Jensen, J.; Løbner-Olesen, A.; Gerdes, K. Dysfunctional MreB inhibits chromosome segregation in Escherichia coli. EMBO J. 2003, 22, 5283–5292. [Google Scholar] [CrossRef] [PubMed]
- Iwai, N.; Nagai, K.; Wachi, M. Novel S-benzylisothiourea compound that induces spherical cells in Escherichia coli probably by acting on a rod-shape-determining protein(s) other than penicillin-binding protein 2. Biosci. Biotechnol. Biochem. 2002, 66, 2658–2662. [Google Scholar] [CrossRef] [PubMed]
- Paul, K.; Erhardt, M.; Hirano, T.; Blair, D.F.; Hughes, K.T. Energy source of flagellar type III secretion. Nature 2008, 451, 489–492. [Google Scholar] [CrossRef] [PubMed]
- Bi, E.F.; Lutkenhaus, J. FtsZ ring structure associated with division in Escherichia coli. Nature 1991, 354, 161–164. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.C.; Margolin, W. FtsZ ring clusters in min and partition mutants: Role of both the Min system and the nucleoid in regulating FtsZ ring localization. Mol. Microbiol. 1999, 32, 315–326. [Google Scholar] [CrossRef] [PubMed]
- Raskin, D.M.; de Boer, P.A. MinDE-dependent pole-to-pole oscillation of division inhibitor MinC in Escherichia coli. J. Bacteriol. 1999, 181, 6419–6424. [Google Scholar] [PubMed]
- Raskin, D.M.; de Boer, P.A. The MinE ring: An FtsZ-independent cell structure required for selection of the correct division site in E. coli. Cell 1997, 91, 685–694. [Google Scholar] [CrossRef]
- Gitai, Z.; Dye, N.; Shapiro, L. An actin-like gene can determine cell polarity in bacteria. Proc. Natl. Acad. Sci. USA 2004, 101, 8643–8648. [Google Scholar] [CrossRef] [PubMed]
- Jones, L.J.; Carballido-Lopez, R.; Errington, J. Control of cell shape in bacteria: Helical, actin-like filaments in Bacillus subtilis. Cell 2001, 104, 913–922. [Google Scholar] [CrossRef]
- Kruse, T.; Bork-Jensen, J.; Gerdes, K. The morphogenetic MreBCD proteins of Escherichia coli form an essential membrane-bound complex. Mol. Microbiol. 2005, 55, 78–89. [Google Scholar] [CrossRef] [PubMed]
- Daniel, R.A.; Errington, J. Control of cell morphogenesis in bacteria: Two distinct ways to make a rod-shaped cell. Cell 2003, 113, 767–776. [Google Scholar] [CrossRef]
- Bean, G.; Flickinger, S.; Westler, W.; McCully, M.; Sept, D.; Weibel, D.; Amann, K.J. A22 disrupts the bacterial actin cytoskeleton by directly binding and inducing a low-affinity state in MreB. Biochemistry 2009, 48, 4852–4857. [Google Scholar] [CrossRef] [PubMed]
- Taneva, S.G.; Moro, F.; Velazquez-Campoy, A.; Muga, A. Energetics of nucleotide-induced DnaK conformational states. Biochemistry 2010, 49, 1338–1345. [Google Scholar] [CrossRef] [PubMed]
- Yamaichi, Y.; Bruckner, R.; Ringgaard, S.; Moll, A.; Cameron, D.E.; Briegel, A.; Jensen, G.J.; Davis, B.M.; Waldor, M.K. A multidomain hub anchors the chromosome segregation and chemotactic machinery to the bacterial pole. Genes Dev. 2012, 26, 2348–2360. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.C.; Ramamurthi, K.S. Macromolecules that prefer their membranes curvy. Mol. Microbiol. 2010, 76, 822–832. [Google Scholar] [CrossRef] [PubMed]
- Laloux, G.; Jacobs-Wagner, C. How do bacteria localize proteins to the cell pole? J. Cell Sci. 2014, 127, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Renner, L.D.; Weibel, D.B. Cardiolipin microdomains localize to negatively curved regions of Escherichia coli membranes. Proc. Natl. Acad. Sci. USA 2011, 108, 6264–6269. [Google Scholar] [CrossRef] [PubMed]
- Lam, H.; Schofield, W.B.; Jacobs-Wagner, C. A landmark protein essential for establishing and perpetuating the polarity of a bacterial cell. Cell 2006, 124, 1011–1023. [Google Scholar] [CrossRef] [PubMed]
- Van der Henst, C.; Charlier, C.; Deghelt, M.; Wouters, J.; Matroule, J.-Y.; Letesson, J.-J.; de Bolle, X. Overproduced Brucella abortus PdhS-mCherry forms soluble aggregates in Escherichia coli, partially associating with mobile foci of IbpA-YFP. BMC Microbiol. 2010, 10, 248. [Google Scholar] [CrossRef] [PubMed]
- Jensen, L.J.; Jensen, T.S.; de Lichtenberg, U.; Brunak, S.; Bork, P. Co-evolution of transcriptional and post-translational cell-cycle regulation. Nature 2006, 443, 594–597. [Google Scholar] [CrossRef] [PubMed]
- Lindas, A.C.; Bernander, R. The cell cycle of archaea. Nat. Rev. Microbiol. 2013, 11, 627–638. [Google Scholar] [CrossRef] [PubMed]
- McAdams, H.H.; Shapiro, L. A bacterial cell-cycle regulatory network operating in time and space. Science 2003, 301, 1874–1877. [Google Scholar] [CrossRef] [PubMed]
- Gorbatyuk, B.; Marczynski, G.T. Regulated degradation of chromosome replication proteins DnaA and CtrA in Caulobacter crescentus. Mol. Microbiol. 2005, 55, 1233–1245. [Google Scholar] [CrossRef] [PubMed]
- Domian, I.J.; Quon, K.C.; Shapiro, L. Cell type-specific phosphorylation and proteolysis of a transcriptional regulator controls the G1-to-S transition in a bacterial cell cycle. Cell 1997, 90, 415–424. [Google Scholar] [CrossRef]
- Bachmann, B.J. Derivations and genotypes of some mutant derivatives of Escherichia coli K-12. In Escherichia coli and Salmonella Typhimurium: Cellular and Molecular Biology; American Society for Microbiology: Washington, DC, USA, 1987; pp. 1190–1219. [Google Scholar]
- Baba, T.; Ara, T.; Hasegawa, M.; Takai, Y.; Okumura, Y.; Baba, M.; Datsenko, K.A.; Tomita, M.; Wanner, B.L.; Mori, H. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: The Keio collection. Mol. Syst. Biol. 2006, 2. [Google Scholar] [CrossRef] [PubMed]
- Chang, A.C.; Cohen, S.N. Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the P15A cryptic miniplasmid. J. Bacteriol. 1978, 134, 1141–1156. [Google Scholar] [PubMed]

© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yao, Y.; Fan, L.; Shi, Y.; Odsbu, I.; Morigen. A Spatial Control for Correct Timing of Gene Expression during the Escherichia coli Cell Cycle. Genes 2017, 8, 1. https://doi.org/10.3390/genes8010001
Yao Y, Fan L, Shi Y, Odsbu I, Morigen. A Spatial Control for Correct Timing of Gene Expression during the Escherichia coli Cell Cycle. Genes. 2017; 8(1):1. https://doi.org/10.3390/genes8010001
Chicago/Turabian StyleYao, Yuan, Lifei Fan, Yixin Shi, Ingvild Odsbu, and Morigen. 2017. "A Spatial Control for Correct Timing of Gene Expression during the Escherichia coli Cell Cycle" Genes 8, no. 1: 1. https://doi.org/10.3390/genes8010001





