Abstract
Numerous sources of evidence suggest that most of the eukaryotic genome is transcribed into protein-coding mRNAs and also into a large number of non-coding RNAs (ncRNAs). Long ncRNAs (lncRNAs), a group consisting of ncRNAs longer than 200 nucleotides, have been found to play critical roles in transcriptional, post-transcriptional, and epigenetic gene regulation across all kingdoms of life. However, lncRNAs and their regulatory roles remain poorly characterized in plants, especially in woody plants. In this paper, we used a computational approach to identify novel lncRNAs from a published RNA-seq data set and analyzed their sequences and expression patterns. In total, 1133 novel lncRNAs were identified in mulberry, and 106 of these lncRNAs displayed a predominant tissue-specific expression in the five major tissues investigated. Additionally, functional predictions revealed that tissue-specific lncRNAs adjacent to protein-coding genes might play important regulatory roles in the development of floral organ and root in mulberry. The pipeline used in this study would be useful for the identification of lncRNAs obtained from other deep sequencing data. Furthermore, the predicted lncRNAs would be beneficial towards an understanding of the variations in gene expression in plants.
1. Introduction
Mulberry (Morus notabilis) belongs to the genus Morus, which comprises 10–13 species and over 1000 cultivars distributed throughout Asia, Africa, Europe, and North America [1,2], and are well known for their important economic and medicinal values [3]. In China, mulberry leaves have been used to feed silkworms for silk production [4], and its fruit is either eaten fresh or widely used in the production of juice, wine, jam and canned food [5]. In addition, the root, bark, branch, leaf, and fruit of mulberry have been used for protecting liver, improving eyesight, treating fever, facilitating urination, and lowering blood pressure due to their high levels of isoprenylated flavonoids, such as sanggenon-type flavanones, Diels-Alder adducts, and flavones [6,7,8]. Previous studies have suggested that secondary metabolism products and some small molecule modulators might play critical roles in plant-herbivore interactions, and mulberry is an ideal research model organism used to study plant-herbivore interaction [9,10]. The genome sequencing of Morus notabilis was completed in 2013, with approximately 29,338 protein-coding genes identified, however, a lot of important information has not been exploited completely [10,11]. Therefore, it is necessary and urgent to identify novel lncRNAs and understand the functions of lncRNAs in Morus notabilis.
Recent advances in DNA sequencing technology and transcriptome analysis have challenged the central dogma of biology. Emerging evidence shows that more than 90% of eukaryotic genomes are transcribed, but only 1%–2% have a protein-coding capacity, and the majority of sequences are transcribed as noncoding RNAs (ncRNAs) [12,13], which play critical roles in regulating gene expression at the transcriptional, post-transcriptional, and epigenetic levels during several biological processes [14,15,16]. Based on their distinct characteristics compared to housekeeping ncRNAs, including rRNAs, tRNAs, and small nucleolar RNAs, ncRNAs can be classified as (1) small RNAs, including microRNAs (miRNAs) and small interfering RNAs (siRNAs); (2) natural antisense transcripts (NATs); and (3) long non-coding RNAs (lncRNAs) [17]. LncRNAs have been defined as non-protein coding RNAs of more than 200 bp in length, distinguishing them from short ncRNAs [18,19].
Since the first report of lncRNAs in humans [20], thousands of lncRNAs have been identified in a number of species. However, genome-wide identifications of lncRNAs have been performed in only a few plant species [17,21]. For instance, vernalization in Arabidopsis is influenced by the lncRNAs COOLAIR and COLDAIR [22,23] and induced by phosphate starvation1 (IPS1), which is a member of the TPS1/Mt4 gene family that acts as a miR399 target mimic in fine tuning of PHO2 (encoding an E2 ubiquitin conjugase-related enzyme) expression and phosphate uptake in Arabidopsis, tomato and Medicago truncatula [24,25]. A large set of Populus RNA-seq data was examined and a total of 504 lncRNAs were found to be drought responsive [26]. A network of interactions among the lncRNAs, miRNAs and mRNAs was constructed with the RNA-seq data of Populu stomentosa, revealing that lncRNAs were involved in the regulation of wood formation [27]. Each of the lncRNA surveys in plants uncovered a substantial number of lncRNAs, which were often expressed at low levels in a tissue-specific manner, as in humans and other mammals, and acted as natural miRNA target mimics, chromatin modifiers, or molecular cargo for protein re-localization [18].
In this study, 1133 lncRNAs were identified for the first time on a genome-wide scale, using a set of published next-generation RNA-seq data from five tissues of mulberry. Furthermore, the structural characteristics and tissue specificity of the predicted lncRNAs were analyzed and compared with the mRNAs. Additionally, the functions of the novel lncRNAs were predicted based on genomic positioning information, which was important for further clarifying the roles of the lncRNAs in the growth and development of woody plants.
2. Experimental Section
2.1. The Pipeline to Identify lncRNAs from RNA-seq Data
A set of Morus notabilis clean RNA-seq data with a length of 90bp and taken from five different tissues was obtained from a published study [28] and downloaded from the NCBI SRA website with the project number SRX504906. The protein-coding genes of RefSeq [29], Ensembl [30], UCSC [31], and Vega [32] were downloaded from the UCSC genome browser and all known noncoding genes from the NONCODE4.0 database [33]. The mulberry reference genome and gene model annotation files were downloaded from the genome website [28], and a pipeline was developed to identify putative lncRNAs (Figure 1).
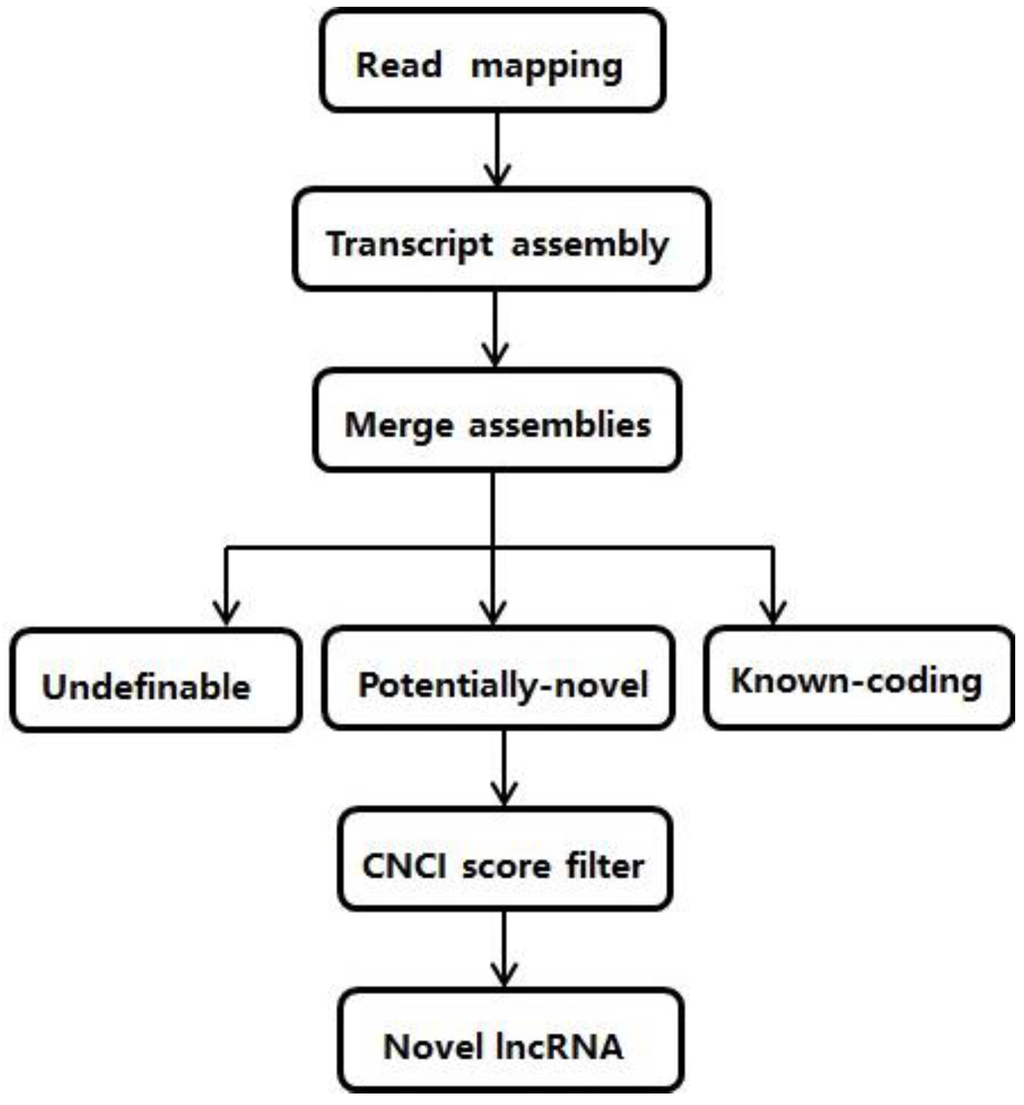
Figure 1.
Pipeline to identify lncRNAs from RNA-seq data.
After filtering out low-quality reads, the spliced read aligner TopHat version 2.0.9 [34] was used to map all clean reads to the mulberry genome. We used two rounds of TopHat mapping to maximize the usage of the splice junction information from all RNA-seq data. In the first round, all reads were mapped with TopHat (parameters: min-anchor = 5, min-isoform-fraction = 0, and other parameters with default values); in the second round of TopHat remapping, all splice junctions produced by the initial mapping were fed into TopHat to map reads (parameters: raw-juncs, no-novel-juncs, and min-anchor = 5, and min-isoform-fraction = 0).
Mapped reads from TopHat for each tissue were assembled for each sample separately by Cufflinks [35]. The cufflinks employed spliced read information to determine exon connectivity. Specifically, it used a probabilistic model approach to assemble and quantify the expression level of a minimal set of isoforms and provided the maximum level of annotation on the expression data for given loci. Cufflinks version 2.1.1 was run with default parameters (except “min-frags-per-transfrag = 0”). The multiple assembled transcript files for different tissues were then merged together to produce a unique transcriptome set using Cuffmerge.
We then used an analysis process to minimize false positives and maximize the number of lncRNAs from the merged transcripts, which included the following steps: (1) compare the merged transcripts with known protein-coding genes and lncRNAs in the public databases; (2) select transcripts that are longer than 200 nt; and (3) filter the putative lncRNA transcripts by coding potential using CNCI software [36], which can be categorized as noncoding (CNCI is a powerful signature tool that profiles adjoining nucleotide triplets to effectively distinguish protein-coding and non-coding sequences independent of known annotations) [37].
2.2. Calculation of lncRNA Conservation
To further demonstrate the reliability of lncRNAs predicted from the RNA-seq data and calculate the conservation of the novel lncRNAs, a set of lncRNAs collected by TAIR [38] and PlncDB [39] was downloaded and then aligned with the sequences of novel mulberry lncRNAs using BLASTN software [40].
2.3. Expression Profiles of Tissue Specific lncRNAs and Functional Predictions
To evaluate the tissue specificity of a transcript, we devised an entropy-based method to quantify the similarity between a transcript’s expression pattern and another predefined pattern, which represented an extreme case where a transcript was expressed in only one tissue [41]. After obtaining the lncRNA dataset with tissue-specific expression, we further searched the genomic location information from the genome comparison results by running a script with Perl, and retrieved the information of coding genes within the scope of its ±10 Kb.
2.4. qRT-PCR Analysis of lncRNAs
Three individuals of mulberry were used as biological replicates. Tissues from bark, root and winter bud were isolated with a sharp chisel and frozen immediately in liquid nitrogen. Total RNA was extracted with Universal Total RNA Kit (BioTeke, Beijing, China). First-strand cDNA synthesis was carried out with approximately 1.0 μg RNA using the Prime Script™ RT Master Mix (Takara, Dalian, China). All primers used in this study are listed in Supplementary Materials Table S1. Real-time qRT-PCR was performed in quadruplicate using the SYBR Premix Ex Taq™ II Kit (Takara, Dalian, China) on a Roche light Cycler 480 (Roche Applied Science, Penzberg, Upper Bavaria, Germany) according to the manufacturer’s instructions. Sample cycle threshold (Ct) values were determined and standardized relative to the endogenous control genes ACTIN3, and the 2–∆∆CT method was used to calculate the relative changes in gene expression based on the qRT-PCR data [42].
3. Results
3.1. Transcripts Reconstruction and Identification of Novel lncRNAs
The RNA-seq data used in this study were downloaded from the NCBI SRA website. These reads were paired and both lengths were 90 nt. Starting from a total of 1.2 billion reads, we performed short read gapped alignment using TopHat [34] and recovered 1.01 billion (84%) mapped reads (Table 1).

Table 1.
RNA-seq data production and alignment results for reads of different tissues.
We then used Cufflinks [35] to de novo reconstruct one set of transcripts for each tissue based on the read-mapping results. Transcripts reconstructed were separately merged into combined sets of transcripts using the Cuffcompare utility provided by Cufflinks. After filtering for exon number, transcript length, and coverage, we obtained 41,042 reliably expressed transcripts (Table 2).

Table 2.
Exon numbers of reconstructed transcripts.
To assess the robustness of these ab initio assemblers, we analyzed their performance on protein-coding genes. The transcripts we reconstructed using Cufflinks covered 70.79% of known mulberry coding genes (Figure 2). These results strongly supported the fact that these assembly approaches could robustly and reliably reconstruct both coding and noncoding transcripts at a global level.
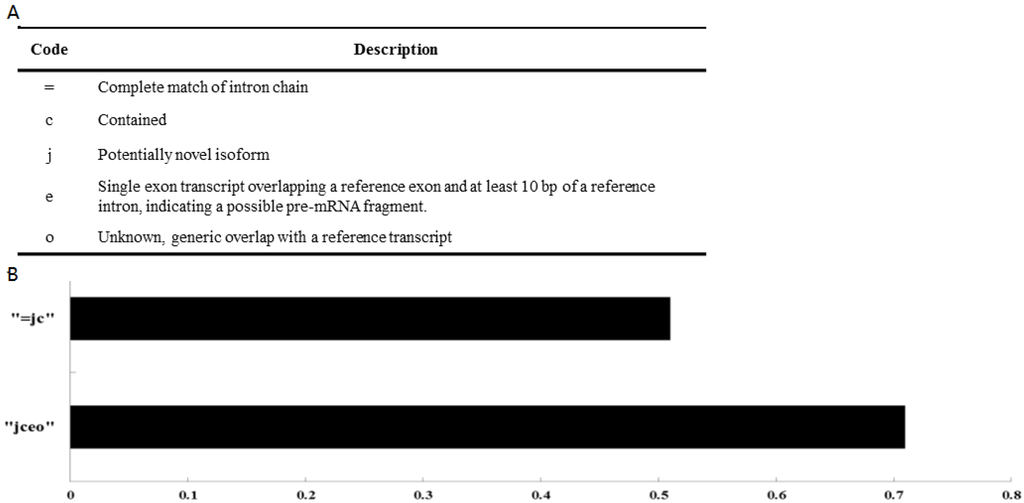
Figure 2.
(A) Different classes of assembled transcripts according to their relative positions with known coding genes; (B) Ratio of the reconstructed transcripts to known coding genes.
Based on the robust transcript reconstruction and broad availability of deep sequencing datasets, we used an analysis process to minimize the false positives and maximize the number of lncRNA transcripts, compared the merged transcripts with known protein-coding genes and lncRNAs in the public databases, and classified the combined transcripts into several different subsets. The majority of the transcripts (53.44%) corresponding to the annotated protein-coding genes, while the rest of the transcripts were undefinable (23.64%), and potentially novel (22.92%). The potentially novel transcripts were then filtered for coding potential based on CNCI software [43], resulting in the identification of 1133 reliably expressed lncRNAs with length >200 nt (Figure 3).
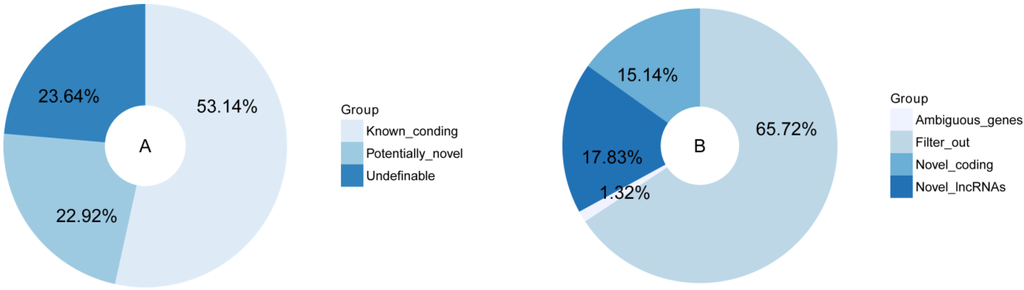
Figure 3.
Classification of reconstructed transcripts (A) and results from CNCI (B).
The identified lncRNAs were classified as intergenic, intronic and antisense lncRNAs based on spatial relationships of their gene loci with protein-coding genes (Figure 4B). The identified lncRNAs were mostly intergenic lncRNAs, with 1092 in total, accounting for 96.4% of the identified lncRNA. There were 38 intronic lncRNAs, accounting for 3.4%, and 3 antisense lncRNAs, accounting for 0.26% (Figure 4A).
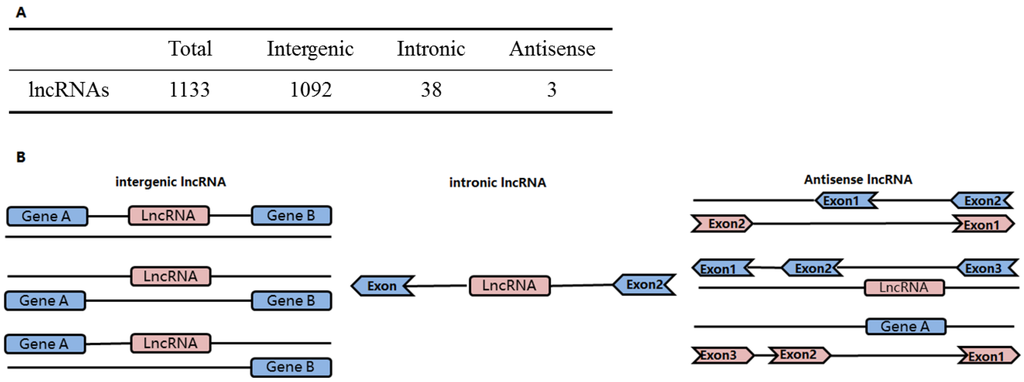
Figure 4.
Classification of the predicted lncRNAs. (A) lncRNAs were classified as intergenic, intronic or antisense lncRNAs based on the spatial relationships of their gene loci with protein-coding genes; (B) Schematic illustration of the classification of lncRNA genes based on their spatial relationship with protein-coding genes.
3.2. Characterization of the Novel lncRNAs
The length distribution results showed that the novel identified 1133 lncRNAs contained 1755 transcripts mainly in the range of 200–1200 bp. The lengths of 25,902 transcripts from known coding genes were greater than that of the lncRNAs, mostly above 800 bp. The distribution results of exon numbers revealed that there were 982 single exons (3.79%) and 24,920 multi-exons (96.21%) in the 25,902 transcripts from known coding genes. There were 75 single exons (4.27%) and 1680 multi-exons (95.73%) in the lncRNA 1755 transcripts, revealing a similar proportion of multi-exons to the known coding genes (Figure 5).
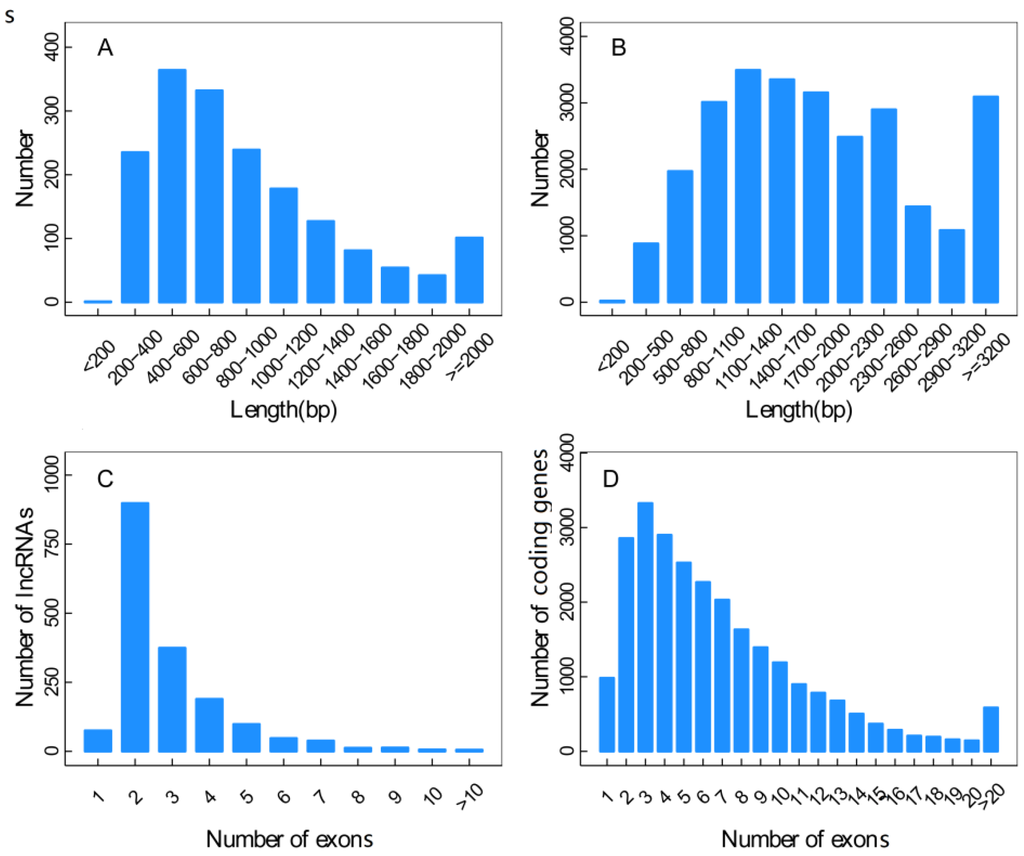
Figure 5.
Sequence features of lncRNAs. (A) and (B) represented the length distributions of the transcripts from novel lncRNAs (A) and known coding genes (B); (C) and (D) represented the exon numbers of the transcripts of lncRNAs (C) and known coding genes (D).
In combination with all known lncRNAs, we established a comprehensive catalog of 1133 transcribed lncRNA genes. Based on the Fragments Per Kilobase of transcript per Million mapped reads (FPKM) of each transcript, calculated by “Cufflinks” “abundance estimation mode” across the five tissues, we compared the expression differences between lncRNAs and protein-coding genes. The average expression levels of lncRNAs were lower than those for protein coding genes, but lncRNAs showed a wider range of abundance, with a subset of them equally abundant to mRNAs (Figure 6).
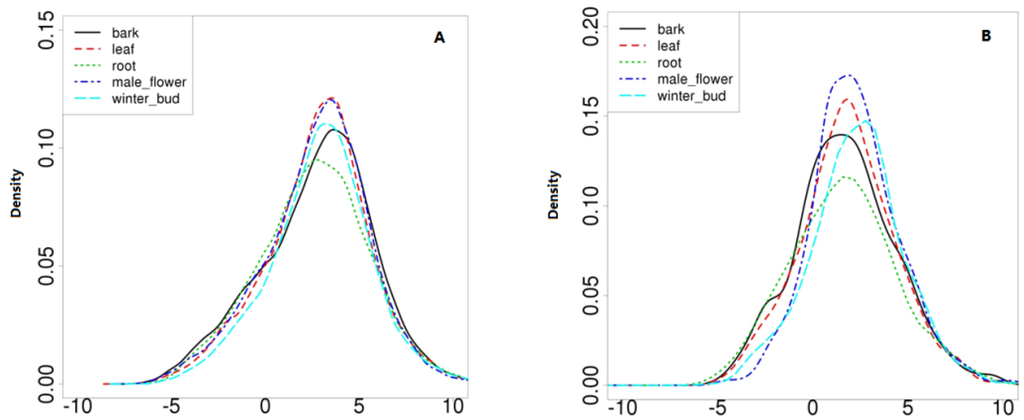
Figure 6.
Comparison of expression levels between known coding genes and lncRNAs. (A) represents the density of expression of known-coding genes N = 26965, Bandwidth = 0.3673; and (B) represents the lncRNAs, N = 1005, Bandwidth = 0.5891.
Through conservation analysis we found that 112 lncRNAs from the 1133 newly identified genes had homologies in the Arabidopsis database, while only 9 lncRNAs had homologies in the poplar database (Supplementary Table S2). The homology comparison results of the novel lncRNAs of mulberry with the mapped poplar lncRNAs confirmed the high level of homology between two sequences as 41.31% (Figure 7).
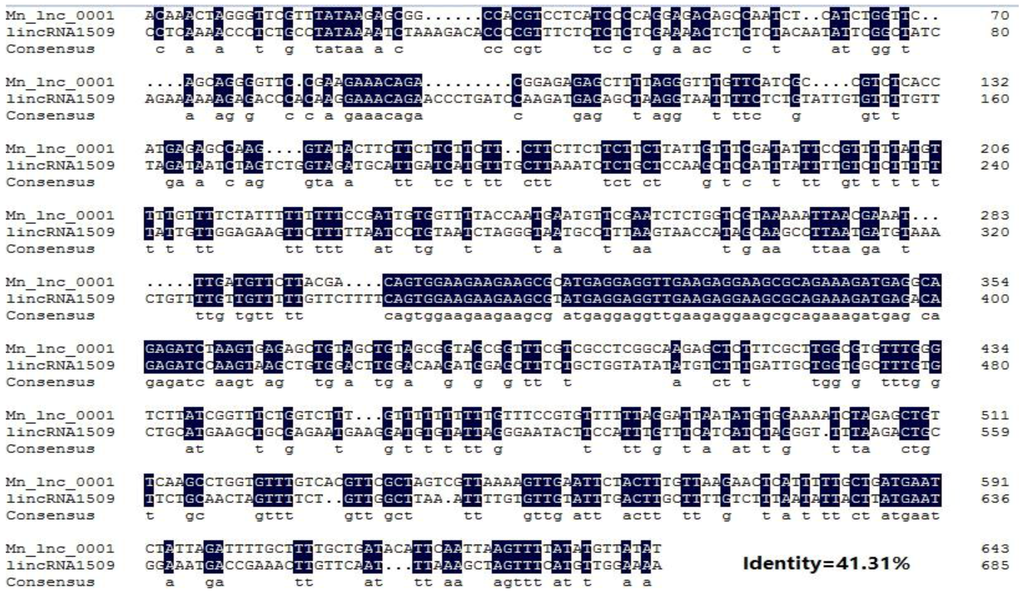

Figure 7.
Alignment of the nucleotide sequences of Mn_lnc_0001 and lincRNA1509. Black and white backgrounds indicate conserved and non-conserved residues, respectively.
3.3. Expression Profiles of Tissue Specific lncRNAs and Functional Predictions
To assess the tissue specificity of mulberry lncRNA expression, we calculated the Jensen-Shannon tissue specificity score (JS score) [40] for each transcript using the established procedure. Using a JS score = 0.9 as a cutoff, we demonstrated that only 9.35% of the lncRNAs were tissue-specific (Figure 8). Thus, some of the lncRNA expressions of mulberry were clearly subject to tissue dependent regulation, either at the level of transcription or degradation.
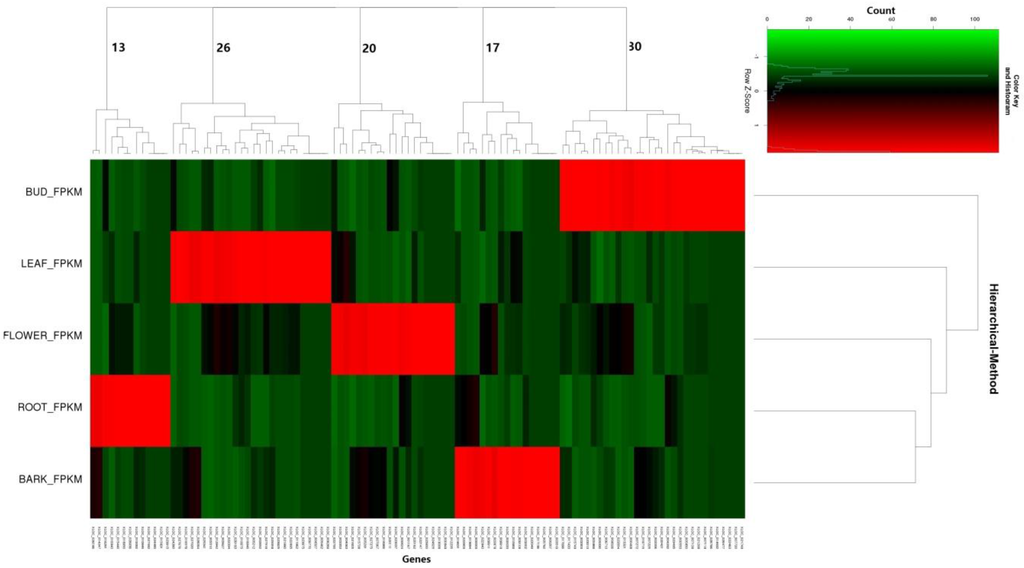
Figure 8.
Tissue specificity of lncRNAs from winter bud, leaf, male flower, root, and bark of mulberry.
Comparing their genomic locations with those of known mulberry coding genes, we found that among the 1133 lncRNAs, 106 (9.35%) were tissue-specific, including 82 lncRNAs adjacent (±10 kb) to protein-coding genes (Supplementary Materials Table S3–S7). The functions annotated to the protein-coding genes mainly involved hormone signal recognition and transduction, plant secondary metabolite synthesis, energy metabolism, etc. The lncRNAs Mn_lnc_0132, Mn_lnc_0521, and Mn_lnc_0782 were specifically expressed in male flower: Mn_lnc_0132 was located near the protein-coding gene EXB28594.1 (Protein PROLIFERA). Protein PROLIFERA, a highly conserved protein, was found in all eukaryotes, and specifically expressed in populations of dividing cells in sporophytic tissues of the plant body, such as the palisade layer of the leaf and founder cells of initiating flower primordia [44]. Mn_lnc_0521 was located near EXB81017.1 (Serine/threonine-protein phosphatase PP). The PP1s were shown to play key roles in many aspects of plant growth and development, such as pollination and pollen tube development [45,46,47]. It was found that Mn_lnc_0782 was located near EXC20310.1 (Phosphoenolpyruvate/phosphate translocator PPT). Located in the plastid, PPT played a pivotal role in the regulation of leaf color, florescence, and female and male gametophyte formation [48,49,50,51]. A sulfate transporter, Mn_lnc_0714, was specifically expressed in root and located near EXC06697.1 (Sulfate transporter 1.3). It was tissue-specifically expressed and was crucial for root development and symbiotic nitrogen fixation in root nodules [52,53,54].
To validate RNA-seq results, qRT-PCR were performed for 10 randomly selected tissue-specific lncRNAs in bark. As a result, all 10 reactions generated sequence products. Remarkable higher relative quantitative expressions of the 10 lncRNA were observed in bark. However, only 2 and 4 of the 10 lncRNAs expressed in winter bud and root, respectively, but their expression levels were quite low, ranging from 1.4% to 15.4% of the expression level in bark (Figure 9).
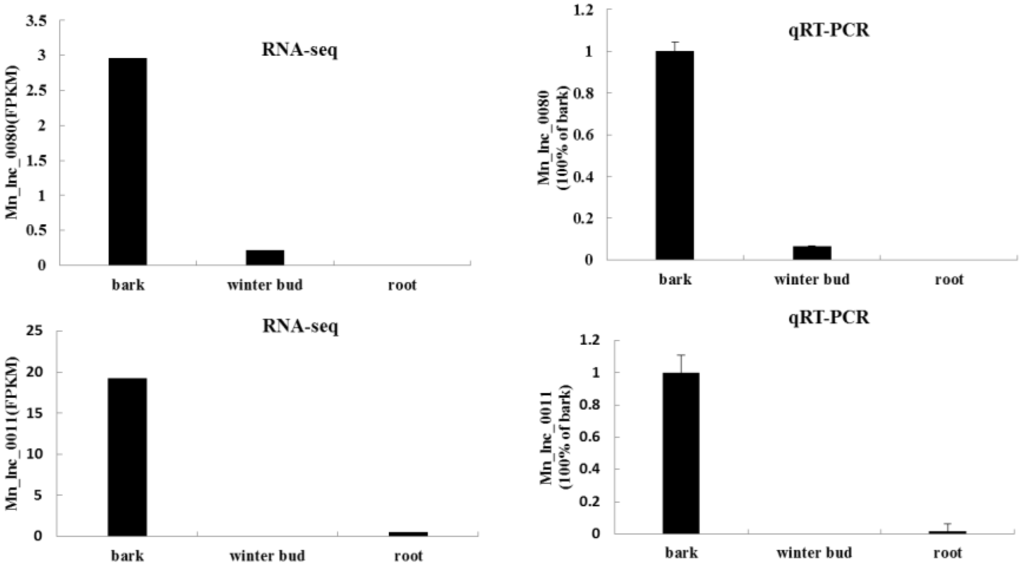
Figure 9.
Typical result of qRT-PCR verification. Transcripts abundance based on RNA-seq (left) and qRT-PCR (right) is shown for lncRNAs identified form RNA-seq data.
4. Discussion
An avalanche of RNA-seq data emerged as powerful high-throughput sequencing technologies became more pervasive and user-friendly. However, systematic identification of lncRNAs was limited to only a few plant species [21,26,27,55,56], leaving most plant transcriptome sequencing data not fully explored, even though these novel molecules play important roles in a wide range of biological processes [15]. Because lncRNAs are generated by the same transcriptional machinery as mRNAs [57], no defining biochemical features could be exclusively ascribed to lncRNAs, such as a 5′ cap, 3′ polyadenylated tail, and splicing [58]. Defining lncRNAs simply on the basis of size and lack of protein-coding capability was intellectually far from satisfying. In this paper, we designed a strict computational pipeline and identified 1133 novel lncRNAs from the entire genome using a set of published mulberry next-generation RNA-seq data. The pipeline used in this study can be easily adapted to other organisms, especially for species that have not been well studied to date.
The expression levels of the novel mulberry lncRNAs in root, leaf, bark, bud, and male flower were below the expression levels of mRNAs, which was consistent with findings in other species [59,60,61]. Conservation analysis found that among the 1133 lncRNAs, 112 (9.4%) had homology in the Arabidopsis database, and 9 (0.8%) had homology in the poplar lncRNA database. The low levels of conservation might be caused by the incomplete lncRNA databases of plants. The results also reflected the less restrictive factors on the evolution of lncRNAs, and thus the low conservation levels of lncRNA sequences among species, factors that reduce the possibility of forming a large family with homologous genes. Moreover, qRT-PCR was performed, and the RNA-seq results were consistent with the qRT-PCR data, providing further proof that the prediction accuracy was sufficient.
Numerous studies have shown that lncRNAs with tissue-specific expression usually had special functions [62], and the lncRNAs of higher species primarily played the biological role of cis-regulation of the adjacent genes [63,64,65]. In the analysis of tissue-specific expression, we found that 106 lncRNAs from our 1133 newly identified genes were expressed specifically in five separate tissues, among which 82 had known protein-coding genes in the range of ±10 Kb. We therefore predicted the functions of these lncRNAs by analysis of the tissue-specific expressions and the functions of adjacent coding genes. Further analysis showed that three male flower-specific lncRNAs were located adjacent to coding genes, which are related to development of floral organs. One root-specific lncRNA was located adjacent to a coding gene, which is crucial for root development and symbiotic nitrogen fixation. These results suggest that these novel lncRNAs might play important regulatory roles in the development of floral organs and root in mulberry.
Regarding the important functions of lncRNAs in plant growth and development, their identification within plant-wide genomes is rapidly developing. By contrast, the functional characterization of lncRNAs for plants is far behind that of other species. So far, the commonly used methods for lncRNA functional prediction are based on co-expression networks [57], miRNA regulation [66], protein binding [67], epigenetic modification [68], and adjacent gene functions. In this study, due to the influence of sequencing data (insufficient sample size) we cannot make functional predictions through the methods of co-expression networks and others. These methods are only based on the functional predictions of bioinformatics, so the accurate assignment of functions of lncRNAs still requires verification through biological experiments. However, with the development of biotechnology and more information becoming known about lncRNAs, their important functions in plant growth and development will be uncovered gradually.
Supplementary Materials
The following are available online at http://www.mdpi.com/2073-4425/7/3/11/s1. Supplementary Table S1–S7.
Acknowledgments
This study was supported by grants from the Special Research for Public Welfare in Forestry Industry (No.201304712) and the National Natural Science Foundation of China (Grant No.31171933).
Author Contributions
Dong Pei and Yi Zhao conceived and designed the research. Xiaobo Song collected the experimental data, performed the data analysis, and drafted the earlier versions of the manuscript. Liang Sun, Haitao Luo and Qingguo Ma involved the data analysis and partially revised the manuscript. Dong Pei and Yi Zhao partially revised the manuscript. All authors read, reviewed and approved the final manuscript. All the authors agreed on the contents of the paper and posted no conflicting interest.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Berg, C.C. Moraceae diversity in a global perspective. Biol. Skr. 2005, 55, 423–440. [Google Scholar]
- Nepal, M.P.; Ferguson, C.J. Phylogenetics of Morus (Moraceae) inferred from ITS and trnL-trnF sequence data. Syst. Bot. 2012, 37, 442–450. [Google Scholar] [CrossRef]
- Wang, M.; Gao, L.X.; Wang, J.; Li, J.; Yu, M.; Li, J.; Hou, A. Diels-Alder adducts with PTP1B inhibition from Morus Notabilis. Phytochem. 2015, 109, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Jia, L.; Zhang, D.; Qi, X.; Ma, B.; Xiang, Z.; He, N. Identification of the conserved and novel miRNAs in mulberry by high-throughput sequencing. PloS ONE 2014, 9, e104409. [Google Scholar]
- Ning, D.; Lu, B.; Zhang, Y. The processing technology of mulberry series product. China Fruit Veg. Process 2005, 5, 38–40. [Google Scholar]
- Nomura, T.; Fukai, T.; Hano, Y. Chemistry and biological activities of isoprenylated flavonoids from medicinal plants (moraceous plants and Glycyrrhiza species). Stud. Nat. Prod. Chem. 2003, 28, 199–256. [Google Scholar]
- Darias-Martín, J.; Lobo-Rodrigo, G.; Hernández-Cordero, J.; Díaz-Díaz, E.; Díaz-Romero, C. Alcoholic beverages obtained from black mulberry. Food Technol. Biotechnol. 2003, 41, 173–176. [Google Scholar]
- Venkatesh, K.R.; Chauhan, S. Mulberry: Life enhancer. J. Med. Plants Res. 2008, 2, 271–278. [Google Scholar]
- Yang, C.; Fang, X.; Wu, X.; Mao, Y.; Wang, L.; Chen, X. Transcriptional regulation of plant secondary metabolism. J. Integr. Plant Biol. 2012, 54, 703–712. [Google Scholar] [CrossRef] [PubMed]
- He, N.; Zhang, C.; Qi, X.; Zhao, S.; Tao, Z.; Yang, G.; Lee, T.H.; Wang, X.; Cai, Q.; Li, D.; et al. Draft genome sequence of the mulberry tree Morus notabilis. Nat. Commun. 2013. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.; Luo, Y.; Jia, L.; Qi, X.; Zeng, Q.; Xiang, Z.; He, N. Genome-wide identification and expression analyses of cytochrome P450 genes in mulberry (Morus notabilis). J. Integr. Plant Biol. 2014, 56, 887–901. [Google Scholar] [CrossRef] [PubMed]
- Carninci, P.; Kasukawa, T.; Katayama, S.; Gough, J.; Frith, M.C.; Maeda, N.; Oyama, R.; Racasi, T.; Lenhard, B.; Wells, C.; et al. The transcriptional landscape of the mammalian genome. Science 2005, 309, 1559–1563. [Google Scholar] [PubMed]
- Cheng, J.; Kapranov, P.; Drenkow, J.; Dike, S.; Brubaker, S.; Patel, S.; Long, J.; Stern, D.; Tammana, H.; Helt, G. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science 2005, 308, 1149–1154. [Google Scholar] [CrossRef] [PubMed]
- Wapinski, O.; Chang, H.Y. Long noncoding RNAs and human disease. Trends cell Biol. 2011, 21, 354–361. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.D.; Sung, S. Long noncoding RNA: Unveiling hidden layer of gene regulatory networks. Trends Plant Sci. 2012, 17, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Hangauer, M.J.; Vaughn, I.W.; McManus, M.T. Pervasive transcription of the human genome produces thousands of previously unidentified long intergenic noncoding RNAs. PLoS Genet. 2013. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Jung, C.; Xu, J.; Wang, H.; Deng, S.; Bernad, L.; Arenas-Huertero, C.; Chua, N. Genome-wide analysis uncovers regulation of long intergenic noncoding RNAs in Arabidopsis. Plant Cell 2012, 24, 4333–4345. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.H.; Wang, M.B. Molecular functions of long non-coding RNAs in plants. Genes 2012, 3, 176–190. [Google Scholar] [CrossRef] [PubMed]
- Rinn, J.L.; Chang, H.Y. Genome regulation by long noncoding RNAs. Ann. Rev. Biochem. 2012, 81, 145–166. [Google Scholar] [CrossRef] [PubMed]
- Lukiw, W.J.; Handley, P.; Wong, L.; McLachlan, D. BC200 RNA in normal human neocortex, non-Alzheimer dementia (NAD), and senile dementia of the Alzheimer type (AD). Neurochem. Res. 1992, 17, 591–597. [Google Scholar] [CrossRef] [PubMed]
- Boerner, S.; McGinnis, K.M. Computational identification and functional predictions of long noncoding RNA in Zea mays. PLoS ONE 2012. [Google Scholar] [CrossRef] [PubMed]
- Swiezewski, S.; Liu, F.; Magusin, A.; Dean, C. Cold-induced silencing by long antisense transcripts of an Arabidopsis Polycomb target. Nature 2009, 462, 799–802. [Google Scholar] [CrossRef] [PubMed]
- Heo, J.B.; Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 2011, 331, 76–79. [Google Scholar] [CrossRef] [PubMed]
- Franco-Zorrilla, J.M.; Valli, A.; Todesco, M.; Mateos, I.; Puga, M.I.; Rubio-Somoza, I.; Leyva, A.; Weigel, D.; Garcia, J.A.; Paz-Ares, J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 2007, 39, 1033–1037. [Google Scholar] [CrossRef] [PubMed]
- Rymarquis, L.A.; Kastenmayer, J.P.; Hüttenhofer, A.G.; Green, P.J. Diamonds in the rough: mRNA-like non-coding RNAs. Trends Plant Sci. 2008, 13, 329–334. [Google Scholar] [CrossRef] [PubMed]
- Shuai, P.; Liang, D.; Tang, S.; Zhang, Z.; Ye, C.; Su, Y.; Xia, X.; Yin, W. Genome-wide identification and functional prediction of novel and drought-responsive lincRNAs in Populus trichocarpa. J. Exp. Bot. 2014. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Quan, M.; Zhang, D. Genome-wide identification of novel long non-coding RNAs in Populus tomentosa tension wood, opposite wood and normal wood xylem by RNA-seq. Planta 2015, 241, 125–143. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Qi, X.; Zeng, Q.; Xiang, Z.; He, N. MorusDB: A resource for mulberry genomics and genome biology. Database 2014. [Google Scholar] [CrossRef] [PubMed]
- Pruitt, K.D.; Tatusova, T.; Maglott, D.R. NCBI Reference Sequence (RefSeq): A curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 2005, 33, 501–504. [Google Scholar] [CrossRef] [PubMed]
- Flicek, P.; Amode, M.R.; Barrell, D.; Beal, K.; Billis, K.; Brent, S.; Carvalho-Silva, D.; Clapham, P.; Coates, G.; Fizgerald, S. Ensembl 2014. Nucleic Acids Res. 2013. [Google Scholar] [CrossRef]
- Rosenbloom, K.R.; Armstrong, J.; Barber, G.P.; Casper, J.; Clawson, H.; Diekhans, M.; Dreszer, T.R.; Fujita, P.A.; Guruvadoo, L.; Haeussler, M. The UCSC genome browser database: 2015 Update. Nucleic Acids Res. 2015, 43, 670–681. [Google Scholar] [CrossRef] [PubMed]
- Wilming, L.G.; Gilbert, J.G.R.; Howe, K.; Trevanion, S.; Hubbard, T.; Harrow, J.L. The vertebrate genome annotation (Vega) database. Nucleic Acids Res. 2008, 36, D753–D760. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Yuan, J.; Li, H.; Li, M.; Zhao, G.; Bu, D.; Zhu, W.; Wu, W.; Chen, R.; Zhao, Y. NONCODEv4: Exploring the world of long non-coding RNA genes. Nucleic Acids Res. 2014, 42, D98–D103. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Pachter, L.; Salzberg, S.L. TopHat: Discovering splice junctions with RNA-seq. Bioinformatics 2009, 25, 1105–1111. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; Van Baren, M.J.; Salzberg, S.L.; Wold, B.L.; Pachter, L. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Luo, H.T.; Liao, Q.; Bu, D.; Zhao, G.; Liu, C.; Liu, Y.; Zhao, Y. Systematic study of human long intergenic non-coding RNAs and their impact on cancer. Sci. China Life Sci. 2013, 56, 324–334. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Sun, L.; Li, P.; Bu, D.; Cao, H.; Zhao, Y. Comprehensive characterization of 10,571 mouse large intergenic noncoding RNAs from whole transcriptome sequencing. PLoS ONE. 2013, 8, e70835. [Google Scholar] [CrossRef] [PubMed]
- Swarbreck, D.; Wilks, C.; Lamesch, P.; Berardini, T.Z.; Garcia-Hernandez, M.; Foerster, H.; Li, D.; Meyer, T.; Muller, R.; Ploeta, L. The Arabidopsis information resource (TAIR): Gene structure and function annotation. Nucleic Acids Res. 2008, 36, 1009–1014. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Liu, J.; Wang, H.; Wong, L.; Chua, N.H. PLncDB: Plant long noncoding RNA database. Bioinformatics 2013, 29, 1068–1071. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myser, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Cabili, M.N.; Trapnell, C.; Goff, L.; Koziol, M.; Tazon-Vega, B.; Regev, A.; Rinn, J.L. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 2011, 25, 1915–1927. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Luo, H.T.; Bu, D.; Zhao, G.; Yu, K.; Zhang, C.; Liu, Y.; Chen, R.; Zhao, Y. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 2013, 41, e166. [Google Scholar] [CrossRef] [PubMed]
- Springer, P.S.; Holding, D.R.; Groover, A.; Yordan, C.; Martienssen, R.A. The essential Mcm7 protein PROLIFERA is localized to the nucleus of dividing cells during the G (1) phase and is required maternally for early Arabidopsis development. Development 2000, 127, 1815–1822. [Google Scholar] [PubMed]
- Lin, Q.; Buckler, E.S.; Muse, S.V.; Walker, J.C. Molecular evolution of type 1 serine/threonine protein phosphatases. Mol. Phylogenet. Evolut. 1999, 12, 57–66. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Smith, R.D.; Walker, J.C. Plant protein phosphatases. Annu. Rev. Plant Biol. 1996, 47, 101–125. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Wang, M.; Wang, Q.; Wang, X.; Lin, J. Protein phosphatases 1 and 2A and the regulation of calcium uptake and pollen tube development in Picea wilsonii. Tree Physiol. 2006, 26, 1001–1012. [Google Scholar] [CrossRef] [PubMed]
- Voll, L.; Häusler, R.E.; Hecker, R.; Weber, A.; Weissenböck, G.; Fiene, G.; Waffenschmidt, S.; Flügge, U.I. The phenotype of the Arabidopsis cue1 mutant is not simply caused by a general restriction of the shikimate pathway. Plant J. 2003, 36, 301–317. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Tang, R.H.; Hao, Y.; Stevens, R.D.; Cook, C.W.; Ahn, S.M.; Jin, L.; Yang, Z.; Chen, L.; Guo, F. Nitric oxide represses the Arabidopsis floral transition. Science 2004, 305, 1968–1971. [Google Scholar] [CrossRef] [PubMed]
- Knappe, S.; Löttgert, T.; Schneider, A.; Voll, L.; Flügge, U.; Fischer, K. Characterization of two functional phosphoenolpyruvate/phosphate translocator (PPT) genes in Arabidopsis–AtPPT1 may be involved in the provision of signals for correct mesophyll development. Plant J. 2003, 36, 411–420. [Google Scholar] [CrossRef] [PubMed]
- Prabhakar, V.; Löttgert, T.; Geimer, S.; Dörmann, P.; Krüger, S.; Vijayakumar, V.; Schreiber, L.; Göbel, C.; Feussner, K.; Feussner, I. Phosphoenolpyruvate provision to plastids is essential for gametophyte and sporophyte development in Arabidopsis thaliana. Plant Cell 2010, 22, 2594–2617. [Google Scholar] [CrossRef] [PubMed]
- Shibagaki, N.; Rose, A.; McDermott, J.P.; Fujiwara, T.; Hayashi, H.; Yoneyama, T.; Davies, J.P. Selenate-resistant mutants of Arabidopsis thaliana identify Sultr1; 2, a sulfate transporter required for efficient transport of sulfate into roots. Plant J. 2002, 29, 475–486. [Google Scholar] [CrossRef] [PubMed]
- Buchner, P.; Parmar, S.; Kriegel, A.; Carpentier, M.; Hawkesford, M.J. The sulfate transporter family in wheat: Tissue-specific gene expression in relation to nutrition. Mol. Plant 2010, 3, 374–389. [Google Scholar] [CrossRef] [PubMed]
- Krusell, L.; Krause, K.; Ott, T.; Desbrosses, T.; Krämer, U.; Sato, S.; Nakamura, Y.; Tabata, S.; James, E.K.; Sandal, N. The sulfate transporter SST1 is crucial for symbiotic nitrogen fixation in Lotus japonicus root nodules. Plant Cell 2005, 17, 1625–1636. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.J.; Wang, Z.M.; Wang, M.; Wang, X.J. Widespread long noncoding RNAs as endogenous target mimics for microRNAs in plants. Plant Physiol. 2013, 161, 1875–1884. [Google Scholar] [CrossRef] [PubMed]
- Xin, M.; Wang, Y.; Yao, Y.; Song, N.; Hu, Z.; Qin, D.; Xie, C.; Peng, H.; Ni, Z.; Sun, Q. Identification and characterization of wheat long non-protein coding RNAs responsive to powdery mildew infection and heat stress by using microarray analysis and SBS sequencing. BMC Plant Biol. 2011. [Google Scholar] [CrossRef] [PubMed]
- Guttman, M.; Amit, I.; Garber, M.; French, C.; Lin, M.F.; Feldser, D.; Huarte, M.; Zuk, Q.; Carey, B.W.; Cassady, J.P. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 2009, 458, 223–227. [Google Scholar] [CrossRef] [PubMed]
- Du, T.A. Non-coding RNA: RNA stability control by Pol II. Nat. Rev. Mol. Cell Biol. 2013, 14, 128–129. [Google Scholar]
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martin, D.; Merkel, A.; Knowles, D.G. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 2012, 22, 1775–1789. [Google Scholar] [CrossRef] [PubMed]
- Liao, Q.; Shen, J.; Liu, J.; Sun, X.; Zhao, G.; Chang, Y.; Xu, L.; Li, X.; Zhao, Y.; Zheng, H. Genome-wide identification and functional annotation of Plasmodium falciparum long noncoding RNAs from RNA-seq data. Parasitol. Res. 2014, 113, 1269–1281. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.C.; Liao, J.Y.; Li, Z.Y.; Yu, Y.; Zhang, J.; Li, Q.; Qu, L.; Shu, W.; Chen, Y. Genome-wide screening and functional analysis identify a large number of long noncoding RNAs involved in the sexual reproduction of rice. Genome biology 2014. [Google Scholar] [CrossRef] [PubMed]
- Grote, P.; Wittler, L.; Hendrix, D.; Koch, F.; Wahrisch, S.; Beisaw, A.; Macura, K.; Blass, G.; Kellis, M.; Werber, M.; et al. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev. Cell 2013, 24, 206–214. [Google Scholar] [CrossRef] [PubMed]
- Katayama, S.; Tomaru, Y.; Kasukawa, T.; Waki, K.; Nakanishi, M.; Nakamura, M.; Nishida, N.; Yap, C.C.; Suzuki, M.; Kawai, K.; et al. Antisense transcription in the mammalian transcriptome. Science 2005, 309, 1564–1566. [Google Scholar] [PubMed]
- Dinger, M.E.; Amaral, P.P.; Mercer, T.R.; Pang, K.C.; Bruce, S.J.; Gardiner, B.B.; Askarian-Amiri, M.E.; Ru, K.; Soldà, G.; Simons, C. Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation. Genome Res. 2008, 18, 1433–1445. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Dinger, M.E.; Sunkin, S.M.; Mehler, M.F.; Mattick, J.S. Specific expression of long noncoding RNAs in the mouse brain. Proc. Natl. Acad. Sci. 2008, 105, 716–721. [Google Scholar] [CrossRef] [PubMed]
- Keniry, A.; Oxley, D.; Monnier, P.; Kyba, M.; Dandolo, L.; Smits, G.; Reik, W. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat. Biol. 2012, 14, 659–665. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.H.; Li, J.H.; Jiang, S.; Zhou, H.; Qu, L.H. ChIPBase: A database for decoding the transcriptional regulation of long non-coding RNA and microRNA genes from ChIP-Seq data. Nucleic Acids Rese. 2013, 41, D177–D187. [Google Scholar] [CrossRef] [PubMed]
- Sati, S.; Ghosh, S.; Jain, V.; Scaria, V.; Sengupta, S. Genome-wide analysis reveals distinct patterns of epigenetic features in long non-coding RNA loci. Nucleic Acids Res. 2012, 40, 10018–10031. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).