Regulation of mRNA Trafficking by Nuclear Pore Complexes
Abstract
:1. Introduction
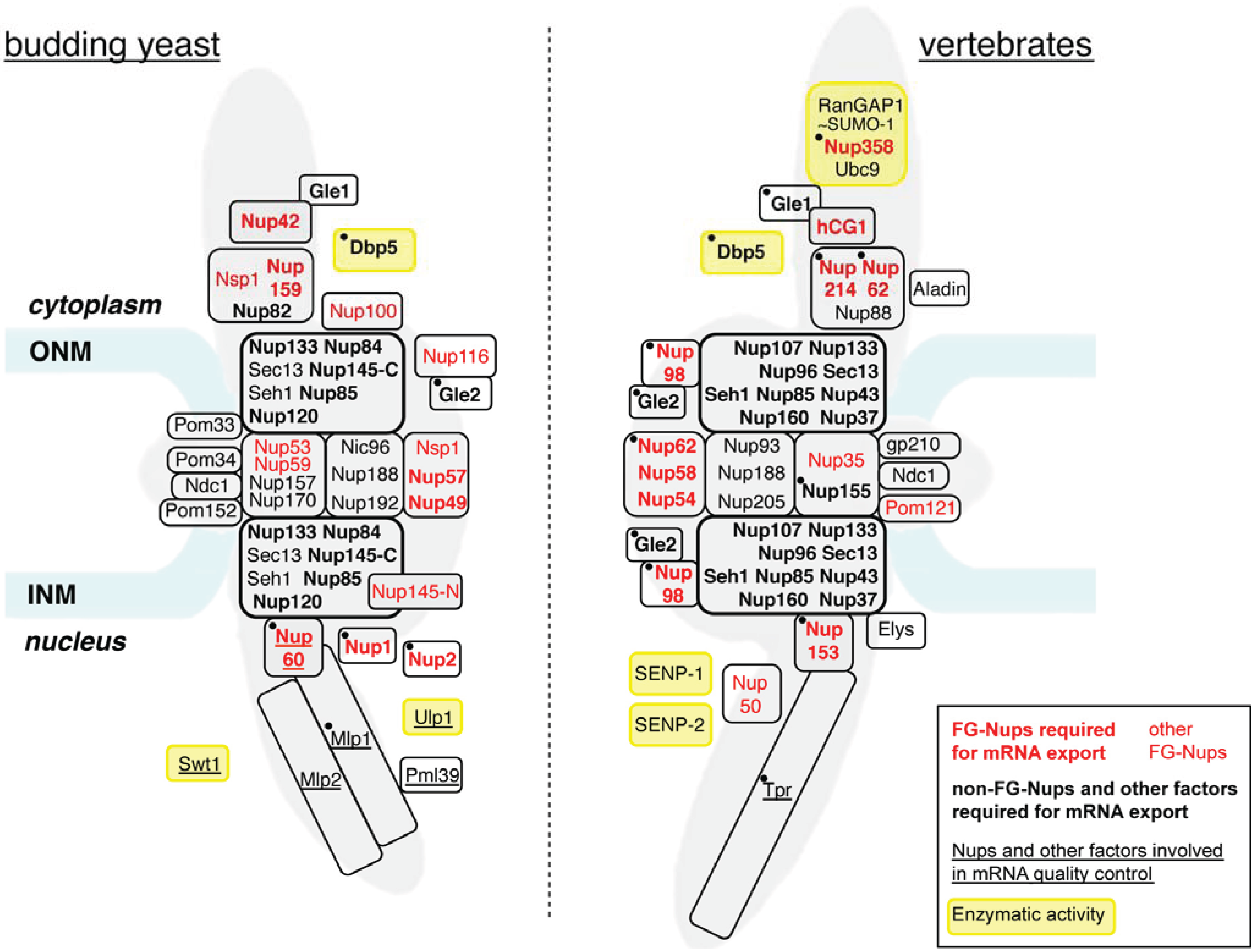
2. Role of NPC Components in Constitutive mRNA Export
2.1. FG-Nucleoporins Are Critical for mRNP Transport
| Phenotypes Observed upon Inactivation | ||||
|---|---|---|---|---|
| FG Repeats | Viability | mRNA Export | References | |
| Symmetric Nups | ||||
| Nup53 | FG | V | n.d. | |
| Nup59 | FG | V | n.d. | |
| Nup157 | V | + | ||
| Nup170 | V | + | ||
| Nup133 | ts | − | [32,33] | |
| Nup84 | ts | − | [11] | |
| Nup145-C | ts | − | [34,35,36] | |
| Nup85 | ts | − | [37] | |
| Nup120 | ts | − | [38,39] | |
| Sec13 | L a | + | ||
| Seh1 | cs | + | ||
| Nic96 | L | + | ||
| Nup188 | V | + | ||
| Nup192 | L | + | ||
| Nsp1 | FG, FxFG | L | + | |
| Nup57 | GLFG | L | − b | [28] |
| Nup49 | GLFG | L | − | [28,32] |
| Nup145-N | GLFG | V | + | |
| Nup116 | FG, GLFG | L/ts e | − c | [31] |
| Gle2 | ts | − | [40] | |
| Nup100 | GLFG | V | + | |
| Ndc1 | L | n.d. | ||
| Pom34 | V | n.d. | ||
| Pom152 | V | + | ||
| Pom33 | V | + | ||
| Asymmetric Nups | ||||
| Nup82 | L | − | [41,42] | |
| Nup159 | FG | L | − | [43] |
| Nup42 | FG | V | − d | [44,45,46] |
| Gle1 | L | − | [47] | |
| Dbp5 | L | − | [48,49] | |
| Nup60 | FxF | V | − | [50] |
| Nup1 | FxFG | L/ts e | − | [28,51,52] |
| Nup2 | FxFG | V | − b | [28] |
| Mlp1 | V | + | ||
| Mlp2 | V | + | ||
2.2. Non-FG-Nucleoporins May Provide Additional mRNP Binding Sites
2.3. NPC-Associated Factors Control mRNP Docking and Release
3. mRNA Quality Control at the Nuclear Basket of NPCs
3.1. Mlp1-2, Central Players in mRNA QC in Yeast
3.2. Multiple NPC-Associated Factors Contribute to mRNA QC in Yeast
| Localization | Molecular function | Inactivation | Overexpression | |||
|---|---|---|---|---|---|---|
| Pre-mRNA Leakage | mRNP Mutants Rescued | Nuclear mRNP Accumulation | ||||
| Mlp1 | Nuclear basket a | mRNP docking | + | ∆N-nab2 | + d,e | [85,92,94,99] |
| yra1-KR | ||||||
| Mlp2 | Nuclear basket a | mRNP docking | − | ∆N-nab2 | − d | [85,92,94,99] |
| yra1-8 | ||||||
| yra1-KR | ||||||
| tom1∆ | ||||||
| Pml39 | Nuclear basket a | mRNP docking ? | + | yra1-8 | + e,f | [96] |
| ∆N-nab2 | ||||||
| Nup60 | NPC | Nuclear basket assembly ? | + | n.d. | n.d. | [94] |
| Esc1 | Inner nuclear membrane- associated | Nuclear basket assembly ? | + | n.d. | n.d. | [100] |
| Ulp1 | NPC a,b | SUMO deconjugation | + | n.d. | n.d. | [100] |
| Swt1 | NPC c | RNA degradation | + | n.d. | + d | [98] |
3.3. mRNA QC at Nuclear Pores: A Conserved Pathway in Mammalian Cells?
4. Role of NPCs in the Control of mRNA Export during Cell Cycle, Development and Stress
4.1. Modifications of NPCs and Cell Cycle-Dependent Changes in mRNA Export
4.2. NPCs and Cell Cycle-Dependent Cytoplasmic Targeting of mRNAs
4.3. NPC-Associated Proteins and mRNA Export during Development
4.4. NPCs and Regulation of mRNA Export upon Stress
5. NPCs and Dysregulation of mRNA Export in Disease Situations
5.1. Alteration of the NPC Scaffold and Dysregulation of mRNA Export
5.2. Alteration of the Cytoplasmic Filaments and Dysregulation of mRNA Export
5.3. FG-Nucleoporins Targeted by Viruses and Inhibition of Cellular mRNA Export
6. Conclusions
Acknowledgments
Conflicts of Interest
References
- Wente, S.R.; Rout, M.P. The nuclear pore complex and nuclear transport. Cold Spring Harb. Perspect. Biol. 2010, 2, a000562. [Google Scholar] [CrossRef] [PubMed]
- Floch, A.G.; Palancade, B.; Doye, V. Fifty years of nuclear pores and nucleocytoplasmic transport studies: Multiple tools revealing complex rules. Methods Cell Biol. 2014, 122, 1–40. [Google Scholar] [PubMed]
- Terry, L.J.; Wente, S.R. Flexible gates: Dynamic topologies and functions for FG nucleoporins in nucleocytoplasmic transport. Eukaryot. Cell 2009, 8, 1814–1827. [Google Scholar] [CrossRef] [PubMed]
- Nino, C.A.; Herissant, L.; Babour, A.; Dargemont, C. mRNA nuclear export in yeast. Chem. Rev. 2013, 113, 8523–8545. [Google Scholar] [CrossRef] [PubMed]
- Oeffinger, M.; Zenklusen, D. To the pore and through the pore: A story of mRNA export kinetics. Biochim. Biophys. Acta 2012, 1819, 494–506. [Google Scholar]
- Tutucci, E.; Stutz, F. Keeping mRNPs in check during assembly and nuclear export. Nat. Rev. Mol. Cell Biol. 2011, 12, 377–384. [Google Scholar] [CrossRef] [PubMed]
- Culjkovic-Kraljacic, B.; Borden, K.L. Aiding and abetting cancer: mRNA export and the nuclear pore. Trends Cell Biol. 2013, 23, 328–335. [Google Scholar]
- Natalizio, B.J.; Wente, S.R. Postage for the messenger: Designating routes for nuclear mRNA export. Trends Cell Biol. 2013, 23, 365–373. [Google Scholar] [CrossRef] [PubMed]
- Bjork, P.; Wieslander, L. Mechanisms of mRNA export. Semin. Cell Dev. Biol. 2014, 32, 47–54. [Google Scholar] [CrossRef]
- Wozniak, R.; Burke, B.; Doye, V. Nuclear transport and the mitotic apparatus: An evolving relationship. Cell Mol. Life Sci. 2010, 67, 2215–2230. [Google Scholar] [CrossRef] [PubMed]
- Siniossoglou, S.; Wimmer, C.; Rieger, M.; Doye, V.; Tekotte, H.; Weise, C.; Emig, S.; Segref, A.; Hurt, E.C. A novel complex of nucleoporins, wich includes Sec13p and a Sec13p homolog, is essential for normal nuclear pores. Cell 1996, 84, 265–275. [Google Scholar] [CrossRef] [PubMed]
- Boehmer, T.; Enninga, J.; Dales, S.; Blobel, G.; Zhong, H. Depletion of a single nucleoporin, Nup107, prevents the assembly of a subset of nucleoporins into the nuclear pore complex. Proc. Natl. Acad. Sci. USA 2003, 100, 981–985. [Google Scholar] [CrossRef] [PubMed]
- Walther, T.C.; Alves, A.; Pickersgill, H.; Loiodice, I.; Hetzer, M.; Galy, V.; Hulsmann, B.B.; Kocher, T.; Wilm, M.; Allen, T.; et al. The conserved Nup107-160 complex is critical for nuclear pore complex assembly. Cell 2003, 113, 195–206. [Google Scholar] [CrossRef] [PubMed]
- Vasu, S.; Shah, S.; Orjalo, A.; Park, M.; Fischer, W.H.; Forbes, D.J. Novel vertebrate nucleoporins Nup133 and Nup160 play a role in mRNA export. J. Cell Biol. 2001, 155, 339–354. [Google Scholar] [CrossRef] [PubMed]
- Amberg, D.C.; Goldstein, A.L.; Cole, C.N. Isolation and characterization of RAT1: An essential gene of Saccharomyces cerevisiae required for the efficient nucleocytoplasmic trafficking of mRNA. Genes Dev. 1992, 6, 1173–1189. [Google Scholar] [CrossRef] [PubMed]
- Doye, V.; Hurt, E. From nucleoporins to nuclear pore complexes. Curr. Opin. Cell Biol. 1997, 9, 401–411. [Google Scholar] [CrossRef] [PubMed]
- Sheinberger, J.; Shav-Tal, Y. The dynamic pathway of nuclear RNA in eukaryotes. Nucleus 2013, 4, 195–205. [Google Scholar] [CrossRef] [PubMed]
- Kiseleva, E.; Goldberg, M.W.; Allen, T.D.; Akey, C.W. Active nuclear pore complexes in Chironomus: Visualization of transporter configurations related to mRNP export. J. Cell Sci. 1998, 111, 223–236. [Google Scholar] [PubMed]
- Mehlin, H.; Daneholt, B.; Skoglund, U. Structural interaction between the nuclear pore complex and a specific translocating RNP particle. J. Cell Biol. 1995, 129, 1205–1216. [Google Scholar] [CrossRef] [PubMed]
- Grunwald, D.; Singer, R.H. In vivo imaging of labelled endogenous beta-actin mRNA during nucleocytoplasmic transport. Nature 2010, 467, 604–607. [Google Scholar] [CrossRef] [PubMed]
- Mor, A.; Suliman, S.; Ben-Yishay, R.; Yunger, S.; Brody, Y.; Shav-Tal, Y. Dynamics of single mRNP nucleocytoplasmic transport and export through the nuclear pore in living cells. Nat. Cell Biol. 2010, 12, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Siebrasse, J.P.; Kaminski, T.; Kubitscheck, U. Nuclear export of single native mRNA molecules observed by light sheet fluorescence microscopy. Proc. Natl. Acad. Sci. USA 2012, 109, 9426–9431. [Google Scholar] [CrossRef] [PubMed]
- Katahira, J.; Strasser, K.; Podtelejnikov, A.; Mann, M.; Jung, J.U.; Hurt, E. The Mex67p-mediated nuclear mRNA export pathway is conserved from yeast to human. EMBO J. 1999, 18, 2593–2609. [Google Scholar] [CrossRef] [PubMed]
- Strasser, K.; Bassler, J.; Hurt, E. Binding of the Mex67p/Mtr2p heterodimer to FXFG, GLFG, and FG repeat nucleoporins is essential for nuclear mRNA export. J. Cell Biol. 2000, 150, 695–706. [Google Scholar] [CrossRef] [PubMed]
- Bachi, A.; Braun, I.C.; Rodrigues, J.P.; Pante, N.; Ribbeck, K.; von Kobbe, C.; Kutay, U.; Wilm, M.; Gorlich, D.; Carmo-Fonseca, M.; et al. The C-terminal domain of TAP interacts with the nuclear pore complex and promotes export of specific CTE-bearing RNA substrates. RNA 2000, 6, 136–158. [Google Scholar] [CrossRef] [PubMed]
- Strawn, L.A.; Shen, T.; Wente, S.R. The GLFG regions of Nup116p and Nup100p serve as binding sites for both Kap95p and Mex67p at the nuclear pore complex. J. Biol. Chem. 2001, 276, 6445–6452. [Google Scholar] [CrossRef]
- Levesque, L.; Guzik, B.; Guan, T.; Coyle, J.; Black, B.E.; Rekosh, D.; Hammarskjold, M.L.; Paschal, B.M. RNA export mediated by tap involves NXT1-dependent interactions with the nuclear pore complex. J. Biol. Chem. 2001, 276, 44953–44962. [Google Scholar] [CrossRef] [PubMed]
- Terry, L.J.; Wente, S.R. Nuclear mRNA export requires specific FG nucleoporins for translocation through the nuclear pore complex. J. Cell Biol. 2007, 178, 1121–1132. [Google Scholar] [CrossRef] [PubMed]
- Strawn, L.A.; Shen, T.; Shulga, N.; Goldfarb, D.S.; Wente, S.R. Minimal nuclear pore complexes define FG repeat domains essential for transport. Nat. Cell Biol. 2004, 6, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Giaever, G.; Chu, A.M.; Ni, L.; Connelly, C.; Riles, L.; Veronneau, S.; Dow, S.; Lucau-Danila, A.; Anderson, K.; Andre, B.; et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 2002, 418, 387–391. [Google Scholar] [CrossRef] [PubMed]
- Wente, S.R.; Blobel, G. A temperature-sensitive NUP116 null mutant forms a nuclear envelope seal over the yeast nuclear pore complex thereby blocking nucleocytoplasmic traffic. J. Cell Biol. 1993, 123, 275–284. [Google Scholar] [CrossRef] [PubMed]
- Doye, V.; Wepf, R.; Hurt, E.C. A novel nuclear pore protein Nup133p with distinct roles in poly(A)+ RNA transport and nuclear pore distribution. EMBO J. 1994, 13, 6062–6075. [Google Scholar] [PubMed]
- Li, O.; Heath, C.V.; Amberg, D.C.; Dockendorff, T.C.; Copeland, C.S.; Snyder, M.; Cole, C.N. Mutation or deletion of the Saccharomyces cerevisiae RAT3/NUP133 gene causes temperature-dependent nuclear accumulation of poly(A)+ RNA and constitutive clustering of nuclear pore complexes. Mol. Biol. Cell 1995, 6, 401–417. [Google Scholar] [CrossRef]
- Fabre, E.; Boelens, W.C.; Wimmer, C.; Mattaj, I.W.; Hurt, E.C. Nup145p is required for nuclear export of mRNA and binds homopolymeric RNA in vitro via a novel conserved motif. Cell 1994, 78, 275–289. [Google Scholar] [CrossRef] [PubMed]
- Dockendorff, T.C.; Heath, C.V.; Goldstein, A.L.; Snay, C.A.; Cole, C.N. C-Terminal truncations of the yeast nucleoporin Nup145p produce a rapid temperature-conditional mRNA export defect and alterations to nuclear structure. Mol. Cell. Biol. 1997, 17, 906–920. [Google Scholar] [PubMed]
- Teixeira, M.T.; Siniossoglou, S.; Podtelejnikov, S.; Benichou, J.C.; Mann, M.; Dujon, B.; Hurt, E.; Fabre, E. Two functionally distinct domains generated by in vivo cleavage of Nup145p: A novel biogenesis pathway for nucleoporins. EMBO J. 1997, 16, 5086–5097. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, A.L.; Snay, C.A.; Heath, C.V.; Cole, C.N. Pleiotropic nuclear defects associated with a conditional allele of the novel nucleoporin Rat9p/Nup85p. Mol. Biol. Cell 1996, 7, 917–934. [Google Scholar] [CrossRef] [PubMed]
- Aitchison, J.D.; Blobel, G.; Rout, M.P. Nup120p: A yeast nucleoporin required for NPC distribution and mRNA transport. J. Cell Biol. 1995, 131, 1659–1675. [Google Scholar] [CrossRef] [PubMed]
- Heath, C.V.; Copeland, C.S.; Amberg, D.C.; del Priore, V.; Snyder, M.; Cole, C.N. Nuclear pore complex clustering and nuclear accumulation of poly(A)+ RNA associated with mutation of the Saccharomyces cerevisiae RAT2/NUP120 gene. J. Cell Biol. 1995, 131, 1677–1697. [Google Scholar] [CrossRef] [PubMed]
- Murphy, R.; Watkins, J.L.; Wente, S.R. GLE2, a Saccharomyces cerevisiae homologue of the Schizosaccharomyces pombe export factor RAE1, is required for nuclear pore complex structure and function. Mol. Biol. Cell 1996, 7, 1921–1937. [Google Scholar] [CrossRef] [PubMed]
- Grandi, P.; Emig, S.; Weise, C.; Hucho, F.; Pohl, T.; Hurt, E.C. A novel nuclear pore protein Nup82p which specifically binds to a fraction of Nsp1p. J. Cell Biol. 1995, 130, 1263–1273. [Google Scholar] [CrossRef] [PubMed]
- Hurwitz, M.E.; Blobel, G. NUP82 is an essential yeast nucleoporin required for poly(A)+ RNA export. J. Cell Biol. 1995, 130, 1275–1281. [Google Scholar] [CrossRef] [PubMed]
- Gorsch, L.C.; Dockendorff, T.C.; Cole, C.N. A conditional allele of the novel repeat-containing yeast nucleoporin RAT7/NUP159 causes both rapid cessation of mRNA export and reversible clustering of nuclear pore complexes. J. Cell Biol. 1995, 129, 939–955. [Google Scholar] [CrossRef] [PubMed]
- Saavedra, C.A.; Hammell, C.M.; Heath, C.V.; Cole, C.N. Yeast heat shock mRNAs are exported through a distinct pathway defined by Rip1p. Genes Dev. 1997, 11, 2845–2856. [Google Scholar] [CrossRef] [PubMed]
- Stutz, F.; Kantor, J.; Zhang, D.; McCarthy, T.; Neville, M.; Rosbash, M. The yeast nucleoporin rip1p contributes to multiple export pathways with no essential role for its FG-repeat region. Genes Dev. 1997, 11, 2857–2868. [Google Scholar] [CrossRef] [PubMed]
- Vainberg, I.E.; Dower, K.; Rosbash, M. Nuclear export of heat shock and non-heat-shock mRNA occurs via similar pathways. Mol. Cell. Biol. 2000, 20, 3996–4005. [Google Scholar] [CrossRef] [PubMed]
- Murphy, R.; Wente, S.R. An RNA-export mediator with an essential nuclear export signal. Nature 1996, 383, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Snay-Hodge, C.A.; Colot, H.V.; Goldstein, A.L.; Cole, C.N. Dbp5p/Rat8p is a yeast nuclear pore-associated DEAD-box protein essential for RNA export. EMBO J. 1998, 17, 2663–2676. [Google Scholar]
- Tseng, S.S.; Weaver, P.L.; Liu, Y.; Hitomi, M.; Tartakoff, A.M.; Chang, T.H. Dbp5p, a cytosolic RNA helicase, is required for poly(A)+ RNA export. EMBO J. 1998, 17, 2651–2662. [Google Scholar] [CrossRef] [PubMed]
- Fischer, T.; Strasser, K.; Racz, A.; Rodriguez-Navarro, S.; Oppizzi, M.; Ihrig, P.; Lechner, J.; Hurt, E. The mRNA export machinery requires the novel Sac3p-Thp1p complex to dock at the nucleoplasmic entrance of the nuclear pores. EMBO J. 2002, 21, 5843–5852. [Google Scholar] [CrossRef] [PubMed]
- Bogerd, A.M.; Hoffman, J.A.; Amberg, D.C.; Fink, G.R.; Davis, L.I. Nup1 mutants exhibit pleiotropic defects in nuclear pore complex function. J. Cell Biol. 1994, 127, 319–332. [Google Scholar] [CrossRef] [PubMed]
- Schlaich, N.L.; Hurt, E.C. Analysis of nucleocytoplasmic transport and nuclear envelope structure in yeast disrupted for the gene encoding the nuclear pore protein Nup1p. Eur. J. Cell Biol. 1995, 67, 8–14. [Google Scholar] [PubMed]
- Bastos, R.; Lin, A.; Enarson, M.; Burke, B. Targeting and function in mRNA export of nuclear pore complex protein Nup153. J. Cell Biol. 1996, 134, 1141–1156. [Google Scholar] [CrossRef] [PubMed]
- Ullman, K.S.; Shah, S.; Powers, M.A.; Forbes, D.J. The nucleoporin nup153 plays a critical role in multiple types of nuclear export. Mol. Biol. Cell 1999, 10, 649–664. [Google Scholar] [CrossRef] [PubMed]
- Dargemont, C.; Schmidt-Zachmann, M.S.; Kuhn, L.C. Direct interaction of nucleoporin p62 with mRNA during its export from the nucleus. J. Cell Sci. 1995, 108, 257–263. [Google Scholar] [PubMed]
- Powers, M.A.; Forbes, D.J.; Dahlberg, J.E.; Lund, E. The vertebrate GLFG nucleoporin, Nup98, is an essential component of multiple RNA export pathways. J. Cell Biol. 1997, 136, 241–250. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, C.E.; Fornerod, M.; Kasper, L.H.; van Deursen, J.M. RAE1 is a shuttling mRNA export factor that binds to a GLEBS-like NUP98 motif at the nuclear pore complex through multiple domains. J. Cell Biol. 1999, 145, 237–254. [Google Scholar] [CrossRef] [PubMed]
- Adams, R.L.; Terry, L.J.; Wente, S.R. Nucleoporin FG domains facilitate mRNP remodeling at the cytoplasmic face of the nuclear pore complex. Genetics 2014, 197, 1213–1224. [Google Scholar] [CrossRef] [PubMed]
- Van Deursen, J.; Boer, J.; Kasper, L.; Grosveld, G. G2 arrest and impaired nucleocytoplasmic transport in mouse embryos lacking the proto-oncogene CAN/Nup214. EMBO J. 1996, 15, 5574–5583. [Google Scholar]
- Forler, D.; Rabut, G.; Ciccarelli, F.D.; Herold, A.; Kocher, T.; Niggeweg, R.; Bork, P.; Ellenberg, J.; Izaurralde, E. RanBP2/Nup358 provides a major binding site for NXF1-p15 dimers at the nuclear pore complex and functions in nuclear mRNA export. Mol. Cell. Biol. 2004, 24, 1155–1167. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Aguilera, C.; Askjaer, P. Dissecting the NUP107 complex: Multiple components and even more functions. Nucleus 2012, 3, 340–348. [Google Scholar]
- Yao, W.; Lutzmann, M.; Hurt, E. A versatile interaction platform on the Mex67-Mtr2 receptor creates an overlap between mRNA and ribosome export. EMBO J. 2008, 27, 6–16. [Google Scholar] [CrossRef] [PubMed]
- Santos-Rosa, H.; Moreno, H.; Simos, G.; Segref, A.; Fahrenkrog, B.; Pante, N.; Hurt, E. Nuclear mRNA export requires complex formation between Mex67p and Mtr2p at the nuclear pores. Mol. Cell. Biol. 1998, 18, 6826–6838. [Google Scholar]
- Ball, J.R.; Dimaano, C.; Bilak, A.; Kurchan, E.; Zundel, M.T.; Ullman, K.S. Sequence preference in RNA recognition by the nucleoporin Nup153. J. Biol. Chem. 2007, 282, 8734–8740. [Google Scholar] [CrossRef] [PubMed]
- Handa, N.; Kukimoto-Niino, M.; Akasaka, R.; Kishishita, S.; Murayama, K.; Terada, T.; Inoue, M.; Kigawa, T.; Kose, S.; Imamoto, N.; et al. The crystal structure of mouse Nup35 reveals atypical RNP motifs and novel homodimerization of the RRM domain. J. Mol. Biol. 2006, 363, 114–124. [Google Scholar] [CrossRef] [PubMed]
- Kassube, S.A.; Stuwe, T.; Lin, D.H.; Antonuk, C.D.; Napetschnig, J.; Blobel, G.; Hoelz, A. Crystal structure of the N-terminal domain of Nup358/RanBP2. J. Mol. Biol. 2012, 423, 752–765. [Google Scholar] [CrossRef]
- Devos, D.; Dokudovskaya, S.; Williams, R.; Alber, F.; Eswar, N.; Chait, B.T.; Rout, M.P.; Sali, A. Simple fold composition and modular architecture of the nuclear pore complex. Proc. Natl. Acad. Sci. USA 2006, 103, 2172–2177. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Seo, H.S.; Blobel, G.; Hoelz, A. Structural and functional analysis of the interaction between the nucleoporin Nup98 and the mRNA export factor Rae1. Proc. Natl. Acad. Sci. USA 2010, 107, 10406–10411. [Google Scholar] [CrossRef] [PubMed]
- Kraemer, D.; Blobel, G. mRNA binding protein mrnp 41 localizes to both nucleus and cytoplasm. Proc. Natl Acad. Sci. USA 1997, 94, 9119–9124. [Google Scholar] [CrossRef] [PubMed]
- Bharathi, A.; Ghosh, A.; Whalen, W.A.; Yoon, J.H.; Pu, R.; Dasso, M.; Dhar, R. The human RAE1 gene is a functional homologue of Schizosaccharomyces pombe rae1 gene involved in nuclear export of Poly(A)+ RNA. Gene 1997, 198, 251–258. [Google Scholar] [CrossRef] [PubMed]
- Satterly, N.; Tsai, P.L.; van Deursen, J.; Nussenzveig, D.R.; Wang, Y.; Faria, P.A.; Levay, A.; Levy, D.E.; Fontoura, B.M. Influenza virus targets the mRNA export machinery and the nuclear pore complex. Proc. Natl. Acad. Sci. USA 2007, 104, 1853–1858. [Google Scholar] [PubMed]
- Jani, D.; Lutz, S.; Marshall, N.J.; Fischer, T.; Kohler, A.; Ellisdon, A.M.; Hurt, E.; Stewart, M. Sus1, Cdc31, and the Sac3 CID region form a conserved interaction platform that promotes nuclear pore association and mRNA export. Mol. Cell 2009, 33, 727–737. [Google Scholar] [CrossRef] [PubMed]
- Jani, D.; Lutz, S.; Hurt, E.; Laskey, R.A.; Stewart, M.; Wickramasinghe, V.O. Functional and structural characterization of the mammalian TREX-2 complex that links transcription with nuclear messenger RNA export. Nucleic Acids Res. 2012, 40, 4562–4573. [Google Scholar] [CrossRef] [PubMed]
- Umlauf, D.; Bonnet, J.; Waharte, F.; Fournier, M.; Stierle, M.; Fischer, B.; Brino, L.; Devys, D.; Tora, L. The human TREX-2 complex is stably associated with the nuclear pore basket. J. Cell Sci. 2013, 126, 2656–2667. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, C.; von Kobbe, C.; Bachi, A.; Pante, N.; Rodrigues, J.P.; Boscheron, C.; Rigaut, G.; Wilm, M.; Seraphin, B.; Carmo-Fonseca, M.; et al. Dbp5, a DEAD-box protein required for mRNA export, is recruited to the cytoplasmic fibrils of nuclear pore complex via a conserved interaction with CAN/Nup159p. EMBO J. 1999, 18, 4332–4347. [Google Scholar] [CrossRef] [PubMed]
- Lund, M.K.; Guthrie, C. The DEAD-box protein Dbp5p is required to dissociate Mex67p from exported mRNPs at the nuclear rim. Mol. Cell 2005, 20, 645–651. [Google Scholar] [CrossRef] [PubMed]
- Tran, E.J.; Zhou, Y.; Corbett, A.H.; Wente, S.R. The DEAD-box protein Dbp5 controls mRNA export by triggering specific RNA: Protein remodeling events. Mol. Cell 2007, 28, 850–859. [Google Scholar] [CrossRef] [PubMed]
- Alcazar-Roman, A.R.; Tran, E.J.; Guo, S.; Wente, S.R. Inositol hexakisphosphate and Gle1 activate the DEAD-box protein Dbp5 for nuclear mRNA export. Nat. Cell Biol. 2006, 8, 711–716. [Google Scholar]
- Weirich, C.S.; Erzberger, J.P.; Flick, J.S.; Berger, J.M.; Thorner, J.; Weis, K. Activation of the DExD/H-box protein Dbp5 by the nuclear-pore protein Gle1 and its coactivator InsP6 is required for mRNA export. Nat. Cell Biol. 2006, 8, 668–676. [Google Scholar] [CrossRef] [PubMed]
- Von Moeller, H.; Basquin, C.; Conti, E. The mRNA export protein DBP5 binds RNA and the cytoplasmic nucleoporin NUP214 in a mutually exclusive manner. Nat. Struct. Mol. Biol. 2009, 16, 247–254. [Google Scholar]
- Noble, K.N.; Tran, E.J.; Alcazar-Roman, A.R.; Hodge, C.A.; Cole, C.N.; Wente, S.R. The Dbp5 cycle at the nuclear pore complex during mRNA export II: Nucleotide cycling and mRNP remodeling by Dbp5 are controlled by Nup159 and Gle1. Genes Dev. 2011, 25, 1065–1077. [Google Scholar] [CrossRef] [PubMed]
- Montpetit, B.; Thomsen, N.D.; Helmke, K.J.; Seeliger, M.A.; Berger, J.M.; Weis, K. A conserved mechanism of DEAD-box ATPase activation by nucleoporins and InsP6 in mRNA export. Nature 2011, 472, 238–242. [Google Scholar] [CrossRef] [PubMed]
- Hodge, C.A.; Tran, E.J.; Noble, K.N.; Alcazar-Roman, A.R.; Ben-Yishay, R.; Scarcelli, J.J.; Folkmann, A.W.; Shav-Tal, Y.; Wente, S.R.; Cole, C.N. The Dbp5 cycle at the nuclear pore complex during mRNA export I: Dbp5 mutants with defects in RNA binding and ATP hydrolysis define key steps for Nup159 and Gle1. Genes Dev. 2011, 25, 1052–1064. [Google Scholar] [CrossRef]
- Schmid, M.; Jensen, T.H. Transcription-associated quality control of mRNP. Biochim. Biophys. Acta 2013, 1829, 158–168. [Google Scholar]
- Kosova, B.; Pante, N.; Rollenhagen, C.; Podtelejnikov, A.; Mann, M.; Aebi, U.; Hurt, E. Mlp2p, a component of nuclear pore attached intranuclear filaments, associates with nic96p. J. Biol. Chem. 2000, 275, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Rajanala, K.; Nandicoori, V.K. Localization of nucleoporin Tpr to the nuclear pore complex is essential for Tpr mediated regulation of the export of unspliced RNA. PLoS One 2012, 7, e29921. [Google Scholar] [CrossRef] [PubMed]
- Bangs, P.; Burke, B.; Powers, C.; Craig, R.; Purohit, A.; Doxsey, S. Functional analysis of Tpr: Identification of nuclear pore complex association and nuclear localization domains and a role in mRNA export. J. Cell Biol. 1998, 143, 1801–1812. [Google Scholar] [CrossRef] [PubMed]
- Green, D.M.; Johnson, C.P.; Hagan, H.; Corbett, A.H. The C-terminal domain of myosin-like protein 1 (Mlp1p) is a docking site for heterogeneous nuclear ribonucleoproteins that are required for mRNA export. Proc. Natl. Acad. Sci. USA 2003, 100, 1010–1015. [Google Scholar] [CrossRef] [PubMed]
- Niepel, M.; Molloy, K.R.; Williams, R.; Farr, J.C.; Meinema, A.C.; Vecchietti, N.; Cristea, I.M.; Chait, B.T.; Rout, M.P.; Strambio-De-Castillia, C. The nuclear basket proteins Mlp1p and Mlp2p are part of a dynamic interactome including Esc1p and the proteasome. Mol. Biol. Cell 2013, 24, 3920–3938. [Google Scholar] [CrossRef] [PubMed]
- Bretes, H.; Rouviere, J.O.; Leger, T.; Oeffinger, M.; Devaux, F.; Doye, V.; Palancade, B. Sumoylation of the THO complex regulates the biogenesis of a subset of mRNPs. Nucleic Acids Res. 2014, 42, 5043–5058. [Google Scholar] [CrossRef] [PubMed]
- Fasken, M.B.; Stewart, M.; Corbett, A.H. Functional significance of the interaction between the mRNA-binding protein, Nab2, and the nuclear pore-associated protein, Mlp1, in mRNA export. J. Biol. Chem. 2008, 283, 27130–27143. [Google Scholar] [CrossRef] [PubMed]
- Vinciguerra, P.; Iglesias, N.; Camblong, J.; Zenklusen, D.; Stutz, F. Perinuclear Mlp proteins downregulate gene expression in response to a defect in mRNA export. EMBO J. 2005, 24, 813–823. [Google Scholar] [CrossRef] [PubMed]
- Hector, R.E.; Nykamp, K.R.; Dheur, S.; Anderson, J.T.; Non, P.J.; Urbinati, C.R.; Wilson, S.M.; Minvielle-Sebastia, L.; Swanson, M.S. Dual requirement for yeast hnRNP Nab2p in mRNA poly(A) tail length control and nuclear export. EMBO J. 2002, 21, 1800–1810. [Google Scholar] [CrossRef] [PubMed]
- Galy, V.; Gadal, O.; Fromont-Racine, M.; Romano, A.; Jacquier, A.; Nehrbass, U. Nuclear retention of unspliced mRNAs in yeast is mediated by perinuclear Mlp1. Cell 2004, 116, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Lo, C.W.; Kaida, D.; Nishimura, S.; Matsuyama, A.; Yashiroda, Y.; Taoka, H.; Ishigami, K.; Watanabe, H.; Nakajima, H.; Tani, T.; et al. Inhibition of splicing and nuclear retention of pre-mRNA by spliceostatin A in fission yeast. Biochem. Biophys. Res. Commun. 2007, 364, 573–577. [Google Scholar] [CrossRef] [PubMed]
- Palancade, B.; Zuccolo, M.; Loeillet, S.; Nicolas, A.; Doye, V. Pml39, a novel protein of the nuclear periphery required for nuclear retention of improper messenger ribonucleoparticles. Mol. Biol. Cell 2005, 16, 5258–5268. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Wu, C.Y.; Blobel, G. Mlp-dependent anchorage and stabilization of a desumoylating enzyme is required to prevent clonal lethality. J. Cell Biol. 2004, 167, 605–611. [Google Scholar] [CrossRef] [PubMed]
- Skruzny, M.; Schneider, C.; Racz, A.; Weng, J.; Tollervey, D.; Hurt, E. An endoribonuclease functionally linked to perinuclear mRNP quality control associates with the nuclear pore complexes. PLoS Biol. 2009, 7, e8. [Google Scholar] [CrossRef] [PubMed]
- Iglesias, N.; Tutucci, E.; Gwizdek, C.; Vinciguerra, P.; von Dach, E.; Corbett, A.H.; Dargemont, C.; Stutz, F. Ubiquitin-mediated mRNP dynamics and surveillance prior to budding yeast mRNA export. Genes Dev. 2010, 24, 1927–1938. [Google Scholar] [CrossRef] [PubMed]
- Lewis, A.; Felberbaum, R.; Hochstrasser, M. A nuclear envelope protein linking nuclear pore basket assembly, SUMO protease regulation, and mRNA surveillance. J. Cell Biol. 2007, 178, 813–827. [Google Scholar] [CrossRef] [PubMed]
- Dziembowski, A.; Ventura, A.P.; Rutz, B.; Caspary, F.; Faux, C.; Halgand, F.; Laprevote, O.; Seraphin, B. Proteomic analysis identifies a new complex required for nuclear pre-mRNA retention and splicing. EMBO J. 2004, 23, 4847–4856. [Google Scholar] [CrossRef] [PubMed]
- Hackmann, A.; Wu, H.; Schneider, U.M.; Meyer, K.; Jung, K.; Krebber, H. Quality control of spliced mRNAs requires the shuttling SR proteins Gbp2 and Hrb1. Nat. Commun. 2014, 5, 3123. [Google Scholar] [CrossRef] [PubMed]
- Kuss, S.K.; Mata, M.A.; Zhang, L.; Fontoura, B.M. Nuclear imprisonment: Viral strategies to arrest host mRNA nuclear export. Viruses 2013, 5, 1824–1849. [Google Scholar] [CrossRef] [PubMed]
- Coyle, J.H.; Bor, Y.C.; Rekosh, D.; Hammarskjold, M.L. The Tpr protein regulates export of mRNAs with retained introns that traffic through the Nxf1 pathway. RNA 2011, 17, 1344–1356. [Google Scholar] [CrossRef] [PubMed]
- Hase, M.E.; Cordes, V.C. Direct interaction with nup153 mediates binding of Tpr to the periphery of the nuclear pore complex. Mol. Biol. Cell 2003, 14, 1923–1940. [Google Scholar] [CrossRef]
- Yap, K.; Lim, Z.Q.; Khandelia, P.; Friedman, B.; Makeyev, E.V. Coordinated regulation of neuronal mRNA steady-state levels through developmentally controlled intron retention. Genes Dev. 2012, 26, 1209–1223. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, P.; Wang, Y.; Wei, J.H.; van Deursen, J.; Yu, H.; Malureanu, L.; Dasso, M.; Forbes, D.J.; Levy, D.E.; Seemann, J.; et al. Nucleoporin levels regulate cell cycle progression and phase-specific gene expression. Dev. Cell 2008, 15, 657–667. [Google Scholar] [CrossRef] [PubMed]
- Wozniak, R.W.; Goldfarb, D.S. Cyclin-like oscillations in levels of the nucleoporin Nup96 control G1/S progression. Dev. Cell 2008, 15, 643–644. [Google Scholar] [CrossRef] [PubMed]
- Glavy, J.S.; Krutchinsky, A.N.; Cristea, I.M.; Berke, I.C.; Boehmer, T.; Blobel, G.; Chait, B.T. Cell-cycle-dependent phosphorylation of the nuclear pore Nup107-160 subcomplex. Proc. Natl. Acad. Sci. USA 2007, 104, 3811–3816. [Google Scholar] [CrossRef] [PubMed]
- Laurell, E.; Beck, K.; Krupina, K.; Theerthagiri, G.; Bodenmiller, B.; Horvath, P.; Aebersold, R.; Antonin, W.; Kutay, U. Phosphorylation of Nup98 by multiple kinases is crucial for NPC disassembly during mitotic entry. Cell 2011, 144, 539–550. [Google Scholar] [CrossRef] [PubMed]
- Makhnevych, T.; Lusk, C.P.; Anderson, A.M.; Aitchison, J.D.; Wozniak, R.W. Cell cycle regulated transport controlled by alterations in the nuclear pore complex. Cell 2003, 115, 813–823. [Google Scholar] [CrossRef] [PubMed]
- Brickner, D.G.; Brickner, J.H. Cdk phosphorylation of a nucleoporin controls localization of active genes through the cell cycle. Mol. Biol. Cell 2010, 21, 3421–3432. [Google Scholar] [CrossRef] [PubMed]
- Capelson, M.; Doucet, C.; Hetzer, M.W. Nuclear pore complexes: Guardians of the nuclear genome. Cold Spring Harb. Symp. Quant. Biol. 2010, 75, 585–597. [Google Scholar] [PubMed]
- Colon-Ramos, D.A.; Salisbury, J.L.; Sanders, M.A.; Shenoy, S.M.; Singer, R.H.; Garcia-Blanco, M.A. Asymmetric distribution of nuclear pore complexes and the cytoplasmic localization of beta2-tubulin mRNA in Chlamydomonas reinhardtii. Dev. Cell 2003, 4, 941–952. [Google Scholar]
- Powrie, E.A.; Zenklusen, D.; Singer, R.H. A nucleoporin, Nup60p, affects the nuclear and cytoplasmic localization of ASH1 mRNA in S. cerevisiae. RNA 2011, 17, 134–144. [Google Scholar]
- Meier, I. mRNA export and sumoylation-Lessons from plants. Biochim. Biophys. Acta 2012, 1819, 531–537. [Google Scholar]
- MacGregor, D.R.; Gould, P.; Foreman, J.; Griffiths, J.; Bird, S.; Page, R.; Stewart, K.; Steel, G.; Young, J.; Paszkiewicz, K.; et al. High expression of osmotically responsive genes1 is required for circadian periodicity through the promotion of nucleo-cytoplasmic mRNA export in Arabidopsis. Plant Cell 2013, 25, 4391–4404. [Google Scholar] [CrossRef] [PubMed]
- Faria, A.M.; Levay, A.; Wang, Y.; Kamphorst, A.O.; Rosa, M.L.; Nussenzveig, D.R.; Balkan, W.; Chook, Y.M.; Levy, D.E.; Fontoura, B.M. The nucleoporin Nup96 is required for proper expression of interferon-regulated proteins and functions. Immunity 2006, 24, 295–304. [Google Scholar] [CrossRef] [PubMed]
- Saavedra, C.; Tung, K.S.; Amberg, D.C.; Hopper, A.K.; Cole, C.N. Regulation of mRNA export in response to stress in Saccharomyces cerevisiae. Genes Dev. 1996, 10, 1608–1620. [Google Scholar] [CrossRef] [PubMed]
- Gallouzi, I.E.; Brennan, C.M.; Stenberg, M.G.; Swanson, M.S.; Eversole, A.; Maizels, N.; Steitz, J.A. HuR binding to cytoplasmic mRNA is perturbed by heat shock. Proc. Natl. Acad. Sci. USA 2000, 97, 3073–3078. [Google Scholar] [CrossRef] [PubMed]
- Rollenhagen, C.; Hodge, C.A.; Cole, C.N. Following temperature stress, export of heat shock mRNA occurs efficiently in cells with mutations in genes normally important for mRNA export. Eukaryot. Cell 2007, 6, 505–513. [Google Scholar] [CrossRef] [PubMed]
- Carmody, S.R.; Tran, E.J.; Apponi, L.H.; Corbett, A.H.; Wente, S.R. The mitogen-activated protein kinase Slt2 regulates nuclear retention of non-heat shock mRNAs during heat shock-induced stress. Mol. Cell. Biol. 2010, 30, 5168–5179. [Google Scholar] [PubMed]
- Rajanala, K.; Sarkar, A.; Jhingan, G.D.; Priyadarshini, R.; Jalan, M.; Sengupta, S.; Nandicoori, V.K. Phosphorylation of nucleoporin Tpr governs its differential localization and is required for its mitotic function. J. Cell Sci. 2014, 127, 3505–3520. [Google Scholar] [CrossRef] [PubMed]
- Skaggs, H.S.; Xing, H.; Wilkerson, D.C.; Murphy, L.A.; Hong, Y.; Mayhew, C.N.; Sarge, K.D. HSF1-TPR interaction facilitates export of stress-induced HSP70 mRNA. J. Biol. Chem. 2007, 282, 33902–33907. [Google Scholar] [CrossRef] [PubMed]
- Regot, S.; de Nadal, E.; Rodriguez-Navarro, S.; Gonzalez-Novo, A.; Perez-Fernandez, J.; Gadal, O.; Seisenbacher, G.; Ammerer, G.; Posas, F. The Hog1 stress-activated protein kinase targets nucleoporins to control mRNA export upon stress. J. Biol. Chem. 2013, 288, 17384–17398. [Google Scholar] [CrossRef] [PubMed]
- Izawa, S.; Takemura, R.; Inoue, Y. Gle2p is essential to induce adaptation of the export of bulk poly(A)+ mRNA to heat shock in Saccharomyces cerevisiae. J. Biol. Chem. 2004, 279, 35469–35478. [Google Scholar] [CrossRef] [PubMed]
- Takemura, R.; Inoue, Y.; Izawa, S. Stress response in yeast mRNA export factor: Reversible changes in Rat8p localization are caused by ethanol stress but not heat shock. J. Cell Sci. 2004, 117, 4189–4197. [Google Scholar] [CrossRef] [PubMed]
- Basel-Vanagaite, L.; Muncher, L.; Straussberg, R.; Pasmanik-Chor, M.; Yahav, M.; Rainshtein, L.; Walsh, C.A.; Magal, N.; Taub, E.; Drasinover, V.; et al. Mutated nup62 causes autosomal recessive infantile bilateral striatal necrosis. Ann. Neurol. 2006, 60, 214–222. [Google Scholar]
- Le Sage, V.; Mouland, A.J. Viral subversion of the nuclear pore complex. Viruses 2013, 5, 2019–2042. [Google Scholar]
- Simon, D.N.; Rout, M.P. Cancer and the nuclear pore complex. Adv. Exp. Med. Biol. 2014, 773, 285–307. [Google Scholar] [PubMed]
- Zhang, X.; Chen, S.; Yoo, S.; Chakrabarti, S.; Zhang, T.; Ke, T.; Oberti, C.; Yong, S.L.; Fang, F.; Li, L.; et al. Mutation in nuclear pore component NUP155 leads to atrial fibrillation and early sudden cardiac death. Cell 2008, 135, 1017–1027. [Google Scholar] [CrossRef] [PubMed]
- Kendirgi, F.; Rexer, D.J.; Alcazar-Roman, A.R.; Onishko, H.M.; Wente, S.R. Interaction between the shuttling mRNA export factor Gle1 and the nucleoporin hCG1: A conserved mechanism in the export of Hsp70 mRNA. Mol. Biol. Cell 2005, 16, 4304–4315. [Google Scholar] [CrossRef] [PubMed]
- Nousiainen, H.O.; Kestila, M.; Pakkasjarvi, N.; Honkala, H.; Kuure, S.; Tallila, J.; Vuopala, K.; Ignatius, J.; Herva, R.; Peltonen, L. Mutations in mRNA export mediator GLE1 result in a fetal motoneuron disease. Nat. Genet. 2008, 40, 155–157. [Google Scholar] [CrossRef] [PubMed]
- Jao, L.E.; Appel, B.; Wente, S.R. A zebrafish model of lethal congenital contracture syndrome 1 reveals Gle1 function in spinal neural precursor survival and motor axon arborization. Development 2012, 139, 1316–1326. [Google Scholar] [CrossRef] [PubMed]
- Folkmann, A.W.; Collier, S.E.; Zhan, X.; Aditi; Ohi, M.D.; Wente, S.R. Gle1 functions during mRNA export in an oligomeric complex that is altered in human disease. Cell 2013, 155, 582–593. [Google Scholar] [CrossRef] [PubMed]
- Folkmann, A.W.; Dawson, T.R.; Wente, S.R. Insights into mRNA export-linked molecular mechanisms of human disease through a Gle1 structure-function analysis. Adv. Biol. Regul. 2014, 54, 74–91. [Google Scholar] [CrossRef] [PubMed]
- Borden, K.L.; Culjkovic-Kraljacic, B. Ribavirin as an anti-cancer therapy: Acute myeloid leukemia and beyond? Leuk. Lymphoma 2010, 51, 1805–1815. [Google Scholar] [CrossRef]
- Culjkovic-Kraljacic, B.; Baguet, A.; Volpon, L.; Amri, A.; Borden, K.L. The oncogene eIF4E reprograms the nuclear pore complex to promote mRNA export and oncogenic transformation. Cell Rep. 2012, 2, 207–215. [Google Scholar]
- Castello, A.; Izquierdo, J.M.; Welnowska, E.; Carrasco, L. RNA nuclear export is blocked by poliovirus 2A protease and is concomitant with nucleoporin cleavage. J. Cell Sci. 2009, 122, 3799–3809. [Google Scholar] [CrossRef] [PubMed]
- Gustin, K.E.; Sarnow, P. Inhibition of nuclear import and alteration of nuclear pore complex composition by rhinovirus. J. Virol. 2002, 76, 8787–8796. [Google Scholar] [CrossRef] [PubMed]
- Lidsky, P.V.; Hato, S.; Bardina, M.V.; Aminev, A.G.; Palmenberg, A.C.; Sheval, E.V.; Polyakov, V.Y.; van Kuppeveld, F.J.; Agol, V.I. Nucleocytoplasmic traffic disorder induced by cardioviruses. J. Virol. 2006, 80, 2705–2717. [Google Scholar] [CrossRef] [PubMed]
- Porter, F.W.; Bochkov, Y.A.; Albee, A.J.; Wiese, C.; Palmenberg, A.C. A picornavirus protein interacts with Ran-GTPase and disrupts nucleocytoplasmic transport. Proc. Natl. Acad. Sci. USA 2006, 103, 12417–12422. [Google Scholar] [CrossRef] [PubMed]
- Park, N.; Skern, T.; Gustin, K.E. Specific cleavage of the nuclear pore complex protein Nup62 by a viral protease. J. Biol. Chem. 2010, 285, 28796–28805. [Google Scholar] [CrossRef] [PubMed]
- Bardina, M.V.; Lidsky, P.V.; Sheval, E.V.; Fominykh, K.V.; van Kuppeveld, F.J.; Polyakov, V.Y.; Agol, V.I. Mengovirus-induced rearrangement of the nuclear pore complex: Hijacking cellular phosphorylation machinery. J. Virol. 2009, 83, 3150–3161. [Google Scholar] [CrossRef] [PubMed]
- Porter, F.W.; Palmenberg, A.C. Leader-induced phosphorylation of nucleoporins correlates with nuclear trafficking inhibition by cardioviruses. J. Virol. 2009, 83, 1941–1951. [Google Scholar] [CrossRef] [PubMed]
- Ricour, C.; Delhaye, S.; Hato, S.V.; Olenyik, T.D.; Michel, B.; van Kuppeveld, F.J.; Gustin, K.E.; Michiels, T. Inhibition of mRNA export and dimerization of interferon regulatory factor 3 by Theiler’s virus leader protein. J. Gen. Virol. 2009, 90, 177–186. [Google Scholar] [CrossRef] [PubMed]
- Gustin, K.E.; Sarnow, P. Effects of poliovirus infection on nucleo-cytoplasmic trafficking and nuclear pore complex composition. EMBO J. 2001, 20, 240–249. [Google Scholar] [CrossRef] [PubMed]
- Her, L.S.; Lund, E.; Dahlberg, J.E. Inhibition of Ran guanosine triphosphatase-dependent nuclear transport by the matrix protein of vesicular stomatitis virus. Science 1997, 276, 1845–1848. [Google Scholar] [CrossRef] [PubMed]
- Petersen, J.M.; Her, L.S.; Dahlberg, J.E. Multiple vesiculoviral matrix proteins inhibit both nuclear export and import. Proc. Natl. Acad. Sci. USA 2001, 98, 8590–8595. [Google Scholar] [CrossRef] [PubMed]
- Von Kobbe, C.; van Deursen, J.M.; Rodrigues, J.P.; Sitterlin, D.; Bachi, A.; Wu, X.; Wilm, M.; Carmo-Fonseca, M.; Izaurralde, E. Vesicular stomatitis virus matrix protein inhibits host cell gene expression by targeting the nucleoporin Nup98. Mol. Cell 2000, 6, 1243–1252. [Google Scholar]
- Chakraborty, P.; Seemann, J.; Mishra, R.K.; Wei, J.H.; Weil, L.; Nussenzveig, D.R.; Heiber, J.; Barber, G.N.; Dasso, M.; Fontoura, B.M. Vesicular stomatitis virus inhibits mitotic progression and triggers cell death. EMBO Rep. 2009, 10, 1154–1160. [Google Scholar] [CrossRef] [PubMed]
- Rajani, K.R.; Pettit Kneller, E.L.; McKenzie, M.O.; Horita, D.A.; Chou, J.W.; Lyles, D.S. Complexes of vesicular stomatitis virus matrix protein with host Rae1 and Nup98 involved in inhibition of host transcription. PLoS Pathog. 2012, 8, e1002929. [Google Scholar] [CrossRef] [PubMed]
- Quan, B.; Seo, H.S.; Blobel, G.; Ren, Y. Vesiculoviral matrix (M) protein occupies nucleic acid binding site at nucleoporin pair (Rae1*Nup98). Proc. Natl Acad. Sci. USA 2014, 111, 9127–9132. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Huang, S.; Chen, Z. Human cellular protein nucleoporin hNup98 interacts with influenza A virus NS2/nuclear export protein and overexpression of its GLFG repeat domain can inhibit virus propagation. J. Gen. Virol. 2010, 91, 2474–2484. [Google Scholar] [CrossRef] [PubMed]
- Enninga, J.; Levy, D.E.; Blobel, G.; Fontoura, B.M. Role of nucleoporin induction in releasing an mRNA nuclear export block. Science 2002, 295, 1523–1525. [Google Scholar] [CrossRef] [PubMed]
- Faria, P.A.; Chakraborty, P.; Levay, A.; Barber, G.N.; Ezelle, H.J.; Enninga, J.; Arana, C.; van Deursen, J.; Fontoura, B.M. VSV disrupts the Rae1/mrnp41 mRNA nuclear export pathway. Mol. Cell 2005, 17, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Hayakawa, A.; Babour, A.; Sengmanivong, L.; Dargemont, C. Ubiquitylation of the nuclear pore complex controls nuclear migration during mitosis in S. cerevisiae. J. Cell Biol. 2012, 196, 19–27. [Google Scholar]
- Vassileva, M.T.; Matunis, M.J. SUMO modification of heterogeneous nuclear ribonucleoproteins. Mol. Cell. Biol. 2004, 24, 3623–3632. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Bonnet, A.; Palancade, B. Regulation of mRNA Trafficking by Nuclear Pore Complexes. Genes 2014, 5, 767-791. https://doi.org/10.3390/genes5030767
Bonnet A, Palancade B. Regulation of mRNA Trafficking by Nuclear Pore Complexes. Genes. 2014; 5(3):767-791. https://doi.org/10.3390/genes5030767
Chicago/Turabian StyleBonnet, Amandine, and Benoit Palancade. 2014. "Regulation of mRNA Trafficking by Nuclear Pore Complexes" Genes 5, no. 3: 767-791. https://doi.org/10.3390/genes5030767
APA StyleBonnet, A., & Palancade, B. (2014). Regulation of mRNA Trafficking by Nuclear Pore Complexes. Genes, 5(3), 767-791. https://doi.org/10.3390/genes5030767





