Integrating Transcriptomics, Network Pharmacology, and Machine Learning to Reveal Transglutaminase 2 (TGM2) as a Key Target Mediating Taurocholate Efficacy in Colitis
Abstract
1. Introduction
2. Materials and Methods
2.1. Acquisition of Relevant Targets of TCA
2.2. Acquisition of UC-Related Targets
2.3. Potential Target Prediction of TCA in UC Treatment
2.4. Functional Enrichment Analysis
2.5. Determination of Hub Genes with Machine Learning
2.6. Construction of Artificial Neural Network Model
2.7. Gene Set Enrichment Analysis (GSEA)
2.8. Methods for Immune Infiltration Analysis
2.9. Chromosome Localization
2.10. Molecular Docking Verification
2.11. Validation of Experimental Protocols in Mice
2.12. Statistical Analysis
3. Results
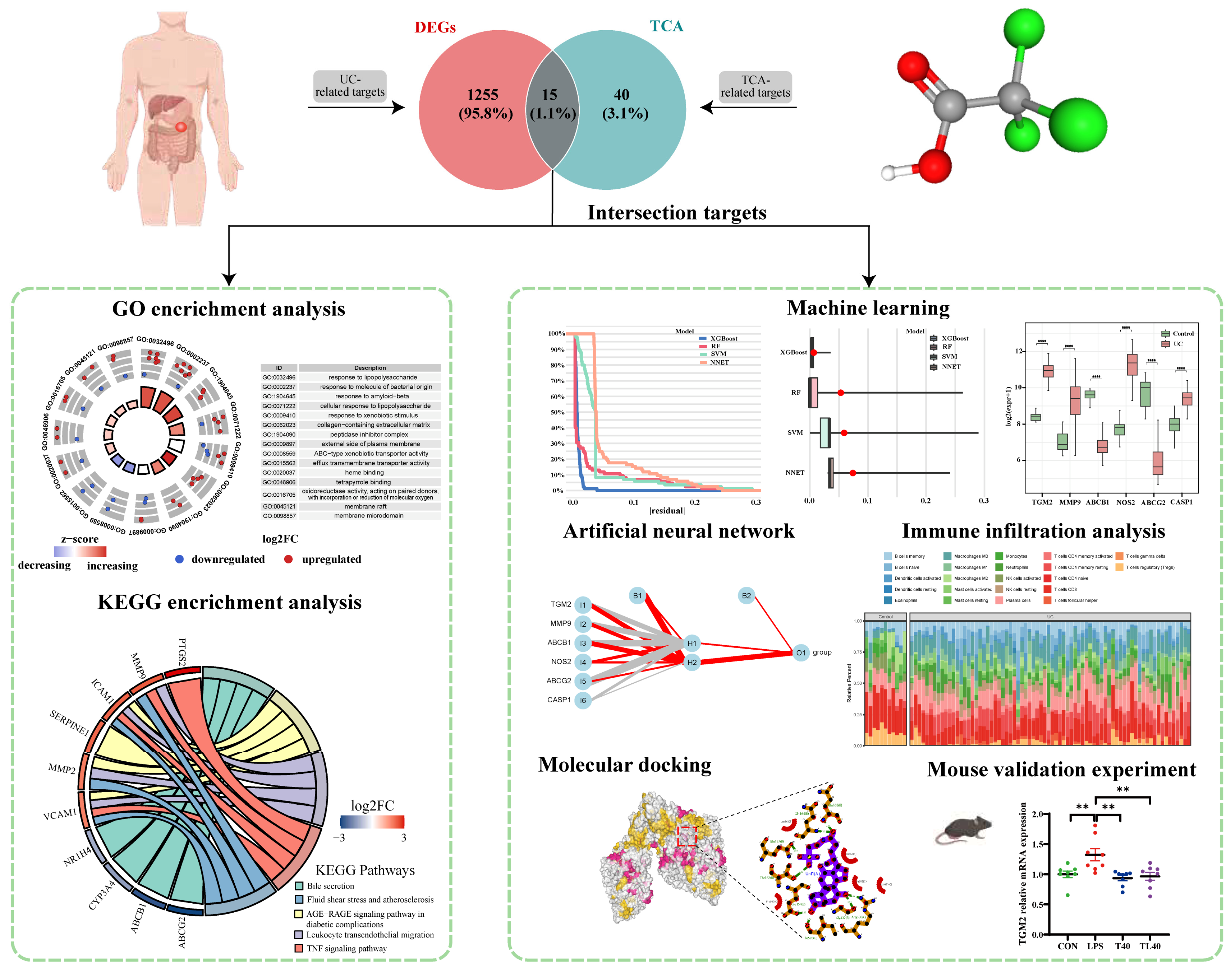
3.1. Acquisition of TCA Targets and Disease Targets and Functional Enrichment Analysis of Intersecting Genes
3.2. Determination of Target Hub Genes with Machine Learning
3.3. Evaluation of Artificial Neural Network Performance
3.4. Enrichment Analysis of Key Genes
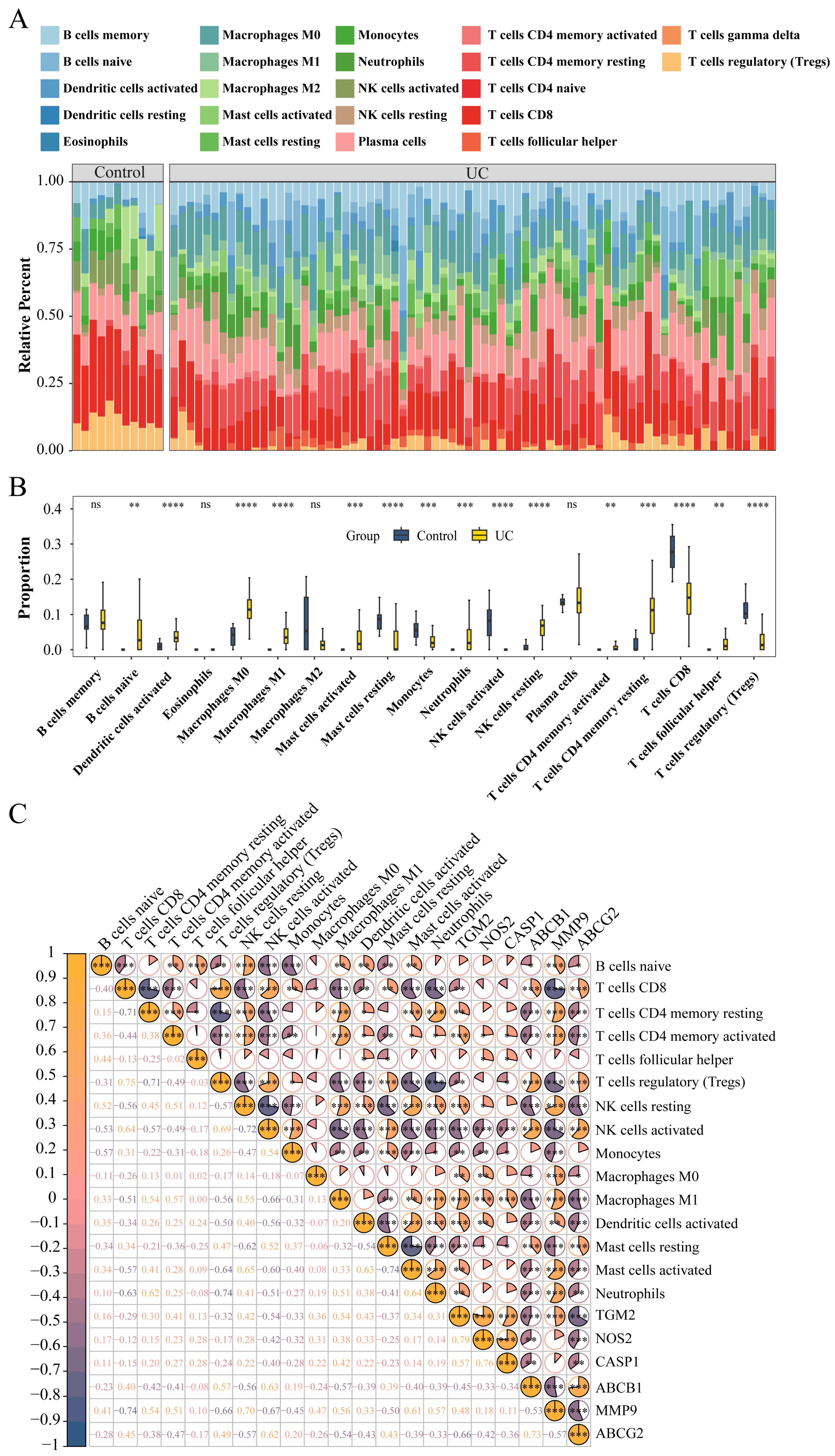
3.5. Evaluation of Immune Cell Infiltration
3.6. Chromosome Localization Analysis
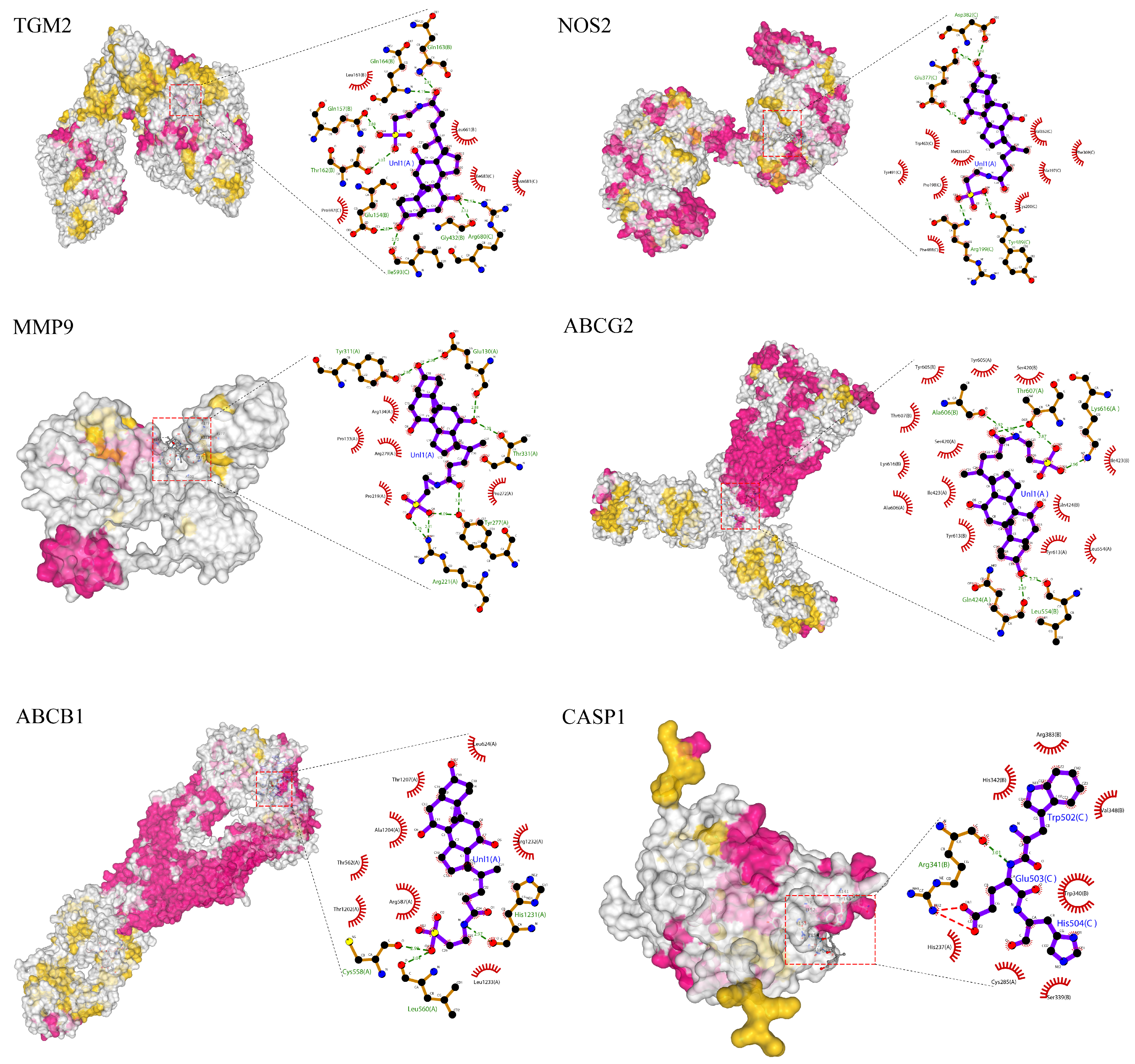
3.7. Molecular Docking of the Key Compounds and Core Targets
3.8. Validation Experiment of the Key Compounds and Core Targets
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Danpanichkul, P.; Duangsonk, K.; Ho, A.H.; Laoveeravat, P.; Vuthithammee, C.; Dejvajara, D.; Prasitsumrit, V.; Auttapracha, T.; Songtanin, B.; Wetwittayakhlang, P.; et al. Disproportionately Increasing Incidence of Inflammatory Bowel Disease in Females and the Elderly: An Update Analysis from the Global Burden of Disease Study 2021. Am. J. Gastroenterol. 2024, 120, 675–680. [Google Scholar] [CrossRef]
- Kobayashi, T.; Siegmund, B.; Le Berre, C.; Wei, S.C.; Ferrante, M.; Shen, B.; Bernstein, C.N.; Danese, S.; Peyrin-Biroulet, L.; Hibi, T. Ulcerative colitis. Nat. Rev. Dis. Prim. 2020, 6, 74. [Google Scholar] [CrossRef]
- Vermeire, S.; Solitano, V.; Peyrin-Biroulet, L.; Tilg, H.; Danese, S.; Ehrlich, H.; Scherrer, D.; Gineste, P.; d’Agay, L.; Sands, B.E. Obefazimod: A First-in-class Drug for the Treatment of Ulcerative Colitis. J. Crohns Colitis 2023, 17, 1689–1697. [Google Scholar] [CrossRef] [PubMed]
- Qi, S.; Liang, X.; Wang, Z.; Jin, H.; Zou, L.; Yang, J. Potential Mechanism of Tibetan Medicine Liuwei Muxiang Pills against Colorectal Cancer: Network Pharmacology and Bioinformatics Analyses. Pharmaceuticals 2024, 17, 429. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Lyu, X.; Markhard, A.L.; Fu, S.; Mardjuki, R.E.; Cavanagh, P.E.; Zeng, X.; Rajniak, J.; Lu, N.; Xiao, S.; et al. Pter is a N-Acetyltaurine Hydrolase That Regulates Feeding and Obesity. Nature 2024, 633, 182–188. [Google Scholar] [CrossRef]
- Singh, P.; Gollapalli, K.; Mangiola, S.; Schranner, D.; Yusuf, M.A.; Chamoli, M.; Shi, S.L.; Lopes Bastos, B.; Nair, T.; Riermeier, A.; et al. Taurine Deficiency as a Driver of Aging. Science 2023, 380, eabn9257. [Google Scholar] [CrossRef]
- Cao, T.; Zhang, W.; Wang, Q.; Wang, C.; Ma, W.; Zhang, C.; Ge, M.; Tian, M.; Yu, J.; Jiao, A.; et al. Cancer Slc6a6-Mediated Taurine Uptake Transactivates Immune Checkpoint Genes and Induces Exhaustion in Cd8(+) T cells. Cell 2024, 187, 2288–2304.e27. [Google Scholar] [CrossRef]
- Mishra, S.; Kell, P.; Scherrer, D.; Dietzen, D.J.; Vite, C.H.; Berry-Kravis, E.; Davidson, C.; Cologna, S.M.; Porter, F.D.; Ory, D.S.; et al. Accumulation of Alkyl-Lysophosphatidylcholines in Niemann-Pick Disease Type C1. J. Lipid Res. 2024, 65, 100600. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Liang, J.; Zhu, S.; Zhao, W.; Zhang, Y.; Sun, W. Protective Effect of Taurohyodeoxycholic Acid from Pulvis Fellis Suis on Trinitrobenzene Sulfonic Acid Induced Ulcerative Colitis in Mice. Eur. J. Pharmacol. 2011, 670, 229–235. [Google Scholar] [CrossRef]
- Martínez-Moya, P.; Romero-Calvo, I.; Requena, P.; Hernández-Chirlaque, C.; Aranda, C.J.; González, R.; Zarzuelo, A.; Suárez, M.D.; Martínez-Augustin, O.; Marín, J.J.; et al. Dose-Dependent Antiinflammatory Effect of Ursodeoxycholic Acid in Experimental Colitis. Int. Immunopharmacol. 2013, 15, 372–380. [Google Scholar] [CrossRef]
- Haldar, S.; Yhome, N.; Muralidaran, Y.; Rajagopal, S.; Mishra, P. Nanoplastics Toxicity Specific to Liver in Inducing Metabolic Dysfunction—a Comprehensive Review. Genes 2023, 14, 590. [Google Scholar] [CrossRef]
- Yang, Y.; He, J.; Suo, Y.; Zheng, Z.; Wang, J.; Lv, L.; Huo, C.; Wang, Z.; Li, J.; Sun, W.; et al. Tauroursodeoxycholate Improves 2,4,6-Trinitrobenzenesulfonic Acid-Induced Experimental Acute Ulcerative Colitis in Mice. Int. Immunopharmacol. 2016, 36, 271–276. [Google Scholar] [CrossRef] [PubMed]
- Izhar, R.; Borriello, M.; La Russa, A.; Di Paola, R.; De, A.; Capasso, G.; Ingrosso, D.; Perna, A.F.; Simeoni, M. Fabry Disease in Women: Genetic Basis, Available Biomarkers, and Clinical Manifestations. Genes 2024, 15, 37. [Google Scholar] [CrossRef] [PubMed]
- Sinha, D.; Maurya, A.K.; Abdi, G.; Majeed, M.; Agarwal, R.; Mukherjee, R.; Ganguly, S.; Aziz, R.; Bhatia, M.; Majgaonkar, A.; et al. Integrated Genomic Selection for Accelerating Breeding Programs of Climate-Smart Cereals. Genes 2023, 14, 1484. [Google Scholar] [CrossRef]
- Kim, E.; Yang, S.-M.; Ham, J.-H.; Lee, W.; Jung, D.-H.; Kim, H.-Y. Integration of Maldi-Tof Ms and Machine Learning to Classify Enterococci: A Comparative Analysis of Supervised Learning Algorithms for Species Prediction. Food Chem. 2025, 462, 140931. [Google Scholar] [CrossRef]
- Wang, Y.; Zeng, L.; Deng, W.; Wang, J.; Zhang, J. The Molecular Reactive Pathway between Lipoxygenase and Lipase and Reactive Species Generated in Dielectric Barrier Discharge Atmospheric Cold Plasma: An Investigation Using Molecular Docking. Food Chem. 2025, 465, 141973. [Google Scholar] [CrossRef]
- Davis, A.P.; Wiegers, T.C.; Sciaky, D.; Barkalow, F.; Strong, M.; Wyatt, B.; Wiegers, J.; McMorran, R.; Abrar, S.; Mattingly, C.J. Comparative Toxicogenomics Database’s 20th Anniversary: Update 2025. Nucleic Acids Res. 2024, 53, D1328–D1334. [Google Scholar] [CrossRef] [PubMed]
- Yao, Z.-J.; Dong, J.; Che, Y.-J.; Zhu, M.-F.; Wen, M.; Wang, N.-N.; Wang, S.; Lu, A.-P.; Cao, D.-S. Targetnet: A Web Service for Predicting Potential Drug–Target Interaction Profiling Via Multi-Target Sar Models. J. Comput. Mol. Des. 2016, 30, 413–424. [Google Scholar] [CrossRef]
- UniProt Consortium. Uniprot: A Worldwide Hub of Protein Knowledge. Nucleic Acids Res. 2019, 47, D506–D515. [Google Scholar] [CrossRef]
- Vancamelbeke, M.; Vanuytsel, T.; Farré, R.; Verstockt, S.; Ferrante, M.; Van Assche, G.; Rutgeerts, P.; Schuit, F.; Vermeire, S.; Arijs, I.; et al. Genetic and Transcriptomic Bases of Intestinal Epithelial Barrier Dysfunction in Inflammatory Bowel Disease. Inflamm. Bowel Dis. 2017, 23, 1718–1729. [Google Scholar] [CrossRef]
- Arijs, I.; De Hertogh, G.; Lemmens, B.; Van Lommel, L.; de Bruyn, M.; Vanhove, W.; Cleynen, I.; Machiels, K.; Ferrante, M.; Schuit, F.; et al. Effect of Vedolizumab (Anti-A4β7-Integrin) Therapy on Histological Healing and Mucosal Gene Expression in Patients with Uc. Gut 2018, 67, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Strauss, R.; Ouahed, J.; Chan, D.; Telesco, S.E.; Shouval, D.S.; Canavan, J.B.; Brodmerkel, C.; Snapper, S.B.; Friedman, J.R. Molecular Comparison of Adult and Pediatric Ulcerative Colitis Indicates Broad Similarity of Molecular Pathways in Disease Tissue. J. Pediatr. Gastroenterol. Nutr. 2018, 67, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Guan, X. Regulation of Porcine Intestinal Barrier Function by Taurine-Conjugated Bile Acids and Validation in Mice. Ph.D. Thesis, Yunnan Agricultural University, Kunming, China, 2024. [Google Scholar]
- Kang, M.J.; Kim, S.Y.; Na, D.L.; Kim, B.C.; Yang, D.W.; Kim, E.J.; Na, H.R.; Han, H.J.; Lee, J.H.; Kim, J.H.; et al. Prediction of Cognitive Impairment Via Deep Learning Trained with Multi-Center Neuropsychological Test Data. BMC Med. Inf. Decis. Mak. 2019, 19, 231. [Google Scholar] [CrossRef]
- Gajendran, M.; Loganathan, P.; Catinella, A.P.; Hashash, J.G. A Comprehensive Review and Update on Crohn’s Disease. Dis. Mon. 2018, 64, 20–57. [Google Scholar] [CrossRef]
- Wu, Z.; Huang, S.; Li, T.; Li, N.; Han, D.; Zhang, B.; Xu, Z.Z.; Zhang, S.; Pang, J.; Wang, S.; et al. Gut Microbiota from Green Tea Polyphenol-Dosed Mice Improves Intestinal Epithelial Homeostasis and Ameliorates Experimental Colitis. Microbiome 2021, 9, 184. [Google Scholar] [CrossRef]
- Cheng, H.; Liu, J.; Zhang, D.; Wang, J.; Tan, Y.; Feng, W.; Peng, C. Ginsenoside Rg1 Alleviates Acute Ulcerative Colitis by Modulating Gut Microbiota and Microbial Tryptophan Metabolism. Front. Immunol. 2022, 13, 817600. [Google Scholar] [CrossRef]
- Vandewalle, J.; Luypaert, A.; De Bosscher, K.; Libert, C. Therapeutic Mechanisms of Glucocorticoids. Trends Endocrinol. Metab. 2018, 29, 42–54. [Google Scholar] [CrossRef]
- Murray, A.; Nguyen, T.M.; Parker, C.E.; Feagan, B.G.; MacDonald, J.K. Oral 5-Aminosalicylic Acid for Maintenance of Remission in Ulcerative Colitis. Cochrane Database Syst. Rev. 2020, 8, Cd000544. [Google Scholar] [CrossRef]
- Wei, M.; Lyu, P.; Li, P.; Hu, J.; Wu, R.; Ouyang, Q.; Guo, K. Baolier Capsule’s Secret Weapon: Piperine Boosts Cholesterol Excretion to Combat Atherosclerosis. Drug Des. Dev. Ther. 2024, 18, 6427–6446. [Google Scholar] [CrossRef]
- Gorenko, Z.A.; Grinchenko, O.A.; Veselsky, S.P.; Baban, V.M. Composition of Gastric Juice and Bile in rats at the Experimental Chronic Pancreatitis. Fiziolohichnyi Zhurnal 2015, 61, 86–95. [Google Scholar] [CrossRef] [PubMed]
- Jagdale, A.D.; Angal, M.M.; Patil, R.S.; Tupe, R.S. Exploring the Glycation Association with Dyslipidaemia: Novel Approach for Diabetic Nephropathy. Biochem. Pharmacol. 2024, 229, 116513. [Google Scholar] [CrossRef]
- Wang, H.; Wang, Y.; Du, Q.; Lu, P.; Fan, H.; Lu, J.; Hu, R. Inflammasome-Independent Nlrp3 is Required for Epithelial-Mesenchymal Transition in Colon Cancer Cells. Exp. Cell Res. 2016, 342, 184–192. [Google Scholar] [CrossRef]
- Wang, D.; Li, J.; Sun, Y.; Ding, X.; Zhang, X.; Liu, S.; Han, B.; Wang, H.; Duan, X.; Sun, T. A Machine Learning Model for Accurate Prediction of Sepsis in Icu Patients. Front. Public Health 2021, 9, 754348. [Google Scholar] [CrossRef]
- Hong, W.; Zhang, Y.; Wang, S.; Zheng, D.; Hsu, S.; Zhou, J.; Fan, J.; Zeng, Z.; Wang, N.; Ding, Z.; et al. Deciphering the Immune Modulation through Deep Transcriptomic Profiling and Therapeutic Implications of DNA Damage Repair Pattern in Hepatocellular Carcinoma. Cancer Lett. 2024, 582, 216594. [Google Scholar] [CrossRef]
- Yang, P.; Yu, D.; Zhou, J.; Zhuang, S.; Jiang, T. TGM2 Interference Regulates the Angiogenesis and Apoptosis of Colorectal Cancer Via Wnt/β-Catenin Pathway. Cell Cycle 2019, 18, 1122–1134. [Google Scholar] [CrossRef]
- Regalado, C.R.; Balogh, M. Mmp9: Link between Neuropathy and Colorectal Cancer? Front. Mol. Biosci. 2024, 11, 1451611. [Google Scholar] [CrossRef] [PubMed]
- Eshelman, M.A.; Matthews, S.M.; Schleicher, E.M.; Fleeman, R.M.; Kawasawa, Y.I.; Stumpo, D.J.; Blackshear, P.J.; Koltun, W.A.; Ishmael, F.T.; Yochum, G.S. Tristetraprolin Targets Nos2 Expression in the Colonic Epithelium. Sci. Rep. 2019, 9, 14413. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.; Zhu, N.; Wang, D.; Zhou, Y.; Liu, Y. Comprehensive Analysis of Prognostic Value and Immune Infiltration of Nlrc4 and Casp1 in Colorectal Cancer. Int. J. Gen. Med. 2022, 15, 5425–5440. [Google Scholar] [CrossRef]
- Valbon, S.F.; Condotta, S.A.; Richer, M.J. Regulation of Effector and Memory CD8(+) T Cell Function by Inflammatory Cytokines. Cytokine 2016, 82, 16–23. [Google Scholar] [CrossRef] [PubMed]
- Valle-Mendiola, A.; Gutiérrez-Hoya, A.; Soto-Cruz, I. Jak/Stat Signaling and Cervical Cancer: From the Cell Surface to the Nucleus. Genes 2023, 14, 1141. [Google Scholar] [CrossRef]
- Wallace, K.L.; Zheng, L.B.; Kanazawa, Y.; Shih, D.Q. Immunopathology of Inflammatory Bowel Disease. World J. Gastroenterol. 2014, 20, 6–21. [Google Scholar] [CrossRef]
- Steinbach, E.C.; Plevy, S.E. The Role of Macrophages and Dendritic Cells in the Initiation of Inflammation in Ibd. Inflamm. Bowel Dis. 2014, 20, 166–175. [Google Scholar] [CrossRef]
- Cornice, J.; Verzella, D.; Arboretto, P.; Vecchiotti, D.; Capece, D.; Zazzeroni, F.; Franzoso, G. Nf-Κb: Governing Macrophages in Cancer. Genes 2024, 15, 197. [Google Scholar] [CrossRef]
- Hart, G.R.; Roffman, D.A.; Decker, R.; Deng, J. A Multi-Parameterized Artificial Neural Network for Lung Cancer Risk Prediction. PLoS ONE 2018, 13, e0205264. [Google Scholar] [CrossRef]
- Chakrabortty, A.; Patton, D.J.; Smith, B.F.; Agarwal, P. Mirnas: Potential as Biomarkers and Therapeutic Targets for Cancer. Genes 2023, 14, 1375. [Google Scholar] [CrossRef] [PubMed]
- Saviana, M.; Le, P.; Micalo, L.; Del Valle-Morales, D.; Romano, G.; Acunzo, M.; Li, H.; Nana-Sinkam, P. Crosstalk between Mirnas and DNA Methylation in Cancer. Genes 2023, 14, 1075. [Google Scholar] [CrossRef]
- Guo, W.; Gao, G.; Dai, J.; Sun, Q. Prediction of Lung Infection during Palliative Chemotherapy of Lung Cancer Based on Artificial Neural Network. Comput. Math. Methods Med. 2022, 2022, 4312117. [Google Scholar] [CrossRef]
- Lu, J.; Xue, Z.; Xu, B.B.; Wu, D.; Zheng, H.L.; Xie, J.W.; Wang, J.B.; Lin, J.X.; Chen, Q.Y.; Li, P.; et al. Application of an Artificial Neural Network for Predicting the Potential Chemotherapy Benefit of Patients with Gastric Cancer after Radical Surgery. Surgery 2022, 171, 955–965. [Google Scholar] [CrossRef] [PubMed]
- Lai, T.S.; Davies, C.; Greenberg, C. Tissue Transglutaminase Enhances Fibrin-Dependent Angiogenesis and Extracellular Matrix Formation during Tissue Repair by Altering Gene Expression and Is Inhibited by Aspirin. Blood 2009, 114, 3055. [Google Scholar] [CrossRef]
- Su, T.; Qin, X.-Y.; Furutani, Y. Transglutaminase 2 as a Marker for Inflammation and Therapeutic Target in Sepsis. Int. J. Mol. Sci. 2021, 22, 1897. [Google Scholar] [CrossRef]
- Zhang, M.; Liu, T.; Luo, L.; Zhang, Y.; Chen, Q.; Wang, F.; Xie, Y. Common Diagnostic Biomarkers and Molecular Mechanisms of Helicobacter Pylori Infection and Inflammatory Bowel Disease. Front. Immunol. 2024, 15, 1492810. [Google Scholar] [CrossRef]
- Yao, Z.; Fan, Y.; Lin, L.; Kellems, R.E.; Xia, Y. Tissue Transglutaminase: A Multifunctional and Multisite Regulator in Health and Disease. Physiol. Rev. 2024, 104, 281–325. [Google Scholar] [CrossRef] [PubMed]
- Englund, G.; Jacobson, A.; Rorsman, F.; Artursson, P.; Kindmark, A.; Rönnblom, A. Efflux Transporters in Ulcerative Colitis: Decreased Expression of Bcrp (Abcg2) and Pgp (Abcb1). Inflamm. Bowel Dis. 2007, 13, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Chiang, J.Y.L.; Ferrell, J.M. Bile Acid Receptors Fxr and Tgr5 Signaling in Fatty Liver Diseases and Therapy. Am. J. Physiol. Gastrointest. Liver Physiol. 2020, 318, G554–G573. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; He, J.; Suo, Y.; Lv, L.; Wang, J.; Huo, C.; Zheng, Z.; Wang, Z.; Li, J.; Sun, W.; et al. Anti-Inflammatory Effect of Taurocholate on Tnbs-Induced Ulcerative Colitis in Mice. Biomed. Pharmacother. 2016, 81, 424–430. [Google Scholar] [CrossRef]
- Calzadilla, N.; Comiskey, S.M.; Dudeja, P.K.; Saksena, S.; Gill, R.K.; Alrefai, W.A. Bile Acids as Inflammatory Mediators and Modulators of Intestinal Permeability. Front. Immunol. 2022, 13, 1021924. [Google Scholar] [CrossRef]
- Zhou, G.; Zhang, M.; Zheng, S.; Yang, G.; Li, L.; Huang, S.; Zeng, Z.; Chen, R.; Zhang, S.; Chen, M. Transglutaminase 2 Modulates Inflammatory Angiogenesis via Vascular Endothelial Growth Factor Receptor 2 Pathway in Inflammatory Bowel Disease. J. Adv. Res. 2025. [Google Scholar] [CrossRef]






Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, J.; Jia, H.; Yi, L.; Song, G.; Fu, P.; Cheng, W.; Xie, Y.; Shi, W.; Zhao, S. Integrating Transcriptomics, Network Pharmacology, and Machine Learning to Reveal Transglutaminase 2 (TGM2) as a Key Target Mediating Taurocholate Efficacy in Colitis. Genes 2025, 16, 1024. https://doi.org/10.3390/genes16091024
Zhu J, Jia H, Yi L, Song G, Fu P, Cheng W, Xie Y, Shi W, Zhao S. Integrating Transcriptomics, Network Pharmacology, and Machine Learning to Reveal Transglutaminase 2 (TGM2) as a Key Target Mediating Taurocholate Efficacy in Colitis. Genes. 2025; 16(9):1024. https://doi.org/10.3390/genes16091024
Chicago/Turabian StyleZhu, Junhong, Huijin Jia, Lanlan Yi, Guangyao Song, Pengfei Fu, Wenjie Cheng, Yuxiao Xie, Wenzhe Shi, and Sumei Zhao. 2025. "Integrating Transcriptomics, Network Pharmacology, and Machine Learning to Reveal Transglutaminase 2 (TGM2) as a Key Target Mediating Taurocholate Efficacy in Colitis" Genes 16, no. 9: 1024. https://doi.org/10.3390/genes16091024
APA StyleZhu, J., Jia, H., Yi, L., Song, G., Fu, P., Cheng, W., Xie, Y., Shi, W., & Zhao, S. (2025). Integrating Transcriptomics, Network Pharmacology, and Machine Learning to Reveal Transglutaminase 2 (TGM2) as a Key Target Mediating Taurocholate Efficacy in Colitis. Genes, 16(9), 1024. https://doi.org/10.3390/genes16091024






